Genome-Wide Identification, Characterization, and Expression Analysis under Abiotic Stresses of the UBP Gene Family in Rice (Oryza sativa L.)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Identification of the OsUBP Family Genes
2.2. Phylogenetic Relationship, Protein Domain, and Gene Structure Analysis
2.3. Chromosomal Location, Synteny, and Ka/Ks Analysis
2.4. Cis-Element, GO Annotation, and Protein Interaction Analysis
2.5. Expression Analysis
2.6. Real-Time PCR Analysis
3. Results
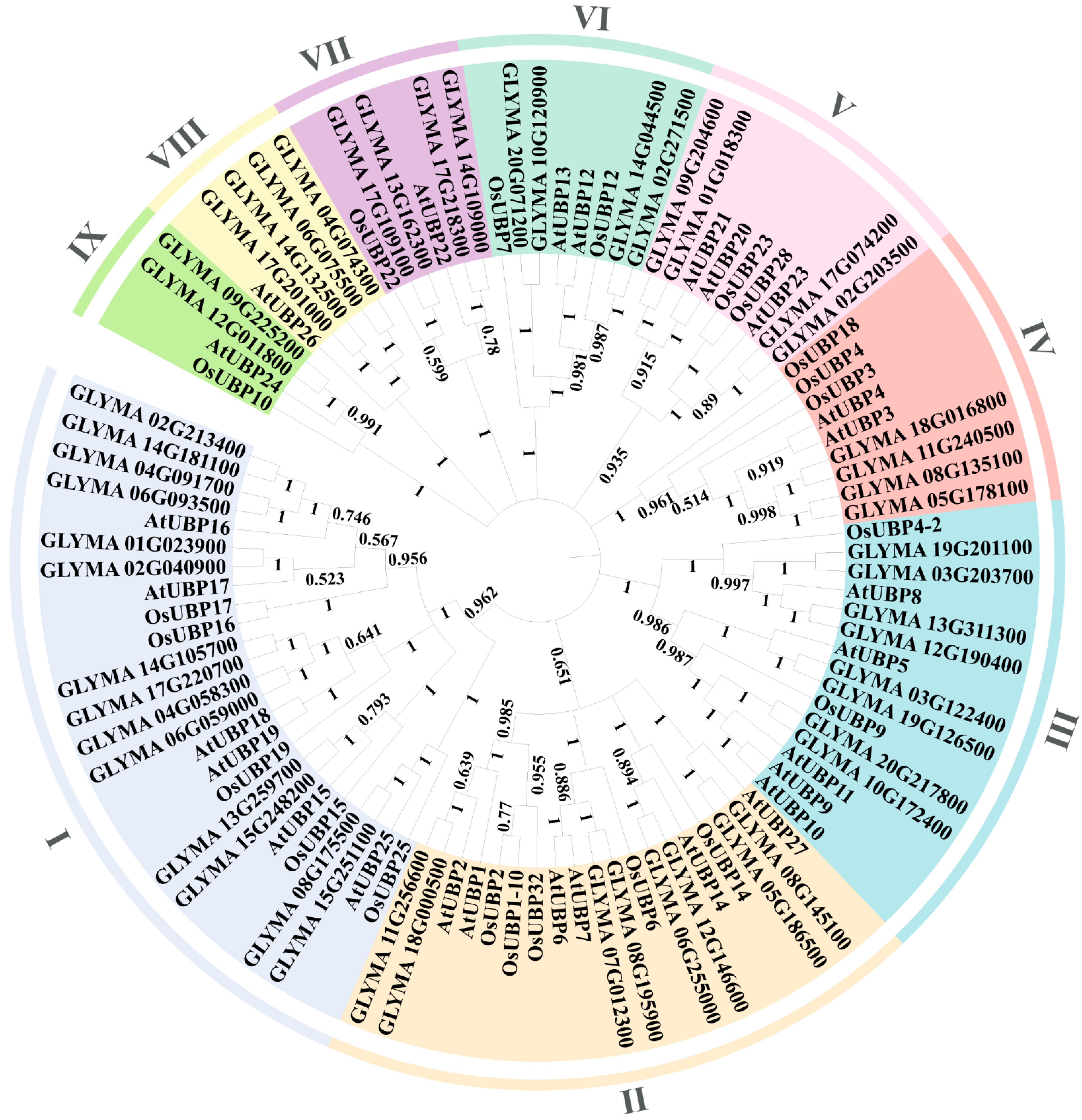
3.1. Characteristics and Phylogenetic Analysis of OsUBPs
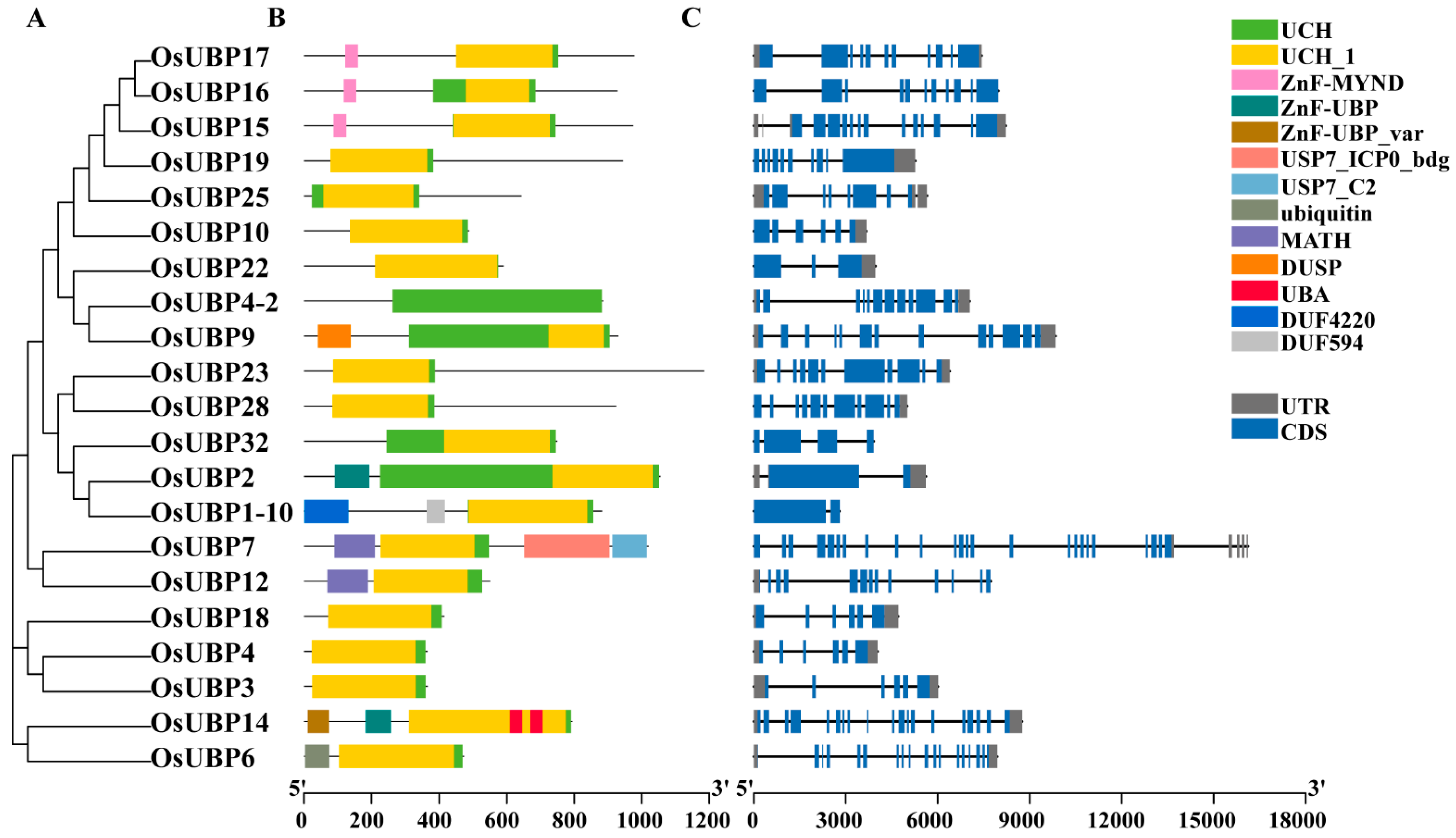
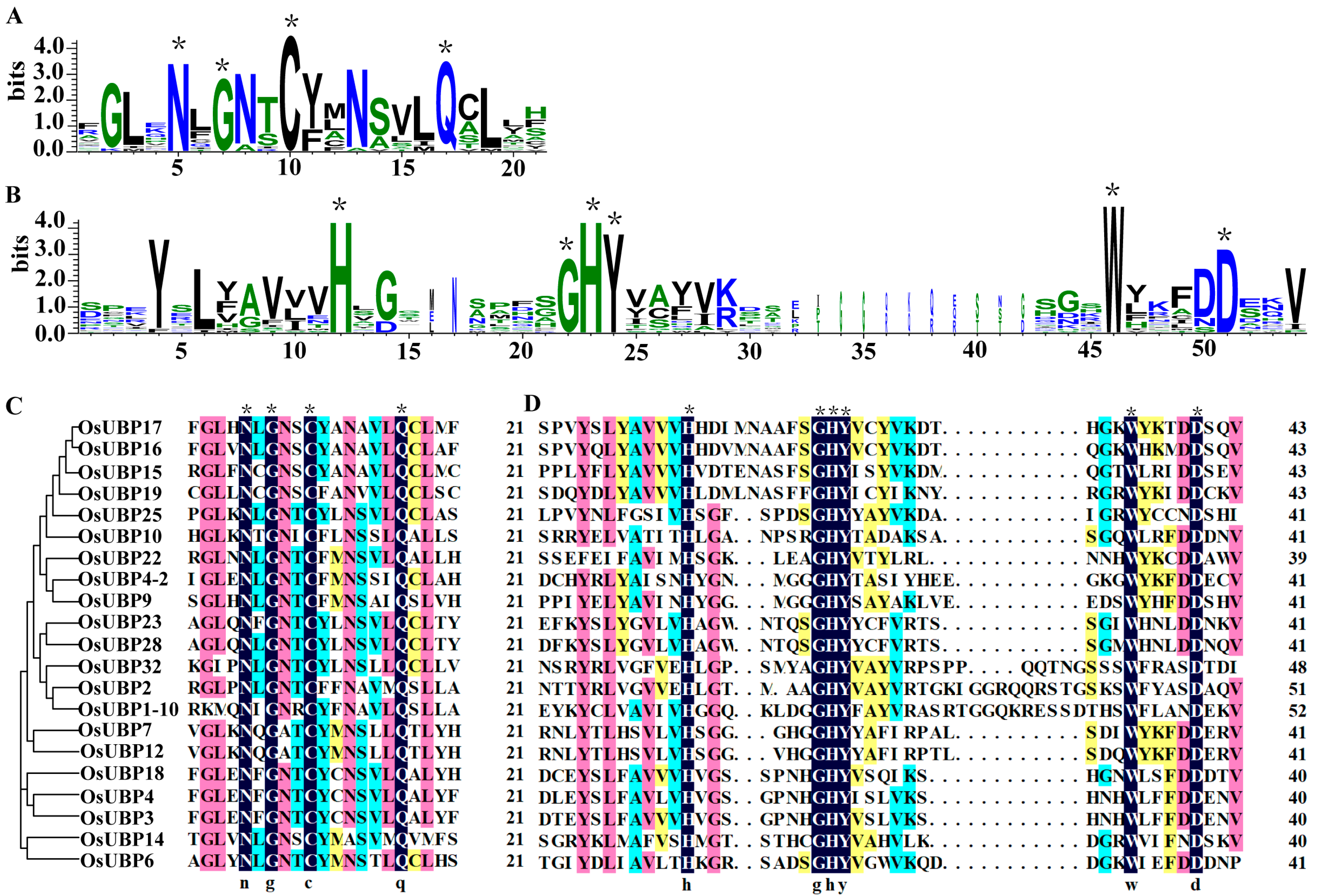
3.2. Conserved Domains and Gene Structure Analysis of OsUBPs
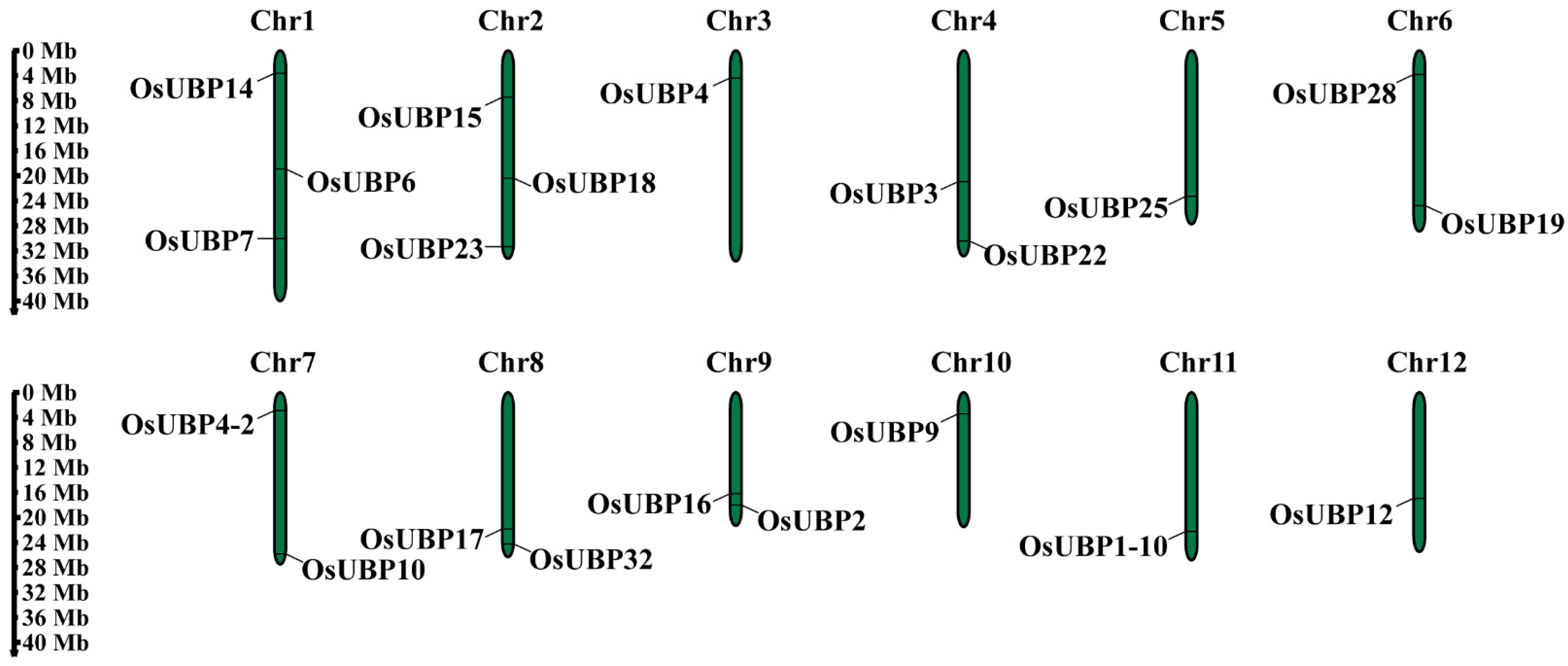
3.3. Chromosomal Location and Synteny Analysis of OsUBPs
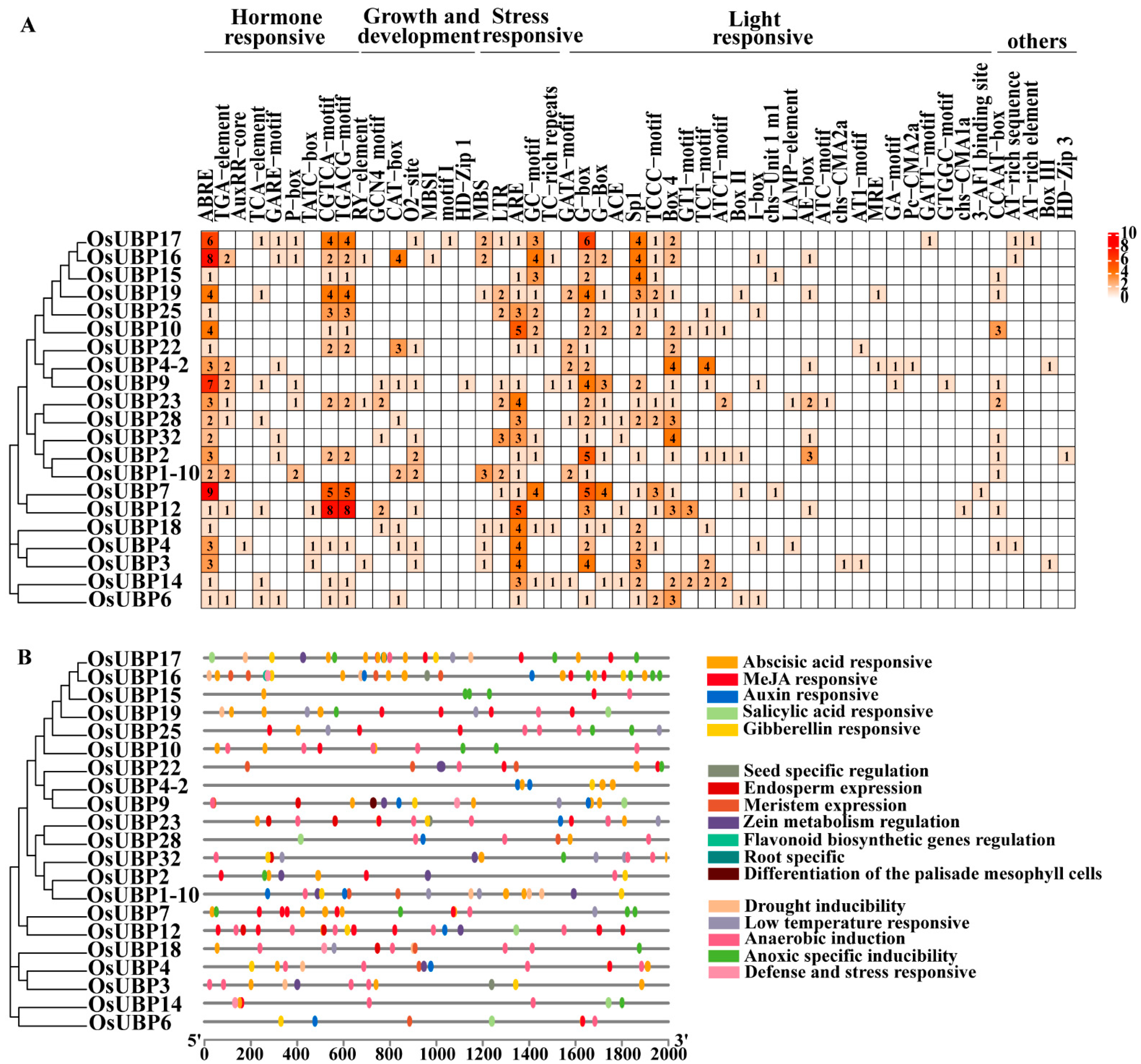
3.4. Cis-Element Analysis of OsUBPs
3.5. GO Annotation and Protein–Protein Interaction Network of OsUBPs
3.6. Expression Patterns of OsUBPs in Different Tissues
3.7. Expression Patterns of OsUBPs under Abiotic Stresses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Ahmad, S.; Jeridi, M.; Siddiqui, S.; Shah, A.Z.; Ali, S. Genome-wide identification, characterization, and expression analysis of the Chalcone Synthase gene family in Oryza sativa under Abiotic Stresses. Plant Stress 2023, 9, 100201. [Google Scholar] [CrossRef]
- Zhu, J.K. Abiotic Stress Signaling and Responses in Plants. Cell. 2016, 167, 313–324. [Google Scholar] [CrossRef]
- Lyzenga, W.J.; Stone, S.L. Abiotic stress tolerance mediated by protein ubiquitination. J. Exp. Bot. 2012, 63, 599–616. [Google Scholar] [CrossRef] [PubMed]
- Neutzner, M.; Neutzner, A. Enzymes of ubiquitination and deubiquitination. Essays Biochem. 2012, 52, 37–50. [Google Scholar] [PubMed]
- Zhou, H.; Zhao, J.; Cai, J.; Patil, S.B. Ubiquitin-specific proteases function in plant development and stress responses. Plant Mol. Biol. 2017, 94, 565–576. [Google Scholar] [CrossRef] [PubMed]
