Predicting the Potential Distribution of the Endangered Plant Eucommia ulmoides in China under the Background of Climate Change
Abstract
:1. Introduction
2. Materials and Methods
2.1. Research Area
2.2. Species Occurrence Data
2.3. Environmental Variable Screening and Data Processing
2.4. Model Establishment, Optimization, and Evaluation
2.5. Classification of Suitable Areas and Modeling Accuracy Evaluation
2.6. Analysis of Low Impact Areas
2.7. Analysis of Spatial Pattern Change
2.8. Centroid Transfer Analysis
3. Results
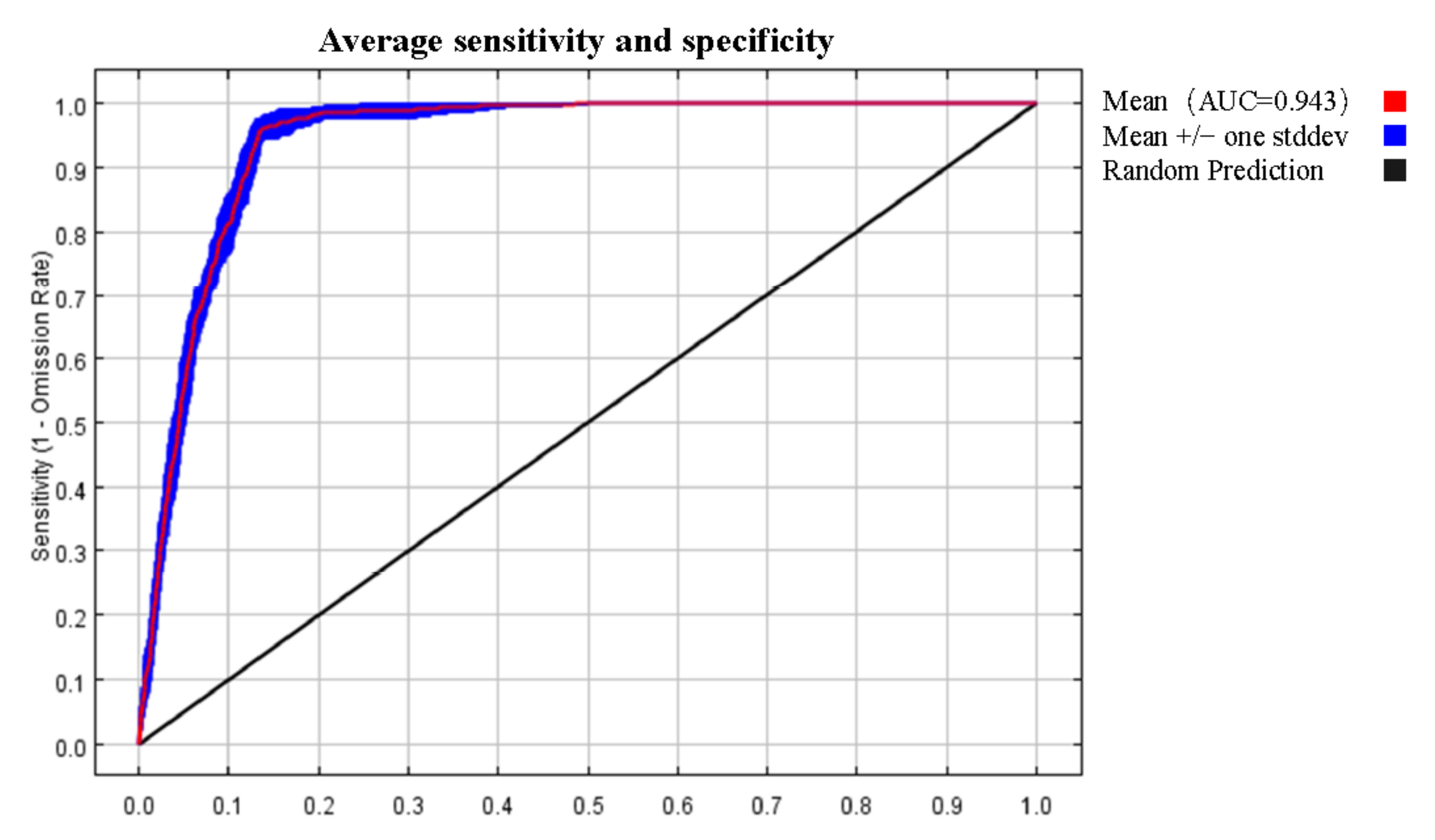
3.1. Analysis of the Model Accuracy and Classification of Suitable Regions
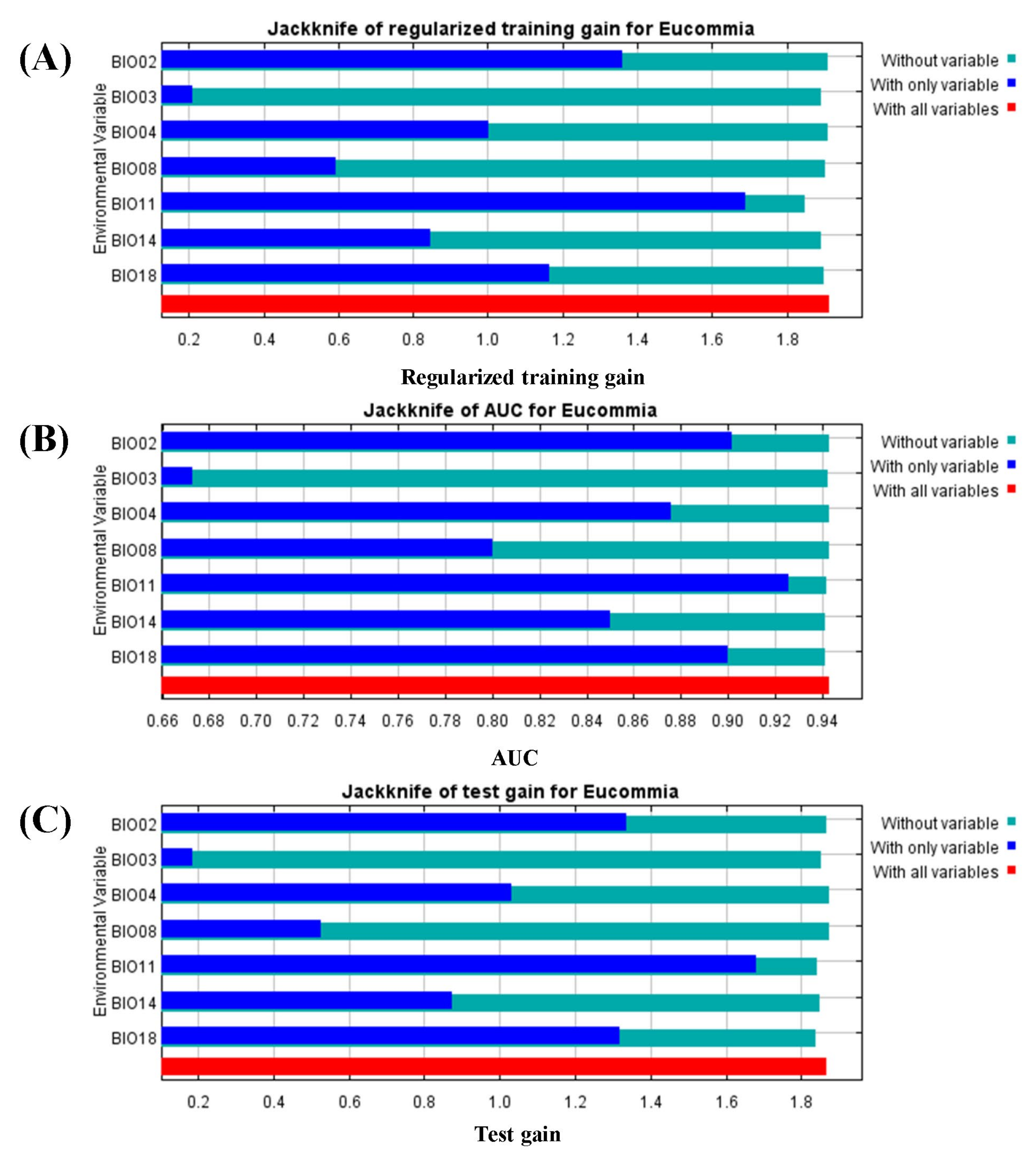
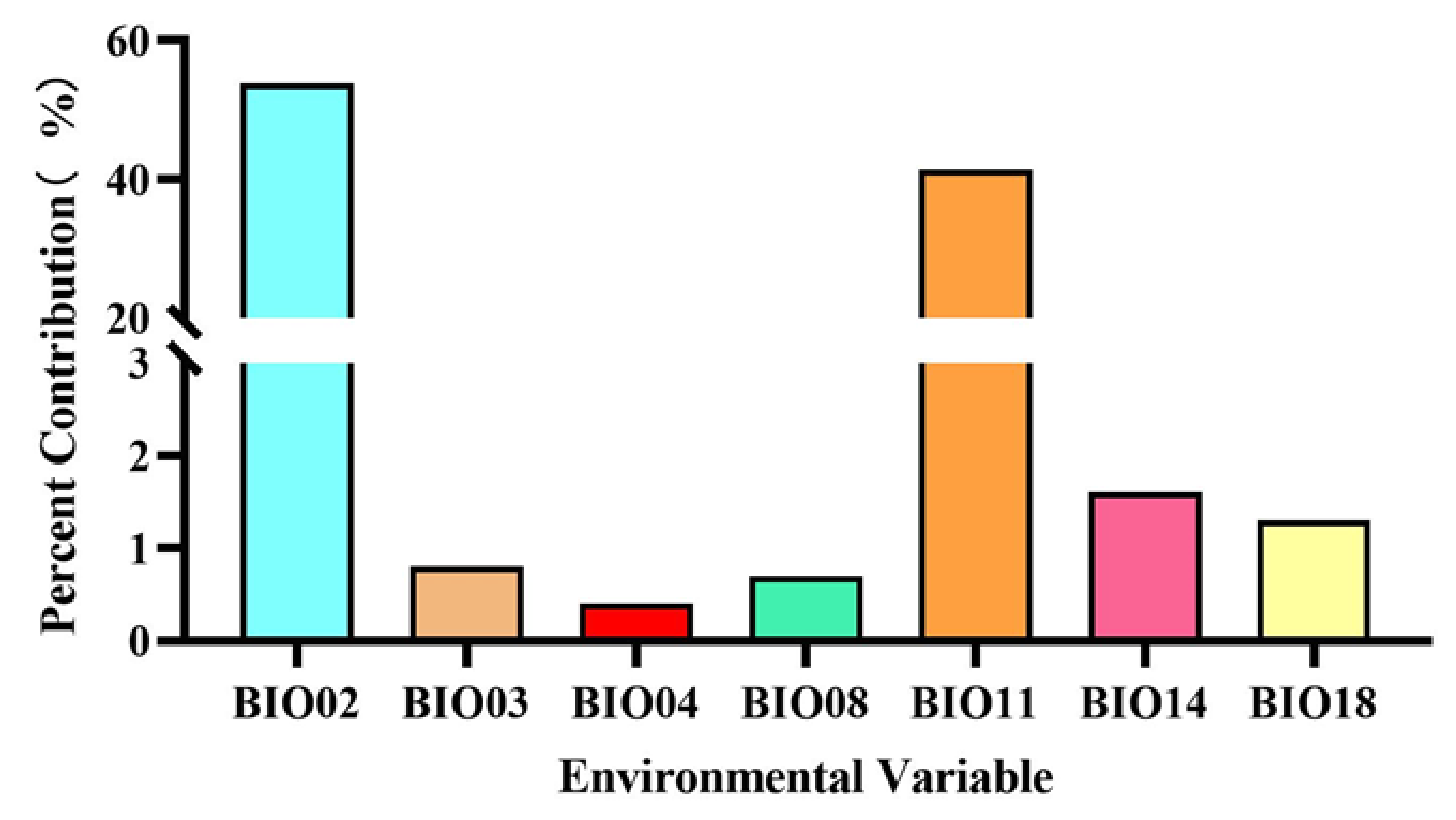
3.2. Environmental Variables Contribution Analysis
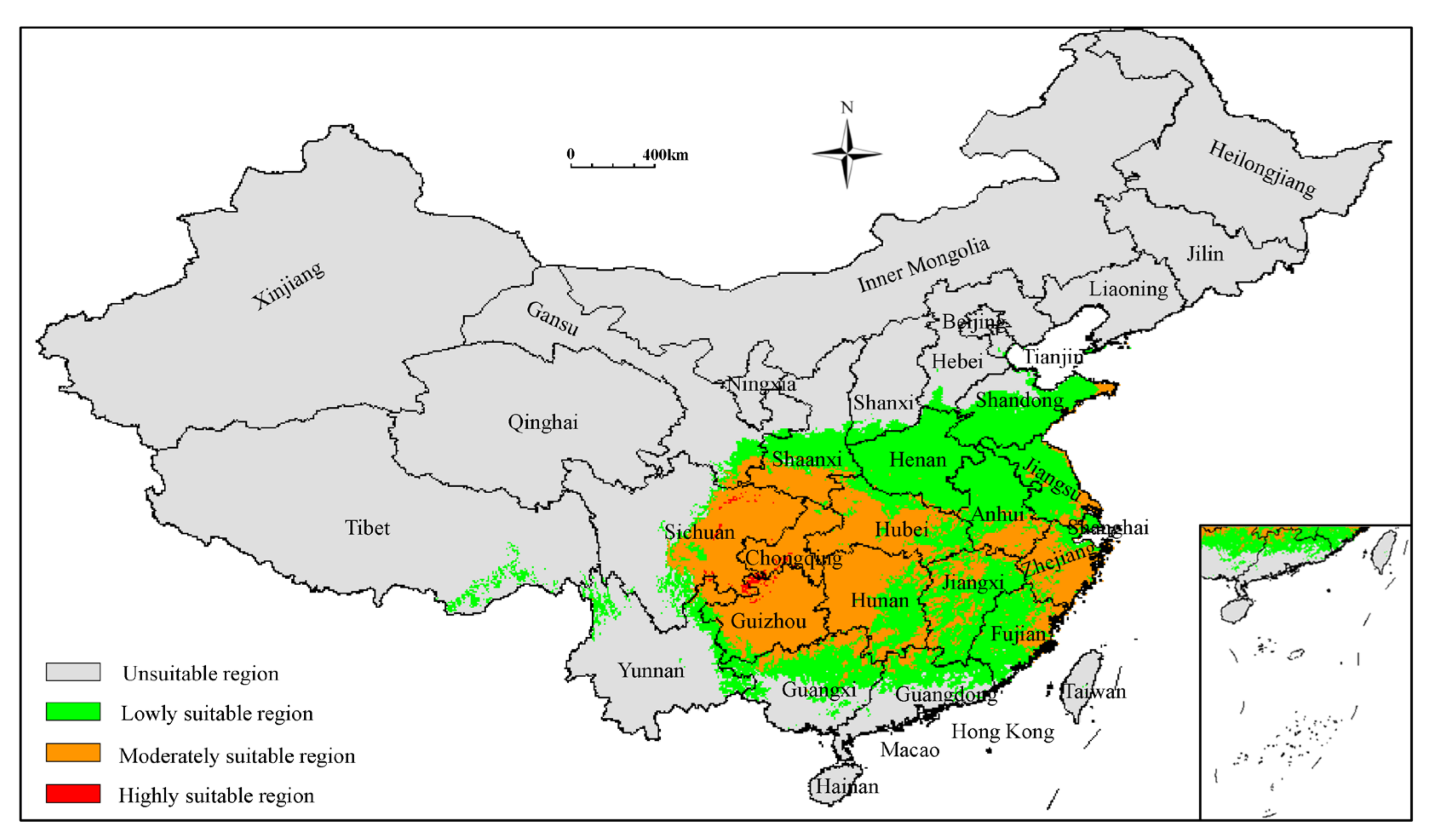
3.3. Analysis of Current Potentially Suitable Regions
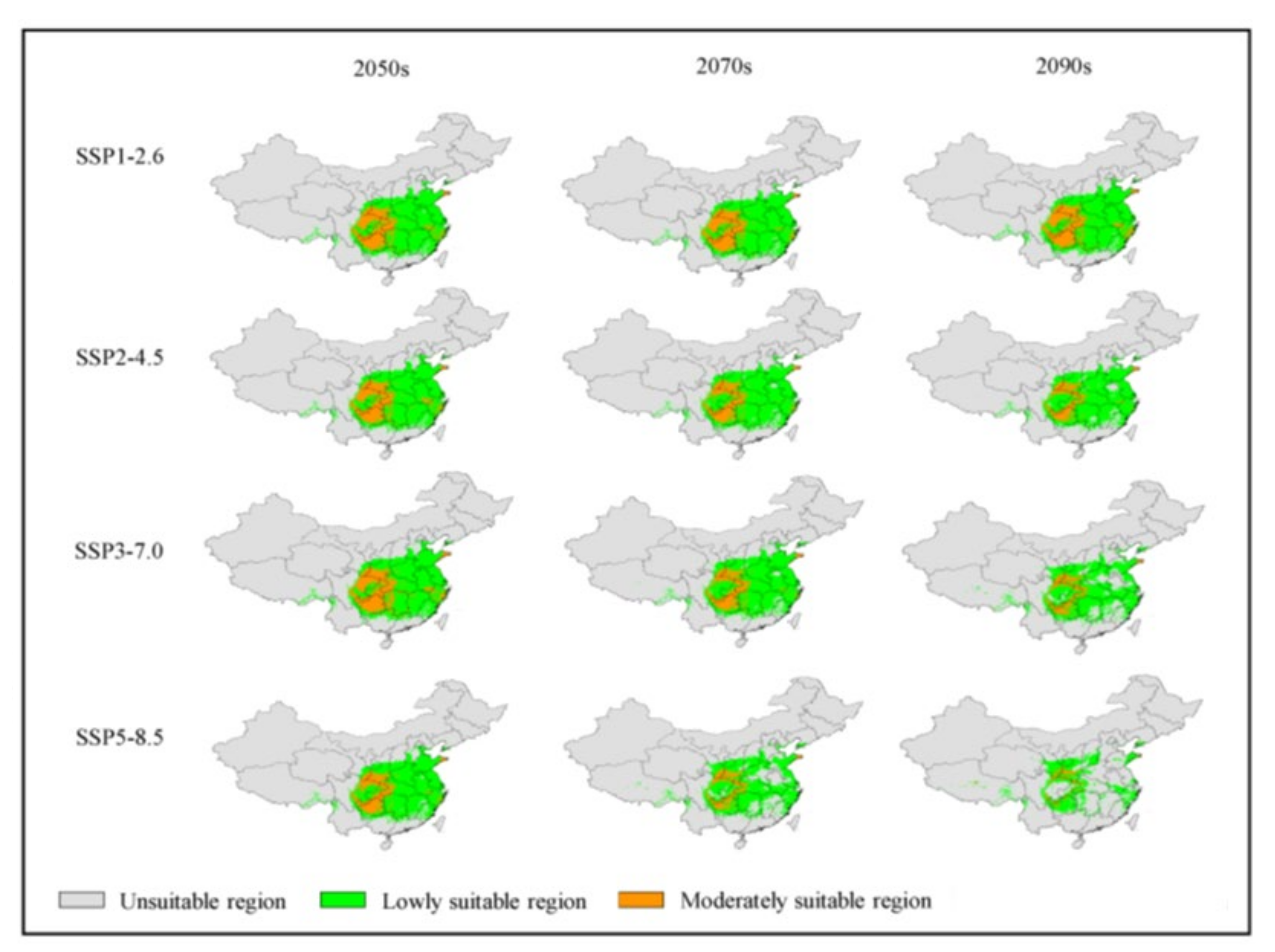
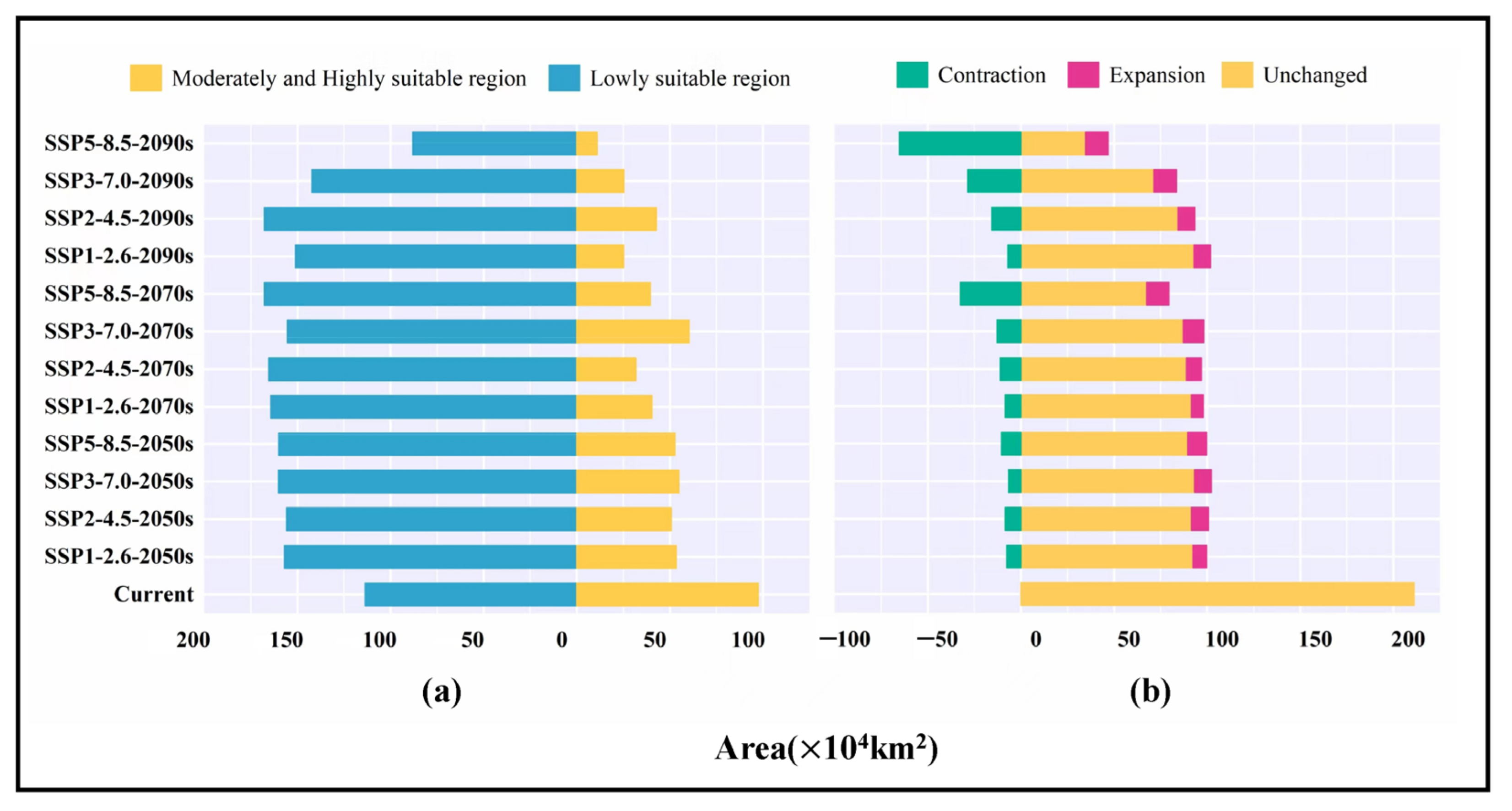
3.4. Analysis of Future Potentially Suitable Regions
3.5. Analysis of Low Impact Areas
3.6. Shift in Distribution Center of the Suitable Region
4. Discussion
4.1. Model Accuracy Analysis
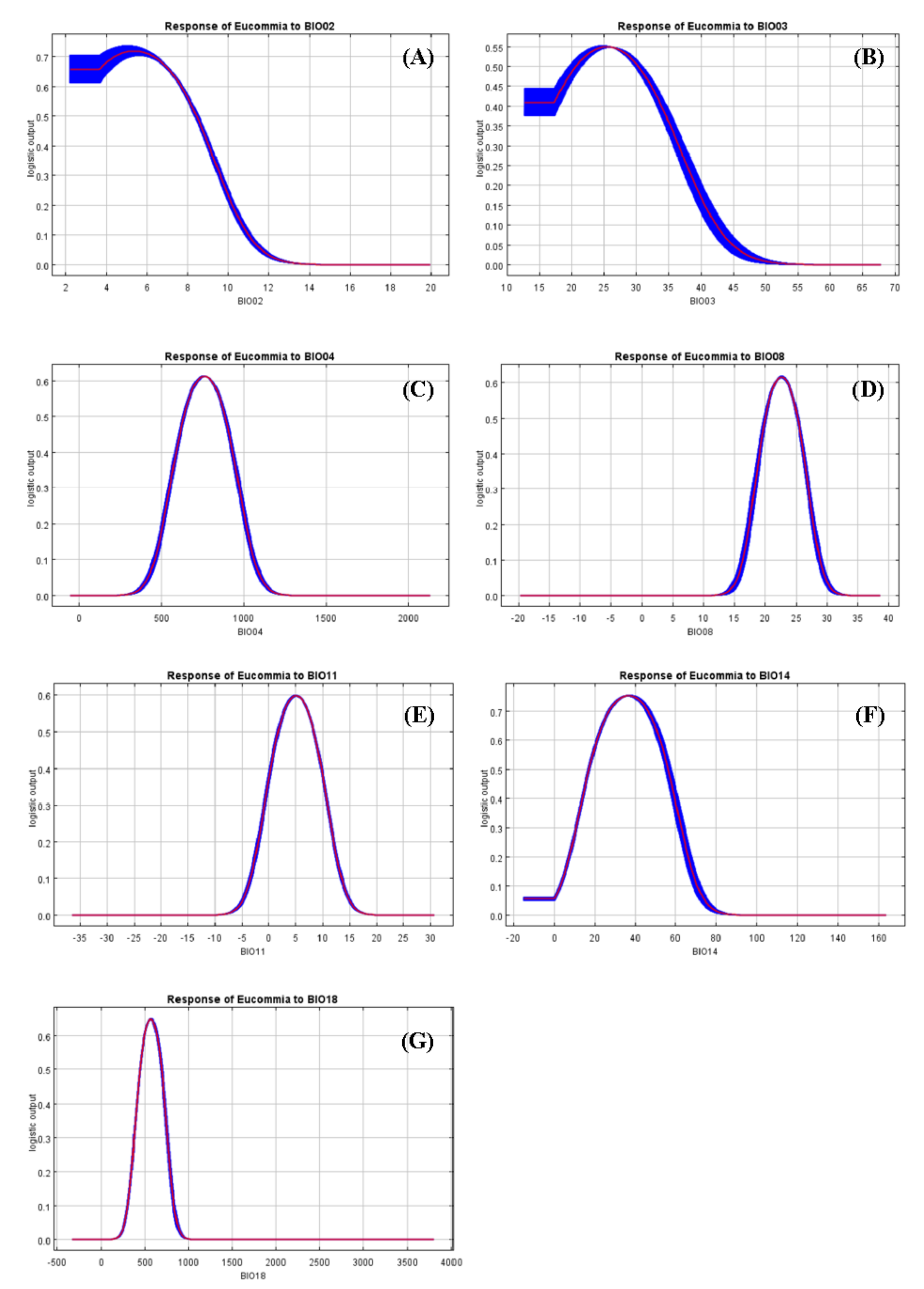
4.2. Impacts of Environmental Variables on Species Distribution
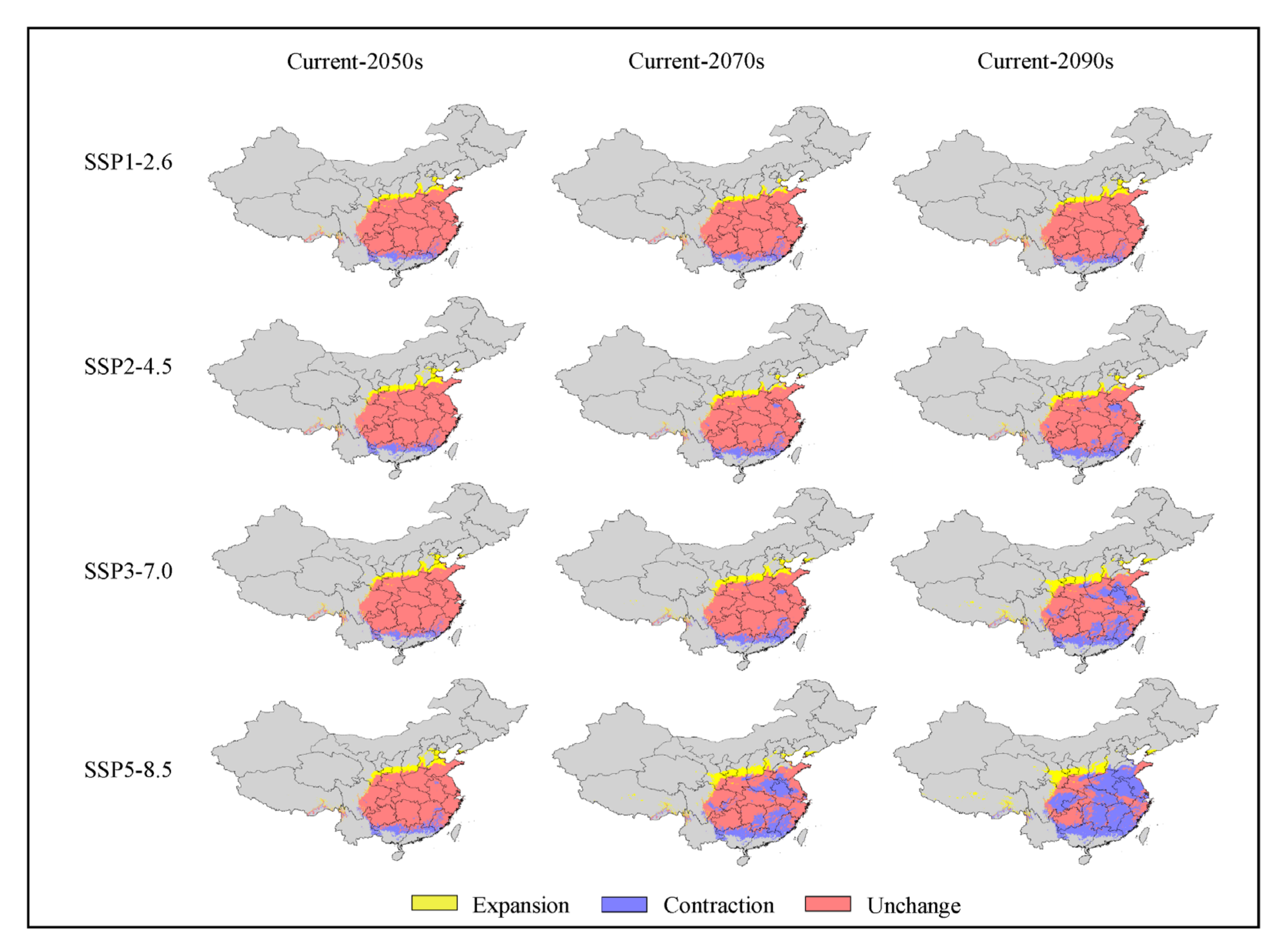
4.3. Spatial Dynamics of Potentially Suitable Areas
4.4. Suggestions on Resource Conservation and Development
4.5. Study Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Species | Lon 1 | Lat 1 | Species | Lon 1 | Lat 1 | Species | Lon 1 | Lat 1 |
|---|---|---|---|---|---|---|---|---|
| Eucommia ulmoides | 100.3541 | 30.8958 | Eucommia ulmoides | 105.3542 | 30.1042 | Eucommia ulmoides | 107.1042 | 30.1042 |
| Eucommia ulmoides | 102.0208 | 29.8958 | Eucommia ulmoides | 105.3542 | 32.2292 | Eucommia ulmoides | 107.1042 | 31.1458 |
| Eucommia ulmoides | 102.0625 | 29.6042 | Eucommia ulmoides | 105.3958 | 28.1042 | Eucommia ulmoides | 107.2292 | 24.2292 |
| Eucommia ulmoides | 102.0625 | 31.4792 | Eucommia ulmoides | 105.5625 | 26.2708 | Eucommia ulmoides | 107.2292 | 29.1458 |
| Eucommia ulmoides | 102.1458 | 29.6458 | Eucommia ulmoides | 105.5625 | 28.1875 | Eucommia ulmoides | 107.2708 | 25.2708 |
| Eucommia ulmoides | 102.5625 | 27.3958 | Eucommia ulmoides | 105.5625 | 28.3125 | Eucommia ulmoides | 107.3125 | 30.1458 |
| Eucommia ulmoides | 102.6875 | 28.9375 | Eucommia ulmoides | 105.5625 | 30.6042 | Eucommia ulmoides | 107.3125 | 30.2708 |
| Eucommia ulmoides | 102.7292 | 29.3542 | Eucommia ulmoides | 105.5625 | 32.1875 | Eucommia ulmoides | 107.3542 | 30.2708 |
| Eucommia ulmoides | 102.8542 | 30.2708 | Eucommia ulmoides | 105.6875 | 24.5625 | Eucommia ulmoides | 107.3542 | 28.9375 |
| Eucommia ulmoides | 103.1042 | 29.8125 | Eucommia ulmoides | 105.9792 | 29.6875 | Eucommia ulmoides | 107.3542 | 29.1042 |
| Eucommia ulmoides | 103.2708 | 27.6875 | Eucommia ulmoides | 106.0208 | 28.4375 | Eucommia ulmoides | 107.3542 | 30.2292 |
| Eucommia ulmoides | 103.3542 | 30.1458 | Eucommia ulmoides | 106.0625 | 26.0625 | Eucommia ulmoides | 107.3958 | 29.3542 |
| Eucommia ulmoides | 103.3958 | 29.6042 | Eucommia ulmoides | 106.0625 | 26.4375 | Eucommia ulmoides | 107.4792 | 29.3542 |
| Eucommia ulmoides | 103.6458 | 28.2708 | Eucommia ulmoides | 106.1042 | 25.5625 | Eucommia ulmoides | 107.6042 | 29.3125 |
| Eucommia ulmoides | 103.8125 | 30.6875 | Eucommia ulmoides | 106.1042 | 28.5625 | Eucommia ulmoides | 107.6042 | 29.4375 |
| Eucommia ulmoides | 104.0208 | 28.7708 | Eucommia ulmoides | 106.1042 | 33.5625 | Eucommia ulmoides | 107.6042 | 32.1042 |
| Eucommia ulmoides | 104.3125 | 25.5625 | Eucommia ulmoides | 106.1875 | 28.1458 | Eucommia ulmoides | 107.6875 | 26.0625 |
| Eucommia ulmoides | 104.3125 | 33.1042 | Eucommia ulmoides | 106.2292 | 29.1458 | Eucommia ulmoides | 107.8958 | 25.9792 |
| Eucommia ulmoides | 104.3958 | 26.3542 | Eucommia ulmoides | 106.2292 | 29.1458 | Eucommia ulmoides | 108.0208 | 26.3542 |
| Eucommia ulmoides | 104.4375 | 27.1458 | Eucommia ulmoides | 106.2292 | 32.2292 | Eucommia ulmoides | 108.1042 | 28.2292 |
| Eucommia ulmoides | 104.5208 | 32.4375 | Eucommia ulmoides | 106.2292 | 32.4375 | Eucommia ulmoides | 108.1458 | 25.5625 |
| Eucommia ulmoides | 104.6458 | 27.1042 | Eucommia ulmoides | 106.2708 | 29.1875 | Eucommia ulmoides | 108.1458 | 27.3125 |
| Eucommia ulmoides | 104.6875 | 29.5208 | Eucommia ulmoides | 106.3125 | 28.3542 | Eucommia ulmoides | 108.2708 | 24.8542 |
