Synovial Fibroblast Extracellular Vesicles Induce Inflammation via Delivering miR-21-5p in Osteoarthritis
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Samples
2.2. Chondrocytes and Synovial Fibroblasts Cultures
2.3. Small Extracellular Vesicles (sEVs) Isolation
2.4. sEVs Protein Quantification and Western Blot
2.5. Transmission Electron Microscopy
2.6. Nanoparticle Tracking Analysis (NTA)
2.7. Microarrays for FS_OA_sEVs miRNA Profiling and Expression Analysis
2.8. Enrichment Analysis and miRNAs’ Target Genes Prediction
2.9. sEVs Labeling and Uptake Assay
2.10. Scratch Assay
2.11. Transfection Experiments of C_OAmax with miR-21-5p Inhibitor and KLF6 siRNA
2.12. Transfection of C_OAmin Chondrocytes with FS_OA_sEVs and miR-21-5p Inhibitors
2.13. RNA Extraction, cDNA Synthesis, and Quantitative Real-Time PCR (qRT-PCR)
2.14. ELISA Assay
2.15. Statistical Analysis
3. Results
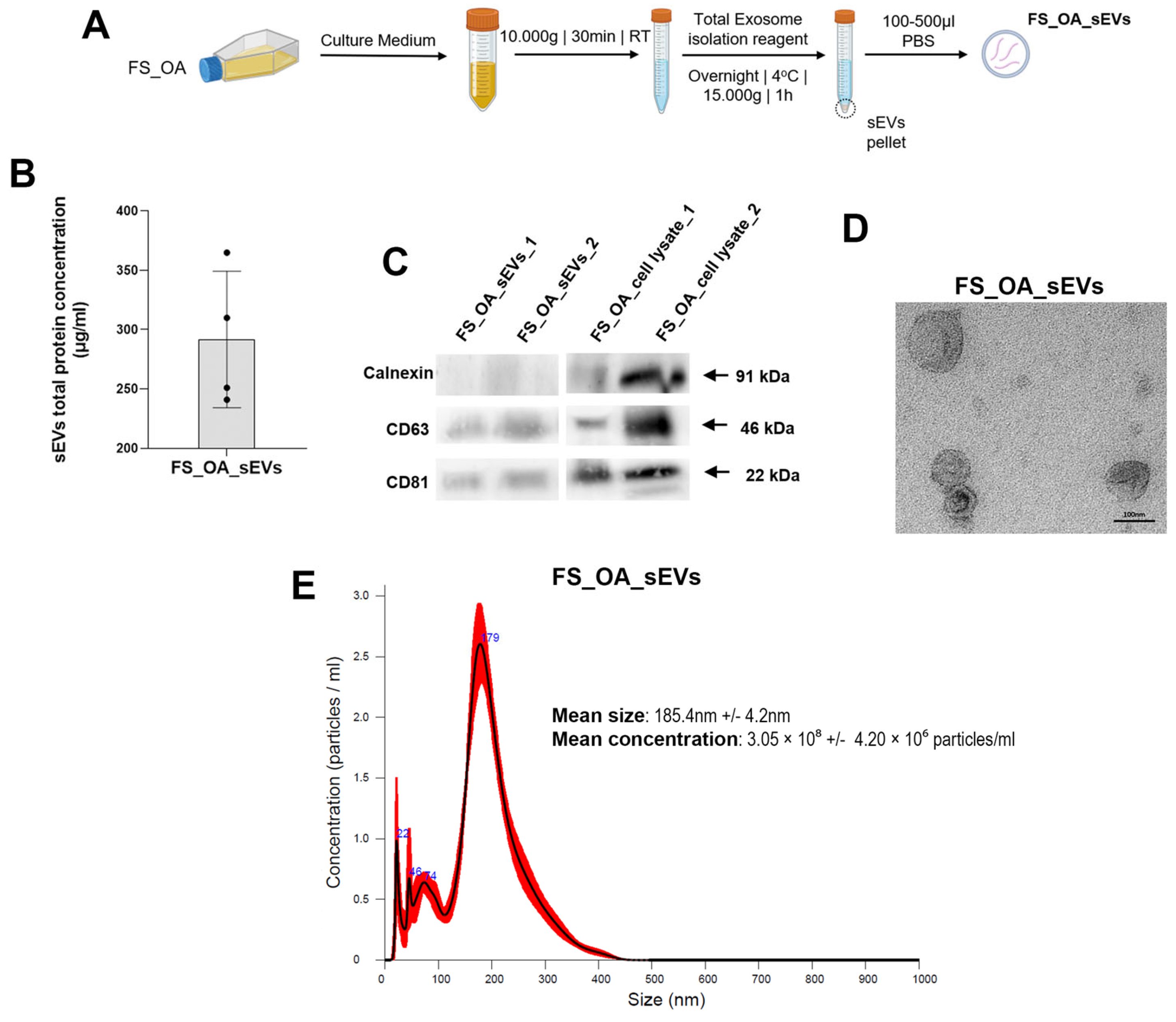
3.1. Isolation and Characterization of Small EVs Derived from OA Synovial Fibroblasts
3.2. FS_OA_sEVs Affect Chondrocytes’ Anabolic and Catabolic OA Gene Markers and Reduce Cell Motility
3.3. miRNA Pattern of FS_OA_sEVs
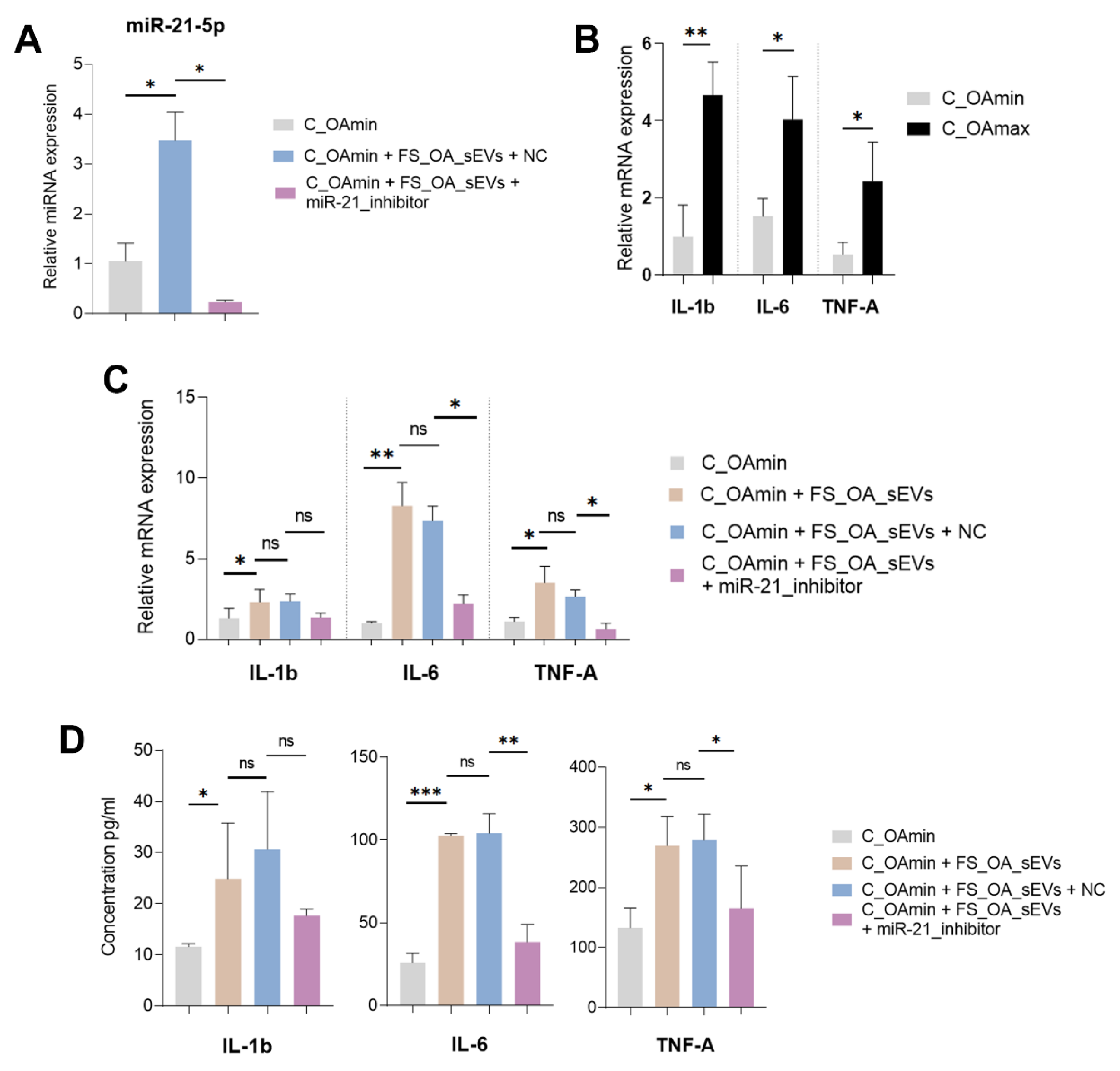
3.4. FS_OA_ sEVs-Derived-miR-21-5p Induces Inflammation-Related Genes in C_OAmin Chondrocytes
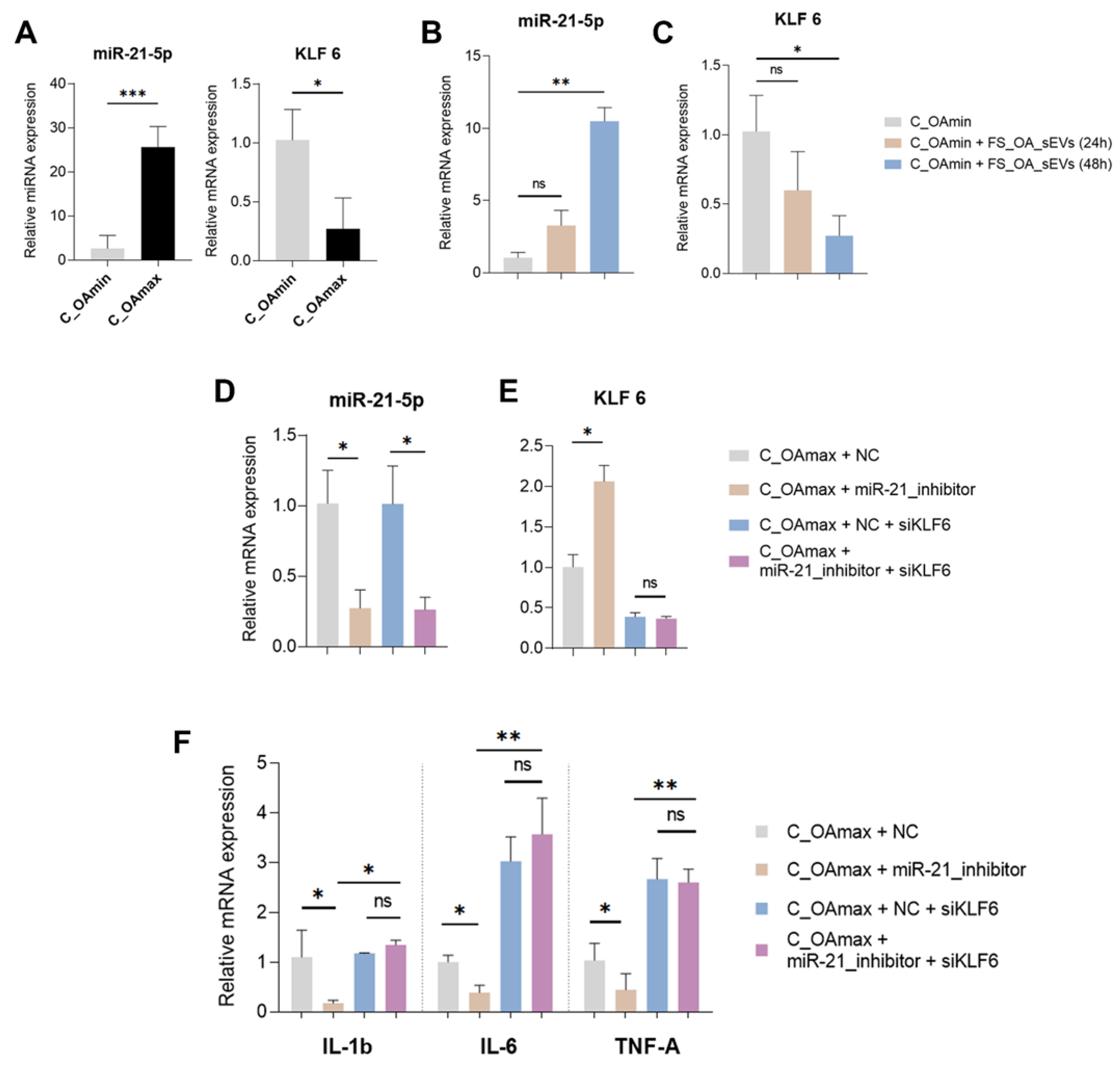
3.5. FS_OA_ sEVs-Derived miR-21 Inhibits KLF6 Expression and Promotes IL-1β, IL-6, and TNF-α Expressions in C_OAmax Chondrocytes
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Long, H.; Liu, Q.; Yin, H.; Wang, K.; Diao, N.; Zhang, Y.; Lin, J.; Guo, A. Prevalence Trends of Site-Specific Osteoarthritis From 1990 to 2019: Findings from the Global Burden of Disease Study 2019. Arthritis Rheumatol. 2022, 74, 1172–1183. [Google Scholar] [CrossRef] [PubMed]
- Stanaway, J.D.; Afshin, A.; Gakidou, E.; Lim, S.S.; Abate, D.; Abate, K.H.; Abbafati, C.; Abbasi, N.; Abbastabar, H.; Abd-Allah, F.; et al. Global, Regional, and National Comparative Risk Assessment of 84 Behavioural, Environmental and Occupational, and Metabolic Risks or Clusters of Risks for 195 Countries and Territories, 1990–2017: A Systematic Analysis for the Global Burden of Disease Study 2017. Lancet 2018, 392, 1923–1994. [Google Scholar] [CrossRef]
