Abstract
1-Amino-3-nitroguanidine (ANQ, 1) was synthesized by hydrazinolysis of nitroguanidine (NQ) with hydrazine hydrate. Four different amino-nitroguanidinium salts (chloride (2), bromide (3), iodide (4) and sulfate (5)) were synthesized and structurally characterized by low-temperature X-ray diffraction. The halides 2–4 could only be obtained crystalline as monohydrates. In addition, they were characterized by NMR and vibrational spectroscopy, elemental analysis and the sensitivities towards impact, friction and electrostatic discharge were determined. The compounds can be used in silver (AgX, X = Cl, Br, I) and barium (BaSO4) based metathesis reactions in order to form more complex salts of 1-amino-nitroguanidine.
1. Introduction
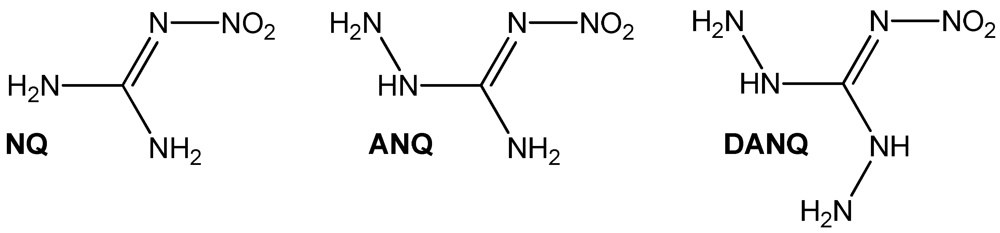
1-Amino-3-nitroguanidine (1, ANQ) can be described as the aminated sister compound of the famous 1-nitroguanidine (NQ) [1]. The next step in the preparation of mixed aminonitroguanidines would be the synthesis of 1,3-diamino-5-nitroguanidine (DANQ), which is completely unknown yet (Figure 1). Nitroguanidine is used in airbags and many other double- and triple- based propellant applications [2]. Interestingly, only very little correspondence on aminonitroguanidine is found in the literature [3,4,5]. We recently investigated the use of ANQ in energetic materials in its neutral as well as protonated form [6]. For example, highly energetic 1-amino-3-nitroguanidinium dinitramide has been described [6]. Because dinitraminic acid (HN(NO2)2) and concentrated aqueous solutions can explode spontaneously, the reaction pathway using a metathesis reaction of amino-nitroguanidinium chloride with silver dinitramide was chosen. This example shows the utility of the herein described compounds as precursor materials for the synthesis of ionic energetic materials containing the 1-amino-3-nitro-guanidinium cation when using metathesis reaction protocols to precipitate low soluble silver halides in case of reacting the ANQ halides with the corresponding silver salts of the used acids or to precipitate BaSO4 in case the corresponding Ba-salts are reacted with the herein described sulfate salt of ANQ. Here we present the synthesis and characterization of three amino-nitroguanidinium halides and bis(amino-nitroguanidinium) sulfate.
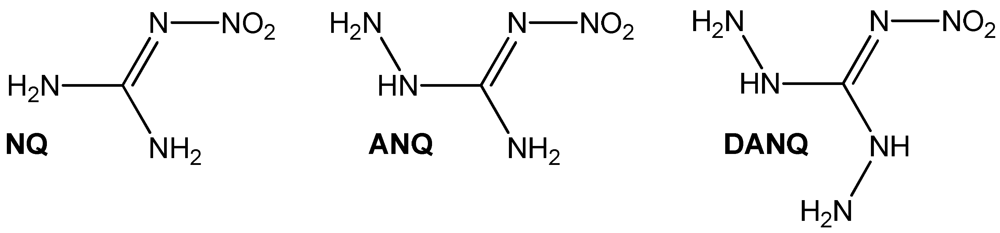
Figure 1.
Chemical structures of amino-nitroguanidines: NQ: 1-nitroguanidine, ANQ: 1-amino-3-nitroguanidine, DANQ: 1,3-Diamino-5-nitroguanidine (not known yet).
Figure 1.
Chemical structures of amino-nitroguanidines: NQ: 1-nitroguanidine, ANQ: 1-amino-3-nitroguanidine, DANQ: 1,3-Diamino-5-nitroguanidine (not known yet).
2. Results and Discussion
2.1. Synthesis
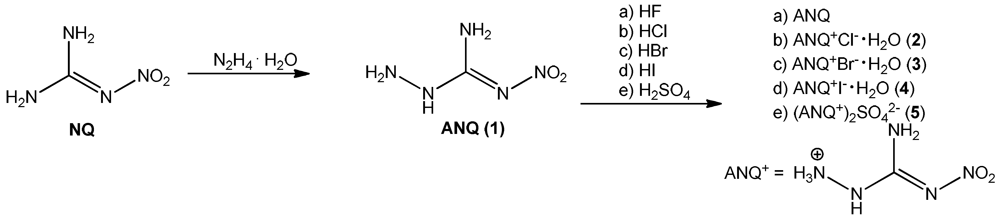
The synthesis of 1-amino-3-nitroguanidine (1: ANQ) was achieved starting from commercially available nitroguanidine (NQ) by treatment with hydrazine hydrate. In a hydrazinolysis reaction, the aminated nitroguanidine is formed after the elimination of ammonia [5]. Unprotonated ANQ shows rather poor solubility in water, so that it can be isolated from the reaction mixture after neutralization by suction filtration. Unlike in water or buffered neutral solutions, in acidic media ANQ is dissolved comparatively easily upon the formation of the protonated ANQ+ species. Therefore, the halides as well as the sulfate salt can be prepared by dissolving ANQ in dilute aqueous solutions of the respective mineral acids HCl, HBr, HI and H2SO4. To fully synthesize and characterize all halides (F−, Cl−, Br−, I−) of ANQ, it was also dissolved in 40% aqueous HF. Unfortunately it was only possible to isolate unprotonated ANQ from this reaction mixture (Scheme 1). The reason for this behavior is easily found in the pKa values of the used mineral acids. ANQ, due to the presence of the electron withdrawing character of the nitro group, is a comparatively weak base and requires strong mineral acids for protonation in aqueous media. Since the acidic strength of the hydrohalides in aqueous solution decreases in the order HI > HBr > HCl > HF, the latter one is not able to protonate ANQ in aqueous solution any more. However, 40% aqueous HF proved to be an excellent solvent for the recrystallization of the poorly water soluble ANQ, especially if single crystals, e.g., for single crystal X-ray diffraction, are needed.
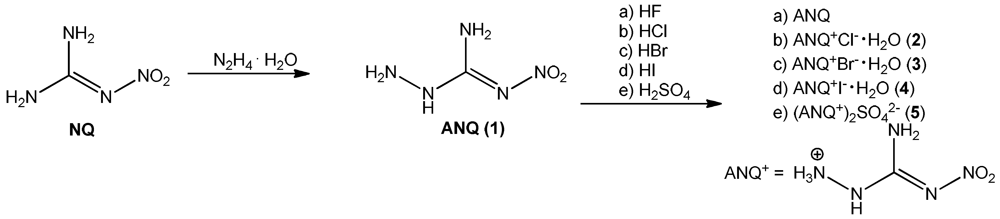
Scheme 1.
Synthesis of 1 (ANQ) and investigated amino-nitroguanidinium salts 2–5.
Scheme 1.
Synthesis of 1 (ANQ) and investigated amino-nitroguanidinium salts 2–5.
The storage stability of the halides decreases with increasing molecular weight of the anion. Whereas the hydrochloride remains a colorless crystalline material even after several months, the hydrobromide discolors slightly and the hydroiodide turns completely dark after the release of I2 indicating a decomposition of the material.
2.2. Single Crystal X-ray Structure Analysis
The low temperature determination of the crystal structures of 2–5 was performed on an Oxford Xcalibur3 diffractometer with a Spellman generator (voltage 50 kV, current 40 mA) and a KappaCCD detector. The data collection and reduction was carried out using the CRYSALISPRO software [7]. The structures were solved either with SHELXS-97 [8] or SIR-92 [9], refined with SHELXL-97 [10] and finally checked using the PLATON [11] software integrated in the WINGX [12] software suite. The non-hydrogen atoms were refined anisotropically and the hydrogen atoms were located and freely refined. The absorptions were corrected with a Scale3 Abspack multi-scan method [13]. Selected data and parameters of the X-ray determinations are given in Table 1. Crystallographic data for the structures have been deposited with the Cambridge Crystallographic Data Centre [14].

Table 1.
X-ray data and parameters of 2–5.
| 2 | 3 | 4 | 5 | |
|---|---|---|---|---|
| Formula | CH6N5O2, H2O, Cl | CH6N5O2, H2O, Br | CH6N5O2, H2O, I | 2(CH6N5O2), SO4 |
| FW [g mol−1] | 173.56 | 218.03 | 265.02 | 336.29 |
| Crystal system | Monoclinic | Monoclinic | Triclinic | Orthorhombic |
| Space Group | P21/n | P21/c | P-1 | Fdd2 |
| Color/Habit | Colorless block | Colorless plate | Colorless block | Colorless block |
| Size [mm] | 0.37 × 0.44 × 0.49 | 0.04 × 0.21 × 0.50 | 0.38 × 0.42 × 0.50 | 0.14 × 0.22 × 0.26 |
| a [Å] | 7.6616(6) | 8.0261(5) | 6.6931(3) | 11.0638(8) |
| b [Å] | 12.7248(7) | 12.7412(7) | 7.0540(4) | 30.380(2) |
| c [Å] | 7.8116(8) | 7.9296(5) | 9.1359(5) | 6.7531(5) |
| α [°] | 90 | 90 | 97.754(5) | 90 |
| β [°] | 119.012(11) | 118.051(9) | 96.942(4) | 90 |
| γ [°] | 90 | 90 | 115.961(5) | 90 |
| V [Å3] | 666.01(12) | 715.64(7) | 376.35(4) | 2269.8(3) |
| Z | 4 | 4 | 2 | 8 |
| ρcalc. [g cm−3] | 1.731 | 2.024 | 2.339 | 1.968 |
| μ [mm−1] | 0.535 | 5.708 | 4.221 | 0.359 |
| F(000) | 360 | 432 | 252 | 1392 |
| λMoKα [Å] | 0.71073 | 0.71073 | 0.71073 | 0.71073 |
| T [K] | 173 | 173 | 173 | 173 |
| θ Min–Max [°] | 4.4, 28.6 | 4.3, 33.6 | 4.4, 32.4 | 4.6, 30.1 |
| Dataset | −10:7; −17:9; −8:9 | −9:9; −15:10; −9:9 | −9:9; −10:10; −13:13 | −15:15; −42:42; −9:9 |
| Reflections collected | 2525 | 5748 | 5175 | 10,853 |
| Independent refl. | 1437 | 2172 | 2457 | 1670 |
| Rint | 0.017 | 0.047 | 0.023 | 0.055 |
| Observed reflections | 1298 | 1395 | 2286 | 1392 |
| Parameters | 116 | 123 | 116 | 120 |
| R1 (obs) | 0.0292 | 0.0411 | 0.0220 | 0.0283 |
| wR2 (all data) | 0.0807 | 0.0697 | 0.0554 | 0.0607 |
| S(GooF) | 1.09 | 1.04 | 1.08 | 0.95 |
| Resd. Dens. [e Å−3] | −0.38, 0.43 | −0.77, 0.68 | −1.56, 0.76 | −0.35, 0.25 |
| Device type | Oxford Xcalibur3 | Oxford Xcalibur3 | Oxford Xcalibur3 | Oxford Xcalibur3 |
| CCD | CCD | CCD | CCD | |
| Solution | SIR-92 | SHELXS-97 | SIR-92 | SHELXS-97 |
| Refinement | SHELXL-97 | SHELXL-97 | SHELXL-97 | SHELXL-97 |
| CCDC | 858928 | 858930 | 858931 | 858929 |
The structure of the 1-amino-3-nitroguanidinium cation is similar in all four structures investigated in this work. The cation is almost planar, controlled by the intramolecular hydrogen bond N5–H5···O1 (D···A: 2.589(2) Å–2.605(3) Å) [graph set [15]: 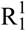 (6)] but also by the strong angulated (~90°) hydrogen bond N4–H4x···N1 (D···A: 2.547(5) Å–2.576(7) Å) [graph set:
(6)] but also by the strong angulated (~90°) hydrogen bond N4–H4x···N1 (D···A: 2.547(5) Å–2.576(7) Å) [graph set: 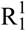 (5)]. The C–N bond distances are between typical C–N single and C=N double bonds showing the delocalization of the positive charge. However, in all structures the bond length between C1 and N5 is observed to be the shortest one. Also the bonds N2–O1, N2–O2 and N1–N2 are significantly shorter than single bonds. These structural details are in agreement with our previously described investigations of this cation in the literature [6]. Selected bond lengths and angles of the cation of all structurally investigated compounds 2–5 are gathered in Table 2. The halides, which were obtained crystalline under their monohydrated form, crystallize in common space groups (2: P21/n, 3: P21/c, 4: P-1) and follow the trend of rising densities (2: 1.73 < 3: 2.02 < 4: 2.34 g cm–3). All three structures are dominated by many hydrogen bonds [16,17,18] involving the crystal water molecules. Figure 2 shows the molecular structure of 2.