- Amerik, A.Y.; Hochstrasser, M. Mechanism and function of deubiquitinating enzymes. BBA Mol. Cell Res. 2004, 1695, 189–207. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.; Zheng, W.; Tan, J.; Sammer, R.; Du, L.; Lu, C. Protein partners of plant ubiquitin-specific proteases (UBPs). Plant Physiol. Biochem. 2019, 145, 227–236. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, F.; Zhang, H.; He, H.; Ma, L.; Deng, X.W. Functional characterization of the Arabidopsis ubiquitin-specific protease gene family reveals specific role and redundancy of individual members in development. Plant J. 2008, 55, 844–856. [Google Scholar] [CrossRef]
- Bonnet, J.; Romier, C.; Tora, L.; Devys, D. Zinc-finger UBPs: Regulators of deubiquitylation. Trends Biochem. Sci. 2008, 33, 369–375. [Google Scholar] [CrossRef]
- Yan, N.; Doelling, J.H.; Falbel, T.G.; Durski, A.M.; Vierstra, R.D. The ubiquitin-specific protease family from Arabidopsis. AtUBP1 and 2 are required for the resistance to the amino acid analog canavanine. Plant Physiol. 2000, 124, 1828–1843. [Google Scholar] [CrossRef]
- Wang, D.H.; Song, W.; Wei, S.W.; Zheng, Y.F.; Chen, Z.S.; Han, J.D.; Zhang, H.T.; Luo, J.C.; Qin, Y.M.; Xu, Z.H.; et al. Characterization of the Ubiquitin C-Terminal Hydrolase and Ubiquitin-Specific Protease Families in Rice (Oryza sativa). Front. Plant Sci. 2018, 9, 1636. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Jin, P.; Liu, T.; Gao, S.; Zhang, T.; Zhang, F.; Han, X.; He, L.; Chen, J.; Yang, J. Genome-wide identification and characterization of UBP gene family in wheat (Triticum aestivum L.). PeerJ 2021, 9, e11594. [Google Scholar] [CrossRef] [PubMed]
- Wu, R.; Shi, Y.; Zhang, Q.; Zheng, W.; Chen, S.; Du, L.; Lu, C. Genome-Wide Identification and Characterization of the UBP Gene Family in Moso Bamboo (Phyllostachys edulis). Int. J. Mol. Sci. 2019, 20, 4309. [Google Scholar] [CrossRef] [PubMed]
- Karamat, U.; Tabusam, J.; Khan, M.K.U.; Awan, M.J.A.; Zulfiqar, S.; Du, W.; Farooq, M.A. Genome-Wide Identification, Characterization, and Expression Profiling of Eukaryotic-Specific UBP Family Genes in Brassica rapa. J. Plant Growth Regul. 2023, 42, 3552–3567. [Google Scholar] [CrossRef]
- Zhou, H.; Zhao, J.; Yang, Y.; Chen, C.; Liu, Y.; Jin, X.; Chen, L.; Li, X.; Deng, X.W.; Schumaker, K.S.; et al. Ubiquitin-specific protease16 modulates salt tolerance in Arabidopsis by regulating Na+/H+ antiport activity and serine hydroxymethyltransferase stability. Plant Cell. 2012, 24, 5106–5122. [Google Scholar] [CrossRef]
