Estimating Olive Tree Density in Delimited Areas Using Sentinel-2 Images
Abstract
1. Introduction
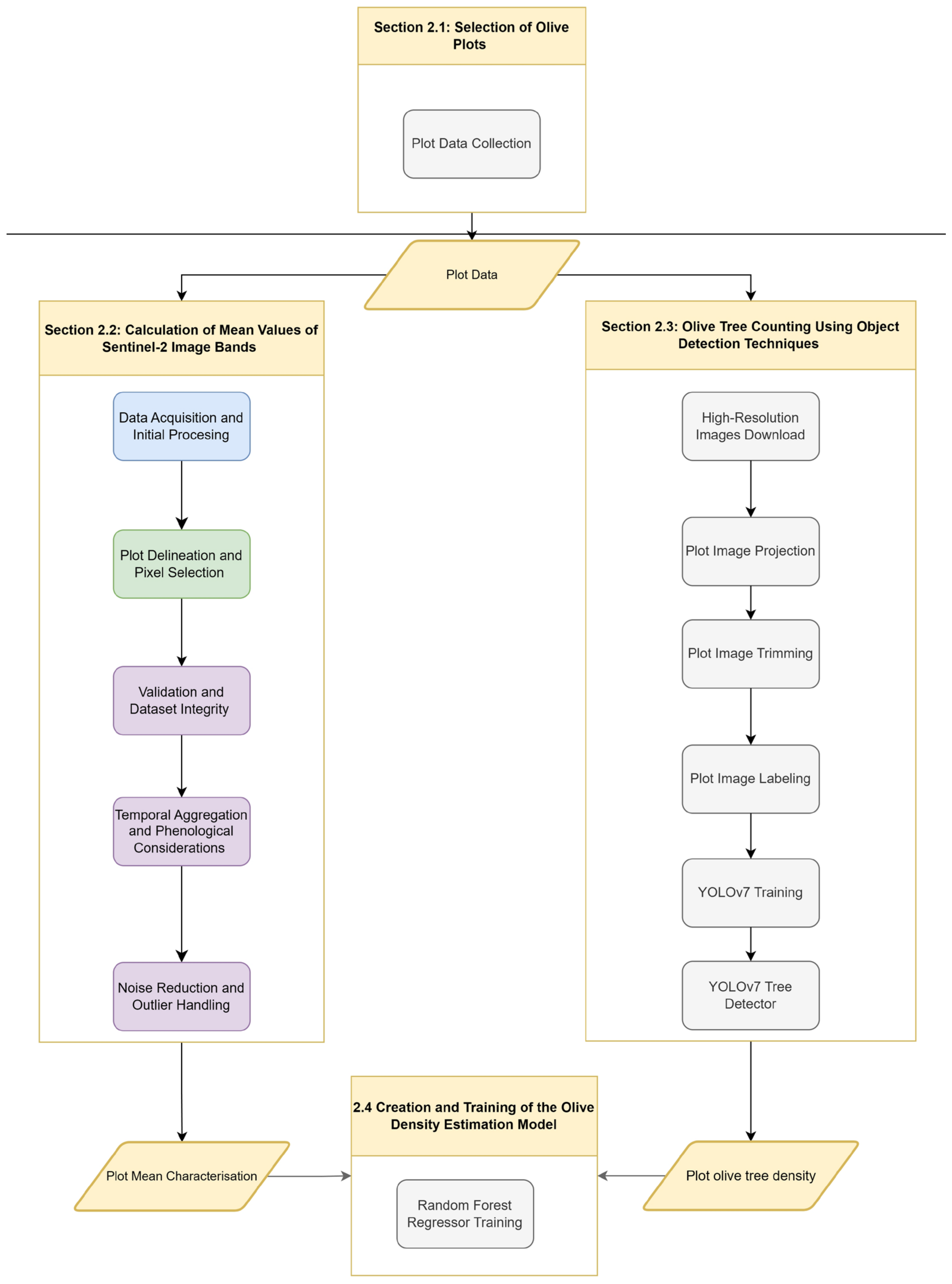
2. Materials and Methods
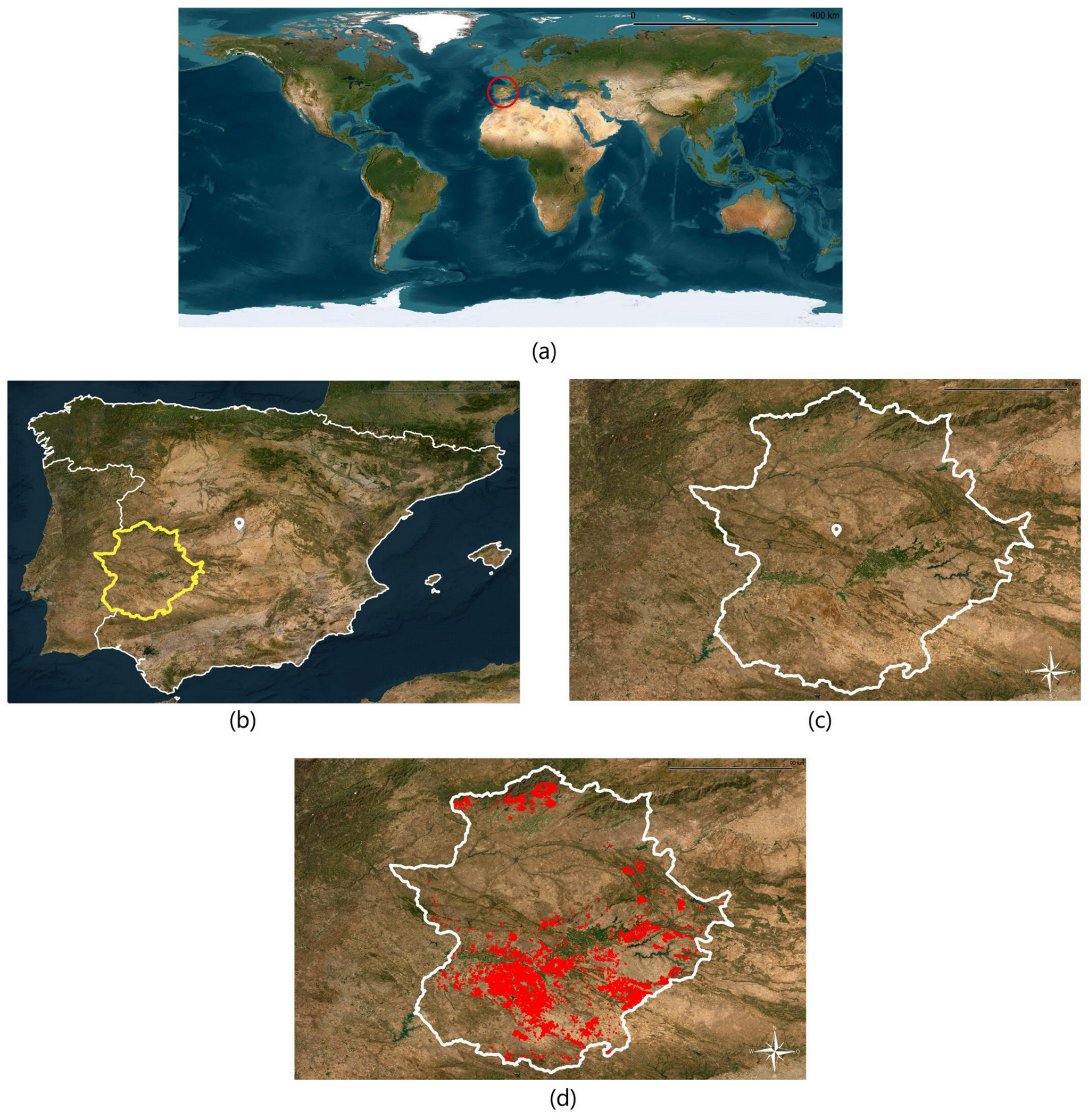
2.1. Selection of Olive Plots
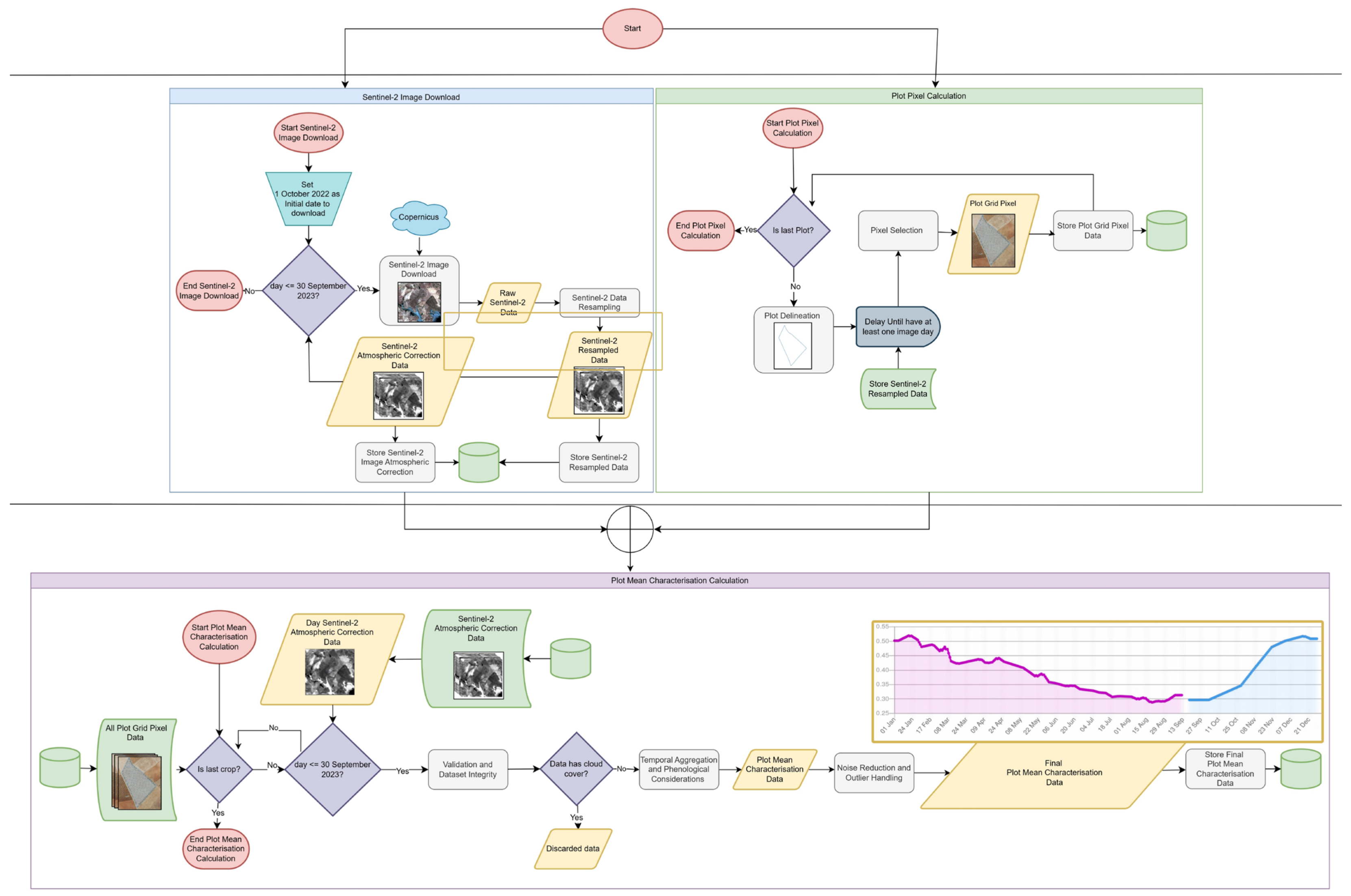
2.2. Calculation of Mean Values of Sentinel-2 Image Bands
- (1)
