Knockdown of the Ribosomal Protein eL29 in Mammalian Cells Leads to Significant Changes in Gene Expression at the Transcription Level
Abstract
:1. Introduction
2. Materials and Methods
2.1. Preparation of siRNAs
2.2. Cell Culture and eL29 Knockdown
2.3. Determination of eL29 and rRNA Contents in eL29-Knocked Down Cells
2.4. Estimation of the Effect of eL29 Knockdown on Cellular Surveillance and Proliferation
2.5. Analysis of the Content of Ribosomal Proteins in the Lysate and Polysome Profile Fractions of Knockdown Cells
2.6. RNA-Seq Analysis
2.7. Validation of RNA-Seq Data by Reverse Transcription-qPCR
2.8. Bioinformatic Analysis
3. Results
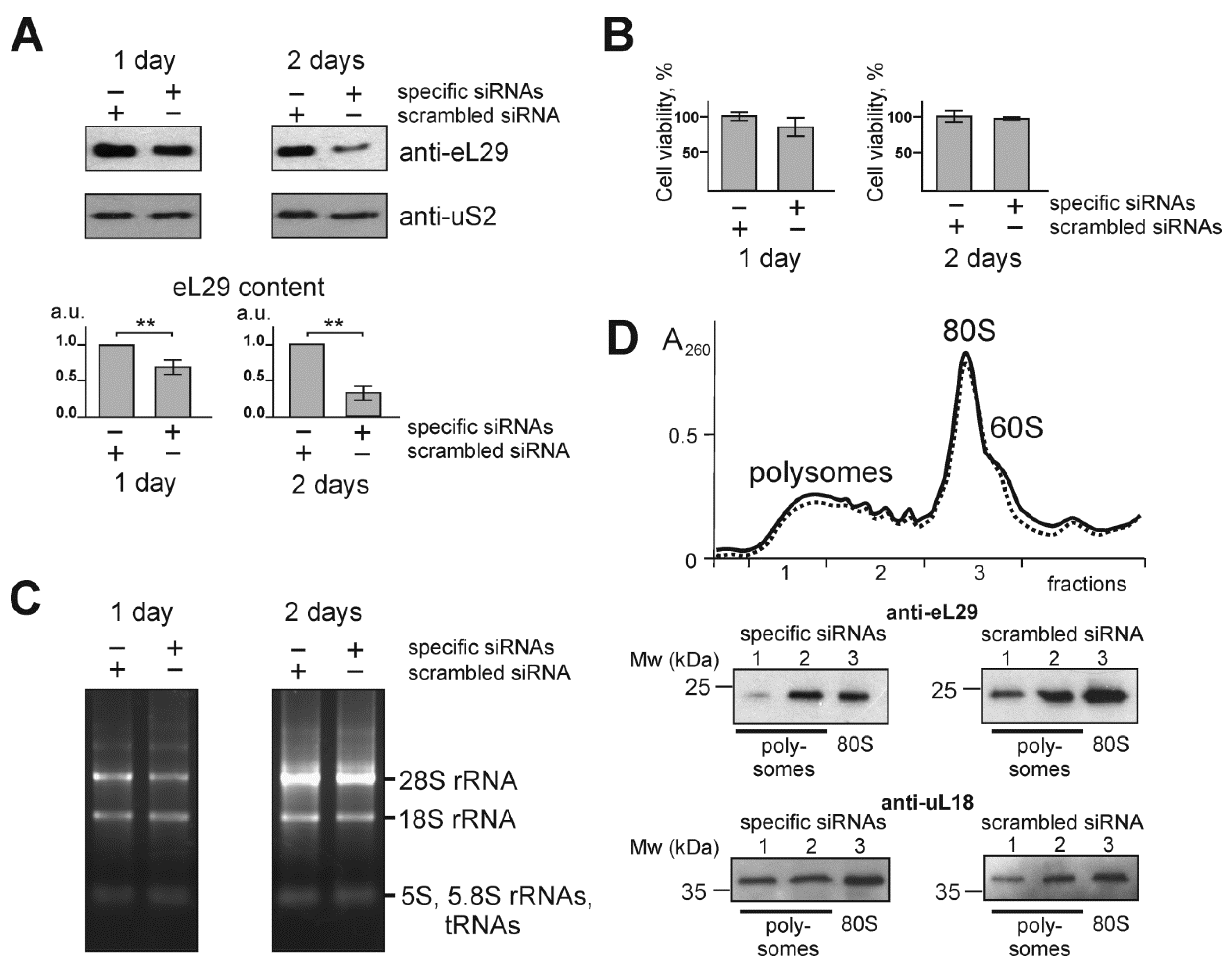
3.1. Knockdown of the Ribosomal Protein eL29 in HEK293 Cells
3.2. RNA-Seq Analysis of eL29-Knocked Down HEK293 Cells
3.3. Cellular Processes Associated with eL29-Dependent DEGs
3.4. DEGs as Targets for p53 and c-Myc
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Yusupova, G.; Yusupov, M. High-resolution structure of the eukaryotic 80S ribosome. Ann. Rev. Biochem. 2014, 83, 467–486. [Google Scholar] [CrossRef] [PubMed]
- Graifer, D.; Karpova, G. Roles of ribosomal proteins in the functioning of translational machinery of eukaryotes. Biochimie 2015, 109, 1–17. [Google Scholar] [CrossRef] [PubMed]
- de la Cruz, J.; Karbstein, K.; Woolford, J.L., Jr. Functions of ribosomal proteins in assembly of eukaryotic ribosomes in vivo. Ann. Rev. Biochem. 2015, 84, 93–129. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gabut, M.; Bourdelais, F.; Durand, S. Ribosome and translational control in stem cells. Cells 2020, 9, 497. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gilles, A.; Frechin, L.; Natchiar, K.; Biondani, G.; Loeffelholz, O.V.; Holvec, S.; Malaval, J.L.; Winum, J.Y.; Klaholz, B.P.; Peyron, J.F. Targeting the human 80S ribosome in cancer: From structure to function and drug design for innovative adjuvant therapeutic strategies. Cells 2020, 9, 629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Robledo, S.; Idol, R.A.; Crimmins, D.L.; Ladenson, J.H.; Mason, P.J.; Bessler, M. The role of human ribosomal proteins in the maturation of rRNA and ribosome production. RNA 2008, 14, 1918–1929. [Google Scholar] [CrossRef] [Green Version]
- Ferreira-Cerca, S.; Pöll, G.; Gleizes, P.E.; Tschochner, H.; Milkereit, P. Roles of eukaryotic ribosomal proteins in maturation and transport of pre-18S rRNA and ribosome function. Mol. Cell 2005, 20, 263–275. [Google Scholar] [CrossRef]
- O’Donohue, M.F.; Choesmel, V.; Faubladier, M.; Fichant, G.; Gleizes, P.E. Functional dichotomy of ribosomal proteins during the synthesis of mammalian 40S ribosomal subunits. J. Cell Biol. 2010, 190, 853–866. [Google Scholar] [CrossRef] [Green Version]
