Global Regulator PhoP is Necessary for Motility, Biofilm Formation, Exoenzyme Production, and Virulence of Xanthomonas citri Subsp. citri on Citrus Plants
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains, Culture Media, and Growth Conditions
2.2. Biochemical Reagents, Primers and Other Materials
2.3. Gene Knockout and Complementation of phoP Gene
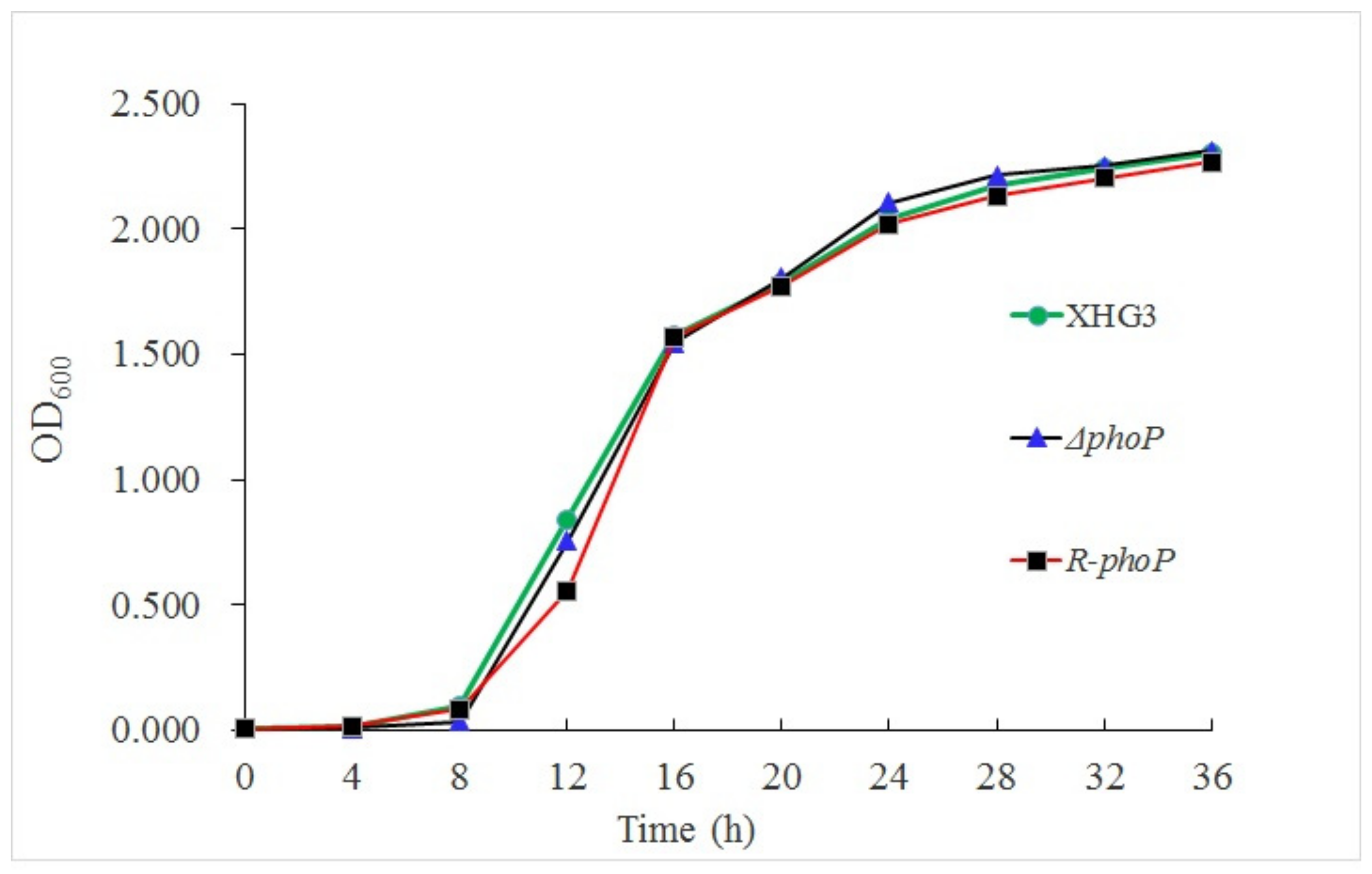
2.4. Bacterial Growth Curve
2.5. Motility and Biofilm-Formation Assay
2.6. Determination of Extracellular Enzymes Activity
2.7. Pathogenicity Assay
2.8. High-throughput RNA Sequencing (RNA-seq) and Data Analysis
2.9. Quantitative Real-time PCR (qRT-PCR) Analysis
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Schaad, N.W.; Postnikova, E.; Lacy, G.; Sechler, A.; Agarkova, L.; Stromberg, P.E.; Stromberg, V.K.; Vidaver, A.K. Emended classification of Xanthomonad pathogens on citrus. Syst. Appl. Microbiol. 2006, 29, 690–695. [Google Scholar] [CrossRef]
- Cubero, J.; Graham, J.H. Genetic relationship among worldwide strains of Xanthomonas causing canker in citrus species and design of new primers for their identification by PCR. Appl. Environ. Microbiol. 2002, 68, 1257–1264. [Google Scholar] [CrossRef] [PubMed]
- Das, A.K. Citrus canker—A review. J. Appl. Hortic. 2003, 5, 52–60. [Google Scholar]
- Graham, J.H.; Gottwald, T.R.; Cubero, J.; Achor, D.S. Xanthomonas axonopodis pv citri: Factors affecting successful eradication of citrus canker. Mol. Plant. Pathol. 2004, 5, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Buttner, D.; Bonas, U. Regulation and secretion of Xanthomonas virulence factors. FEMS Microbiol. Rev. 2010, 34, 107–133. [Google Scholar] [CrossRef]
- Yan, Q.; Wang, N. The ColR/ColS two-component system plays multiple roles in the pathogenicity of the citrus canker pathogen Xanthomonas citri subsp. citri. J. Bacteriol. 2011, 193, 1590–1599. [Google Scholar] [CrossRef]
- Da Silva, A.C.R.; Ferro, J.A.; Reinach, F.C.; Farah, C.S.; Furlan, L.R.; Quaggio, R.B.; Monteiro-Vitorello, C.B.; Van Sluys, M.A.; Almeida, N.F.; Alves, L.M.C.; et al. Comparison of the genomes of two Xanthomonas pathogens with differing host specificities. Nature 2002, 417, 459–463. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.P.; Figueiredo, F.; Jones, J.; Wang, N. HrpG and HrpX play global roles in coordinating different virulence traits of Xanthomonas axonopodis pv. citri. Mol. Plant. Microbe Inter. 2011, 24, 649–661. [Google Scholar] [CrossRef]