- Hunter, D.J.; March, L.; Chew, M. Osteoarthritis in 2020 and beyond: A Lancet Commission. Lancet 2020, 396, 1711–1712. [Google Scholar] [CrossRef] [PubMed]
- Martel-Pelletier, J.; Barr, A.J.; Cicuttini, F.M.; Conaghan, P.G.; Cooper, C.; Goldring, M.B.; Goldring, S.R.; Jones, G.; Teichtahl, A.J.; Pelletier, J.P. Osteoarthritis. Nat. Rev. Dis. Primers 2016, 2, 16072. [Google Scholar] [CrossRef]
- Oo, W.M.; Yu, S.P.C.; Daniel, M.S.; Hunter, D.J. Disease-Modifying Drugs in Osteoarthritis: Current Understanding and Future Therapeutics. Expert. Opin. Emerg. Drugs 2018, 23, 331–347. [Google Scholar] [CrossRef]
- Oo, W.M.; Little, C.; Duong, V.; Hunter, D.J. The Development of Disease-Modifying Therapies for Osteoarthritis (DMOADs): The Evidence to Date. Drug Des. Dev. Ther. 2021, 15, 2921–2945. [Google Scholar] [CrossRef]
- Cross, M.; Smith, E.; Hoy, D.; Nolte, S.; Ackerman, I.; Fransen, M.; Bridgett, L.; Williams, S.; Guillemin, F.; Hill, C.L.; et al. The Global Burden of Hip and Knee Osteoarthritis: Estimates from the Global Burden of Disease 2010 Study. Ann. Rheum. Dis. 2014, 73, 1323–1330. [Google Scholar] [CrossRef]
- Pas, H.I.; Winters, M.; Haisma, H.J.; Koenis, M.J.; Tol, J.L.; Moen, M.H. Stem Cell Injections in Knee Osteoarthritis: A Systematic Review of the Literature. Br. J. Sports Med. 2017, 51, 1125–1133. [Google Scholar] [CrossRef]
- Hunter, D.J.; Bierma-Zeinstra, S. Osteoarthritis. Lancet 2019, 393, 1745–1759. [Google Scholar] [CrossRef]
- Li, Z.; Huang, Z.; Bai, L. Cell Interplay in Osteoarthritis. Front. Cell Dev. Biol. 2021, 9, 720477. [Google Scholar] [CrossRef]
- Chou, C.H.; Jain, V.; Gibson, J.; Attarian, D.E.; Haraden, C.A.; Yohn, C.B.; Laberge, R.M.; Gregory, S.; Kraus, V.B. Synovial Cell Cross-Talk with Cartilage Plays a Major Role in the Pathogenesis of Osteoarthritis. Sci. Rep. 2020, 10, 10868. [Google Scholar] [CrossRef] [PubMed]
- Sondag, G.R.; Haqqi, T.M. The Role of MicroRNAs and Their Targets in Osteoarthritis. Curr. Rheumatol. Rep. 2016, 18, 56. [Google Scholar] [CrossRef] [PubMed]