- Zhao, J.; Zhou, H.; Zhang, M.; Gao, Y.; Li, L.; Gao, Y.; Li, M.; Yang, Y.; Guo, Y.; Li, X. Ubiquitin-specific protease 24 negatively regulates abscisic acid signalling in Arabidopsis thaliana. Plant Cell Environ. 2016, 39, 427–440. [Google Scholar] [CrossRef] [PubMed]
- Shi, C.; Ren, Y.; Liu, L.; Wang, F.; Zhang, H.; Tian, P.; Pan, T.; Wang, Y.; Jing, R.; Liu, T.; et al. Ubiquitin Specific Protease 15 Has an Important Role in Regulating Grain Width and Size in Rice. Plant Physiol. 2019, 180, 381–391. [Google Scholar] [CrossRef] [PubMed]
- Jiang, R.; Zhou, S.; Da, X.; Chen, T.; Xu, J.; Yan, P.; Mo, X. Ubiquitin-Specific Protease 2 (OsUBP2) Negatively Regulates Cell Death and Disease Resistance in Rice. Plants 2022, 11, 2568. [Google Scholar] [CrossRef]
- Zou, T.; Li, G.; Liu, M.; Liu, R.; Yang, S.; Wang, K.; Lu, L.; Ye, Q.; Liu, J.; Liang, J.; et al. A ubiquitin-specific protease functions in regulating cell death and immune responses in rice. Plant Cell Environ. 2023, 46, 1312–1326. [Google Scholar] [CrossRef]
- Sun, J.; Song, W.; Chang, Y.; Wang, Y.; Lu, T.; Zhang, Z. OsLMP1, Encoding a Deubiquitinase, Regulates the Immune Response in Rice. Front. Plant Sci. 2022, 12, 814465. [Google Scholar] [CrossRef]
- Moon, Y.K.; Hong, J.P.; Cho, Y.C.; Yang, S.J.; An, G.; Kim, W.T. Structure and expression of OsUBP6, an ubiquitin-specific protease 6 homolog in rice (Oryza sativa L.). Mol. Cells 2009, 28, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.; Jin, J.; Dong, Q.; Qiu, J.; Li, Y.; Yang, Y.; Shi, Y.; Si, W.; Gu, L.; Yang, F.; et al. Maize factors ZmUBP15, ZmUBP16 and ZmUBP19 play important roles for plants to tolerance the cadmium stress and salt stress. Plant Sci. 2019, 280, 77–89. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Chen, J.; Xia, J.; Hong, D. BnaUBP15s positively regulates seed size and seed weight in Brassica napus. Oil Crop Sci. 2023, 8, 149–155. [Google Scholar] [CrossRef]
- Park, S.H.; Jeong, J.S.; Zhou, Y.; Mustafa, N.F.B.; Chua, N.H. Deubiquitination of BES1 by UBP12/UBP13 promotes brassinosteroid signaling and plant growth. Plant Commun. 2022, 3, 100348. [Google Scholar] [CrossRef] [PubMed]