- Landry, D.M.; Hertz, M.I.; Thompson, S.R. RPS25 is essential for translation initiation by the Dicistroviridae and hepatitis C viral IRESs. Genes Dev. 2009, 23, 2753–2764. [Google Scholar] [CrossRef] [Green Version]
- Briones, E.; Briones, C.; Remacha, M.; Ballesta, J.P. The GTPase center protein L12 is required for correct ribosomal stalk assembly but not for Saccharomyces cerevisiae viability. J. Biol. Chem. 1998, 273, 31956–31961. [Google Scholar] [CrossRef] [Green Version]
- Chantrel, Y.; Gaisne, M.; Lions, C.; Verdiere, J. The transcriptional regulator Hap1p (Cyp1p) is essential for anaerobic or heme-deficient growth of Saccharomyces cerevisiae: Genetic and molecular characterization of an extragenic suppressor that encodes a WD repeat protein. Genetics 1998, 148, 559–569. [Google Scholar] [PubMed]
- DeLabre, M.L.; Kessl, J.; Karamanou, S.; Trumpower, B.L. RPL29 codes for a non-essential protein of the 60S ribosomal subunit in Saccharomyces cerevisiae and exhibits synthetic lethality with mutations in genes for proteins required for subunit coupling. Biochim. Biophys. Acta 2002, 1574, 255–261. [Google Scholar] [CrossRef]
- Kim, S.J.; Strich, R. Rpl22 is required for IME1 mRNA translation and meiotic induction in S. cerevisiae. Cell Div. 2016, 11, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, X.; Warner, J.R. Expression of a micro-protein. J. Biol. Chem. 2001, 276, 33821–33825. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mitrovich, Q.M.; Anderson, P. Unproductively spliced ribosomal protein mRNAs are natural targets of mRNA surveillance in C. elegans. Genes Dev. 2000, 14, 2173–2184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoke, D.E.; LaBrenz, S.R.; Hook, M.; Carson, D.D. Multiple domains contribute to heparin/heparan sulfate binding by human HIP/L29. Biochemistry 2000, 39, 15686–15694. [Google Scholar] [CrossRef]
- Chang, B.Y.; Harte, R.A.; Cartwright, C.A. RACK1: A novel substrate for the Src protein-tyrosine kinase. Oncogene 2002, 21, 7619–7629. [Google Scholar] [CrossRef] [Green Version]
- Dresios, J.; Panopoulos, P.; Synetos, D. Eukaryotic ribosomal proteins lacking a eubacterial counterpart: Important players in ribosomal function. Mol. Microbiol. 2006, 59, 1651–1663. [Google Scholar] [CrossRef]
- Raboudi, N.; Julian, J.; Rohde, L.H.; Carson, D.D. Identification of cell-surface heparin/heparan sulfate-binding proteins of a human uterine epithelial cell line (RL95). J. Biol. Chem. 1992, 267, 11930–11939. [Google Scholar]
- Hoke, D.E.; Regisford, E.G.; Julian, J.; Amin, A.; Begue-Kirn, C.; Carson, D.D. Murine HIP/L29 is a heparin-binding protein with a restricted pattern of expression in adult tissues. J. Biol. Chem. 1998, 273, 25148–25157. [Google Scholar] [CrossRef] [Green Version]