- Albanese, M.; Chen, Y.F.A.; Hüls, C.; Gärtner, K.; Tagawa, T.; Mejias-Perez, E.; Keppler, O.T.; Göbel, C.; Zeidler, R.; Shein, M.; et al. MicroRNAs Are Minor Constituents of Extracellular Vesicles That Are Rarely Delivered to Target Cells. PLoS Genet. 2021, 17, e1009951. [Google Scholar] [CrossRef]
- Otahal, A.; Kuten-Pella, O.; Kramer, K.; Neubauer, M.; Lacza, Z.; Nehrer, S.; De Luna, A. Functional Repertoire of EV-Associated MiRNA Profiles after Lipoprotein Depletion via Ultracentrifugation and Size Exclusion Chromatography from Autologous Blood Products. Sci. Rep. 2021, 11, 5823. [Google Scholar] [CrossRef]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding Light on the Cell Biology of Extracellular Vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Wu, Y.; Li, J.; Zeng, Y.; Pu, W.; Mu, X.; Sun, K.; Peng, Y.; Shen, B. Exosomes Rewire the Cartilage Microenvironment in Osteoarthritis: From Intercellular Communication to Therapeutic Strategies. Int. J. Oral. Sci. 2022, 14, 40. [Google Scholar] [CrossRef]
- Esa, A.; Connolly, K.D.; Williams, R.; Archer, C.W. Extracellular Vesicles in the Synovial Joint: Is There a Role in the Pathophysiology of Osteoarthritis? Malays. Orthop. J. 2019, 13, 1–7. [Google Scholar] [CrossRef]
- Pakdaman Kolour, S.S.; Nematollahi, S.; Dehbozorgi, M.; Fattahi, F.; Movahed, F.; Esfandiari, N.; Kahrizi, M.S.; Ghavamikia, N.; Hajiagha, B.S. Extracecellulr Vesicles (EVs) MicroRNAs (MiRNAs) Derived from Mesenchymal Stem Cells (MSCs) in Osteoarthritis (OA); Detailed Role in Pathogenesis and Possible Therapeutics. Heliyon 2025, 11, e42258. [Google Scholar] [CrossRef]
- Withrow, J.; Murphy, C.; Liu, Y.; Hunter, M.; Fulzele, S.; Hamrick, M.W. Extracellular Vesicles in the Pathogenesis of Rheumatoid Arthritis and Osteoarthritis. Arthritis Res. Ther. 2016, 18, 286. [Google Scholar] [CrossRef]
- Ludwig, A.K.; Giebel, B. Exosomes: Small Vesicles Participating in Intercellular Communication. Int. J. Biochem. Cell Biol. 2012, 44, 11–15. [Google Scholar] [CrossRef]
- Mao, G.; Hu, S.; Zhang, Z.; Wu, P.; Zhao, X.; Lin, R.; Liao, W.; Kang, Y. Exosomal MiR-95-5p Regulates Chondrogenesis and Cartilage Degradation via Histone Deacetylase 2/8. J. Cell. Mol. Med. 2018, 22, 5354–5366. [Google Scholar] [CrossRef] [PubMed]