(5)]. The C–N bond distances are between typical C–N single and C=N double bonds showing the delocalization of the positive charge. However, in all structures the bond length between C1 and N5 is observed to be the shortest one. Also the bonds N2–O1, N2–O2 and N1–N2 are significantly shorter than single bonds. These structural details are in agreement with our previously described investigations of this cation in the literature [6]. Selected bond lengths and angles of the cation of all structurally investigated compounds 2–5 are gathered in Table 2. The halides, which were obtained crystalline under their monohydrated form, crystallize in common space groups (2: P21/n, 3: P21/c, 4: P-1) and follow the trend of rising densities (2: 1.73 < 3: 2.02 < 4: 2.34 g cm–3). All three structures are dominated by many hydrogen bonds [16,17,18] involving the crystal water molecules. Figure 2 shows the molecular structure of 2.
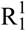 (6)] but also by the strong angulated (~90°) hydrogen bond N4–H4x···N1 (D···A: 2.547(5) Å–2.576(7) Å) [graph set:
(6)] but also by the strong angulated (~90°) hydrogen bond N4–H4x···N1 (D···A: 2.547(5) Å–2.576(7) Å) [graph set: 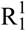 (5)]. The C–N bond distances are between typical C–N single and C=N double bonds showing the delocalization of the positive charge. However, in all structures the bond length between C1 and N5 is observed to be the shortest one. Also the bonds N2–O1, N2–O2 and N1–N2 are significantly shorter than single bonds. These structural details are in agreement with our previously described investigations of this cation in the literature [6]. Selected bond lengths and angles of the cation of all structurally investigated compounds 2–5 are gathered in Table 2. The halides, which were obtained crystalline under their monohydrated form, crystallize in common space groups (2: P21/n, 3: P21/c, 4: P-1) and follow the trend of rising densities (2: 1.73 < 3: 2.02 < 4: 2.34 g cm–3). All three structures are dominated by many hydrogen bonds [16,17,18] involving the crystal water molecules. Figure 2 shows the molecular structure of 2.
(5)]. The C–N bond distances are between typical C–N single and C=N double bonds showing the delocalization of the positive charge. However, in all structures the bond length between C1 and N5 is observed to be the shortest one. Also the bonds N2–O1, N2–O2 and N1–N2 are significantly shorter than single bonds. These structural details are in agreement with our previously described investigations of this cation in the literature [6]. Selected bond lengths and angles of the cation of all structurally investigated compounds 2–5 are gathered in Table 2. The halides, which were obtained crystalline under their monohydrated form, crystallize in common space groups (2: P21/n, 3: P21/c, 4: P-1) and follow the trend of rising densities (2: 1.73 < 3: 2.02 < 4: 2.34 g cm–3). All three structures are dominated by many hydrogen bonds [16,17,18] involving the crystal water molecules. Figure 2 shows the molecular structure of 2. 
Table 2.
Selected bond lengths [Å] of compounds 2–5.
| bond lengths [Å] | 2 | 3 | 4 | 5 |
|---|---|---|---|---|
| O1–N2 | 1.235(2) | 1.241(4) | 1.236(3) | 1.239(2) |
| O2–N2 | 1.234(2) | 1.229(4) | 1.242(3) | 1.255(2) |
| N1–N2 | 1.361(2) | 1.366(4) | 1.345(3) | 1.330(2) |
| N3–N4 | 1.423(2) | 1.433(4) | 1.419(3) | 1.417(2) |
| N1–C1 | 1.339(2) | 1.337(4) | 1.348(3) | 1.356(3) |
| N3–C1 | 1.368(2) | 1.374(4) | 1.355(4) | 1.349(3) |
| N5–C1 | 1.317(2) | 1.318(4) | 1.315(3) | 1.311(2) |
| bond angles [°] | 2 | 3 | 4 | 5 |
| C1–N1–N2 | 117.7(1) | 117.5(3) | 118.0(2) | 117.5(1) |
| C1–N3–N4 | 117.8(1) | 116.6(3) | 118.1(2) | 119.4(2) |
| O2–N2–O1 | 121.8(1) | 121.8(3) | 121.2(2) | 120.7(1) |
| O2–N2–N1 | 114.2(1) | 114.3(3) | 114.4(2) | 115.7(1) |
| O1–N2–N1 | 124.0(2) | 123.8(3) | 124.5(2) | 123.6(1) |
| N5–C1–N1 | 130.5(2) | 130.9(3) | 130.6(2) | 130.0(2) |
| N5–C1–N3 | 116.0(1) | 115.1(3) | 116.7(2) | 118.0(2) |
| N1–C1–N3 | 113.5(1) | 114.0(3) | 112.7(2) | 111.8(2) |
| Torsion angles [°] | 2 | 4 | 5 | |
| N2–N1–C1–N5 | 0.5(2) | −0.8(5) | 4.6(3) | 10.9(3) |
| N4–N3–C1–N5 | 171.1(1) | 174.0(3) | −177.1(2) | −174.0(2) |
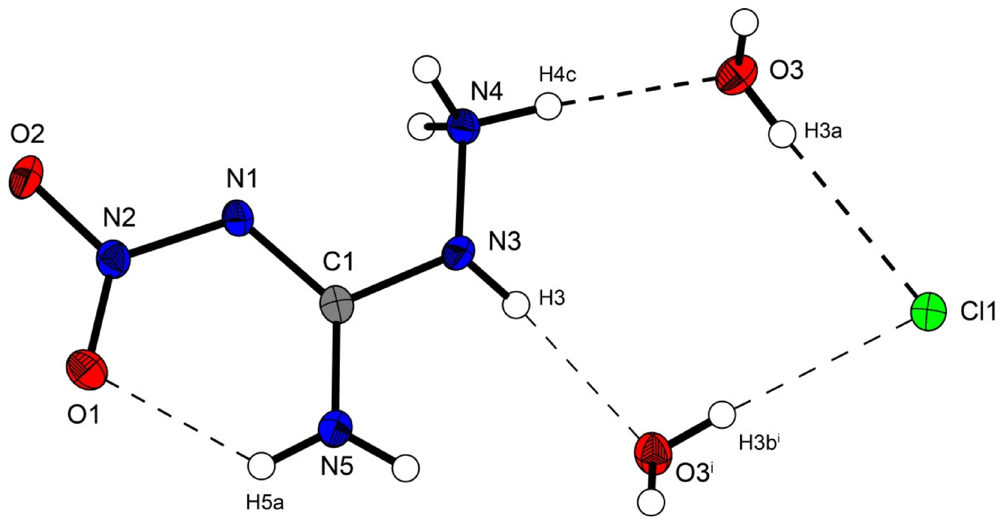
Figure 2.
Molecular structure of amino-nitro-guanidinium chloride monohydrate (2). Ellipsoids are drawn at the 50% probability level. Symmetry code: (i) 0.5 + x, 0.5 − y, 0.5 + z.
Figure 2.
Molecular structure of amino-nitro-guanidinium chloride monohydrate (2). Ellipsoids are drawn at the 50% probability level. Symmetry code: (i) 0.5 + x, 0.5 − y, 0.5 + z.
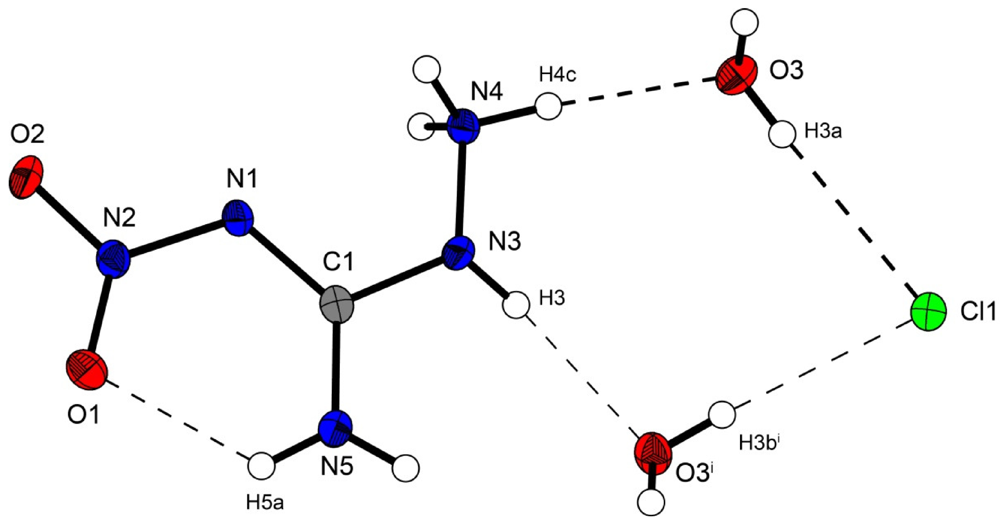
In the crystal structure the ions are held together by a three dimensional network of moderate N–H···O (d(N–O) ~2.83–3.04 Å), N–H···Cl (d(N–Cl) ~3.24–3.30 Å) and O–H···Cl (d(O–Cl ~3.08–3.14 Å) hydrogen bonds. The distorted pseudo-octahedral coordination sphere of the chloride anions is shown in Figure 3. The chloride anion is coordinated by two water molecules and four cations (two times by the amine group and two times by the ammonium group). The bond lengths and angles of selected hydrogen bonds are listed in Table 3 and Table 4. The water molecules bridge between the chloride anions and 1-amino-3-nitroguanidinium cation, and act both as acceptor and donor. The cations build a chain via the hydrogen bridge N3vi–H3vi···O2 along the b-axis [graph set: C(6)]. Two cations build dimers via the hydrogen bridge N4ii–H4cii···O2 parallel to the c-axis [graph set: 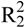 (14)]. The oxygen atom (O2) of the nitro group acts as bifurcated acceptor.
(14)]. The oxygen atom (O2) of the nitro group acts as bifurcated acceptor.
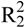 (14)]. The oxygen atom (O2) of the nitro group acts as bifurcated acceptor.