| Eucommia ulmoides | 104.7292 | 31.8542 | Eucommia ulmoides | 106.3542 | 28.4375 | Eucommia ulmoides | 108.3542 | 25.2292 |
| Eucommia ulmoides | 104.9792 | 28.4792 | Eucommia ulmoides | 106.3542 | 29.9375 | Eucommia ulmoides | 108.3542 | 27.6042 |
| Eucommia ulmoides | 104.9792 | 28.6042 | Eucommia ulmoides | 106.4375 | 26.4792 | Eucommia ulmoides | 108.3958 | 29.3542 |
| Eucommia ulmoides | 105.0208 | 25.1458 | Eucommia ulmoides | 106.4375 | 27.3125 | Eucommia ulmoides | 108.4375 | 28.0625 |
| Eucommia ulmoides | 105.1042 | 24.3542 | Eucommia ulmoides | 106.5208 | 26.4792 | Eucommia ulmoides | 108.4375 | 28.2292 |
| Eucommia ulmoides | 105.1042 | 24.5625 | Eucommia ulmoides | 106.5625 | 26.2708 | Eucommia ulmoides | 108.4792 | 28.3958 |
| Eucommia ulmoides | 105.2292 | 26.1875 | Eucommia ulmoides | 106.5625 | 29.3958 | Eucommia ulmoides | 108.5208 | 27.3542 |
| Eucommia ulmoides | 105.2292 | 27.2708 | Eucommia ulmoides | 106.5625 | 29.5208 | Eucommia ulmoides | 108.5208 | 29.2708 |
| Eucommia ulmoides | 105.2292 | 28.1042 | Eucommia ulmoides | 106.6875 | 25.9375 | Eucommia ulmoides | 108.5208 | 29.3542 |
| Eucommia ulmoides | 105.2292 | 28.3125 | Eucommia ulmoides | 106.6875 | 26.4792 | Eucommia ulmoides | 108.6458 | 27.0208 |
| Eucommia ulmoides | 105.2708 | 24.2292 | Eucommia ulmoides | 106.7292 | 26.6458 | Eucommia ulmoides | 108.6458 | 31.4792 |
| Eucommia ulmoides | 105.2708 | 24.3542 | Eucommia ulmoides | 106.9375 | 31.9792 | Eucommia ulmoides | 108.6875 | 27.9375 |
| Eucommia ulmoides | 105.2708 | 32.5625 | Eucommia ulmoides | 107.0208 | 25.3125 | Eucommia ulmoides | 108.8542 | 27.7292 |
| Eucommia ulmoides | 105.3125 | 25.2708 | Eucommia ulmoides | 107.0625 | 24.5208 | Eucommia ulmoides | 108.8542 | 31.5208 |
| Eucommia ulmoides | 105.3125 | 25.4792 | Eucommia ulmoides | 107.0625 | 29.6458 | Eucommia ulmoides | 108.9792 | 28.3542 |
| Eucommia ulmoides | 105.3125 | 26.5625 | Eucommia ulmoides | 107.0625 | 32.3125 | Eucommia ulmoides | 109.0625 | 26.2708 |
| Eucommia ulmoides | 105.3125 | 28.1458 | Eucommia ulmoides | 107.1042 | 29.0625 | Eucommia ulmoides | 109.1042 | 26.1875 |
| Eucommia ulmoides | 105.3542 | 25.5625 | Eucommia ulmoides | 107.1042 | 29.4792 | Eucommia ulmoides | 109.1042 | 26.3125 |
| Eucommia ulmoides | 109.1458 | 29.3542 | Eucommia ulmoides | 111.1875 | 30.2292 | Eucommia ulmoides | 114.6458 | 29.3125 |
| Eucommia ulmoides | 109.1875 | 32.1875 | Eucommia ulmoides | 111.1875 | 31.5625 | Eucommia ulmoides | 114.8125 | 31.6458 |
| Eucommia ulmoides | 109.2292 | 28.3958 | Eucommia ulmoides | 111.3125 | 27.4792 | Eucommia ulmoides | 114.8542 | 31.6042 |
| Eucommia ulmoides | 109.3958 | 28.6042 | Eucommia ulmoides | 111.3958 | 26.2708 | Eucommia ulmoides | 115.1458 | 29.1875 |
| Eucommia ulmoides | 109.4375 | 30.0208 | Eucommia ulmoides | 111.5208 | 24.4792 | Eucommia ulmoides | 115.1458 | 29.3125 |
| Eucommia ulmoides | 109.4375 | 30.6458 | Eucommia ulmoides | 111.5208 | 28.1458 | Eucommia ulmoides | 115.4375 | 29.6875 |
| Eucommia ulmoides | 109.4792 | 24.3125 | Eucommia ulmoides | 111.6875 | 25.3125 | Eucommia ulmoides | 115.9375 | 29.6042 |
| Eucommia ulmoides | 109.4792 | 32.3542 | Eucommia ulmoides | 111.8542 | 24.8958 | Eucommia ulmoides | 115.9792 | 40.4792 |
| Eucommia ulmoides | 109.5208 | 25.3542 | Eucommia ulmoides | 111.8958 | 28.1458 | Eucommia ulmoides | 116.0625 | 29.5625 |
| Eucommia ulmoides | 109.5625 | 28.3542 | Eucommia ulmoides | 111.9792 | 26.7292 | Eucommia ulmoides | 116.0625 | 39.9792 |
| Eucommia ulmoides | 109.6042 | 29.4792 | Eucommia ulmoides | 112.3958 | 33.7708 | Eucommia ulmoides | 116.2292 | 29.2708 |
| Eucommia ulmoides | 109.6042 | 31.3958 | Eucommia ulmoides | 112.4792 | 37.7292 | Eucommia ulmoides | 116.2292 | 39.9792 |
| Eucommia ulmoides | 109.6042 | 34.3542 | Eucommia ulmoides | 112.6042 | 37.8125 | Eucommia ulmoides | 116.5208 | 27.0208 |
| Eucommia ulmoides | 109.7292 | 31.8542 | Eucommia ulmoides | 112.7292 | 27.2292 | Eucommia ulmoides | 116.6042 | 35.4375 |
| Eucommia ulmoides | 109.7292 | 34.3542 | Eucommia ulmoides | 113.0625 | 25.8125 | Eucommia ulmoides | 116.8542 | 27.0625 |
| Eucommia ulmoides | 109.8542 | 25.7292 | Eucommia ulmoides | 113.0625 | 28.1875 | Eucommia ulmoides | 116.9792 | 35.6042 |
| Eucommia ulmoides | 110.0208 | 28.2292 | Eucommia ulmoides | 113.3125 | 35.3125 | Eucommia ulmoides | 117.0208 | 36.2708 |
| Eucommia ulmoides | 110.0625 | 28.3958 | Eucommia ulmoides | 113.5208 | 35.2708 | Eucommia ulmoides | 117.0208 | 36.6458 |