- Asghar, S.; Litherland, G.J.; Cole, J.J.; McInnes, I.B.; Meek, R.M.D.; Lockhart, J.C.; Goodyear, C.S.; Crilly, A. Small Extracellular Vesicles Derived from Synovial Fibroblasts Contain Distinct MiRNA Profiles and Contribute to Chondrocyte Damage in Osteoarthritis. Arthritis Res. Ther. 2024, 26, 167. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Liu, Y.M.; Fu, B.L.; Xu, L.L.; Wang, B. MicroRNA-21: An Emerging Player in Bone Diseases. Front. Pharmacol. 2021, 12, 722804. [Google Scholar] [CrossRef]
- Qin, L.; Yang, J.; Su, X.; Li, X.; Lei, Y.; Dong, L.; Chen, H.; Chen, C.; Zhao, C.; Zhang, H.; et al. The MiR-21-5p Enriched in the Apoptotic Bodies of M2 Macrophage-Derived Extracellular Vesicles Alleviates Osteoarthritis by Changing Macrophage Phenotype. Genes. Dis. 2022, 10, 1114–1129. [Google Scholar] [CrossRef]
- Simopoulou, T.; Malizos, K.N.; Iliopoulos, D.; Stefanou, N.; Papatheodorou, L.; Ioannou, M.; Tsezou, A. Differential Expression of Leptin and Leptin’s Receptor Isoform (Ob-Rb) MRNA between Advanced and Minimally Affected Osteoarthritic Cartilage; Effect on Cartilage Metabolism. Osteoarthr. Cartil. 2007, 15, 872–883. [Google Scholar] [CrossRef]
- Li, F.; Tang, Y.; Song, B.; Yu, M.; Li, Q.; Zhang, C.; Hou, J.; Yang, R. Nomenclature Clarification: Synovial Fibroblasts and Synovial Mesenchymal Stem Cells. Stem Cell Res. Ther. 2019, 10, 260. [Google Scholar] [CrossRef]
- Welsh, J.A.; Goberdhan, D.C.I.; O’Driscoll, L.; Buzas, E.I.; Blenkiron, C.; Bussolati, B.; Cai, H.; Di Vizio, D.; Driedonks, T.A.P.; Erdbrügger, U.; et al. Minimal Information for Studies of Extracellular Vesicles (MISEV2023): From Basic to Advanced Approaches. J. Extracell. Vesicles 2024, 13, e12404. [Google Scholar] [CrossRef]
- Papageorgiou, A.A.; Roussos, A.; Papathanasiou, I.; Balis, C.; Karachalios, T.; Varitimidis, S.E.; Malizos, K.N.; Tsezou, A. MiR-217 Regulates SIRT1 Expression and Promotes Inflammatory and Apoptotic Responses in Osteoarthritis. Genes 2023, 14, 2155. [Google Scholar] [CrossRef]