(14)]. The oxygen atom (O2) of the nitro group acts as bifurcated acceptor. 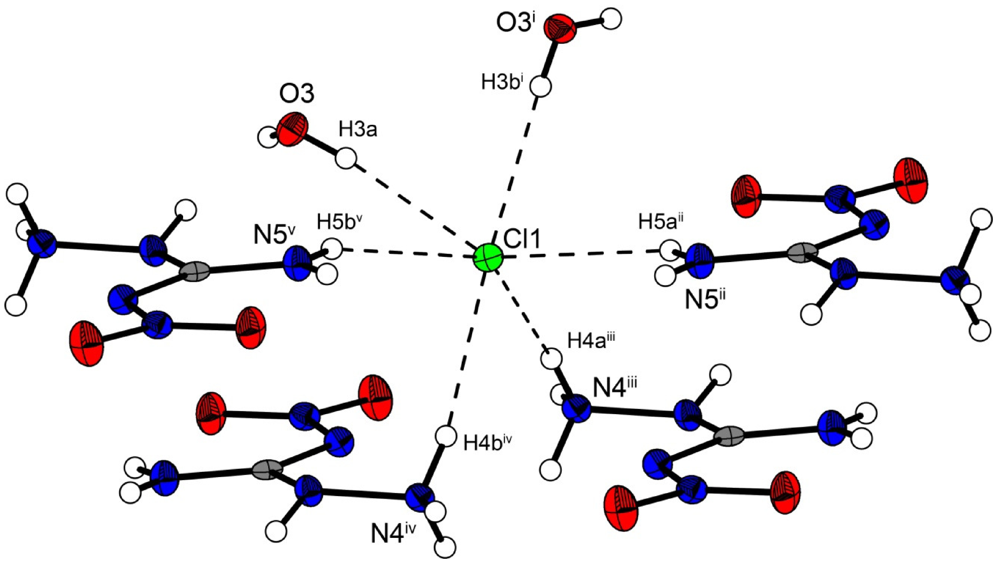
Figure 3.
Coordination sphere of one chloride anion in 2. Symmetry codes: (i) 0.5 + x, 0.5 − y, 0.5 + z; (ii) 1.5 − x, 0.5 + y, 1.5 − z; (iii) −0.5 + x, 0.5 − y, 0.5 + z; (iv) 0.5 − x, 0.5 + y, 0.5 − z; (v) −0.5 + x, 0.5 − y, −0.5 + z.
Figure 3.
Coordination sphere of one chloride anion in 2. Symmetry codes: (i) 0.5 + x, 0.5 − y, 0.5 + z; (ii) 1.5 − x, 0.5 + y, 1.5 − z; (iii) −0.5 + x, 0.5 − y, 0.5 + z; (iv) 0.5 − x, 0.5 + y, 0.5 − z; (v) −0.5 + x, 0.5 − y, −0.5 + z.
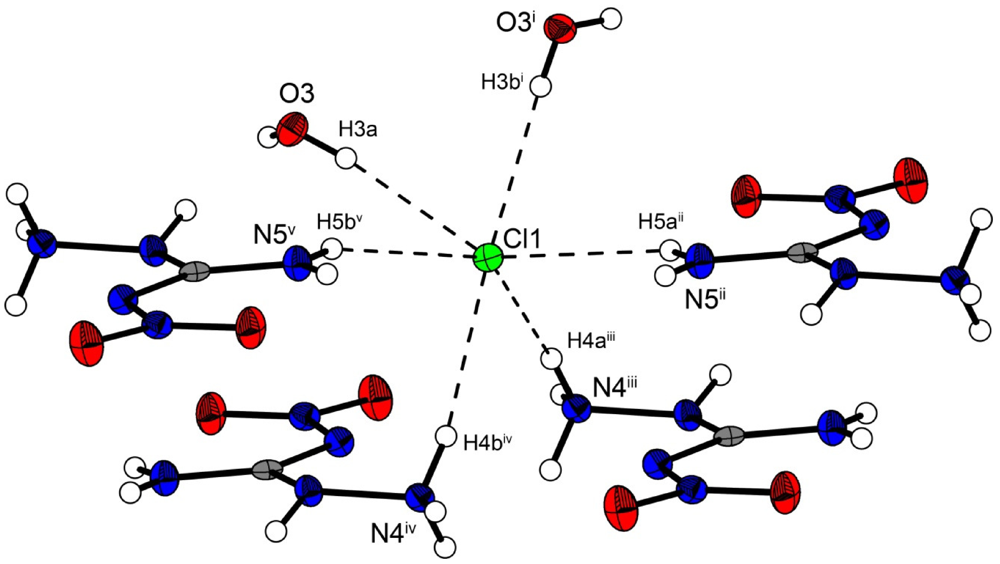

Table 3.
Selected hydrogen bonds in the structure of 2. Symmetry codes: (vi) 1.5 − x, −0.5 + y, 0.5 − z; (vii) 1 − x, −y, −z.
| D–H···A | d(D–H) | d(H···A) | d(D···A) | <(D–H···A) | <(H···A–X) |
|---|---|---|---|---|---|
| N3vi–H3vi···O2 | 0.83(2) | 2.57(2) | 3.043(2) | 117.5(15) | 150.7(5) |
| N4vii–H4cvii···O2 | 0.89(2) | 2.417(18) | 2.881(2) | 112.6(14) | 126.9(5) |

Table 4.
Selected hydrogen bonds in the structure of 2. Symmetry codes: (i) 0.5 + x, 0.5 − y, 0.5 + z; (ii) 1.5 − x, 0.5 + y, 1.5 − z; (iii) −0.5 + x, 0.5 − y, 0.5 + z; (iv) 0.5 − x, 0.5 + y, 0.5 − z; (v) −0.5 + x, 0.5 − y, −0.5 + z.
| D–H···A | d(D–H) | d(H···A) | d(D···A) | <(D–H···A) |
|---|---|---|---|---|
| N4iii–H4aiii···Cl1 | 0.88(2) | 2.42(2) | 3.279(1) | 168.8(2) |
| N4iv–H4biv···Cl1 | 0.88(2) | 2.37(2) | 3.242(1) | 159.1(2) |
| N5ii–H5aii···Cl1 | 0.85(2) | 2.59(2) | 3.291(1) | 141.2(2) |
| N5v–H5bv···Cl1 | 0.82(2) | 2.54(2) | 3.299(1) | 155.5(2) |
| O3–H3a···Cl1 | 0.78(2) | 2.34(2) | 3.084(1) | 160(2) |
| O3i–H3bi···Cl1 | 0.80(2) | 2.33(2) | 3.135(1) | 175(2) |
Figure 4 shows the molecular moiety of 3 while Figure 5 shows the same pseudo-octahedral coordination sphere of the bromide anion as we could observe in the chloride structure. The bond lengths and angles of selected hydrogen bonds are listed in Table 5, Table 6. Also in this structure the ions are held together by a three dimensional network of moderate N–H···O (d(N–O) ~2.82–3.08 Å), N–H···Cl (d(N–Br) ~3.36–3.49 Å) and O–···Cl (d(O–Cl ~3.20–3.28 Å) hydrogen bonds. The crystal packing is identical to the chloride structure 2. The N–Br and O–Br hydrogen bonds are approximately 0.15 Å longer than those in the chloride structure 2.
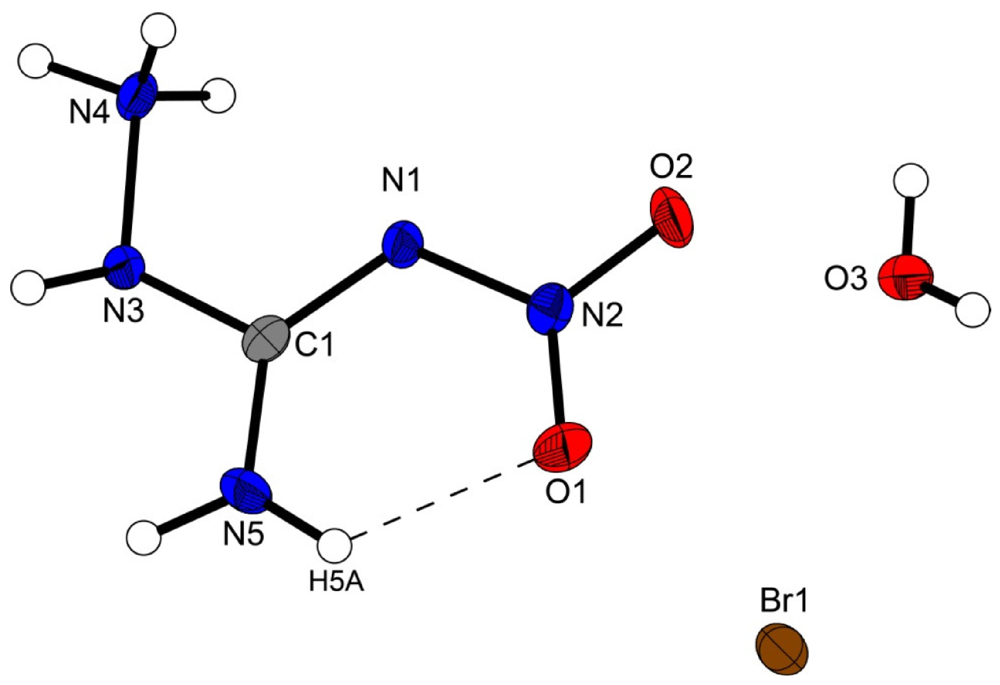
Figure 4.
Molecular structure of amino-nitro-guanidinium bromide monohydrate (3). Ellipsoids are drawn at the 50% probability level.
Figure 4.
Molecular structure of amino-nitro-guanidinium bromide monohydrate (3). Ellipsoids are drawn at the 50% probability level.