- Data Acquisition and Initial Processing
- (2)
- Plot Delineation and Pixel Selection
- (3)
- Validation and Dataset Integrity
- (4)
- Temporal Aggregation and Phenological Considerations
- (5)
- Noise Reduction and Outlier Handling
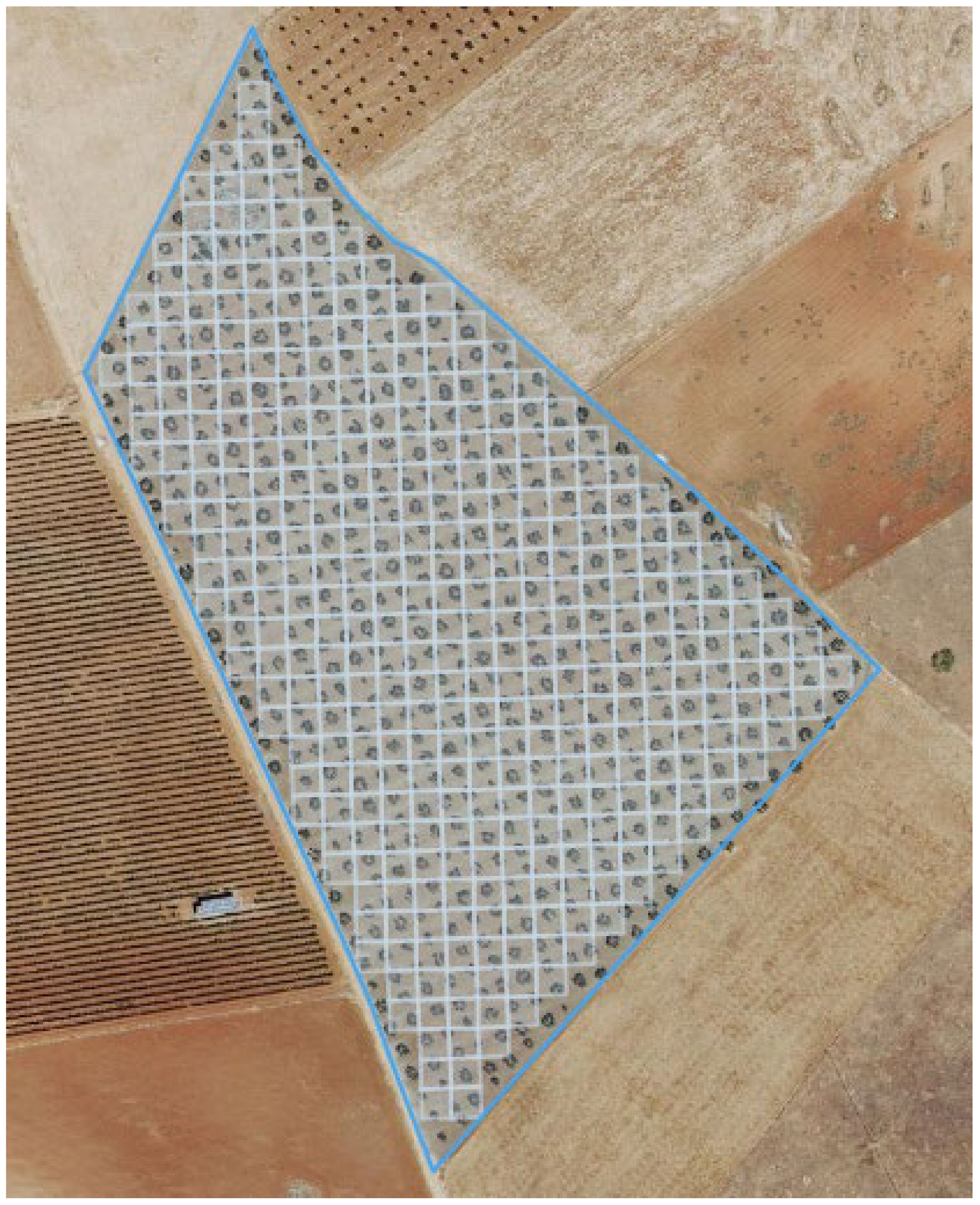
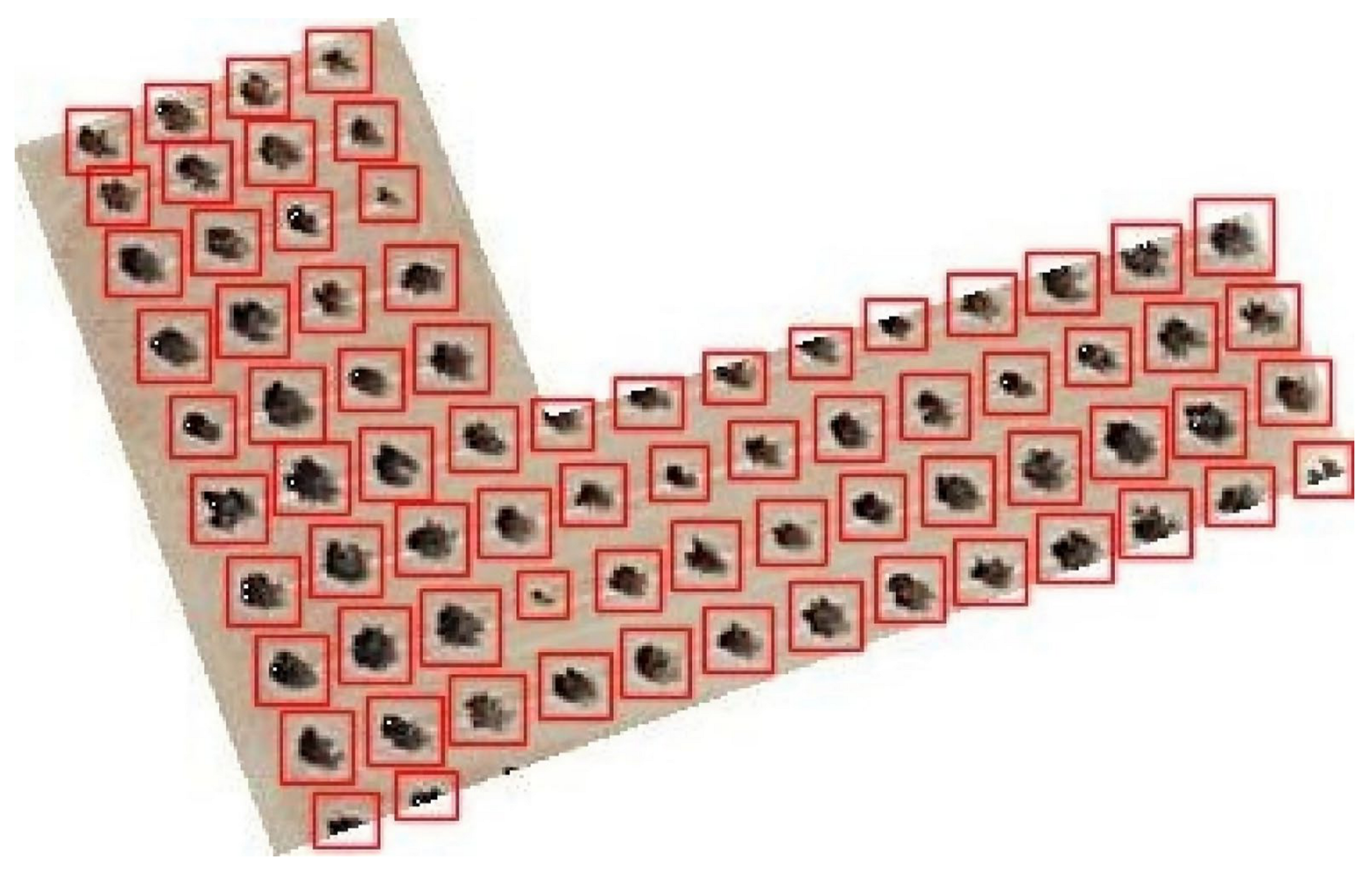
2.3. Olive Tree Counting Using Object Detection Techniques