- Papathanasiou, I.; Balis, C.; Destounis, D.; Mourmoura, E.; Tsezou, A. NEAT1-Mediated MiR-150-5p Downregulation Regulates b-Catenin Expression in OA Chondrocytes. Funct. Integr. Genom. 2023, 23, 246. [Google Scholar] [CrossRef]
- Konteles, V.; Papathanasiou, I.; Tzetis, M.; Goussetis, E.; Trachana, V.; Mourmoura, E.; Balis, C.; Malizos, K.; Tsezou, A. Integration of Transcriptome and MicroRNA Profile Analysis of IMSCs Defines Their Rejuvenated State and Conveys Them into a Novel Resource for Cell Therapy in Osteoarthritis. Cells 2023, 12, 1756. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma Powers Differential Expression Analyses for RNA-Sequencing and Microarray Studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Jia, J.; Yang, S.; Liu, X.; Ye, S.; Tian, H. MicroRNA-21 Controls the Development of Osteoarthritis by Targeting GDF-5 in Chondrocytes. Exp. Mol. Med. 2014, 46, e79. [Google Scholar] [CrossRef] [PubMed]
- Hoshikawa, N.; Sakai, A.; Takai, S.; Suzuki, H. Targeting Extracellular MiR-21-TLR7 Signaling Provides Long-Lasting Analgesia in Osteoarthritis. Mol. Ther. Nucleic Acids 2020, 19, 199–207. [Google Scholar] [CrossRef]
- Słomka, A.; Urban, S.K.; Lukacs-Kornek, V.; Żekanowska, E.; Kornek, M. Large Extracellular Vesicles: Have We Found the Holy Grail of Inflammation? Front. Immunol. 2018, 9, 2723. [Google Scholar] [CrossRef]
- Wang, C.; Zhang, C.; Liu, L.; Xi, A.; Chen, B.; Li, Y.; Du, J. Macrophage-Derived Mir-155-Containing Exosomes Suppress Fibroblast Proliferation and Promote Fibroblast Inflammation during Cardiac Injury. Mol. Ther. 2017, 25, 192–204. [Google Scholar] [CrossRef]
- Liu, J.; Jiang, M.; Deng, S.; Lu, J.; Huang, H.; Zhang, Y.; Gong, P.; Shen, X.; Ruan, H.; Jin, M.; et al. MiR-93-5p-Containing Exosomes Treatment Attenuates Acute Myocardial Infarction-Induced Myocardial Damage. Mol. Ther. Nucleic Acids 2018, 11, 103–115. [Google Scholar] [CrossRef]
- Witwer, K.W.; Soekmadji, C.; Hill, A.F.; Wauben, M.H.; Buzás, E.I.; Di Vizio, D.; Falcon-Perez, J.M.; Gardiner, C.; Hochberg, F.; Kurochkin, I.V.; et al. Updating the MISEV Minimal Requirements for Extracellular Vesicle Studies: Building Bridges to Reproducibility. J. Extracell. Vesicles 2017, 6, 1396823. [Google Scholar] [CrossRef]