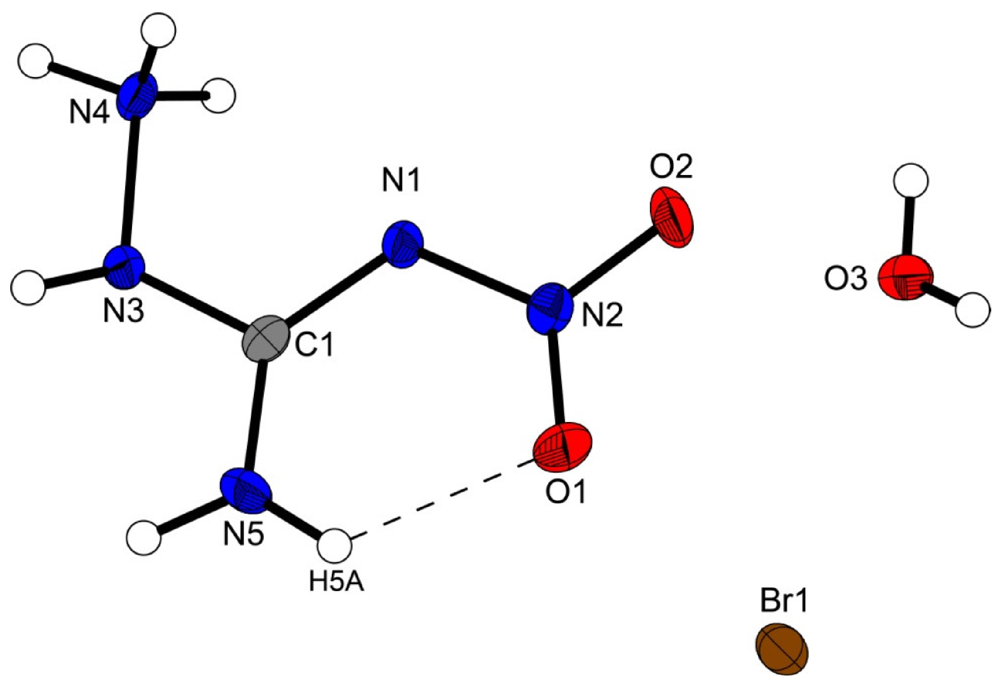
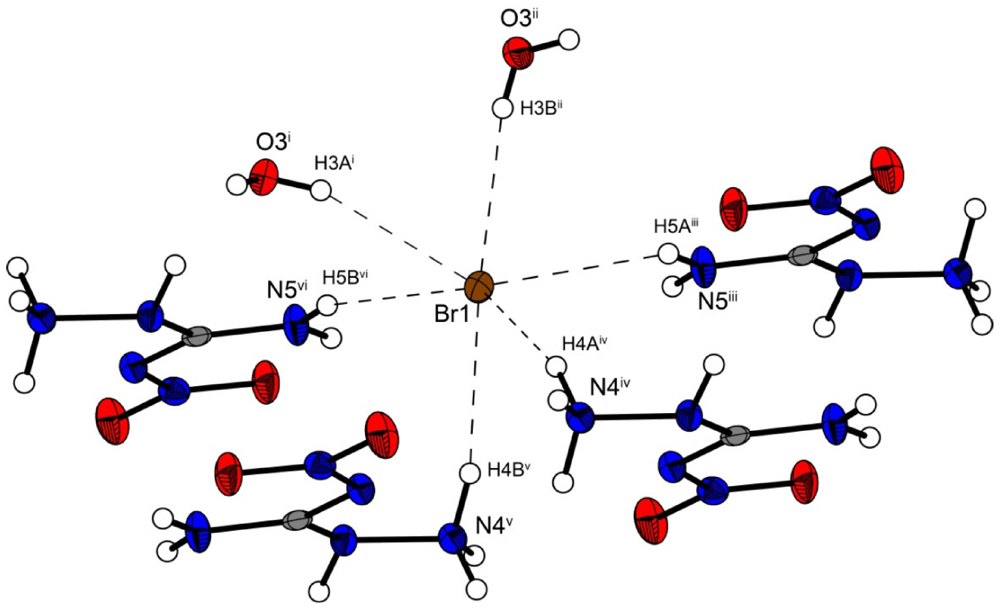
Figure 5.
Coordination sphere of one bromide anion in 3. Symmetry codes: (i) −1 + x, 0.5 − y, −0.5 + z; (ii) −1 + x, y, z; (iii) x, 0.5 − y, 0.5 + z; (iv) 1 − x, 0.5 + y, 1.5 − z; (v) x, 0.5 − y, −0.5 + z, (vi) −x, 0.5 + y, 0.5 − z.
Figure 5.
Coordination sphere of one bromide anion in 3. Symmetry codes: (i) −1 + x, 0.5 − y, −0.5 + z; (ii) −1 + x, y, z; (iii) x, 0.5 − y, 0.5 + z; (iv) 1 − x, 0.5 + y, 1.5 − z; (v) x, 0.5 − y, −0.5 + z, (vi) −x, 0.5 + y, 0.5 − z.
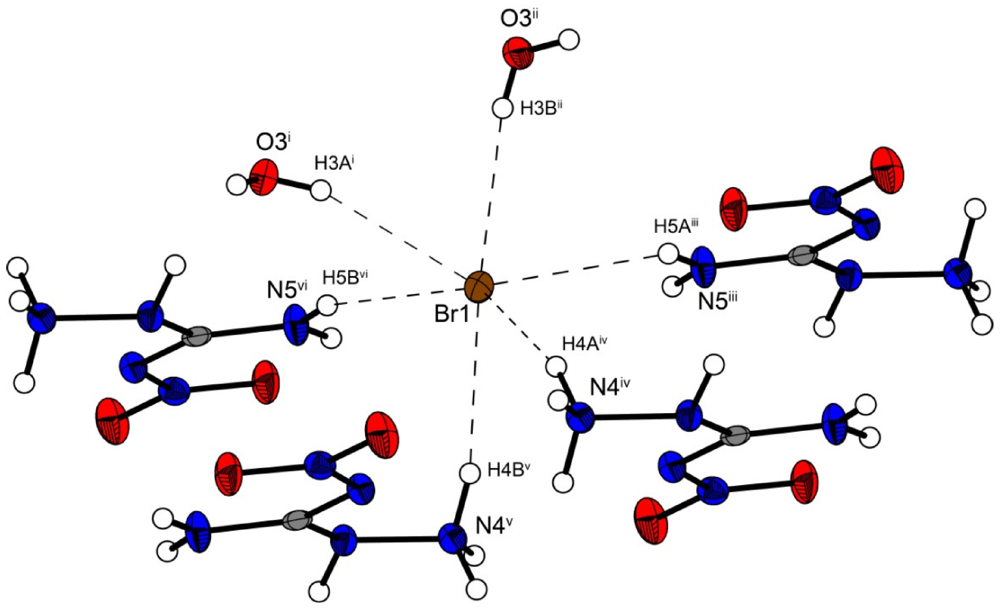

Table 5.
Selected hydrogen bonds in the structure of 3. Symmetry codes: (vi) 1 − x, 0.5 + y, 1.5 − z; (vii) 1 − x, −y, 2 − z.
| D–H···A | d(D–H) | d(H···A) | d(D···A) | <(D–H···A) | <(H···A–X) |
|---|---|---|---|---|---|
| N3vi–H3vi···O2 | 0.89(1) | 2.52(4) | 3.075(4) | 121(3) | 151.1(7) |
| N4vii–H4cvii···O2 | 0.90(4) | 2.40(4) | 2.871(4) | 113(3) | 128.0(10) |

Table 6.
Selected hydrogen bonds in the structure of 3.
| D–H···A | d(D–H) | d(H···A) | d(D···A) | <(D–H···A) |
|---|---|---|---|---|
| N4iv–H4aiv···Br1 | 0.91(4) | 2.57(4) | 3.427(3) | 156(3) |
| N4v–H4bv··· Br1 | 0.92(4) | 2.47(4) | 3.360(3) | 163(3) |
| N5iii–H5aiii··· Br1 | 0.90(1) | 2.71(3) | 3.428(3) | 138(3) |
| N5vi–H5bvi··· Br1 | 0.90(1) | 2.65(2) | 3.493(3) | 156(3) |
| O3i–H3ai··· Br1 | 0.78(4) | 2.48(5) | 3.203(3) | 156(4) |
| O3ii–H3bii··· Br1 | 0.78(4) | 2.50(4) | 3.275(3) | 172(4) |
However the iodine structure 4 crystallizes in the triclinic space group P–1 with two formula units in the unit cell. The crystal packing is different from the described chloride (2) and bromide (3) structure. Two molecular moieties of 4 connected to dimers are depicted in Figure 6. These dimers are packed in layers which are linked by the water molecules.
The iodide anions participate in four hydrogen bonds listed in Table 7. The N···I and O···I distances observed reveal ordinary values comparable to those found in 3-Cyano-anilinium iodide monohydrate [19].
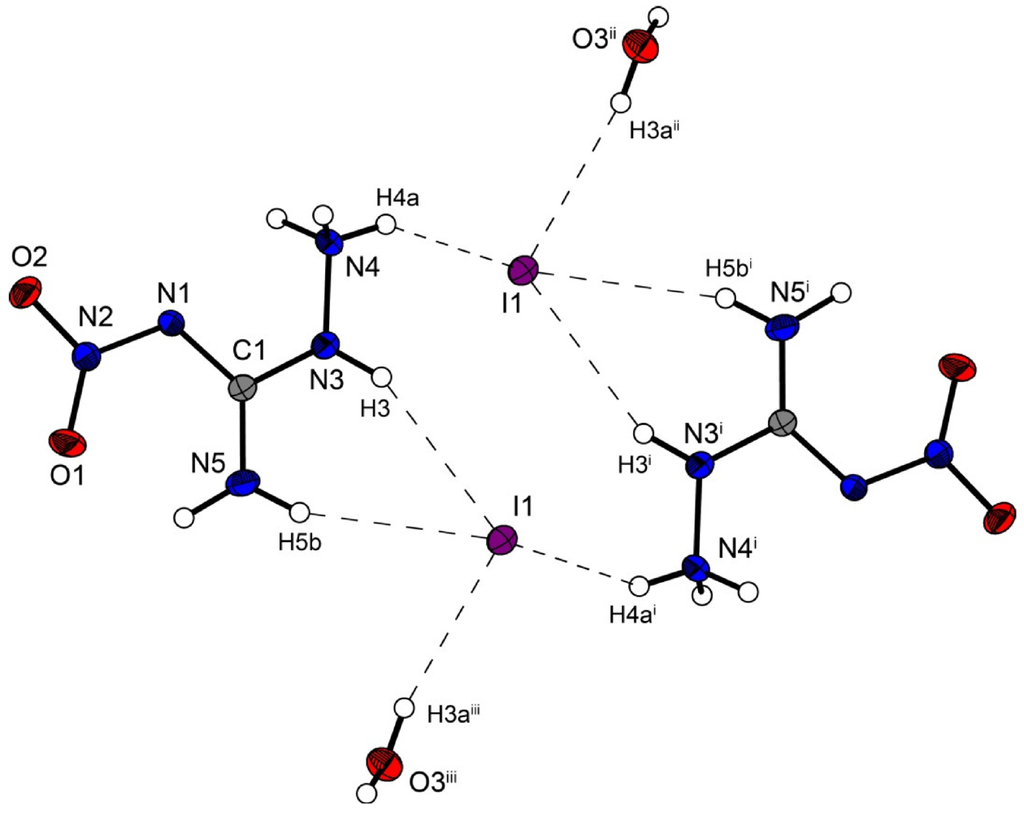
Figure 6.
Molecular structure of amino-nitro-guanidinium iodide monohydrate (4). Ellipsoids are drawn at the 50% probability level. Symmetry codes: (i) 2 − x, 1 − y, 1 − z; (ii) 1 + x, y, z; (iii) 1 − x, 1 − y, 1 − z.
Figure 6.
Molecular structure of amino-nitro-guanidinium iodide monohydrate (4). Ellipsoids are drawn at the 50% probability level. Symmetry codes: (i) 2 − x, 1 − y, 1 − z; (ii) 1 + x, y, z; (iii) 1 − x, 1 − y, 1 − z.