- Kirn-Safran, C.B.; Dayal, S.; Martin-DeLeon, P.A.; Carson, D.D. Cloning, expression, and chromosome mapping of the murine Hip/Rpl29 gene. Genomics 2000, 68, 210–219. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Hoke, D.; Julian, J.; Carson, D.D. Heparin/heparan sulfate (HP/HS) interacting protein (HIP) supports cell attachment and selective, high affinity binding of HP/HS. J. Biol. Chem. 1997, 272, 25856–25862. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Zhou, F.; Hook, M.; Carson, D.D. A heparin-binding synthetic peptide of heparin/heparan sulfate-interacting protein modulates blood coagulation activities. Proc. Natl. Acad. Sci. USA 1997, 94, 1739–1744. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, C.; Ge, M.; Yin, Y.; Luo, M.; Chen, D. Silencing expression of ribosomal protein L26 and L29 by RNA interfering inhibits proliferation of human pancreatic cancer PANC-1 cells. Mol. Cell. Biochem. 2012, 370, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.J.; Huang, B.H.; Zhang, J.; Carson, D.D.; Hooi, S.C. Repression of HIP/RPL29 expression induces differentiation in colon cancer cells. J. Cell. Physiol. 2006, 207, 287–292. [Google Scholar] [CrossRef]
- Jones, D.T.; Lechertier, T.; Reynolds, L.E.; Mitter, R.; Robinson, S.D.; Kirn-Safran, C.B.; Hodivala-Dilke, K.M. Endogenous ribosomal protein L29 (RPL29): A newly identified regulator of angiogenesis in mice. Dis. Model. Mech. 2013, 6, 115–124. [Google Scholar] [CrossRef] [Green Version]
- Kirn-Safran, C.B.; Oristian, D.S.; Focht, R.J.; Parker, S.G.; Vivian, J.L.; Carson, D.D. Global growth deficiencies in mice lacking the ribosomal protein HIP/RPL29. Dev. Dyn. 2007, 236, 447–460. [Google Scholar] [CrossRef]
- Oristian, D.S.; Sloofman, L.G.; Zhou, X.; Wang, L.; Farach-Carson, M.C.; Kirn-Safran, C.B. Ribosomal protein L29/HIP deficiency delays osteogenesis and increases fragility of adult bone in mice. J. Orthop. Res. 2009, 27, 28–35. [Google Scholar] [CrossRef]
- Sloofman, L.G.; Verdelis, K.; Spevak, L.; Zayzafoon, M.; Yamauchi, M.; Opdenaker, L.M.; Farach-Carson, M.C.; Boskey, A.L.; Kirn-Safran, C.B. Effect of HIP/ribosomal protein L29 deficiency on mineral properties of murine bones and teeth. Bone 2010, 47, 93–101. [Google Scholar] [CrossRef] [Green Version]
- Aravindan, R.G.; Kirn-Safran, C.B.; Smith, M.A.; Martin-DeLeon, P.A. Ultrastructural changes and asthenozoospermia in murine spermatozoa lacking the ribosomal protein L29/HIP gene. Asian J. Androl. 2014, 16, 925–926. [Google Scholar]
- McGettigan, P.A. Transcriptomics in the RNA-seq era. Curr. Opin. Chem. Biol. 2013, 17, 4–11. [Google Scholar] [CrossRef] [PubMed]