- Xiong, J.; Yang, F.; Yao, X.; Zhao, Y.; Wen, Y.; Lin, H.; Guo, H.; Yin, Y.; Zhang, D. The deubiquitinating enzymes UBP12 and UBP13 positively regulate recovery after carbon starvation by modulating BES1 stability in Arabidopsis thaliana. Plant Cell 2022, 34, 4516–4530. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Park, S.H.; Chua, N.H. UBP12/UBP13-mediated deubiquitination of salicylic acid receptor NPR3 suppresses plant immunity. Mol. Plant. 2023, 16, 232–244. [Google Scholar] [CrossRef] [PubMed]
- Lindbäck, L.N.; Hu, Y.; Ackermann, A.; Artz, O.; Pedmale, U.V. UBP12 and UBP13 deubiquitinases destabilize the CRY2 blue light receptor to regulate Arabidopsis growth. Curr. Biol. 2022, 32, 3221–3231. [Google Scholar] [CrossRef]
- Luo, Y.; Yasuda, S.; Takagi, J.; Hasegawa, Y.; Chiba, Y.; Yamaguchi, J.; Sato, T. Deubiquitinating enzymes UBP12 and UBP13 regulate carbon/nitrogen-nutrient stress responses by interacting with the membrane-localized ubiquitin ligase ATL31 in Arabidopsis. Biochem. Biophys. Res. Commun. 2022, 636, 55–61. [Google Scholar] [CrossRef]
- Zhou, Y.; Park, S.H.; Soh, M.Y.; Chua, N.H. Ubiquitin-specific proteases UBP12 and UBP13 promote shade avoidance response by enhancing PIF7 stability. Proc. Natl. Acad. Sci. USA 2021, 118, e2103633118. [Google Scholar] [CrossRef]
- Kralemann, L.E.M.; Liu, S.; Trejo-Arellano, M.S.; Muñoz-Viana, R.; Köhler, C.; Hennig, L. Removal of H2Aub1 by ubiquitin-specific proteases 12 and 13 is required for stable Polycomb-mediated gene repression in Arabidopsis. Genome Biol. 2020, 21, 144. [Google Scholar] [CrossRef]
- Finn, R.D.; Clements, J.; Arndt, W.; Miller, B.L.; Wheeler, T.J.; Schreiber, F.; Bateman, A.; Eddy, S.R. HMMER web server: 2015 update. Nucleic Acids Res. 2015, 43, W30–W38. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar] [CrossRef] [PubMed]
- Chou, K.C.; Shen, H.B. Plant-mPLoc: A Top-Down Strategy to Augment the Power for Predicting Plant Protein Subcellular Localization. PLoS ONE 2010, 5, e11335. [Google Scholar] [CrossRef]
- Kumar, S.; Nei, M.; Dudley, J.; Tamura, K. MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 2008, 9, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Chao, J.; Li, Z.; Sun, Y.; Aluko, O.O.; Wu, X.; Wang, Q.; Liu, G. MG2C: A user-friendly online tool for drawing genetic maps. Mol. Hortic. 2021, 1, 16. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.h.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Nei, M.; Gojobori, T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 1986, 3, 418–426. [Google Scholar]