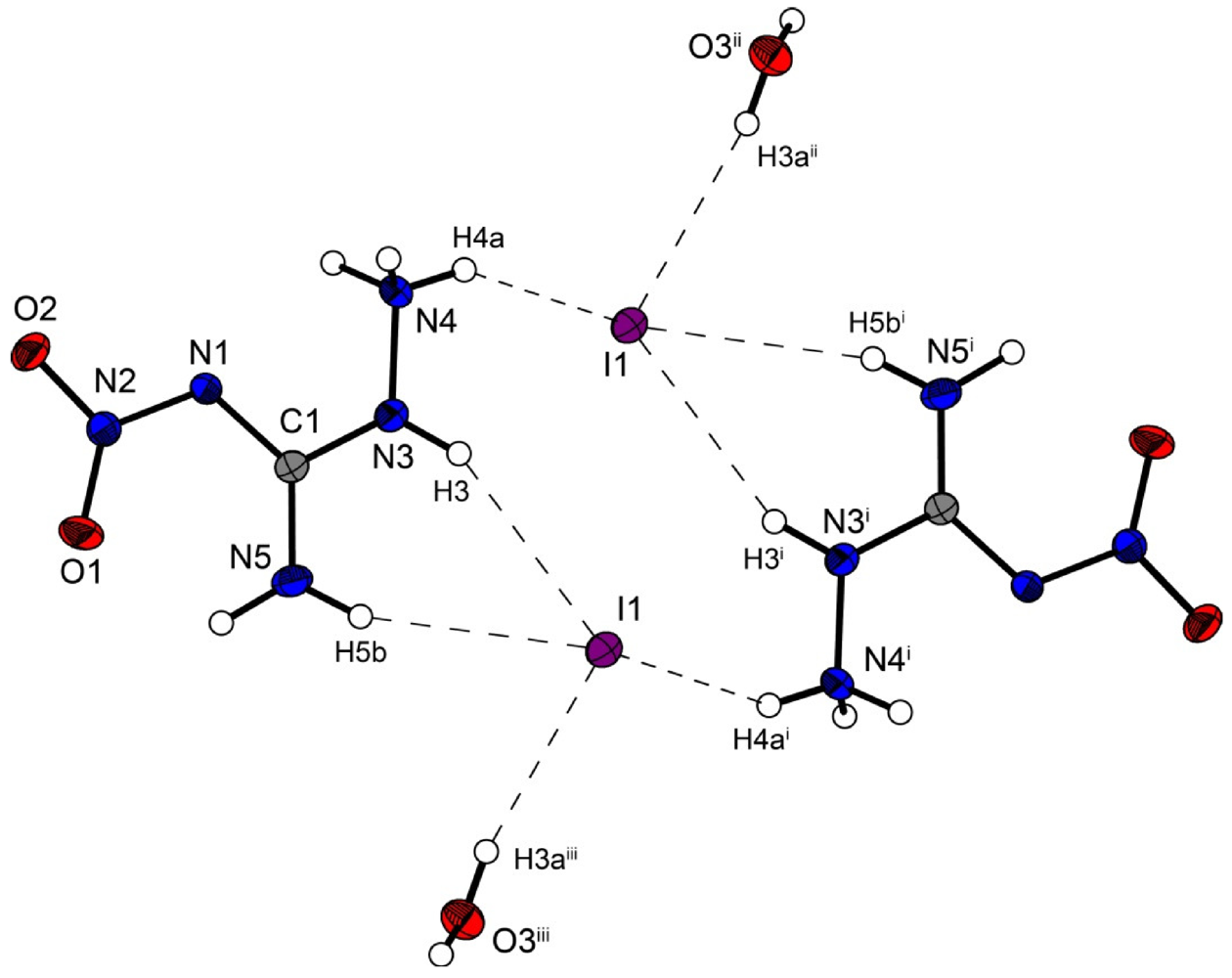

Table 7.
Selected hydrogen bonds in the structure of 4.
| D–H···A | d(D–H) | d(H···A) | d(D···A) | <(D–H···A) |
|---|---|---|---|---|
| N3i–H3i ···I1 | 0.90(3) | 2.80(3) | 3.635(2) | 156(3) |
| N4–H4a···I1 | 0.87(3) | 2.78(3) | 3.501(2) | 141(3) |
| N5i–H5bi ···I1 | 0.88(3) | 2.83(3) | 3.669(2) | 160(3) |
| O3ii–H3aii ···I1 | 0.84(4) | 2.71(4) | 3.540(2) | 169(3) |
Bis(amino-nitroguanidinium) sulfate crystallizes in the non-centrosymmetric orthorhombic space group Fdd2 with eight formula units in the unit cell. The density of 1.97 g cm−3 is slightly lower than that of the bromide structure 3 but higher than that of 2. The molecular structure of 5 is depicted in Figure 7. The sulfur atoms lie on the special position ½,0,z. The S–O bonds are uniform with a length of 1.48 Å, which is in agreement to many sulfate structures in the literature, e.g., that of potassium sulfate [20].
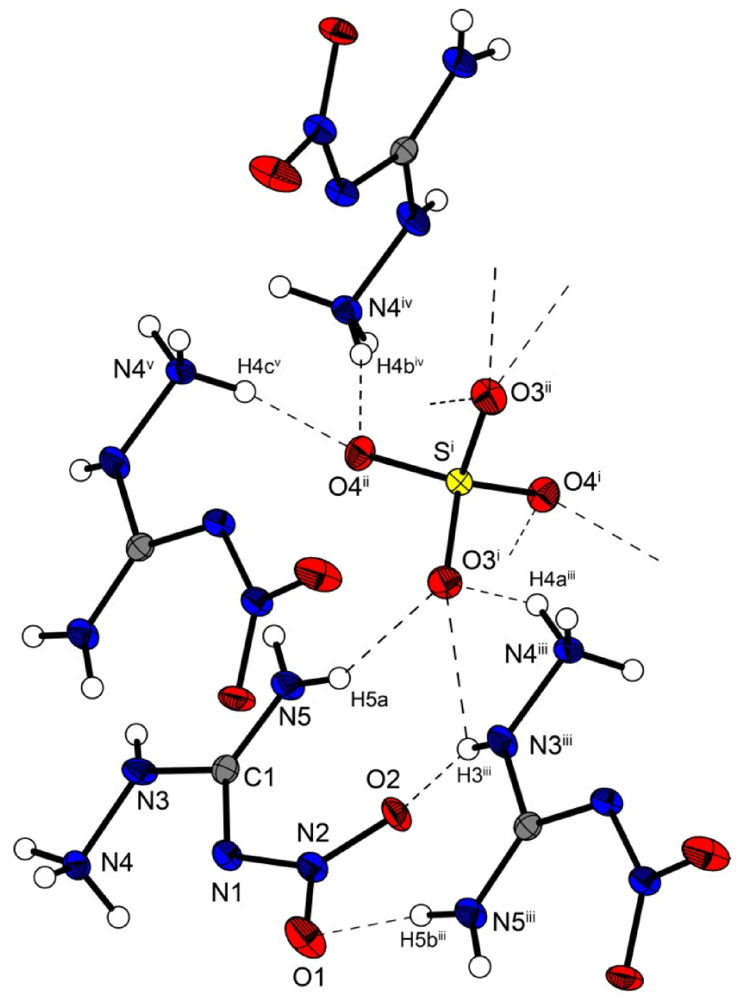
Figure 7.
Molecular structure of bis(amino-nitro-guanidinium) sulfate (5). Ellipsoids are drawn at the 50% probability level. Symmetry codes: (i) 0.75 − x, 0.25 + y, 0.75 + z; (ii) −0.25 + x, 0.25 − y, 0.75 + z; (iii) 0.25 + x, 0.25 − y, −0.75 + z; (iv) 0.25 − x, 0.25 + y, 0.25 + z; (v) −0.25 + x, 0.25 − y, −0.25 + z.
Figure 7.
Molecular structure of bis(amino-nitro-guanidinium) sulfate (5). Ellipsoids are drawn at the 50% probability level. Symmetry codes: (i) 0.75 − x, 0.25 + y, 0.75 + z; (ii) −0.25 + x, 0.25 − y, 0.75 + z; (iii) 0.25 + x, 0.25 − y, −0.75 + z; (iv) 0.25 − x, 0.25 + y, 0.25 + z; (v) −0.25 + x, 0.25 − y, −0.25 + z.
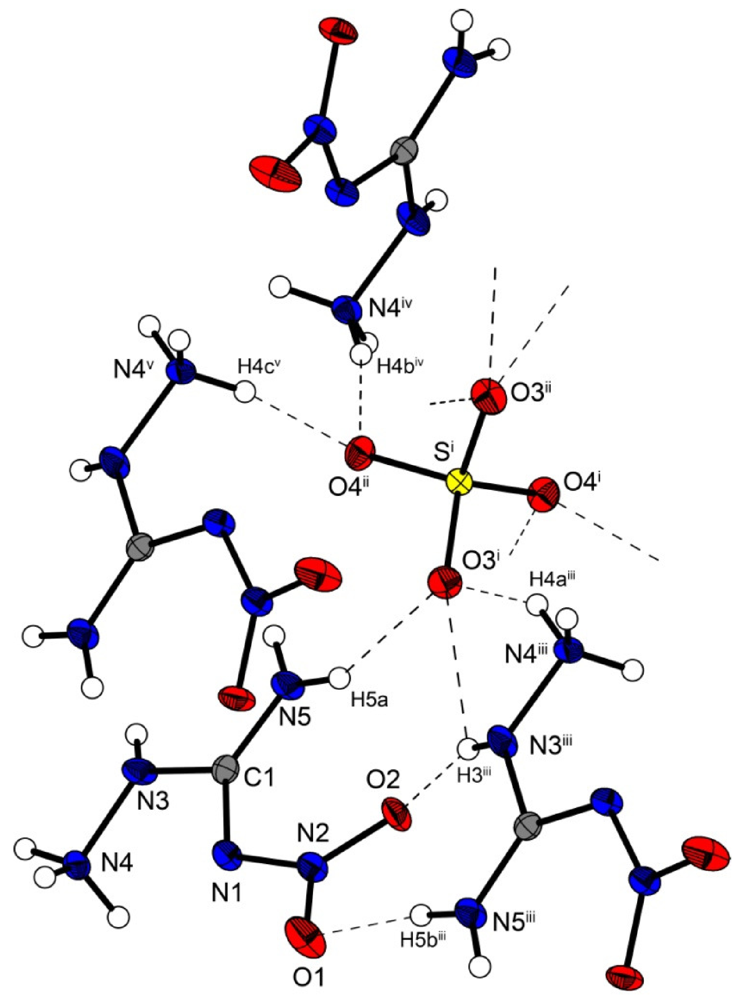
In the crystal structure of 5, the amino-nitro-guanidinium cations and the sulfate anion are held together by a three dimensional network of moderate N–H···O (d(N–O) ~2.60–3.08 Å) hydrogen bonds. Selected bond lengths and angles of the hydrogen bonds are listed in Table 8 In the crystal structure there are two layers of cations which twisted around ~60° against each other. The layers are connected via hydrogen bonds with the sulfate anions. The oxygen atoms of the sulfate anions act as bifurcated acceptors. The oxygen atom O4 bridges two cations via the hydrogen bonds N4iv–H4biv ····O4ii and N4v–H4cv ····O4ii, which form zig-zag chains along the c-axis [graph set: 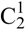 (4)]. Also the oxygen atom O3 bridges two cations via the hydrogen bonds N5–H5a····O3i, N4iii–H4aiii····O3i and N3iii–H3iii····O3i. This hydrogen bonding pattern forms a “chain of rings” along the ab-diagonal [graph set:
(4)]. Also the oxygen atom O3 bridges two cations via the hydrogen bonds N5–H5a····O3i, N4iii–H4aiii····O3i and N3iii–H3iii····O3i. This hydrogen bonding pattern forms a “chain of rings” along the ab-diagonal [graph set: 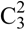 (7)
(7) 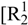 (5)]]. However this chain is also held together by the hydrogen bonds N3iii–H3iii····O2 and N5iii–H5biii····O1 of the nitrate group [graph set:
(5)]]. However this chain is also held together by the hydrogen bonds N3iii–H3iii····O2 and N5iii–H5biii····O1 of the nitrate group [graph set: 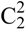 (6)
(6) 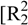 (8)]]. In the crystal structure one observes a short O···O contact (O1···O4: 2.733(2) Å), which is generated by the existing hydrogen bonds. Such a short O···O contact has also been observed in the literature [21,22].
(8)]]. In the crystal structure one observes a short O···O contact (O1···O4: 2.733(2) Å), which is generated by the existing hydrogen bonds. Such a short O···O contact has also been observed in the literature [21,22].