- Hrdlickova, R.; Toloue, M.; Tian, B. RNA-Seq methods for transcriptome analysis. Wiley Interdiscip. Rev. RNA 2017, 8, e1364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cai, R.; Tang, G.; Zhang, Q.; Yong, W.; Zhang, W.; Xiao, J.; Wei, C.; He, C.; Yang, G.; Pang, W. A novel lnc-RNA, named lnc-ORA, is identified by RNA-Seq analysis, and its knockdown inhibits adipogenesis by regulating the PI3K/AKT/mTOR signaling pathway. Cells 2019, 8, e477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Babaylova, E.; Malygin, A.; Gopanenko, A.; Graifer, D.; Karpova, G. Tetrapeptide 60–63 of human ribosomal protein uS3 is crucial for translation initiation. BBA Gene Regul. Mech. 2019, 1862, 194411. [Google Scholar] [CrossRef] [PubMed]
- Leek, J.T.; Scharpf, R.B.; Bravo, H.C.; Simcha, D.; Langmead, B.; Johnson, W.E.; Geman, D.; Baggerly, K.; Irizarry, R.A. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat. Rev. Genet. 2010, 11, 733–739. [Google Scholar] [CrossRef] [Green Version]
- Mi, H.; Muruganujan, A.; Ebert, D.; Huang, X.; Thomas, P.D. PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019, 47, D419–D426. [Google Scholar] [CrossRef]
- Fischer, M. Census and evaluation of p53 target genes. Oncogene 2017, 36, 3943–3956. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.; Lee, J.H.; Iyer, V.R. Global identification of Myc target genes reveals its direct role in mitochondrial biogenesis and its E-box usage in vivo. PLoS ONE 2008, 3, e1798. [Google Scholar] [CrossRef] [Green Version]
- Deisenroth, C.; Franklin, D.A.; Zhang, Y. The evolution of the ribosomal protein-MDM2-p53 pathway. Cold Spring Harb. Perspect. Med. 2016, 6, a026138. [Google Scholar] [CrossRef] [Green Version]
- Zhou, X.; Liao, W.J.; Liao, J.M.; Liao, P.; Lu, H. Ribosomal proteins: Functions beyond the ribosome. J. Mol. Cell Biol. 2015, 7, 92–104. [Google Scholar] [CrossRef] [Green Version]
- Dai, M.S.; Lu, H. Crosstalk between c-Myc and ribosome in ribosomal biogenesis and cancer. J. Cell. Biochem. 2008, 105, 670–677. [Google Scholar] [CrossRef] [Green Version]
- Sulima, S.O.; Kampen, K.R.; De Keersmaecker, K. Cancer biogenesis in ribosomopathies. Cells 2019, 8, e229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Donati, G.; Peddigari, S.; Mercer, C.A.; Thomas, G. 5S ribosomal RNA is an essential component of a nascent ribosomal rrecursor complex that regulates the Hdm2-p53 checkpoint. Cell Rep. 2013, 4, 87–98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koch, M.; Laub, F.; Zhou, P.; Hahn, R.A.; Tanaka, S.; Burgeson, R.E.; Gerecke, D.R.; Ramirez, F.; Gordon, M.K. Collagen XXIV, a vertebrate fibrillar collagen with structural features of invertebrate collagens: Selective expression in developing cornea and bone. J. Biol. Chem. 2003, 278, 43236–43244. [Google Scholar] [CrossRef] [Green Version]