- Wang, W.; Zhao, P.; Zhou, X.M.; Xiong, H.X.; Sun, M.X. Genome-wide identification and characterization of cystatin family genes in rice (Oryza sativa L.). Plant Cell Rep. 2015, 34, 1579–1592. [Google Scholar] [CrossRef]
- Huang, X.; Lei, Y.; Guan, H.; Hao, Y.; Liu, H.; Sun, G.; Chen, R.; Song, S. Transcriptomic analysis of the regulation of stalk development in flowering Chinese cabbage (Brassica campestris) by RNA sequencing. Sci. Rep. 2017, 7, 15517. [Google Scholar] [CrossRef]
- Ma, R.; Chen, J.; Huang, B.; Huang, Z.; Zhang, Z. The BBX gene family in Moso bamboo (Phyllostachys edulis): Identification, characterization and expression profiles. BMC Genom. 2021, 22, 533. [Google Scholar] [CrossRef] [PubMed]
- Sakai, H.; Lee, S.S.; Tanaka, T.; Numa, H.; Kim, J.; Kawahara, Y.; Wakimoto, H.; Yang, C.C.; Iwamoto, M.; Abe, T.; et al. Rice Annotation Project Database (RAP-DB): An Integrative and Interactive Database for Rice Genomics. Plant Cell Physiol. 2013, 54, e6. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Enany, S. Structural and functional analysis of hypothetical and conserved proteins of Clostridium tetani. J. Infect. Public Health 2014, 7, 296–307. [Google Scholar] [CrossRef] [PubMed]
- Ikai, A. Thermostability and aliphatic index of globular proteins. J. Biochem. 1980, 88, 1895–1898. [Google Scholar] [PubMed]
- Chang, K.Y.; Yang, J.R. Analysis and Prediction of Highly Effective Antiviral Peptides Based on Random Forests. PLoS ONE 2013, 8, e70166. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Moore, B.M.; Panchy, N.L.; Meng, F.; Lehti-Shiu, M.D.; Shiu, S.H. Factors Influencing Gene Family Size Variation Among Related Species in a Plant Family, Solanaceae. Genome Biol. Evol. 2018, 10, 2596–2613. [Google Scholar] [CrossRef]
- Liu, H.; Yang, W.; Zhao, X.; Kang, G.; Li, N.; Xu, H. Genome-wide analysis and functional characterization of CHYR gene family associated with abiotic stress tolerance in bread wheat (Triticum aestivum L.). BMC Plant Biol. 2022, 22, 204. [Google Scholar] [CrossRef]
- Saleem, B.; Farooq, U.; Rehman, O.U.; Aqeel, M.; Farooq, M.S.; Naeem, M.K.; Inam, S.; Ajmal, W.; Rahim, A.A.; Chen, M.; et al. Genome-wide and molecular characterization of the DNA replication helicase 2 (DNA2) gene family in rice under drought and salt stress. Front. Genet. 2022, 13, 1039548. [Google Scholar] [CrossRef]