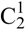 (4)]. Also the oxygen atom O3 bridges two cations via the hydrogen bonds N5–H5a····O3i, N4iii–H4aiii····O3i and N3iii–H3iii····O3i. This hydrogen bonding pattern forms a “chain of rings” along the ab-diagonal [graph set:
(4)]. Also the oxygen atom O3 bridges two cations via the hydrogen bonds N5–H5a····O3i, N4iii–H4aiii····O3i and N3iii–H3iii····O3i. This hydrogen bonding pattern forms a “chain of rings” along the ab-diagonal [graph set: 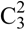 (7)
(7) 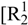 (5)]]. However this chain is also held together by the hydrogen bonds N3iii–H3iii····O2 and N5iii–H5biii····O1 of the nitrate group [graph set:
(5)]]. However this chain is also held together by the hydrogen bonds N3iii–H3iii····O2 and N5iii–H5biii····O1 of the nitrate group [graph set: 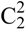 (6)
(6) 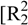 (8)]]. In the crystal structure one observes a short O···O contact (O1···O4: 2.733(2) Å), which is generated by the existing hydrogen bonds. Such a short O···O contact has also been observed in the literature [21,22].
(8)]]. In the crystal structure one observes a short O···O contact (O1···O4: 2.733(2) Å), which is generated by the existing hydrogen bonds. Such a short O···O contact has also been observed in the literature [21,22]. 
Table 8.
Selected hydrogen bonds in the structure of 5.
| D–H···A | d(D–H) | d(H···A) | d(D···A) | <(D–H···A) |
|---|---|---|---|---|
| N5–H5a····O3i | 0.89(3) | 2.17(2) | 2.979(2) | 152(2) |
| N4iii–H4aiii····O3i | 0.88(2) | 1.95(2) | 2.731(2) | 149(2) |
| N3iii–H3iii····O3i | 0.74(2) | 2.57(2) | 2.964(2) | 115(2) |
| N3iii–H3iii····O2 | 0.74(2) | 2.15(2) | 2.869(2) | 163(2) |
| N5iii–H5biii····O1 | 0.83(2) | 2.09(2) | 2.916(2) | 173(2) |
| N4iv–H4biv ····O4ii | 0.87(3) | 2.23(3) | 3.081(2) | 167(2) |
| N4v–H4cv ····O4ii | 0.89(2) | 1.81(2) | 2.688(2) | 168(2) |
2.3. Analytical Characterization
All investigated compounds were characterized using vibrational (IR and Raman) and NMR (1H, 13C) spectroscopy. Also mass spectrometry and elemental analysis were employed to identify the corresponding materials. For analytical details of the respective compounds see the Experimental Section. The decomposition temperatures were determined by differential scanning calorimetry on a Linseis PT 10 DSC [23].
2.3.1. Infrared Spectroscopy
The strongest observed absorptions in the IR spectra of the ANQ+-species are the symmetric N–H-valence vibrations of the –NH3+ functionality at 3379–3414 cm−1. In the IR spectrum of neutral ANQ, an additional strong absorption at 3551 cm−1 is observed. Here, the symmetric N–H-valence vibrations are found at higher energy since the amine group is not protonated. Further strong absorptions indicate the presence of the nitramine moiety found in all investigated compounds, which reveals the antisymmetric N–O-valence vibration of the nitro group at 1632–1639 cm−1 and the symmetric N–O-valence vibration at 1278–1280 cm−1. The IR spectrum of the sulfate salt additionally shows the strong antisymmetric S–O-valence vibration of the sulfate anion at 1065 cm−1.
2.3.2. Differential Scanning Calorimetry
Unlike unprotonated ANQ, which decomposes at 180 °C, the protonated ANQ+ salts show thermal stabilities, which are far below that value. The halides decompose at 80 °C (2, 3) and 86 °C (4) respectively, whereas the sulfate salt 5 is stable up to 144 °C.
2.3.3. Sensitivity Testing
Since 1-amino-3-nitroguanidinium salts show enhanced sensitivity towards outer stimuli such as impact and friction, the impact and friction sensitivities were determined and carried out according to STANAG 4489 [24] and 4487 [25] modified instructions [26,27] using a BAM (Bundesanstalt für Materialforschung) drophammer [28,29,30] and a BAM friction tester [28,29,30]. Regarding the sensitivities of the salts, a large difference between the halides 2–4 and the sulfate salt 5 is also observed. Whereas the halides reveal sensitivities of 25 J (impact sensitivity) and 288 N (friction sensitivity), the sulfate salt 5 is much more sensitive, having 6 J (impact sensitivity) and 120 N (friction sensitivity). This can primarily be explained by the formation of monohydrates, which is true for all halides 2–4, whereas the sulfate salt 5 crystallizes water-free. The determined values imply a classification [25] of the tested materials as “sensitive” towards both impact and friction. The higher sensitivities of the protonated species discussed herein compared to the neutral compound ANQ can be correlated to a lowered C–N bond order of the hydrazine moiety of the molecule, which appears upon protonation of the molecule as it was found during the structural investigation of neutral ANQ and its protonated species in reference 4. The same argumentation can also be applied to the thermal stabilities of the compounds, whereas a weaker C–N bond of the hydrazine moiety facilitates the loss of the hydrazine moiety and thus, decomposition of the material.
3. Experimental Section
All reagents and solvents were used as received (Sigma-Aldrich, Fluka, Acros Organics) unless stated otherwise. Melting and decomposition points were measured with a Linseis PT10 DSC using a heating rate of 5 °C min−1, which were checked with a Büchi Melting Point B-450 apparatus. 1H and 13C NMR spectra were measured with a JEOL instrument. All chemical shifts are quoted in ppm relative to TMS (1H, 13C). Infrared spectra were measured with a Perkin-Elmer Spektrum One FT-IR instrument. Raman spectra were measured with a Perkin-Elmer Spektrum 2000R NIR FT-Raman instrument equipped with a Nd:YAG laser (1064 nm). Elemental analyses were performed with a Netsch STA 429 simultaneous thermal analyzer. Friction and Impact sensitivity data were determined using a BAM drophammer and a BAM friction tester [25]. The electrostatic sensitivity tests were carried out using an Electric Spark Tester ESD 2010 EN (OZM Research) operating with the “Winspark 1.15” software package [31].
3.1. 1-Amino-3-nitroguanidinium Chloride Monohydrate (2)
1 (2.38 g, 20.0 mmol) was suspended in 2 M HCl (24 mL, 48 mmol), the mixture was heated to 60 °C and the water was removed in vacuum. The residue was recrystallized from ethanol/water to yield 2.80 g (16.1 mmol, 81%) of 2 as a white powder. Alternatively, the compound can be crystallized by slow evaporation of the water in an open dish in slightly lower yields.
DSC (5 °C min−1, °C): 80 °C (dec. 1); IR (KBr, cm−1): 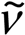 = 3549 (s), 3414 (vs), 3236 (s), 2964 (m), 2719 (m), 2675 (m), 2109 (w), 1636 (s), 1618 (s), 1561 (m), 1488 (m), 1384 (m), 1278 (s), 1223 (m), 1195 (m), 908 (w), 783 (w), 621 (w), 483 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1):
= 3549 (s), 3414 (vs), 3236 (s), 2964 (m), 2719 (m), 2675 (m), 2109 (w), 1636 (s), 1618 (s), 1561 (m), 1488 (m), 1384 (m), 1278 (s), 1223 (m), 1195 (m), 908 (w), 783 (w), 621 (w), 483 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1): 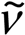 = 3335 (14), 3250 (27), 3159 (6), 2965 (16), 1625 (7), 1584 (6), 1565 (18), 1551 (5), 1478 (15), 1402 (10), 1258 (98), 1194 (16), 992 (12), 913 (26), 800 (32), 616 (30), 432 (7), 345 (15); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.44 (s, 1H, NH), 6.24 (s, 5H, –NH3+, –NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 35.0 [Cl−]; EA (CH5N5O2·H2O·HCl, 173.56) calc.: C 6.92, H 4.65, N 40.35%; found: C 6.82, H 4.51, N 39.82%; BAM drophammer: 40 J; friction tester: 288 N; ESD: 0.15 J (at grain size 100–500 µm).
= 3335 (14), 3250 (27), 3159 (6), 2965 (16), 1625 (7), 1584 (6), 1565 (18), 1551 (5), 1478 (15), 1402 (10), 1258 (98), 1194 (16), 992 (12), 913 (26), 800 (32), 616 (30), 432 (7), 345 (15); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.44 (s, 1H, NH), 6.24 (s, 5H, –NH3+, –NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 35.0 [Cl−]; EA (CH5N5O2·H2O·HCl, 173.56) calc.: C 6.92, H 4.65, N 40.35%; found: C 6.82, H 4.51, N 39.82%; BAM drophammer: 40 J; friction tester: 288 N; ESD: 0.15 J (at grain size 100–500 µm).
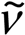 = 3549 (s), 3414 (vs), 3236 (s), 2964 (m), 2719 (m), 2675 (m), 2109 (w), 1636 (s), 1618 (s), 1561 (m), 1488 (m), 1384 (m), 1278 (s), 1223 (m), 1195 (m), 908 (w), 783 (w), 621 (w), 483 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1):
= 3549 (s), 3414 (vs), 3236 (s), 2964 (m), 2719 (m), 2675 (m), 2109 (w), 1636 (s), 1618 (s), 1561 (m), 1488 (m), 1384 (m), 1278 (s), 1223 (m), 1195 (m), 908 (w), 783 (w), 621 (w), 483 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1): 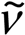 = 3335 (14), 3250 (27), 3159 (6), 2965 (16), 1625 (7), 1584 (6), 1565 (18), 1551 (5), 1478 (15), 1402 (10), 1258 (98), 1194 (16), 992 (12), 913 (26), 800 (32), 616 (30), 432 (7), 345 (15); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.44 (s, 1H, NH), 6.24 (s, 5H, –NH3+, –NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 35.0 [Cl−]; EA (CH5N5O2·H2O·HCl, 173.56) calc.: C 6.92, H 4.65, N 40.35%; found: C 6.82, H 4.51, N 39.82%; BAM drophammer: 40 J; friction tester: 288 N; ESD: 0.15 J (at grain size 100–500 µm).
= 3335 (14), 3250 (27), 3159 (6), 2965 (16), 1625 (7), 1584 (6), 1565 (18), 1551 (5), 1478 (15), 1402 (10), 1258 (98), 1194 (16), 992 (12), 913 (26), 800 (32), 616 (30), 432 (7), 345 (15); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.44 (s, 1H, NH), 6.24 (s, 5H, –NH3+, –NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 35.0 [Cl−]; EA (CH5N5O2·H2O·HCl, 173.56) calc.: C 6.92, H 4.65, N 40.35%; found: C 6.82, H 4.51, N 39.82%; BAM drophammer: 40 J; friction tester: 288 N; ESD: 0.15 J (at grain size 100–500 µm).3.2. 1-Amino-3-nitroguanidinium Bromide Monohydrate (3)
1 (2.38 g, 20.0 mmol) was suspended in 20 mL of water and an aqueous HBr-solution (48% (w/v), 5.0 mL, 44.0 mmol) was added. The mixture was heated to 60 °C until a clear solution is obtained and the water was slowly evaporated in an open dish. 3 crystallizes in colorless blocks in 86% yield (3.73 g, 17.1 mmol).