2.4. Creation and Training of the Olive Density Estimation Model
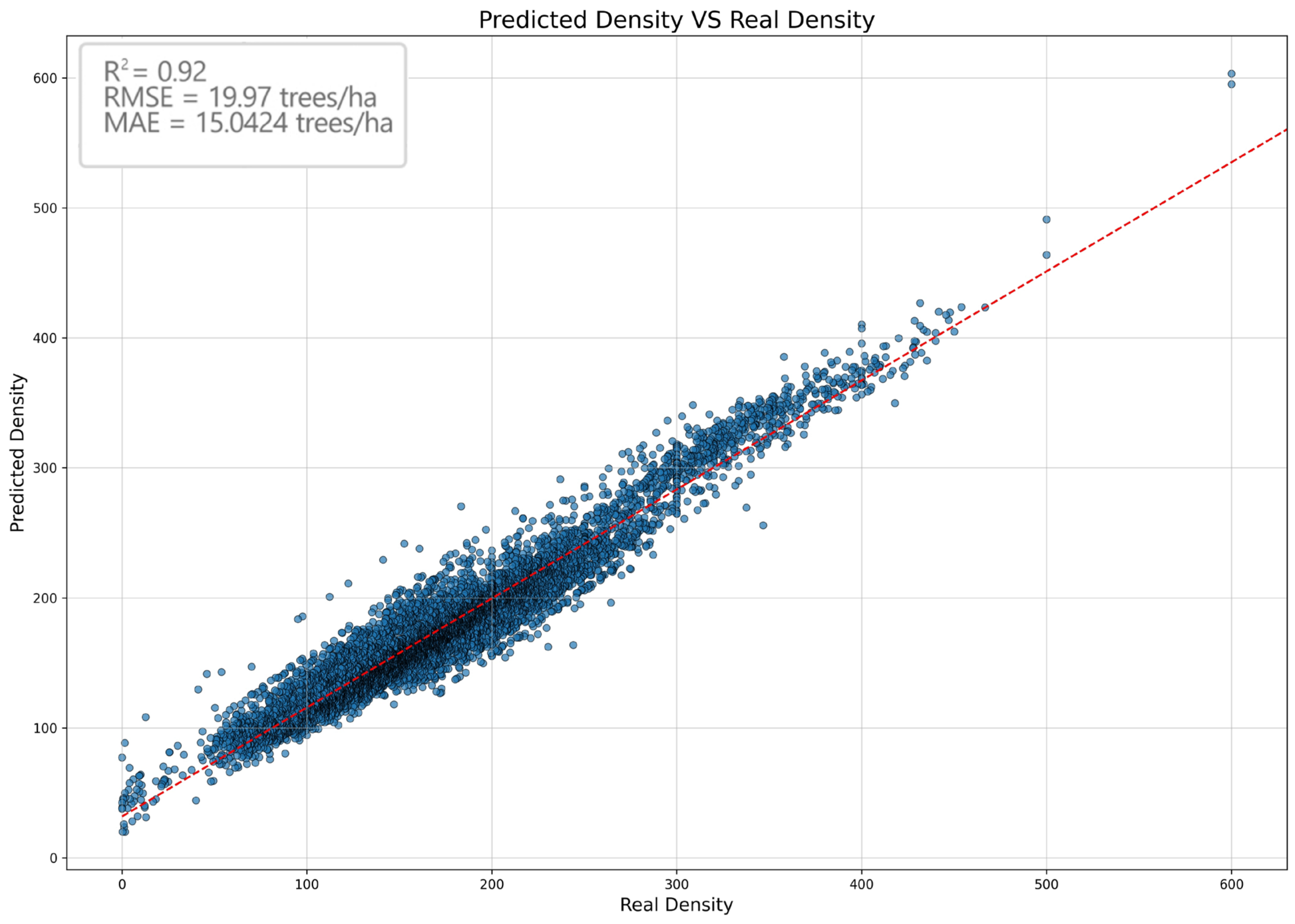
3. Results and Discussion
- -
- Mean Squared Error (MSE): 2283.3388: This metric measures the average of squared errors, that is, the difference between predicted and actual values squared. Squaring the errors magnifies them, giving more weight to larger errors and thus penalising them more heavily.
- -
- Root Mean Squared Error (RMSE): 47.7842: RMSE helps interpret MSE in the same units as the target variable, aiding in understanding the magnitude of errors (by taking the square root of the MSE). Like MSE, RMSE penalises larger errors.
- -
- Mean Absolute Error (MAE): 28.9256: This metric calculates the average of absolute errors, i.e., the absolute difference between the predicted and actual values without penalising large prediction errors. This makes MAE more robust and provides a more realistic interpretation of the overall performance of the model, especially in the presence of outliers.