- Du, L.; Li, N.; Chen, L.; Xu, Y.; Li, Y.; Zhang, Y.; Li, C.; Li, Y. The ubiquitin receptor DA1 regulates seed and organ size by modulating the stability of the ubiquitin-specific protease UBP15/SOD2 in Arabidopsis. Plant Cell. 2014, 26, 665–677. [Google Scholar] [CrossRef]
- Chung, S.; Kwon, H.L.; Yun, H.S.; Lee, J.H. The Function of Deubiquitinating Enzymes in Arabidopsis: Recent Progress of Ubiquitin-Specific Proteases (UBPs). J. Plant Biol. 2023. [Google Scholar] [CrossRef]
- Ewan, R.; Pangestuti, R.; Thornber, S.; Craig, A.; Carr, C.; O’Donnell, L.; Zhang, C.; Sadanandom, A. Deubiquitinating enzymes AtUBP12 and AtUBP13 and their tobacco homologue NtUBP12 are negative regulators of plant immunity. New Phytol. 2011, 191, 92–106. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.W.; Baek, W.; Lee, S.C. Tobacco ubiquitin-specific protease 12 (NbUBP12) positively modulates drought resistance. Plant Signal Behav. 2021, 16, 1974725. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.W.; Baek, W.; Lim, J.; Hong, E.; Lee, S.C. Pepper ubiquitin-specific protease, CaUBP12, positively modulates dehydration resistance by enhancing CaSnRK2.6 stability. Plant J. 2021, 107, 1148–1165. [Google Scholar] [CrossRef] [PubMed]
- Majumdar, P.; Nath, U. De-ubiquitinases on the move: An emerging field in plant biology. Plant Biol. 2020, 22, 563–572. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, C.R.; Seeger, M.; Hartmann-Petersen, R.; Stone, M.; Wallace, M.; Semple, C.; Gordon, C. Proteins containing the UBA domain are able to bind to multi-ubiquitin chains. Nat. Cell Biol. 2001, 3, 939–943. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Qin, H.; Zhou, S.; Wei, P.; Zhang, H.; Zhou, Y.; Miao, Y.; Huang, R. The Ubiquitin-Binding Protein OsDSK2a Mediates Seedling Growth and Salt Responses by Regulating Gibberellin Metabolism in Rice. Plant Cell 2020, 32, 414–428. [Google Scholar] [CrossRef] [PubMed]
- Jacquemin, J.; Laudié, M.; Cooke, R. A recent duplication revisited: Phylogenetic analysis reveals an ancestral duplication highly-conserved throughout the Oryza genus and beyond. BMC Plant Biol. 2009, 9, 146. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Van der Does, D.; Zamioudis, C.; Leon-Reyes, A.; Van Wees, S.C.M. Hormonal Modulation of Plant Immunity. Annu. Rev. Cell Dev. Biol. 2012, 28, 489–521. [Google Scholar] [CrossRef]