DSC (5 °C min−1, °C): 80 °C (dec.); IR (KBr, cm−1): 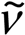 = 3379 (vs), 2996 (s), 2681 (m), 2375 (w), 2086 (w), 1633 (s), 1586 (s), 1557 (m), 1538 (m), 1490 (m), 1475 (m), 1399 (m), 1383 (m), 1323 (m), 1279 (s), 1223 (s), 1190 (m), 1155 (m), 1121 (m), 1096 (m), 1037 (w), 908 (w), 783 (w), 662 (w), 581 (w), 532 (w), 473 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1):
= 3379 (vs), 2996 (s), 2681 (m), 2375 (w), 2086 (w), 1633 (s), 1586 (s), 1557 (m), 1538 (m), 1490 (m), 1475 (m), 1399 (m), 1383 (m), 1323 (m), 1279 (s), 1223 (s), 1190 (m), 1155 (m), 1121 (m), 1096 (m), 1037 (w), 908 (w), 783 (w), 662 (w), 581 (w), 532 (w), 473 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1): 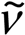 = 3318 (1), 3254 (8), 2974 (11), 1609 (8), 1580 (3), 1567 (2), 1556 (15), 1533 (2), 1485 (10), 1406 (7), 1255 (100), 1186 (13), 1155 (4), 1094 (5), 994 (15), 913 (24), 800 (45), 720 (2), 614 (37), 587 (2), 519 (2), 467 (1), 431 (8); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 9.64 (s, 1H, -NHH), 9.17 (s, 1H, -NHH), 6.31 (s, 4H, NHNH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.8 (C(NNO2)(N2H3)(NH2)), 161.0 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 78.9 [79Br−], 80.9 [81Br−]; EA (CH5N5O2·H2O·HBr, 218.01) calc.: C 5.51, H 3.70, N 32.12%; found: C 5.43, H 3.57, N 31.76 %; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).
= 3318 (1), 3254 (8), 2974 (11), 1609 (8), 1580 (3), 1567 (2), 1556 (15), 1533 (2), 1485 (10), 1406 (7), 1255 (100), 1186 (13), 1155 (4), 1094 (5), 994 (15), 913 (24), 800 (45), 720 (2), 614 (37), 587 (2), 519 (2), 467 (1), 431 (8); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 9.64 (s, 1H, -NHH), 9.17 (s, 1H, -NHH), 6.31 (s, 4H, NHNH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.8 (C(NNO2)(N2H3)(NH2)), 161.0 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 78.9 [79Br−], 80.9 [81Br−]; EA (CH5N5O2·H2O·HBr, 218.01) calc.: C 5.51, H 3.70, N 32.12%; found: C 5.43, H 3.57, N 31.76 %; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).
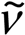 = 3379 (vs), 2996 (s), 2681 (m), 2375 (w), 2086 (w), 1633 (s), 1586 (s), 1557 (m), 1538 (m), 1490 (m), 1475 (m), 1399 (m), 1383 (m), 1323 (m), 1279 (s), 1223 (s), 1190 (m), 1155 (m), 1121 (m), 1096 (m), 1037 (w), 908 (w), 783 (w), 662 (w), 581 (w), 532 (w), 473 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1):
= 3379 (vs), 2996 (s), 2681 (m), 2375 (w), 2086 (w), 1633 (s), 1586 (s), 1557 (m), 1538 (m), 1490 (m), 1475 (m), 1399 (m), 1383 (m), 1323 (m), 1279 (s), 1223 (s), 1190 (m), 1155 (m), 1121 (m), 1096 (m), 1037 (w), 908 (w), 783 (w), 662 (w), 581 (w), 532 (w), 473 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1): 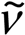 = 3318 (1), 3254 (8), 2974 (11), 1609 (8), 1580 (3), 1567 (2), 1556 (15), 1533 (2), 1485 (10), 1406 (7), 1255 (100), 1186 (13), 1155 (4), 1094 (5), 994 (15), 913 (24), 800 (45), 720 (2), 614 (37), 587 (2), 519 (2), 467 (1), 431 (8); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 9.64 (s, 1H, -NHH), 9.17 (s, 1H, -NHH), 6.31 (s, 4H, NHNH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.8 (C(NNO2)(N2H3)(NH2)), 161.0 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 78.9 [79Br−], 80.9 [81Br−]; EA (CH5N5O2·H2O·HBr, 218.01) calc.: C 5.51, H 3.70, N 32.12%; found: C 5.43, H 3.57, N 31.76 %; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).
= 3318 (1), 3254 (8), 2974 (11), 1609 (8), 1580 (3), 1567 (2), 1556 (15), 1533 (2), 1485 (10), 1406 (7), 1255 (100), 1186 (13), 1155 (4), 1094 (5), 994 (15), 913 (24), 800 (45), 720 (2), 614 (37), 587 (2), 519 (2), 467 (1), 431 (8); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 9.64 (s, 1H, -NHH), 9.17 (s, 1H, -NHH), 6.31 (s, 4H, NHNH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.8 (C(NNO2)(N2H3)(NH2)), 161.0 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 78.9 [79Br−], 80.9 [81Br−]; EA (CH5N5O2·H2O·HBr, 218.01) calc.: C 5.51, H 3.70, N 32.12%; found: C 5.43, H 3.57, N 31.76 %; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).3.3. 1-Amino-3-nitroguanidinium Iodide Monohydrate (4)
1 (2.38 g, 20.0 mmol) was suspended in 20 mL of water and an aqueous HI-solution (57% (w/v), 5.8 mL, 44.0 mmol) was added. The mixture was heated to 60 °C until a clear solution is obtained and the water was slowly evaporated in an open dish. 4 crystallizes in large colorless blocks in 86% yield (3.73 g, 17.1 mmol), which tend to turn yellowish after a few days upon the release of molecular I2.
DSC (5 °C min−1, °C): 86 °C (dec.); IR (KBr, cm−1): 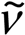 = 3384 (vs), 3278 (s), 3224 (s), 2927 (m), 2732 (m), 2371 (w), 1632 (vs), 1587 (s), 1475 (s), 1458 (m), 1369 (m), 1278 (vs), 1122 (m), 1099 (m), 1037 (w), 1000 (w), 913 (w), 835 (w), 783 (w), 701 (w), 618 (w), 581 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1):
= 3384 (vs), 3278 (s), 3224 (s), 2927 (m), 2732 (m), 2371 (w), 1632 (vs), 1587 (s), 1475 (s), 1458 (m), 1369 (m), 1278 (vs), 1122 (m), 1099 (m), 1037 (w), 1000 (w), 913 (w), 835 (w), 783 (w), 701 (w), 618 (w), 581 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1): 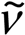 = 3393 (2), 3235 (7), 2978 (2), 1629 (7), 1604 (1), 1563 (44), 1531 (8), 1489 (19), 1456 (2), 1383 (24), 1264 (100), 1162 (21), 1110 (3), 999 (21), 921 (38), 797 (69), 699 (5), 620 (37), 571 (2), 433 (32); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.30 (s, 3H, NH, NH2), 6.47 (s, 3H, -NH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.3 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 126.9 [I−]; EA (CH5N5O2·H2O·HI, 265.01) calc.: C 4.53, H 3.04, N 26.43 %; found: C 4.87, H 2.92, N 26.46%; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).
= 3393 (2), 3235 (7), 2978 (2), 1629 (7), 1604 (1), 1563 (44), 1531 (8), 1489 (19), 1456 (2), 1383 (24), 1264 (100), 1162 (21), 1110 (3), 999 (21), 921 (38), 797 (69), 699 (5), 620 (37), 571 (2), 433 (32); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.30 (s, 3H, NH, NH2), 6.47 (s, 3H, -NH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.3 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 126.9 [I−]; EA (CH5N5O2·H2O·HI, 265.01) calc.: C 4.53, H 3.04, N 26.43 %; found: C 4.87, H 2.92, N 26.46%; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).
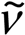 = 3384 (vs), 3278 (s), 3224 (s), 2927 (m), 2732 (m), 2371 (w), 1632 (vs), 1587 (s), 1475 (s), 1458 (m), 1369 (m), 1278 (vs), 1122 (m), 1099 (m), 1037 (w), 1000 (w), 913 (w), 835 (w), 783 (w), 701 (w), 618 (w), 581 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1):
= 3384 (vs), 3278 (s), 3224 (s), 2927 (m), 2732 (m), 2371 (w), 1632 (vs), 1587 (s), 1475 (s), 1458 (m), 1369 (m), 1278 (vs), 1122 (m), 1099 (m), 1037 (w), 1000 (w), 913 (w), 835 (w), 783 (w), 701 (w), 618 (w), 581 (w); Raman (1064 nm, 300 mW, 25 °C, cm−1): 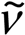 = 3393 (2), 3235 (7), 2978 (2), 1629 (7), 1604 (1), 1563 (44), 1531 (8), 1489 (19), 1456 (2), 1383 (24), 1264 (100), 1162 (21), 1110 (3), 999 (21), 921 (38), 797 (69), 699 (5), 620 (37), 571 (2), 433 (32); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.30 (s, 3H, NH, NH2), 6.47 (s, 3H, -NH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.3 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 126.9 [I−]; EA (CH5N5O2·H2O·HI, 265.01) calc.: C 4.53, H 3.04, N 26.43 %; found: C 4.87, H 2.92, N 26.46%; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).
= 3393 (2), 3235 (7), 2978 (2), 1629 (7), 1604 (1), 1563 (44), 1531 (8), 1489 (19), 1456 (2), 1383 (24), 1264 (100), 1162 (21), 1110 (3), 999 (21), 921 (38), 797 (69), 699 (5), 620 (37), 571 (2), 433 (32); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.30 (s, 3H, NH, NH2), 6.47 (s, 3H, -NH3+); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 159.3 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB−): 126.9 [I−]; EA (CH5N5O2·H2O·HI, 265.01) calc.: C 4.53, H 3.04, N 26.43 %; found: C 4.87, H 2.92, N 26.46%; BAM drophammer: 25 J; friction tester: 288 N; ESD: 0.20 J (at grain size 500–1000 µm).3.4. Bis(1-amino-3-nitroguanidinium) Sulfate (5)
1 (2.38 g, 20.0 mmol) was suspended in 20 mL of water and 1 M H2SO4 (11 mL, 11 mmol) was added. The mixture was heated to 60 °C and the water was removed in vacuum. The residue was again recrystallized from water to yield 2.93 g (8.71 mmol, 87%) of 5 as colorless crystals.
DSC (5 °C min−1, °C): 144 °C (dec.); IR (KBr, cm−1): 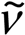 = 3401 (s), 3307 (vs), 3153 (s), 2923 (m), 2647 (m), 2060 (w), 1684 (m), 1659 (m), 1639 (m), 1566 (m), 1485 (m), 1459 (m), 1384 (m), 1280 (s), 1256 (s), 1161 (m), 1124 (m), 1091 (m), 1065 (s), 1009 (m), 976 (m), 924 (w), 788 (w), 728 (w), 647 (w), 614 (m), 457 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1):
= 3401 (s), 3307 (vs), 3153 (s), 2923 (m), 2647 (m), 2060 (w), 1684 (m), 1659 (m), 1639 (m), 1566 (m), 1485 (m), 1459 (m), 1384 (m), 1280 (s), 1256 (s), 1161 (m), 1124 (m), 1091 (m), 1065 (s), 1009 (m), 976 (m), 924 (w), 788 (w), 728 (w), 647 (w), 614 (m), 457 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1): 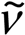 = 3238 (4), 3165 (4), 1643 (10), 1580 (9), 1543 (8), 1485 (11), 1414 (5), 1280 (23), 1161 (19), 1131 (13), 1091 (7), 982 (100), 935 (35), 802 (31), 635 (28), 604 (10), 476 (11), 458 (15), 435 (12), 356 (30), 248 (14), 196 (30), 178 (29), 136 (44), 116 (43); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.17 (s, NH), 7.64 (s, 5H, -NH3+, -NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB–): 96.9 [HSO4–]; EA (C2H12N10O8S, 336.24) calc.: C 7.14, H 3.60, N 41.66%; found: C 7.47, H 3.44, N 41.35%; BAM drophammer: 6 J; friction tester: 120 N; ESD: 0.30 J (at grain size 500–1000 µm).