| Eucommia ulmoides | 110.0625 | 28.6458 | Eucommia ulmoides | 113.5625 | 35.1875 | Eucommia ulmoides | 117.1458 | 36.1875 |
| Eucommia ulmoides | 110.1042 | 26.3958 | Eucommia ulmoides | 113.5625 | 35.3542 | Eucommia ulmoides | 117.2708 | 36.1042 |
| Eucommia ulmoides | 110.1042 | 28.3958 | Eucommia ulmoides | 113.6042 | 35.7292 | Eucommia ulmoides | 117.3125 | 29.0208 |
| Eucommia ulmoides | 110.1458 | 28.4375 | Eucommia ulmoides | 113.8958 | 28.6042 | Eucommia ulmoides | 117.9792 | 35.5625 |
| Eucommia ulmoides | 110.1458 | 29.1042 | Eucommia ulmoides | 113.9792 | 35.3542 | Eucommia ulmoides | 118.0625 | 36.1875 |
| Eucommia ulmoides | 110.1875 | 24.1458 | Eucommia ulmoides | 114.0208 | 26.1042 | Eucommia ulmoides | 118.1042 | 28.5208 |
| Eucommia ulmoides | 110.1875 | 28.8125 | Eucommia ulmoides | 114.0208 | 26.5208 | Eucommia ulmoides | 118.1458 | 29.1875 |
| Eucommia ulmoides | 110.2292 | 26.0208 | Eucommia ulmoides | 114.0208 | 27.3125 | Eucommia ulmoides | 118.1458 | 29.1458 |
| Eucommia ulmoides | 110.2292 | 28.4792 | Eucommia ulmoides | 114.0208 | 27.3958 | Eucommia ulmoides | 118.9792 | 35.9792 |
| Eucommia ulmoides | 110.2292 | 31.2708 | Eucommia ulmoides | 114.0208 | 28.4792 | Eucommia ulmoides | 119.1875 | 36.5625 |
| Eucommia ulmoides | 110.3542 | 24.5625 | Eucommia ulmoides | 114.1042 | 27.1458 | Eucommia ulmoides | 119.4375 | 29.1875 |
| Eucommia ulmoides | 110.3958 | 31.4375 | Eucommia ulmoides | 114.1042 | 31.8125 | Eucommia ulmoides | 119.4375 | 30.3125 |
| Eucommia ulmoides | 110.4375 | 34.7708 | Eucommia ulmoides | 114.1458 | 27.2292 | Eucommia ulmoides | 119.4375 | 32.3958 |
| Eucommia ulmoides | 110.4375 | 34.8125 | Eucommia ulmoides | 114.1458 | 27.3125 | Eucommia ulmoides | 119.9375 | 27.8542 |
| Eucommia ulmoides | 110.4792 | 25.7708 | Eucommia ulmoides | 114.2708 | 25.6042 | Eucommia ulmoides | 120.1042 | 36.9792 |
| Eucommia ulmoides | 110.5208 | 24.5625 | Eucommia ulmoides | 114.2708 | 26.6042 | Eucommia ulmoides | 120.3958 | 36.3125 |
| Eucommia ulmoides | 110.6875 | 26.0208 | Eucommia ulmoides | 114.3125 | 27.6875 | Eucommia ulmoides | 120.5208 | 28.9792 |
| Eucommia ulmoides | 110.8542 | 34.1875 | Eucommia ulmoides | 114.3958 | 28.6458 | Eucommia ulmoides | 120.7708 | 37.5208 |
| Eucommia ulmoides | 110.9375 | 31.0625 | Eucommia ulmoides | 114.4375 | 29.0625 | Eucommia ulmoides | 120.8958 | 31.3125 |
| Eucommia ulmoides | 110.9375 | 33.3125 | Eucommia ulmoides | 114.4792 | 29.0208 | Eucommia ulmoides | 121.2708 | 37.2292 |
| Eucommia ulmoides | 111.0208 | 26.0625 | Eucommia ulmoides | 114.5208 | 28.0208 | Eucommia ulmoides | 121.3542 | 37.5208 |
| Eucommia ulmoides | 111.0625 | 26.7708 | Eucommia ulmoides | 114.5625 | 25.8125 | Eucommia ulmoides | 121.3958 | 31.2292 |
| Eucommia ulmoides | 111.1875 | 28.1042 | Eucommia ulmoides | 114.6458 | 28.9792 | Eucommia ulmoides | 122.1875 | 37.3542 |
| Eucommia ulmoides | 122.2708 | 29.8125 |
References
- Wilczek, A.M.; Cooper, M.D.; Korves, T.M.; Schmitt, J. Lagging adaptation to warming climate in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2014, 111, 7906–7913. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guan, L.; Yang, Y.; Jiang, P.; Mou, Q.; Gou, Y.; Zhu, X.; Xu, Y.W.; Wang, R. Potential distribution of Blumea balsamifera in China using MaxEnt and the ex situ conservation based on its effective components and fresh leaf yield. Environ. Sci. Pollut. Res. 2022, 29, 44003–44019. [Google Scholar] [CrossRef]
- Xu, Y.; Ramanathan, V.; Victor, D.G. Global warming will happen faster than we think. Nature 2018, 564, 30–32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, R.; Ji, M.; Zhang, S. Global warming and obesity: A systematic review. Obes. Rev. 2018, 19, 150–163. [Google Scholar] [CrossRef]
- Lippmann, R.; Babben, S.; Menger, A.; Delker, C.; Quint, M. Development of Wild and Cultivated Plants under Global Warming Conditions. Curr. Biol. 2019, 29, R1326–R1338. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.J.; Hsu, T.W.; Hung, K.H.; Lin, C.J.; Huang, C.C.; Huang, C.C.; Chiang, Y.C.; Chiang, T.Y. Inferring multiple refugia and phylogeographical patterns in Pinus massoniana based on nucleotide sequence variation and DNA fingerprinting. PLoS ONE 2012, 7, e43717. [Google Scholar] [CrossRef] [Green Version]
- Alberto, F.J.; Aitken, S.N.; Alía, R.; González-Martínez, S.C.; Hänninen, H.; Kremer, A.; Lefèvre, F.; Lenormand, T.; Yeaman, S.; Whetten, R.; et al. Potential for evolutionary responses to climate change–evidence from tree populations. Glob. Chang. Biol. 2013, 19, 1645–1661. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shaw, R.G.; Etterson, J.R. Rapid climate change and the rate of adaptation: Insight from experimental quantitative genetics. New Phytol. 2012, 195, 752–765. [Google Scholar] [CrossRef]