- -
- R-squared (R2): 0.8105: This metric measures the proportion of variance in the target variable that can be explained by the independent variables of the model. An R2 value close to 1 indicates that the model effectively expresses the correlation between the input data and the predicted values.
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Saini, R.; Ghosh, S.K. Crop Classification on Single Date Sentinel-2 Imagery Using Random Forest and Support Vector Machine. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2018, XLII-5, 683–688. [Google Scholar] [CrossRef]
- Prins, A.J.; Van Niekerk, A. Crop type mapping using lidar, sentinel-2 and aerial imagery with machine learning algorithms. Geo-Spat. Inf. Sci. 2020, 24, 215–227. [Google Scholar] [CrossRef]
- Tian, S.; Zhang, X.; Tian, J.; Sun, Q. Random Forest Classification of Wetland Landcovers from Multi-Sensor Data in the Arid Region of Xinjiang, China. Remote Sens. 2016, 8, 954. [Google Scholar] [CrossRef]
- Sun, R.; Chen, S.; Su, H.; Mi, C.; Jin, N. The Effect of NDVI Time Series Density Derived from Spatiotemporal Fusion of Multisource Remote Sensing Data on Crop Classification Accuracy. ISPRS Int. J. Geo-Inf. 2019, 8, 502. [Google Scholar] [CrossRef]
- Yi, Z.; Jia, L.; Chen, Q. Crop Classification Using Multi-Temporal Sentinel-2 Data in the Shiyang River Basin of China. Remote Sens. 2020, 12, 4052. [Google Scholar] [CrossRef]
- Siesto, G.; Fernández-Sellers, M.; Lozano-Tello, A. Crop Classification of Satellite Imagery Using Synthetic Multitemporal and Multispectral Images in Convolutional Neural Networks. Remote Sens. 2021, 13, 3378. [Google Scholar] [CrossRef]
- Zhang, T.; Su, J.; Liu, C.; Chen, W.H.; Liu, H.; Liu, G. Band selection in sentinel-2 satellite for agriculture applications. In Proceedings of the 23rd International Conference on Automation and Computing (ICAC), Huddersfield, UK, 7–8 September 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 1–6. [Google Scholar] [CrossRef]
- Huang, Y.; Liu, X.; Li, X.; Yan, Y.; Ou, J. Comparing the Effects of Temporal Features Derived from Synthetic Time-Series NDVI on Fine Land Cover Classification. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2018, 11, 4618–4629. [Google Scholar] [CrossRef]
- Zhang, T.X.; Su, J.Y.; Liu, C.J.; Chen, W.H. Potential Bands of Sentinel-2A Satellite for Classification Problems in Precision Agriculture. Int. J. Autom. Comput. 2018, 16, 16–26. [Google Scholar] [CrossRef]
- Jiao, L.; Zhang, F.; Liu, F.; Yang, S.; Li, L.; Feng, Z.; Qu, R. A Survey of Deep Learning-Based Object Detection. IEEE Access 2019, 7, 128837–128868. [Google Scholar] [CrossRef]
- Shankar, R.; Muthulakshmi, M. Comparing YOLOV3, YOLOV5 & YOLOV7 architectures for underwater marine creatures detection. In Proceedings of the International Conference on Computational Intelligence and Knowledge Economy (ICCIKE), Dubai, United Arab Emirates, 9–10 March 2023; pp. 25–30. [Google Scholar] [CrossRef]
- Chen, Y.; Xu, H.; Zhang, X.; Gao, P.; Xu, Z.; Huang, X. An object detection method for bayberry trees based on an improved YOLO algorithm. Int. J. Digit. Earth 2023, 16, 781–805. [Google Scholar] [CrossRef]
- Xu, S.; Wang, R.; Shi, W.; Wang, X. Classification of Tree Species in Transmission Line Corridors Based on YOLO v7. Forests 2024, 15, 61. [Google Scholar] [CrossRef]
- Mosin, V.; Aguilar, R.; Platonov, A.; Vasiliev, A.; Kedrov, A.; Ivanov, A. Remote sensing and machine learning for tree detection and classification in forestry applications. Image Signal Process. Remote Sens. XXV 2019, 11155, 130–141. [Google Scholar] [CrossRef]
- Nevalainen, O.; Honkavaara, E.; Tuominen, S.; Viljanen, N.; Hakala, T.; Yu, X.; Hyyppä, J.; Saari, H.; Pölönen, I.; Imai, N.N.; et al. Individual Tree Detection and Classification with UAV-Based Photogrammetric Point Clouds and Hyperspectral Imaging. Remote Sens. 2017, 9, 185. [Google Scholar] [CrossRef]
- Ke, Y.; Quackenbush, L. A review of methods for automatic individual tree-crown detection and delineation from passive remote sensing. Int. J. Remote Sens. 2011, 32, 4725–4747. [Google Scholar] [CrossRef]
- Santos, A.A.d.; Marcato Junior, J.; Araújo, M.S.; Di Martini, D.R.; Tetila, E.C.; Siqueira, H.L.; Aoki, C.; Eltner, A.; Matsubara, E.T.; Pistori, H.; et al. Assessment of CNN-Based Methods for Individual Tree Detection on Images Captured by RGB Cameras Attached to UAVs. Sensors 2019, 19, 3595. [Google Scholar] [CrossRef] [PubMed]