= 3238 (4), 3165 (4), 1643 (10), 1580 (9), 1543 (8), 1485 (11), 1414 (5), 1280 (23), 1161 (19), 1131 (13), 1091 (7), 982 (100), 935 (35), 802 (31), 635 (28), 604 (10), 476 (11), 458 (15), 435 (12), 356 (30), 248 (14), 196 (30), 178 (29), 136 (44), 116 (43); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.17 (s, NH), 7.64 (s, 5H, -NH3+, -NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB–): 96.9 [HSO4–]; EA (C2H12N10O8S, 336.24) calc.: C 7.14, H 3.60, N 41.66%; found: C 7.47, H 3.44, N 41.35%; BAM drophammer: 6 J; friction tester: 120 N; ESD: 0.30 J (at grain size 500–1000 µm).
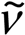 = 3401 (s), 3307 (vs), 3153 (s), 2923 (m), 2647 (m), 2060 (w), 1684 (m), 1659 (m), 1639 (m), 1566 (m), 1485 (m), 1459 (m), 1384 (m), 1280 (s), 1256 (s), 1161 (m), 1124 (m), 1091 (m), 1065 (s), 1009 (m), 976 (m), 924 (w), 788 (w), 728 (w), 647 (w), 614 (m), 457 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1):
= 3401 (s), 3307 (vs), 3153 (s), 2923 (m), 2647 (m), 2060 (w), 1684 (m), 1659 (m), 1639 (m), 1566 (m), 1485 (m), 1459 (m), 1384 (m), 1280 (s), 1256 (s), 1161 (m), 1124 (m), 1091 (m), 1065 (s), 1009 (m), 976 (m), 924 (w), 788 (w), 728 (w), 647 (w), 614 (m), 457 (w); Raman (1064 nm, 200 mW, 25 °C, cm−1): 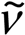 = 3238 (4), 3165 (4), 1643 (10), 1580 (9), 1543 (8), 1485 (11), 1414 (5), 1280 (23), 1161 (19), 1131 (13), 1091 (7), 982 (100), 935 (35), 802 (31), 635 (28), 604 (10), 476 (11), 458 (15), 435 (12), 356 (30), 248 (14), 196 (30), 178 (29), 136 (44), 116 (43); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.17 (s, NH), 7.64 (s, 5H, -NH3+, -NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB–): 96.9 [HSO4–]; EA (C2H12N10O8S, 336.24) calc.: C 7.14, H 3.60, N 41.66%; found: C 7.47, H 3.44, N 41.35%; BAM drophammer: 6 J; friction tester: 120 N; ESD: 0.30 J (at grain size 500–1000 µm).
= 3238 (4), 3165 (4), 1643 (10), 1580 (9), 1543 (8), 1485 (11), 1414 (5), 1280 (23), 1161 (19), 1131 (13), 1091 (7), 982 (100), 935 (35), 802 (31), 635 (28), 604 (10), 476 (11), 458 (15), 435 (12), 356 (30), 248 (14), 196 (30), 178 (29), 136 (44), 116 (43); 1H NMR (DMSO-d6, 25 °C, ppm) δ: 8.17 (s, NH), 7.64 (s, 5H, -NH3+, -NH2); 13C NMR (DMSO-d6, 25 °C, ppm) δ: 160.5 (C(NNO2)(N2H4+)(NH2)); m/z (FAB+): 120.1 [CH6N5O2+]; m/z (FAB–): 96.9 [HSO4–]; EA (C2H12N10O8S, 336.24) calc.: C 7.14, H 3.60, N 41.66%; found: C 7.47, H 3.44, N 41.35%; BAM drophammer: 6 J; friction tester: 120 N; ESD: 0.30 J (at grain size 500–1000 µm).4. Conclusions
From this experimental study the following conclusions can be drawn:
- 1-Amino-3-nitro-guanidine (1) can be protonated with strong acids (pKs < 1). Protonation was accomplished with hydrochloric, hydrobromic, hydroiodic and sulfuric acid. No protonation could be observed with aqueous hydrofluoric acid (40%).
- The structures of the crystalline state of amino-nitroguanidinium chloride monohydrate (2), amino-nitroguanidinium bromide monohydrate (3), amino-nitroguanidinium iodide monohydrate (4) and bis(amino-nitroguanidinium) sulfate (5) were determined by low temperature X-ray diffraction. The salts crystallize in common space group (2: P21/n, 3:P21/c, 4: P–1, 5: Fdd2).
- The good solubility and availability of 2–5 allows the metathesis reaction with different silver and barium salts under precipitation of low-soluble AgCl, AgBr, AgI or BaSO4 for the synthesis of a variety of amino-nitroguanidinium salts as exemplarily shown for 1-amino-3-nitroguanidinium dinitramide [4].
Acknowledgments
Financial support of this work by the Ludwig-Maximilian University of Munich (LMU), the U.S. Army Research Laboratory (ARL), the Armament Research, Development and Engineering Center (ARDEC), the Strategic Environmental Research and Development Program (SERDP) and the Office of Naval Research (ONR).
Conflict of Interest
The authors declare no “conflict of interest”.
References and Notes
- McKay, A.F. Nitroguanidines. Chem. Rev. 1952, 51, 301–346. [Google Scholar] [CrossRef]
- Klapötke, T.M. Chemistry of High-Energy Materials, 1st ed; Walter de Gruyter GmbH & Co. KG: Berlin, Germany; New York, NY, USA, 2011. [Google Scholar]
- Henry, R.A.; Lewis, H.D.; Smith, G.B.L. Hydrazinolysis of nitroguanidine and alkylnitroguanidines. J. Am. Chem. Soc. 1950, 72, 2015–2018. [Google Scholar]
- Phillips, R.; Williams, J.F. Nitroaminoguanidine. J. Am. Chem. Soc. 1928, 50, 2465–2470. [Google Scholar] [CrossRef]
- Castillo-Meléndez, J.A.; Golding, B.T. Optimization of the synthesis of guanidines from amines via nitroguanidines using 3,5-dimethyl-N-nitro-1H-pyrazole-1-carboxamidine. Synthesis 2004, 10, 1655–1663. [Google Scholar] [CrossRef]
- Fischer, N.; Klapötke, T.M.; Stierstorfer, J. 1-Amino-3-nitroguanidine (ANQ) in high performing energetic materials. Z. Naturforsch. B 2012, in press.. [Google Scholar]
- CrysAlisPro, Agilent Technologies, Version 1.171.35.11, Agilent Technologies: Yarnton, UK, 2011.
- Sheldrick, G.M. SHELXS-97, Program for Crystal Structure Solution; University of Göttingen: Göttingen, Germany, 1997. [Google Scholar]
- Altomare, A.; Cascarano, G.; Giacovazzo, C.; Guagliardi, A. Completion and refinement of crystal structures with SIR92. Appl. Cryst. 1993, 26, 343–350. [Google Scholar] [CrossRef]
- Sheldrick, G.M. SHELXL-97, Program for the Refinement of Crystal Structures; University of Göttingen: Göttingen, Germany, 1994. [Google Scholar]
- Spek, A.L. Platon, A Multipurpose Crystallographic Tool; Utrecht University: Utrecht, The Netherlands, 1999. [Google Scholar]
- Farrugia, L. WinGX suite for small-molecule single crystal crystallography. J. Appl. Cryst. 1999, 32, 837–838. [Google Scholar] [CrossRef]
- Empirical Absorption Correction Using Spherical Harmonics, Implemented in Scale3 Abspack Scaling Algorithm, Version 171.33.41, CrysAlisPro Oxford Diffraction Ltd.: Yarnton, UK, 2009.
- Crystallographic data for the structure(s) have been deposited with the Cambridge Crystallographic Data Centre. Copies of the data can be obtained free of charge on application to The Director, CCDC, 12 Union Road, Cambridge CB2 1EZ, UK (Fax: int.code_(1223)336-033; e-mail for inquiry: fileserv@ccdc.cam.ac.uk; e-mail for deposition: deposit-@ccdc.cam.ac.uk).
- Bernstein, J.; Davis, R.E.; Shimoni, L.; Chang, N.-L. Patterns in hydrogen bonding: Functionality and graph set analysis in crystals. Angew. Chem. Int. Ed. 1995, 34, 1555–1573. [Google Scholar] [CrossRef]
- Kollman, P.A.; Allen, L.C. Theory of hydrogen bond. Chem. Rev. 1972, 72, 283–303. [Google Scholar] [CrossRef]
- Arunan, E.; Desiraju, G.R.; Klein, R.A.; Sadlej, J.; Scheiner, S.; Alkorta, I.; Clary, D.C.; Crabtree, R.H.; Dannenberg, J.J.; Hobza, P.; et al. Definition of the hydrogen bond (IUPAC Recommendations 2011). Pure Appl. Chem. 2011, 83, 1637–1641. [Google Scholar] [CrossRef]
- Desiraju, G.R.; Steiner, T. The Weak Hydrogen Bond in Structural Chemistry and Biology; Oxford University Press: Oxford, UK, 1999. [Google Scholar]
- Dai, J.; Chen, X.-Y. 3-Cyanoanilinium iodide monohydrate. Acta Cryst. 2010, E66, o3295. [Google Scholar]
- McGinnety, J.A. Redetermination of the structures of potassium sulfate and potassium chromate. Effect of electrostatic crystal forces upon observed bond lengths. Acta Cryst. 1972, B28, 2845–2852. [Google Scholar]
- Ramesh, P.; Subbiahpandi, A.; Manikannan, R.; Muthusubramanian, S.; Ponnuswamy, M.N. 3-Cyclohexylsulfanyl-2-(4-methylphenyl)-5,7-dinitro-1H-indole. Acta Cryst. 2008, E64, o1890. [Google Scholar]
- Yu, Z.; Kuroda-Sowa, T.; Nabei, A.; Maekawa, M.; Okubo, T. {6,6'-Dimethoxy-2,2´-[naphthalene-2,3-diylbis(nitrilomethylidyne)]diphenolato}thiocyanatocobalt(III) diethyl ether dichloromethane solvate. Acta Cryst. 2009, E65, m257–m258. [Google Scholar]
- Linseis. Available online: http://www.linseis.com (accessed on 26 March 2012).
- NATO, Standardization Agreement (STANAG) on Explosives, Impact Sensitivity Tests, No. 4489, 1st ed; NATO: Brussels, Belgium, 1999.
- NATO, Standardization Agreement (STANAG) on Explosive, Friction Sensitivity Tests, No. 4487, 1st ed; NATO: Brussels, Belgium, 2002.
- WIWEB-Standardarbeitsanweisung, Nr. 4-5.1.02, Ermittlung der Explosionsgefährlichkeit (Impact Sensitivity Tests with a Drophammer); WIWEB: Erding, Germany, 2002.
- WIWEB-Standardarbeitsanweisung, Nr. 4-5.1.03, Ermittlung der Explosionsgefährlichkeit oder der Reibeempfindlichkeit mit dem Reibeapparat; WIWEB: Erding, Germany, 2002.
- Bundesanstalt für Materialforschung und -prüfung. Available online: http://www.bam.de (accessed on 26 March 2012).
- Reichel & Partner GmbH. Available online: http://www.reichel-partner.de (accessed on 26 March 2012).
- Impact: insensitive > 40 J, less sensitive > 35 J, sensitive > 4 J, very sensitive < 3 J. Friction: insensitive > 360 N, less sensitive = 360 N, sensitive < 360 N and > 80 N, very sensitive < 80 N, extremely sensitive < 10 N. According to the UN Recommendations on the Transport of Dangerous Goods, (+) indicates not safe for transport.
- ESD 2008A small-scale electrostatic spark sensitivity test. Available online: http://ozm.cz/en/sensitivity-tests/esd-2008a-small-scale-electrostatic-spark-sensitivity-test/ (accessed on 26 March 2012).
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