- Aguirre-Liguori, J.A.; Ramírez-Barahona, S.; Gaut, B.S. The evolutionary genomics of species’ responses to climate change. Nat. Ecol. Evol. 2021, 5, 1350–1360. [Google Scholar] [CrossRef]
- McGaughran, A.; Laver, R.; Fraser, C. Evolutionary Responses to Warming. Trends Ecol. Evol. 2021, 36, 591–600. [Google Scholar] [CrossRef]
- Reich, P.B.; Oleksyn, J. Climate warming will reduce growth and survival of Scots pine except in the far north. Ecol. Lett. 2008, 11, 588–597. [Google Scholar] [CrossRef] [PubMed]
- Spencer, D.; Yue, Y.R.; Bolin, D.; Ryan, S.; Mejia, A.F. Spatial Bayesian GLM on the cortical surface produces reliable task activations in individuals and groups. Neuroimage 2022, 249, 118908. [Google Scholar] [CrossRef]
- Thameur, B.H.; Soufiène, S.; Haj Ammar, H.; Hammami, S. Spatial distribution and habitat selection of culicoides imicola: The potential vector of bluetongue virus in Tunisia. Onderstepoort. J. Vet. Res. 2021, 88, e1–e9. [Google Scholar] [CrossRef]
- Zhao, Y.; Adjallah, K.H.; Sava, A.; Wang, Z. MaxEnt feature-based reliability model method for real-time detection of early chatter in high-speed milling. ISA Trans. 2021, 113, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Banavar, J.R.; Maritan, A.; Volkov, I. Applications of the principle of maximum entropy: From physics to ecology. J. Phys. Condens. Matter. 2010, 22, 063101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harte, J.; Newman, E.A. Maximum information entropy: A foundation for ecological theory. Trends Ecol. Evol. 2014, 29, 384–389. [Google Scholar] [CrossRef]
- Guo, Y.; Wei, H.; Lu, C.; Gao, B.; Gu, W. Predictions of potential geographical distribution and quality of Schisandra sphenanthera under climate change. PeerJ 2016, 4, e2554. [Google Scholar] [CrossRef] [Green Version]
- Wouyou, H.G.; Lokonon, B.E.; Idohou, R.; Zossou-Akete, A.G.; Assogbadjo, A.E.; Glèlè Kakaï, R. Predicting the potential impacts of climate change on the endangered Caesalpinia bonduc (L.) Roxb in Benin (West Africa). Heliyon 2022, 8, e09022. [Google Scholar] [CrossRef] [PubMed]
- Xian, X.; Zhao, H.; Wang, R.; Qiao, H.; Guo, J.; Zhang, G.; Liu, W.; Wan, F. Ecological Niche Shifts Affect the Potential Invasive Risk of Rapistrum rugosum (L.) All. in China. Front. Plant Sci. 2022, 13, 827497. [Google Scholar] [CrossRef]
- Zhang, L.; Cao, B.; Bai, C.; Li, G.; Mao, M. Predicting suitable cultivation regions of medicinal plants with Maxent modeling and fuzzy logics: A case study of Scutellaria baicalensis in China. Environ. Earth Sci. 2016, 75, 361. [Google Scholar] [CrossRef]
- He, X.; Wang, J.; Li, M.; Hao, D.; Yang, Y.; Zhang, C.; He, R.; Tao, R. Eucommia ulmoides Oliv.: Ethnopharmacology, phytochemistry and pharmacology of an important traditional Chinese medicine. J. Ethnopharmacol. 2014, 151, 78–92. [Google Scholar] [CrossRef]
- Ye, J.; Han, W.; Fan, R.; Liu, M.; Li, L.; Jia, X. Integration of Transcriptomes, Small RNAs, and Degradome Sequencing to Identify Putative miRNAs and Their Targets Related to Eu-Rubber Biosynthesis in Eucommia ulmoides. Genes 2019, 10, 623. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Tan, D.-C.; Peng, B.; Yang, L.; Zhang, S.-Y.; Shi, R.-P.; Chong, C.-M.; Zhong, Z.-F.; Wang, S.-P.; Liang, Q.-L.; et al. Neuroendocrine–Immune Regulatory Network of Eucommia ulmoides Oliver. Molecules 2022, 27, 3697. [Google Scholar] [CrossRef]
- Dong, C.; Yao, T.; Zhang, Z.; Chen, W.; Liang, J.; Han, Y.; Huang, J.; Deshmukh, S.K.; Liang, Z. Structure and Function of Bacterial Microbiota in Eucommia ulmoides Bark. Curr. Microbiol. 2020, 77, 3623–3632. [Google Scholar] [CrossRef] [PubMed]
- Do, M.H.; Hur, J.; Choi, J.; Kim, M.; Kim, M.J.; Kim, Y.; Ha, S.K. Eucommia ulmoides Ameliorates Glucotoxicity by Suppressing Advanced Glycation End-Products in Diabetic Mice Kidney. Nutrients 2018, 10, 265. [Google Scholar] [CrossRef] [Green Version]
- Su, L.; Wang, Q.; Xiang, P.; Yin, D.; Ding, X.; Liu, L.; Zhao, X. Development of nitrile rubber/eucommia ulmoides gum composites for controllable dynamic damping and sound absorption performance. RSC Adv. 2022, 12, 21503–21511. [Google Scholar] [CrossRef]