- Chen, P.; Zhou, J.; Ruan, A.; Guan, H.; Xie, J.; Zeng, L.; Liu, J.; Wang, Q. Synovial Tissue-Derived Extracellular Vesicles Induce Chondrocyte Inflammation and Degradation via NF-ΚB Signalling Pathway: An in Vitro Study. J. Cell. Mol. Med. 2022, 26, 2038–2048. [Google Scholar] [CrossRef]
- Akkiraju, H.; Nohe, A. Role of Chondrocytes in Cartilage Formation, Progression of Osteoarthritis and Cartilage Regeneration. J. Dev. Biol. 2015, 3, 177–192. [Google Scholar] [CrossRef]
- Iliopoulos, D.; Malizos, K.N.; Oikonomou, P.; Tsezou, A. Integrative MicroRNA and Proteomic Approaches Identify Novel Osteoarthritis Genes and Their Collaborative Metabolic and Inflammatory Networks. PLoS ONE 2008, 3, e3740. [Google Scholar] [CrossRef]
- Li, L.; Jia, J.; Liu, X.; Yang, S.; Ye, S.; Yang, W.; Zhang, Y. MicroRNA-16-5p Controls Development of Osteoarthritis by Targeting SMAD3 in Chondrocytes. Curr. Pharm. Des. 2015, 21, 5160–5167. [Google Scholar] [CrossRef] [PubMed]
- Knights, A.J.; Redding, S.J.; Maerz, T. Inflammation in Osteoarthritis: The Latest Progress and Ongoing Challenges. Curr. Opin. Rheumatol. 2023, 35, 128–134. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Lopez, E.; Coras, R.; Torres, A.; Lane, N.E.; Guma, M. Synovial Inflammation in Osteoarthritis Progression. Nat. Rev. Rheumatol. 2022, 18, 258–275. [Google Scholar] [CrossRef]
- Duan, R.; Xie, H.; Liu, Z.Z. The Role of Autophagy in Osteoarthritis. Front. Cell Dev. Biol. 2020, 8, 608388. [Google Scholar] [CrossRef]
- McConnell, B.B.; Yang, V.W. Mammalian Krüppel-like Factors in Health and Diseases. Physiol. Rev. 2010, 90, 1337–1381. [Google Scholar] [CrossRef]
- Liu, C. Screen for autophagy-related biomarkers in osteoarthritis based on bioinformatic analysis. Biocell 2024, 48, 339–351. [Google Scholar] [CrossRef]
- Li, N.; Xue, Y.; Zhu, C.; Chen, N.; Qi, M.; Fang, M.; Huang, S. The Zinc-Finger Transcription Factor KLF6 Regulates Cardiac Fibrosis. Life Sci. 2024, 351, 122805. [Google Scholar] [CrossRef]