- Matsuo, N.; Tanaka, S.; Gordon, M.K.; Koch, M.; Yoshioka, H.; Ramirez, F. CREB-AP1 protein complexes regulate transcription of the collagen XXIV gene (Col24a1) in osteoblasts. J. Biol. Chem. 2006, 281, 5445–5452. [Google Scholar] [CrossRef] [PubMed] [Green Version]



| GO Term | Definition | Fold Enrichment |
|---|---|---|
| For up-regulated DEGs: | ||
| GO:0006515 | protein quality control for misfolded or incompletely synthesized proteins | 6.54 |
| GO:0034976 | response to endoplasmic reticulum stress | 2.76 |
| GO:0007005 | mitochondrion organization | 2.39 |
| GO:0043087 | regulation of GTPase activity | 2.09 |
| GO:0007346 | regulation of mitotic cell cycle | 2.09 |
| For down-regulated DEGs: | ||
| GO:0006413 | translational initiation | 3.37 |
| GO:0022613 | ribonucleoprotein complex biogenesis | 2.87 |
| GO:0008654 | phospholipid biosynthetic process | 2.70 |
| GO:0051169 | nuclear transport | 2.61 |
| GO:0090407 | organophosphate biosynthetic process | 2.52 |
| GO:0006417 | regulation of translation | 2.48 |
| GO:0031346 | positive regulation of cell projection organization | 2.35 |
| GO:0072657 | protein localization to membrane | 2.17 |
| GO:0016032 | viral process | 2.11 |
| GO:0006396 | RNA processing | 2.08 |
| GO:0055086 | nucleobase-containing small molecule metabolic process | 2.08 |
| GO:0031329 | regulation of cellular catabolic process | 2.07 |
| GO:0007015 | actin filament-based process | 2.03 |
| GO:0080135 | regulation of cellular response to stress | 2.01 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gopanenko, A.V.; Kolobova, A.V.; Meschaninova, M.I.; Venyaminova, A.G.; Tupikin, A.E.; Kabilov, M.R.; Malygin, A.A.; Karpova, G.G. Knockdown of the Ribosomal Protein eL29 in Mammalian Cells Leads to Significant Changes in Gene Expression at the Transcription Level. Cells 2020, 9, 1228. https://doi.org/10.3390/cells9051228
Gopanenko AV, Kolobova AV, Meschaninova MI, Venyaminova AG, Tupikin AE, Kabilov MR, Malygin AA, Karpova GG. Knockdown of the Ribosomal Protein eL29 in Mammalian Cells Leads to Significant Changes in Gene Expression at the Transcription Level. Cells. 2020; 9(5):1228. https://doi.org/10.3390/cells9051228
Chicago/Turabian StyleGopanenko, Alexander V., Alena V. Kolobova, Maria I. Meschaninova, Alya G. Venyaminova, Alexey E. Tupikin, Marsel R. Kabilov, Alexey A. Malygin, and Galina G. Karpova. 2020. "Knockdown of the Ribosomal Protein eL29 in Mammalian Cells Leads to Significant Changes in Gene Expression at the Transcription Level" Cells 9, no. 5: 1228. https://doi.org/10.3390/cells9051228