- Suzuki, N.; Uefuji, H.; Nishikawa, T.; Mukai, Y.; Yamashita, A.; Hattori, M.; Ogasawara, N.; Bamba, T.; Fukusaki, E.; Kobayashi, A.; et al. Construction and analysis of EST libraries of the trans-polyisoprene producing plant, Eucommia ulmoides Oliver. Planta 2012, 236, 1405–1417. [Google Scholar] [CrossRef] [PubMed]
- Mwange, K.N.; Hou, H.W.; Cui, K.M. Relationship between endogenous indole-3-acetic acid and abscisic acid changes and bark recovery in Eucommia ulmoides Oliv. after girdling. J. Exp. Bot. 2003, 54, 1899–1907. [Google Scholar] [CrossRef] [Green Version]
- Yu, J.; Wang, Y.; Ru, M.; Peng, L.; Liang, Z. Genetic diversity in intraspecific hybrid populations of Eucommia ulmoides Oliver evaluated from ISSR and SRAP molecular marker analysis. Genet Mol Res 2015, 14, 7417–7425. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.G. China Plant Red Data Book-Rare and Endangered Plants; Science Press: Beijing, China, 1991. (In Chinese) [Google Scholar]
- Jin, C.; Li, Z.; Li, Y.; Wang, S.; Li, L.; Liu, M.; Ye, J. Transcriptome analysis of terpenoid biosynthetic genes and simple sequence repeat marker screening in Eucommia ulmoides. Mol. Biol. Rep. 2020, 47, 1979–1990. [Google Scholar] [CrossRef] [PubMed]
- Li, A.; Yuan, J.F.; Gong, Q.; Zhang, N.; Chen, L.Y.; Luo, Y.Y.; Cui, Y.R.; Wang, H.L.; Liu, R.H. Effects of Eucommia ulmoides extract against renal injury caused by long-term high purine diets in rats. Food Funct. 2021, 12, 5607–5620. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Guo, F.F.; Xiao, J.P.; Wei, J.Y.; Tang, L.Y.; Yang, H.J. Research advances in chemical constituents and pharmacological activities of different parts of Eucommia ulmoides. Chin. J. Chin. Mater. Med. 2020, 45, 497–512. (In Chinese) [Google Scholar] [CrossRef]
- Oikawa, H.; Miyazaki, S.; Kurata, R.; Hattori, M.; Hayashi, N.; Kawaguchi, N.; Hirata, T.; Ueda, T.; Fujikawa, T. Eucommia Leaf Extract Induces BDNF Production in Rat Hypothalamus and Enhances Lipid Metabolism and Aerobic Glycolysis in Rat Liver. Curr. Mol. Pharmacol. 2021, 14, 234–244. [Google Scholar] [CrossRef] [PubMed]
- Qing, J.; Du, Q.; Meng, Y.; Liu, P.; Du, H.; Wang, L. Genome-wide identification and expression pattern analysis of the ribonuclease T2 family in Eucommia ulmoides. Sci. Rep. 2021, 11, 6900. [Google Scholar] [CrossRef]
- Wu, G.H.; Chen, Y.; Zheng, L.H.; Huang, T.; Hao, X.J.; Zhang, J.X. Chemical Constituents of Eucommia ulmoides in Guizhou Province. Chin. Med. Mat. 2015, 38, 980–984. (In Chinese) [Google Scholar]
- Yan, Y.; Zhao, H.; Zou, L.S.; Liu, X.H.; Chai, C.; Wang, S.N.; Hua, Y.J. Difference of chemical constituents in Eucommiae Cortex from different habitats by LC-QTOF MS/MS. Chin. J. Chin. Mater. Med. 2017, 42, 2730–2737. (In Chinese) [Google Scholar] [CrossRef]
- Zhao, X.; Wang, Y.; Nie, Z.; Han, L.; Zhong, X.; Yan, X.; Gao, X. Eucommia ulmoides leaf extract alters gut microbiota composition, enhances short-chain fatty acids production, and ameliorates osteoporosis in the senescence-accelerated mouse P6 (SAMP6) model. Food Sci. Nutr. 2020, 8, 4897–4906. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.F.; Li, C.S.; Collinson, M.E.; Lin, J.; Sun, Q.G. Eucommia (Eucommiaceae), a potential biothermometer for the reconstruction of paleoenvironments. Am. J. Bot. 2003, 90, 1–7. [Google Scholar] [CrossRef]
- Xu, D.; Zhuo, Z.; Wang, R.; Ye, M.; Pu, B. Modeling the distribution of Zanthoxylum armatum in China with MaxEnt modeling. Glob. Ecol. Conserv. 2019, 19, e00691. [Google Scholar] [CrossRef]
- Zhang, L.; Cao, X.; Yao, Z.; Dong, X.; Chen, M.; Xiao, L.; Zhao, S. Identification of risk areas for Orobanche cumana and Phelipanche aegyptiaca in China, based on the major host plant and CMIP6 climate scenarios. Ecol. Evol. 2022, 12, e8824. [Google Scholar] [CrossRef]
- Zhao, Q.; Zhang, Y.; Li, W.-N.; Hu, B.-W.; Zou, J.-B.; Wang, S.-Q.; Niu, J.-F.; Wang, Z.-Z. Predicting the Potential Distribution of Perennial Plant Coptis chinensis Franch. In China under Multiple Climate Change Scenarios. Forests 2021, 12, 1464. [Google Scholar] [CrossRef]
- Nolzen, H.; Brugger, K.; Reichold, A.; Brock, J.; Lange, M.; Thulke, H.H. Model-based extrapolation of ecological systems under future climate scenarios: The example of Ixodes ricinus ticks. PLoS ONE 2022, 17, e0267196. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Sharma, L.K.; Rajkumar, M.; Mohammad, N.; Khan, M.L. Predicting habitat suitability of Litsea glutinosa: A declining tree species, under the current and future climate change scenarios in India. Landsc. Ecol. Eng. 2023, 1–15. [Google Scholar] [CrossRef]