- Yao, L.; Liu, T.; Qin, J.; Lu, N.; Zhou, C. Tree counting with high spatial-resolution satellite imagery based on deep neural networks. Ecol. Indic. 2021, 125, 107591. [Google Scholar] [CrossRef]
- Liu, T.; Yao, L.; Qin, J.; Lu, J.; Lu, N.; Zhou, C. A Deep Neural Network for the Estimation of Tree Density Based on High-Spatial Resolution Image. IEEE Trans. Geosci. Remote Sens. 2021, 60, 4403811. [Google Scholar] [CrossRef]
- Bazi, Y.; Melgani, F.; Al-Sharari, H.D. An automatic method for counting olive trees in very high spatial remote sensing images. In Proceedings of the IEEE International Geoscience and Remote Sensing Symposium (IGARSS), Cape Town, South Africa, 12–17 July 2009; pp. 125–128. [Google Scholar] [CrossRef]
- Persson, M.; Lindberg, E.; Reese, H. Tree Species Classification with Multi-Temporal Sentinel-2 Data. Remote Sens. 2018, 10, 1794. [Google Scholar] [CrossRef]
- Wessel, M.; Brandmeier, M.; Tiede, D. Evaluation of Different Machine Learning Algorithms for Scalable Classification of Tree Types and Tree Species Based on Sentinel-2 Data. Remote Sens. 2018, 10, 1419. [Google Scholar] [CrossRef]
- Mantas, V.; Fonseca, L.; Baltazar, E.; Canhoto, J.; Abrantes, I. Detection of Tree Decline (Pinus pinaster Aiton) in European Forests Using Sentinel-2 Data. Remote Sens. 2022, 14, 2028. [Google Scholar] [CrossRef]
- Polyakova, A.; Mukharamova, S.; Yermolaev, O.; Shaykhutdinova, G. Automated Recognition of Tree Species Composition of Forest Communities Using Sentinel-2 Satellite Data. Remote Sens. 2023, 15, 329. [Google Scholar] [CrossRef]
- Grabska, E.; Frantz, D.; Ostapowicz, K. Evaluation of machine learning algorithms for forest stand species mapping using Sentinel-2 imagery and environmental data in the Polish Carpathians. Remote Sens. Environ. 2020, 251, 112103. [Google Scholar] [CrossRef]
- Lozano-Tello, A.; Fernández-Sellers, M.; Quirós, E.; Fragoso-Campón, L.; García-Martín, A.; Gutiérrez Gallego, J.A.; Mateos, C.; Muñoz, P. Crop identification by massive processing of multiannual satellite imagery for EU common agriculture policy subsidy control. Eur. J. Remote Sens. 2021, 54, 1–12. [Google Scholar] [CrossRef]
- Zhu, X.; Wang, R.; Shi, W.; Liu, X.; Ren, Y.; Xu, S.; Wang, X. Detection of Pine-Wilt-Disease-Affected Trees Based on Improved YOLO v7. Forests 2024, 15, 691. [Google Scholar] [CrossRef]
- Liu, K.; Sun, Q.; Sun, D.; Peng, L.; Yang, M.; Wang, N. Underwater Target Detection Based on Improved YOLOv7. J. Mar. Sci. Eng. 2023, 11, 677. [Google Scholar] [CrossRef]
- Yun, T.; Li, J.; Ma, L.; Zhou, J.; Wang, R.; Eichhorn, M.P.; Zhang, H. Status, advancements and prospects of deep learning methods applied in forest studies. Int. J. Appl. Earth Obs. Geoinf. 2024, 131, 103938. [Google Scholar] [CrossRef]
- Wang, C.Y.; Bochkovskiy, A.; Liao, H.Y.M. YOLOv7: Trainable Bag-of-Freebies Sets New State-of-the-Art for Real-Time Object Detectors. In Proceedings of the 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Vancouver, BC, Canada, 17–24 June 2023; pp. 7464–7475. [Google Scholar] [CrossRef]
- Lankford, S. Effective tuning of regression models using an evolutionary approach: A case study. In Proceedings of the 2020 3rd Artificial Intelligence and Cloud Computing Conference, Kyoto, Japan, 18–20 December 2020; Association for Computing Machinery: New York, NY, USA, 2021; pp. 102–108. [Google Scholar] [CrossRef]
- Rodriguez-Galiano, V.; Sanchez-Castillo, M.; Chica-Olmo, M.; Chica-Rivas, M. Machine learning predictive models for mineral prospectivity: An evaluation of neural networks, random forest, regression trees and support vector machines. Ore Geol. Rev. 2015, 71, 804–818. [Google Scholar] [CrossRef]
- Gzar, D.A.; Mahmood, A.M.; Abbas, M.K. A comparative study of regression machine learning algorithms: Tradeoff between accuracy and computational complexity. MMEP 2022, 9, 1217–1224. [Google Scholar] [CrossRef]
- Lozano-Tello, A.; Siesto, G.; Fernández-Sellers, M.; Caballero-Mancera, A. Evaluation of the Use of the 12 Bands vs. NDVI from Sentinel-2 Images for Crop Identification. Sensors 2023, 23, 7132. [Google Scholar] [CrossRef]
- Muraina, I. Ideal dataset splitting ratios in machine learning algorithms: General concerns for data scientists and data analysts. In Proceedings of the 7th International Mardin Artuklu Scientific Research Conference, Mardin, Turkey, 13–15 December 2022; pp. 496–504. Available online: https://www.researchgate.net/publication/358284895 (accessed on 5 September 2024).