- Holian, J.; Qi, W.; Kelly, D.J.; Zhang, Y.; Mreich, E.; Pollock, C.A.; Chen, X.M. Role of Kruppel-like Factor 6 in Transforming Growth Factor-Beta1-Induced Epithelial-Mesenchymal Transition of Proximal Tubule Cells. Am. J. Physiol. Renal Physiol. 2008, 295, F1388–F1396. [Google Scholar] [CrossRef]
- Goodman, W.A.; Omenetti, S.; Date, D.; Di Martino, L.; De Salvo, C.; Kim, G.D.; Chowdhry, S.; Bamias, G.; Cominelli, F.; Pizarro, T.T.; et al. KLF6 Contributes to Myeloid Cell Plasticity in the Pathogenesis of Intestinal Inflammation. Mucosal Immunol. 2016, 9, 1250–1262. [Google Scholar] [CrossRef]
- Palau, N.; Julià, A.; Ferrándiz, C.; Puig, L.; Fonseca, E.; Fernández, E.; López-Lasanta, M.; Tortosa, R.; Marsal, S. Genome-Wide Transcriptional Analysis of T Cell Activation Reveals Differential Gene Expression Associated with Psoriasis. BMC Genom. 2013, 14, 825. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Ratziu, V.; Choi, S.G.; Lalazar, A.; Theiss, G.; Dang, Q.; Kim, S.J.; Friedman, S.L. Transcriptional Activation of Transforming Growth Factor Beta1 and Its Receptors by the Kruppel-like Factor Zf9/Core Promoter-Binding Protein and Sp1. Potential Mechanisms for Autocrine Fibrogenesis in Response to Injury. J. Biol. Chem. 1998, 273, 33750–33758. [Google Scholar] [CrossRef] [PubMed]
- Halabitska, I.; Babinets, L.; Oksenych, V.; Kamyshnyi, O. Diabetes and Osteoarthritis: Exploring the Interactions and Therapeutic Implications of Insulin, Metformin, and GLP-1-Based Interventions. Biomedicines 2024, 12, 1630. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Lu, K.; Umar, M.; Zhu, Z.; Lu, W.W.; Speakman, J.R.; Chen, Y.; Tong, L.; Chen, D. Risk of Metabolic Abnormalities in Osteoarthritis: A New Perspective to Understand Its Pathological Mechanisms. Bone Res. 2023, 11, 63. [Google Scholar] [CrossRef]
- Gkretsi, V.; Simopoulou, T.; Tsezou, A. Lipid Metabolism and Osteoarthritis: Lessons from Atherosclerosis. Prog. Lipid Res. 2011, 50, 133–140. [Google Scholar] [CrossRef]
- Ma, J.; Chen, L.; Zhu, X.; Li, Q.; Hu, L.; Li, H. Mesenchymal Stem Cell-Derived Exosomal MiR-21a-5p Promotes M2 Macrophage Polarization and Reduces Macrophage Infiltration to Attenuate Atherosclerosis. Acta Biochim. Biophys. Sin. 2021, 53, 1227–1236. [Google Scholar] [CrossRef]