- Zhang, J.; Jiang, F.; Li, G.; Qin, W.; Li, S.; Gao, H.; Cai, Z.; Lin, G.; Zhang, T. Maxent modeling for predicting the spatial distribution of three raptors in the Sanjiangyuan National Park, China. Ecol. Evol. 2019, 9, 6643–6654. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Torii, M.; Masuya, H.; Hattori, T. Temperature Characteristics of Two Fomitiporia Fungi Determine Their Geographical Distributions in Japan. Forests 2021, 12, 1580. [Google Scholar] [CrossRef]
- Guo, Q.; Li, H.; Zheng, W.; Pan, J.; Lu, J.; Li, J.; Zheng, Y. Analysis of genetic diversity and prediction of Larix species distribution in the Qinghai–Tibet Plateau, China. J. For. Res. 2022, 1–11. [Google Scholar] [CrossRef]
- Phillips, S.J.; Anderson, R.P.; Dudík, M.; Schapire, R.E.; Blair, M.E. Opening the black box: An open-source release of Maxent. Ecography 2017, 40, 887–893. [Google Scholar] [CrossRef]
- Anand, V.; Oinam, B.; Singh, I.H. Predicting the current and future potential spatial distribution of endangered Rucervus eldii eldii (Sangai) using MaxEnt model. Environ. Monit. Assess. 2021, 193, 147. [Google Scholar] [CrossRef] [PubMed]
- Cobos, M.E.; Peterson, A.T.; Barve, N.; Osorio-Olvera, L. kuenm: An R package for detailed development of ecological niche models using Maxent. PeerJ 2019, 7, e6281. [Google Scholar] [CrossRef] [Green Version]
- Akpoti, K.; Kabo-bah, A.T.; Dossou-Yovo, E.R.; Groen, T.A.; Zwart, S.J. Mapping suitability for rice production in inland valley landscapes in Benin and Togo using environmental niche modeling. Sci. Total Environ. 2020, 709, 136165. [Google Scholar] [CrossRef] [PubMed]
- Fang, B.; Zhao, Q.; Qin, Q.; Yu, J. Prediction of Potentially Suitable Distribution Areas for Prunus tomentosa in China Based on an Optimized MaxEnt Model. Forests 2022, 13, 381. [Google Scholar] [CrossRef]
- Li, Y.; Shao, W.; Jiang, J. Predicting the potential global distribution of Sapindus mukorossi under climate change based on MaxEnt modelling. Environ. Sci. Pollut. Res. 2022, 29, 21751–21768. [Google Scholar] [CrossRef] [PubMed]
- Kato, R.; Noguchi, H.; Honda, H.; Kobayashi, T. Hidden Markov model-based approach as the first screening of binding peptides that interact with MHC class II molecules. Enzyme Microb. Technol. 2003, 33, 472–481. [Google Scholar] [CrossRef]
- Lin, L.; Jin, L.; Wang, Z.H.; Cui, Z.J.; Ma, Y. Prediction of the potential distribution of Tibetan medicinal Lycium ruthenicum in context of climate change. Chin. J. Chin. Mater. Med. 2017, 42, 2659–2669. (In Chinese) [Google Scholar] [CrossRef]
- Jiménez-Valverde, A.; Lobo, J.M. Threshold criteria for conversion of probability of species presence to either–or presence–absence. Acta Oecol. 2007, 31, 361–369. [Google Scholar] [CrossRef]
- Pan, J.; Fan, X.; Luo, S.; Zhang, Y.; Yao, S.; Guo, Q.; Qian, Z. Predicting the Potential Distribution of Two Varieties of Litsea coreana (Leopard-Skin Camphor) in China under Climate Change. Forests 2020, 11, 1159. [Google Scholar] [CrossRef]
- Li, W.-N.; Zhao, Q.; Guo, M.-H.; Lu, C.; Huang, F.; Wang, Z.-Z.; Niu, J.-F. Predicting the Potential Distribution of the Endangered Plant Cremastra appendiculata (Orchidaceae) in China under Multiple Climate Change Scenarios. Forests 2022, 13, 1504. [Google Scholar] [CrossRef]
- Santos-Hernández, A.F.; Monterroso-Rivas, A.I.; Granados-Sánchez, D.; Villanueva-Morales, A.; Santacruz-Carrillo, M. Projections for Mexico’s Tropical Rainforests Considering Ecological Niche and Climate Change. Forests 2021, 12, 119. [Google Scholar] [CrossRef]
- Brown, J.L.; Bennett, J.R.; French, C.M. SDMtoolbox 2.0: The next generation Python-based GIS toolkit for landscape genetic, biogeographic and species distribution model analyses. PeerJ 2017, 5, e4095. [Google Scholar] [CrossRef] [Green Version]
- Garcia, K.; Lasco, R.; Ines, A.; Lyon, B.; Pulhin, F. Predicting geographic distribution and habitat suitability due to climate change of selected threatened forest tree species in the Philippines. Appl. Geogr. 2013, 44, 12–22. [Google Scholar] [CrossRef]
- Molloy, S.W.; Davis, R.A.; Van Etten, E.J.B. Species distribution modelling using bioclimatic variables to determine the impacts of a changing climate on the western ringtail possum (Pseudocheirus occidentals; Pseudocheiridae). Environ. Conserv. 2014, 41, 176–186. [Google Scholar] [CrossRef]