| Model | Plots | Olives | Precision | Recall | mAP@0.5 | mAP@0.5:0.95 |
|---|---|---|---|---|---|---|
| 1 | 25 | 1102 | 0.131 | 0.279 | 0.0937 | 0.174 |
| 2 | 32 | 2032 | 0.156 | 0.192 | 0.149 | 0.035 |
| 3 | 41 | 3031 | 0.331 | 0.386 | 0.266 | 0.0559 |
| 4 | 50 | 4105 | 0.353 | 0.445 | 0.297 | 0.0614 |
| 5 | 56 | 5090 | 0.466 | 0.626 | 0.511 | 0.134 |
| 6 | 68 | 6082 | 0.488 | 0.651 | 0.515 | 0.134 |
| 7 | 83 | 7249 | 0.518 | 0.622 | 0.525 | 0.145 |
| 8 | 86 | 8055 | 0.546 | 0.596 | 0.544 | 0.181 |
| 9 | 93 | 9042 | 0.645 | 0.527 | 0.545 | 0.183 |
| 10 | 49 | 10,055 | 0.671 | 0.557 | 0.587 | 0.193 |
| 11 | 97 | 11,060 | 0.707 | 0.597 | 0.6155 | 0.226 |
| 12 | 98 | 12,063 | 0.723 | 0.634 | 0.682 | 0.246 |
| 13 | 108 | 13,044 | 0.767 | 0.597 | 0.67 | 0.247 |
| 14 | 113 | 14,085 | 0.804 | 0.721 | 0.698 | 0.258 |
| 15 | 133 | 15,014 | 0.846 | 0.836 | 0.823 | 0.288 |
| 16 | 142 | 16,004 | 0.857 | 0.848 | 0.859 | 0.292 |
| 17 | 159 | 17,013 | 0.865 | 0.851 | 0.896 | 0.315 |
| Real Density Range | Low | Aperture (Q1) | Close (Median) | High Aperture (Q3) | High |
|---|---|---|---|---|---|
| 0–100 | 20.02607737 | 91.53829665 | 100.63791 | 111.7480847 | 185.9161699 |
| 100–200 | 94.37370502 | 129.4377352 | 153.1214327 | 173.1675677 | 270.48398 |
| 200–300 | 154.322933 | 201.6940327 | 217.7084302 | 238.5834046 | 336.4272699 |
| 300–400 | 255.9488633 | 308.8662211 | 325.5536854 | 344.8509101 | 389.2324947 |
| 400–500 | 349.8323968 | 373.2557864 | 382.5139817 | 399.8740504 | 426.781692 |
| 500–600 | 463.8554844 | 470.6864178 | 477.5173513 | 484.3482847 | 491.1792182 |
| 600–700 | 595.0706677 | 597.1009745 | 599.1312812 | 601.161588 | 603.1918948 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lozano-Tello, A.; Luceño, J.; Caballero-Mancera, A.; Clemente, P.J. Estimating Olive Tree Density in Delimited Areas Using Sentinel-2 Images. Remote Sens. 2025, 17, 508. https://doi.org/10.3390/rs17030508
Lozano-Tello A, Luceño J, Caballero-Mancera A, Clemente PJ. Estimating Olive Tree Density in Delimited Areas Using Sentinel-2 Images. Remote Sensing. 2025; 17(3):508. https://doi.org/10.3390/rs17030508
Chicago/Turabian StyleLozano-Tello, Adolfo, Jorge Luceño, Andrés Caballero-Mancera, and Pedro J. Clemente. 2025. "Estimating Olive Tree Density in Delimited Areas Using Sentinel-2 Images" Remote Sensing 17, no. 3: 508. https://doi.org/10.3390/rs17030508
APA StyleLozano-Tello, A., Luceño, J., Caballero-Mancera, A., & Clemente, P. J. (2025). Estimating Olive Tree Density in Delimited Areas Using Sentinel-2 Images. Remote Sensing, 17(3), 508. https://doi.org/10.3390/rs17030508