- Zhao, J.; Gu, Y.; Hou, P. Protective Effect and Molecular Mechanism of Mesenchymal Stem Cell-Derived Extracellular Vesicles in Diabetic Foot Ulcers. Cell. Reprogram. 2025, 27, 33–44. [Google Scholar] [CrossRef]


| Gene Name | Forward Primer (5′-3′) | Reverse Primes (3′-5′) |
|---|---|---|
| COLII A1 | ATGACAATCTGGCTCCCAACACTGC | GACCGGCCCTATGTCCACACCGAAT |
| ACAN | TGAGGAGGGCTGGAACAAGTACC | GGAGGTGGTAATTGCAGGGAACA |
| MMP 3 | CCTCAGGAAGCTTGAACCTG | GGAAACCTAGGGTGTGGAT |
| MMP 13 | TGGCATTGCTGACATCATGA | GCCAGAGGGCCCATCAA |
| IL-1β | ATGGACAAGCTGAGGAAGATG | CCTCGTTATCCCATGTGTCG |
| IL-6 | CAACCTGAACCTTCCAAAGATG | ACCTCAAACTCCAAAAGACCAG |
| TNF-A | CCTGAAAACAACCCTCAGA | AAGAGGCTGAGGAACAAGCA |
| GAPDH | GAGTCAACGGATTTGGTCGT | GACAAGCTTCCCGTTCTCAG |
| KLF6 | CAAGGGAAATGGCGATGCCT | CTTTTCTCCTGTGTGCGTCC |
| hsa-miR-21-5p | GCGGCAACACCAGTCGATG | TGCGTGTCGTGGAGTC |
| U6 | GCTTCGGCAGCACATATACTAAAAT | CTCACACCGTGTCGTTCCA |
| Stem-loop primer sequences | ||
| has-miR-21-5p | 5′-GTCGTATCCAGTGCGTGTCGTGGAGTCGGCAATTGCACT-3′ | |
| U6 | 5′-CACGGAAGCCCTCACACCGTGTCGTTC-3′ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Konteles, V.; Papathanasiou, I.; Tzetis, M.; Kriebardis, A.; Tsezou, A. Synovial Fibroblast Extracellular Vesicles Induce Inflammation via Delivering miR-21-5p in Osteoarthritis. Cells 2025, 14, 519. https://doi.org/10.3390/cells14070519
Konteles V, Papathanasiou I, Tzetis M, Kriebardis A, Tsezou A. Synovial Fibroblast Extracellular Vesicles Induce Inflammation via Delivering miR-21-5p in Osteoarthritis. Cells. 2025; 14(7):519. https://doi.org/10.3390/cells14070519
Chicago/Turabian StyleKonteles, Vasileios, Ioanna Papathanasiou, Maria Tzetis, Anastasios Kriebardis, and Aspasia Tsezou. 2025. "Synovial Fibroblast Extracellular Vesicles Induce Inflammation via Delivering miR-21-5p in Osteoarthritis" Cells 14, no. 7: 519. https://doi.org/10.3390/cells14070519
APA StyleKonteles, V., Papathanasiou, I., Tzetis, M., Kriebardis, A., & Tsezou, A. (2025). Synovial Fibroblast Extracellular Vesicles Induce Inflammation via Delivering miR-21-5p in Osteoarthritis. Cells, 14(7), 519. https://doi.org/10.3390/cells14070519