- Shabani, F.; Kumar, L.; Ahmadi, M. A comparison of absolute performance of different correlative and mechanistic species distribution models in an independent area. Ecol. Evol. 2016, 6, 5973–5986. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kong, W.Y.; Li, X.H.; Zou, H.F. Optimizing MaxEnt model in the prediction of species distribution. J. Appl. Ecol. 2019, 30, 2116–2128. (In Chinese) [Google Scholar] [CrossRef]
- Çoban, H.O.; Örücü, Ö.K.; Arslan, E.S. MaxEnt Modeling for Predicting the Current and Future Potential Geographical Distribution of Quercus libani Olivier. Sustainability 2020, 12, 2671. [Google Scholar] [CrossRef] [Green Version]
- Qin, A.; Liu, B.; Guo, Q.; Bussmann, R.W.; Ma, F.; Jian, Z.; Xu, G.; Pei, S. Maxent modeling for predicting impacts of climate change on the potential distribution of Thuja sutchuenensis Franch., an extremely endangered conifer from southwestern China. Glob. Ecol. Conserv. 2017, 10, 139–146. [Google Scholar] [CrossRef]
- Zhang, K.; Yao, L.; Meng, J.; Tao, J. Maxent modeling for predicting the potential geographical distribution of two peony species under climate change. Sci. Total Environ. 2018, 634, 1326–1334. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Tian, E.; Jim, C.Y.; Liu, D.; Hu, Z. Effects of climate-change scenarios on the distribution patterns of Castanea henryi. Ecol. Evol. 2022, 12, e9597. [Google Scholar] [CrossRef]
- Ding, Y.; Yang, S. Surviving and thriving: How plants perceive and respond to temperature stress. Dev. Cell 2022, 57, 947–958. [Google Scholar] [CrossRef]
- Li, Y.; Kong, D.; Fu, Y.; Sussman, M.R.; Wu, H. The effect of developmental and environmental factors on secondary metabolites in medicinal plants. Plant Physiol. Biochem. 2020, 148, 80–89. [Google Scholar] [CrossRef] [PubMed]
- Zhu, M.Q.; Sun, R.C. Eucommia ulmoides Oliver: A Potential Feedstock for Bioactive Products. J. Agric. Food Chem. 2018, 66, 5433–5438. [Google Scholar] [CrossRef]
- Ramakrishna, A.; Ravishankar, G.A. Influence of abiotic stress signals on secondary metabolites in plants. Plant Signal. Behav. 2011, 6, 1720–1731. [Google Scholar] [CrossRef] [PubMed]
- Qian, C.; Zhang, R.; Li, J.; Huang, Z.; Liu, X.; Yu, L.; Yan, L.; Fu, Y. The characteristics of habitat, functional traits and medicinal components of Eucommia ulmoides from Guizhou. Environ. Sci. Pollut. Res. 2022, 29, 12629–12647. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.; Yao, J.; He, Q.; Chen, J. Changes in precipitation amounts and extremes across Xinjiang (northwest China) and their connection to climate indices. PeerJ 2021, 9, e10792. [Google Scholar] [CrossRef] [PubMed]
- Xing, Y.-F.; He, D.; Wang, Y.; Zeng, W.; Zhang, C.; Lu, Y.; Su, N.; Kong, Y.-H.; Xing, X.-H. Chemical constituents, biological functions and pharmacological effects for comprehensive utilization of Eucommia ulmoides Oliver. Food Sci. Hum. Wellness 2019, 8, 177–188. [Google Scholar] [CrossRef]
- Panfeng, L.; Lu, W.; Qingxin, D.; Lanying, D. Estimation of potential suitable distribution area and the ecological characteristics of Eucommia ulmoides Oliv. in China. Acta Ecol. Sin. 2020, 40, 5674–5684. (In Chinese) [Google Scholar]
- Puchałka, R.; Dyderski, M.K.; Vítková, M.; Sádlo, J.; Klisz, M.; Netsvetov, M.; Prokopuk, Y.; Matisons, R.; Mionskowski, M.; Wojda, T.; et al. Black locust (Robinia pseudoacacia L.) range contraction and expansion in Europe under changing climate. Glob. Chang. Biol. 2021, 27, 1587–1600. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Jiang, P.; Huang, Y.; Yang, Y.; Wang, R.; Yang, Y. Potential geographic distribution of relict plant Pteroceltis tatarinowii in China under climate change scenarios. PLoS ONE 2022, 17, e0266133. [Google Scholar] [CrossRef]
- Chen, I.-C.; Hill, J.K.; Ohlemüller, R.; Roy, D.B.; Thomas, C.D. Rapid Range Shifts of Species Associated with High Levels of Climate Warming. Science 2011, 333, 1024–1026. [Google Scholar] [CrossRef]
- Davis, M.B.; Shaw, R.G. Range Shifts and Adaptive Responses to Quaternary Climate Change. Science 2001, 292, 673–679. [Google Scholar] [CrossRef] [Green Version]
- Call, V.; Dilcher, D. The fossil record of Eucommia (Eucommiaceae) in North America. Am. J. Bot. 1997, 84, 798. [Google Scholar] [CrossRef]
- Zhang, L.; Jing, Z.; Li, Z.; Liu, Y.; Fang, S. Predictive Modeling of Suitable Habitats for Cinnamomum Camphora (L.) Presl Using Maxent Model under Climate Change in China. Int. J. Environ. Res. Pub. He. 2019, 16, 3185. [Google Scholar] [CrossRef] [Green Version]