- Bari, R.; Jones, J.D.G. Role of plant hormones in plant defence responses. Plant Mol. Biol. 2008, 69, 473–488. [Google Scholar] [CrossRef]
- Wei, H.; Xu, H.; Su, C.; Wang, X.; Wang, L. Rice CIRCADIAN CLOCK ASSOCIATED 1 transcriptionally regulates ABA signaling to confer multiple abiotic stress tolerance. Plant Physiol. 2022, 190, 1057–1073. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Wu, H.; Ma, S.; Xiang, D.; Liu, R.; Xiong, L. OsJAZ1 Attenuates Drought Resistance by Regulating JA and ABA Signaling in Rice. Front. Plant Sci. 2017, 8, 2108. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Porras, J.L.; Riaño-Pachón, D.M.; Dreyer, I.; Mayer, J.E.; Mueller-Roeber, B. Genome-wide analysis of ABA-responsive elements ABRE and CE3 reveals divergent patterns in Arabidopsis and rice. BMC Genom. 2007, 8, 260. [Google Scholar] [CrossRef] [PubMed]
- Basyuni, M.; Wati, R.; Sulistiyono, N.; Sumardi; Oku, H.; Baba, S.; Sagami, H. Isolation and analysis of a multifunctional triterpene synthase KcMS promoter region from mangrove plant kandelia candel. IOP Conf. Ser. Earth Environ. Sci. 2018, 130, 012013. [Google Scholar] [CrossRef]
- Sridhar, V.V.; Kapoor, A.; Zhang, K.; Zhu, J.; Zhou, T.; Hasegawa, P.M.; Bressan, R.A.; Zhu, J.K. Control of DNA methylation and heterochromatic silencing by histone H2B deubiquitination. Nature 2007, 447, 735–738. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Jiang, W.; Long, F.; Cheng, S.; Yang, W.; Zhao, Y.; Zhou, D.X. Rice Homeodomain Protein WOX11 Recruits a Histone Acetyltransferase Complex to Establish Programs of Cell Proliferation of Crown Root Meristem. Plant Cell 2017, 29, 1088–1104. [Google Scholar] [CrossRef]
- Zhou, Y.; Ma, B.; Tao, J.J.; Yin, C.C.; Hu, Y.; Huang, Y.H.; Wei, W.; Xin, P.Y.; Chu, J.F.; Zhang, W.K.; et al. Rice EIL1 interacts with OsIAAs to regulate auxin biosynthesis mediated by the tryptophan aminotransferase MHZ10/OsTAR2 during root ethylene responses. Plant Cell 2022, 34, 4366–4387. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Liang, J.; Lou, L.; Tian, M.; Zhang, X.; Liu, L.; Zhao, Q.; Xia, R.; Wu, Y.; Xie, Q.; et al. The deubiquitinases UBP12 and UBP13 integrate with the E3 ubiquitin ligase XBAT35. 2 to modulate VPS23A stability in ABA signaling. Sci. Adv. 2022, 8, eabl5765. [Google Scholar] [CrossRef]