- Qing, J.; Meng, Y.D.; He, F.; Du, Q.X.; Zhong, J.; Du, H.Y.; Liu, P.F.; Du, L.Y.; Wang, L. Whole genome re-sequencing reveals the genetic diversity and evolutionary patterns of Eucommia ulmoides. Mol. Genet. Genom. 2022, 297, 485–494. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Deng, X.; Xiang, W.; Chen, L.; Ouyang, S. Predicting potential suitable habitats of Chinese fir under current and future climatic scenarios based on Maxent model. Ecol. Inform. 2021, 64, 101393. [Google Scholar] [CrossRef]
- Xing, Y.Y.; Wang, J.Y.; Wang, K.; Zhang, Y.; Liu, K.; Chen, X.Y.; Yuan, Y. Inhibition of Rheumatoid Arthritis Using Bark, Leaf, and Male Flower Extracts of Eucommia ulmoides. Evid. Based Complement. Altern. Med. 2020, 2020, 3260278. [Google Scholar] [CrossRef] [PubMed]
- Babu, K.N.; Rajesh, M.K.; Samsudeen, K.; Minoo, D.; Suraby, E.J.; Anupama, K.; Ritto, P. Randomly amplified polymorphic DNA (RAPD) and derived techniques. Methods Mol. Biol. 2014, 1115, 191–209. [Google Scholar] [CrossRef]
- Yuan, Y.; Tang, X.; Liu, M.; Liu, X.; Tao, J. Species Distribution Models of the Spartina alterniflora Loisel in Its Origin and Invasive Country Reveal an Ecological Niche Shift. Front. Plant Sci. 2021, 12, 738769. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Zhang, Y.; Huang, D.; Wang, H.; Cao, Q.; Fan, P.; Yang, N.; Zheng, P.; Wang, R. The effect of climate change on the richness distribution pattern of oaks (Quercus L.) in China. Sci. Total Environ. 2020, 744, 140786. [Google Scholar] [CrossRef]

| Code | Environmental Variable | Percent Contribution (%) |
|---|---|---|
| BIO02 | Mean diurnal range (Mean of monthly (max temp–min temp)) (°C) | 53.8 |
| BIO03 | Isothermality | 0.8 |
| BIO04 | Temperature seasonality (standard deviation × 100) | 0.4 |
| BIO08 | Mean temperature of wettest quarter (°C) | 0.7 |
| BIO11 | Mean temperature of coldest quarter (°C) | 41.4 |
| BIO14 | Precipitation of driest month (mm) | 1.6 |
| BIO18 | Precipitation of warmest quarter (mm) | 1.3 |
| Decades | Predicted Area (×104 km2) and % of the Corresponding Current Area | ||||
|---|---|---|---|---|---|
| Total Suitable Region | Poorly Suitable Region | Moderately Suitable Region | Highly Suitable Region | ||
| 1970–2000 | 211.14 | 113.68 | 96.39 | 1.07 | |
| SSP1-2.6 | 2050s | 210.49 | 156.97 | 53.48 | 0.04 |
| (99.69%) | (138.08%) | (55.48%) | (3.73%) | ||
| 2070s | 206.60 | 155.83 | 50.70 | 0.07 | |
| (99.69%) | (138.08%) | (55.48%) | (3.73%) | ||
| 2090s | 214.96 | 160.11 | 54.79 | 0.06 | |
| (101.81%) | (140.84%) | (56.84%) | (5.36%) | ||
| SSP2-4.5 | 2050s | 212.76 | 160.02 | 52.68 | 0.06 |
| (100.77%) | (140.76%) | (54.66%) | (5.68%) | ||
| 2070s | 204.64 | 164.38 | 40.24 | 0.06 | |
| (96.92%) | (144.59%) | (41.75%) | (1.46%) | ||
| 2090s | 197.05 | 165.42 | 31.62 | 0.01 | |
| (93.33%) | (145.51%) | (32.81%) | (0.81%) | ||
| SSP3-7.0 | 2050s | 215.84 | 155.51 | 60.29 | 0.03 |
| (102.23%) | (136.80%) | (62.55%) | (3.08%) | ||
| 2070s | 207.33 | 167.73 | 39.59 | 0.01 | |
| (98.20%) | (147.55%) | (41.07%) | (1.14%) | ||
| 2090s | 176.23 | 151.00 | 25.22 | 0.01 | |
| (83.47%) | (132.83%) | (26.17%) | (0.81%) | ||
| SSP5-8.5 | 2050s | 210.40 | 167.74 | 42.65 | 0.02 |
| (99.65%) | (147.55%) | (44.25%) | (1.62%) | ||
| 2070s | 167.59 | 142.25 | 25.33 | 0.01 | |
| (79.37%) | (125.13%) | (26.28%) | (0.81%) | ||
| 2090s | 98.87 | 87.99 | 10.87 | 0.01 | |
| (46.83%) | (77.40%) | (11.27%) | (0.65%) | ||
| LIA Statistics | Shared Socio-Economic Pathways (SSPs) | |||
|---|---|---|---|---|
| SSP1-2.6 | SSP2-4.5 | SSP3-7.0 | SSP5-8.5 | |
| Geographic area (×104 km2) | 192.77 | 178.24 | 150.35 | 73.37 |
| Percentage of current suitable area (%) | 91.30 | 84.42 | 71.21 | 34.75 |
| Percentage of SSP1-2.6 area (%) | 100.00 | 92.46 | 77.99 | 38.06 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xie, S.; Si, H.; Sun, H.; Zhao, Q.; Li, X.; Wang, S.; Niu, J.; Wang, Z. Predicting the Potential Distribution of the Endangered Plant Eucommia ulmoides in China under the Background of Climate Change. Sustainability 2023, 15, 5349. https://doi.org/10.3390/su15065349
Xie S, Si H, Sun H, Zhao Q, Li X, Wang S, Niu J, Wang Z. Predicting the Potential Distribution of the Endangered Plant Eucommia ulmoides in China under the Background of Climate Change. Sustainability. 2023; 15(6):5349. https://doi.org/10.3390/su15065349
Chicago/Turabian StyleXie, Siyuan, He Si, Hongxia Sun, Qian Zhao, Xiaodong Li, Shiqiang Wang, Junfeng Niu, and Zhezhi Wang. 2023. "Predicting the Potential Distribution of the Endangered Plant Eucommia ulmoides in China under the Background of Climate Change" Sustainability 15, no. 6: 5349. https://doi.org/10.3390/su15065349