- De Poot, S.A.H.; Tian, G.; Finley, D. Meddling with Fate: The Proteasomal Deubiquitinating Enzymes. J. Mol. Biol. 2017, 429, 3525–3545. [Google Scholar] [CrossRef]
- Sowa, G.; Westrick, E.; Rajasekhar, A.G.; Woods, B.; Leckie, S.; Coelho, P.; Vo, N.; Studer, R.; Kang, J. Identification of Candidate Serum Biomarkers for Intervertebral Disk Degeneration in an Animal Model. PM&R 2009, 1, 536–540. [Google Scholar]
- Wolberger, C. Mechanisms for regulating deubiquitinating enzymes. Protein Sci. 2014, 23, 344–353. [Google Scholar] [CrossRef]
- Zhou, H.; Zhou, M.; Yang, Y.; Li, J.; Zhu, L.; Jiang, D.; Dong, J.; Liu, Q.; Gu, L.; Zhou, L.; et al. RNase ZS1 processes UbL40 mRNAs and controls thermosensitive genic male sterility in rice. Nat. Commun. 2014, 5, 4884. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Sun, S.; Xu, D.; Lan, H.; Sun, H.; Wang, Z.; Bao, Y.; Wang, J.; Tang, H.; Zhang, H. A TFIIIA-type zinc finger protein confers multiple abiotic stress tolerances in transgenic rice (Oryza sativa L.). Plant Mol. Biol. 2012, 80, 337–350. [Google Scholar] [CrossRef] [PubMed]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zou, X.; Li, Y.; Yin, H.; Xu, J.; Li, Z.; Jiang, S.; Chen, F.; Li, Y.; Xiao, W.; Liu, S.; et al. Genome-Wide Identification, Characterization, and Expression Analysis under Abiotic Stresses of the UBP Gene Family in Rice (Oryza sativa L.). Agronomy 2023, 13, 2809. https://doi.org/10.3390/agronomy13112809
Zou X, Li Y, Yin H, Xu J, Li Z, Jiang S, Chen F, Li Y, Xiao W, Liu S, et al. Genome-Wide Identification, Characterization, and Expression Analysis under Abiotic Stresses of the UBP Gene Family in Rice (Oryza sativa L.). Agronomy. 2023; 13(11):2809. https://doi.org/10.3390/agronomy13112809
Chicago/Turabian StyleZou, Xiaoxiao, Yongliang Li, Huangping Yin, Jiajin Xu, Zeqi Li, Shuai Jiang, Fenglin Chen, You Li, Wenjun Xiao, Shucan Liu, and et al. 2023. "Genome-Wide Identification, Characterization, and Expression Analysis under Abiotic Stresses of the UBP Gene Family in Rice (Oryza sativa L.)" Agronomy 13, no. 11: 2809. https://doi.org/10.3390/agronomy13112809