- Dunger, G.; Garofalo, C.G.; Gottig, N.; Garavaglia, B.S.; Rosa, M.C.P.; Farah, C.S.; Orellano, E.G.; Ottado, J. Analysis of three Xanthomonas axonopodis pv. citri effector proteins in pathogenicity and their interactions with host plant proteins. Mol. Plant. Pathol. 2012, 13, 865–876. [Google Scholar] [CrossRef] [PubMed]
- Sgro, G.G.; Ficarra, F.A.; Dunger, G.; Scarpec, T.E.; Valle, E.M.; Cortadi, A.; Orellano, E.G.; Gottig, N.; Ottado, J. Contribution of a harpin protein from Xanthomonas axonopodis pv. citri to pathogen virulence. Mol. Plant. Pathol. 2012, 13, 1047–1059. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Guo, J.; Ma, W.X.; Ji, Z.Y.; Zou, L.F.; Chen, G.Y.; Zou, H.S. Identification of seven novel virulence genes from Xanthomonas citri subsp. citri by Tn5-based random mutagenesis. J. Microbiol. 2015, 53, 330–336. [Google Scholar] [CrossRef] [PubMed]
- Xia, T.; Li, Y.J.; Sun, D.L.; Zhuo, T.; Fan, X.J.; Zou, H.S. Identification of an extracellular endoglucanase that is required for full virulence in Xanthomonas citri subsp. citri. Plos ONE 2016, 11, e0151017. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Castaneda, A.; Zhao, G.; Erdos, G.; Gabriel, D. Expression of a single, host-specific, bacterial pathogenicity gene in plant cells elicits division, enlargement, and cell death. Mol. Plant. Microbe Interact. 1999, 12, 556–560. [Google Scholar] [CrossRef]
- Guo, Y.P.; Sagaram, U.S.; Kim, J.S.; Wang, N. Requirement of the galU gene for polysaccharide production by and pathogenicity and growth in planta of Xanthomonas citri subsp. citri. Appl. Environ. Microbiol. 2010, 76, 2234–2242. [Google Scholar] [CrossRef] [PubMed]
- Moreira, L.M.; Almeida, N.F.; Potnis, N.; Digiampietri, L.A.; Adi, S.S.; Bortolossi, J.C.; da Silva, A.C.; da Silva, A.M.; de Moraes, F.E.; de Oliveira, J.C.; et al. Novel insights into the genomic basis of citrus canker based on the genome sequences of two strains of Xanthomonas fuscans subsp. aurantifolii. BMC Genomics 2010, 11, 238. [Google Scholar] [CrossRef] [PubMed]
- Groisman, E.A. The pleiotropic two-component regulatory system PhoP/PhoQ. J. Bacteriol. 2001, 183, 1835–1842. [Google Scholar] [CrossRef] [PubMed]
- Derzelle, S.; Turlin, E.; Duchaud, E.; Pages, S.; Kunst, F.; Givaudan, A.; Danchin, A. The PhoP/PhoQ two-component regulatory system of Photorhabdus luminescens is essential for virulence in insects. J. Bacteriol. 2004, 186, 1270–1279. [Google Scholar] [CrossRef] [PubMed]
- Peng, B.Y.; Pan, Y.; Li, R.J.; Wei, J.W.; Liang, F.; Wang, L.; Wang, F.F.; Qian, W. An essential regulatory system originating from polygenic transcriptional rewiring of PhoP-PhoQ of Xanthomonas campestris. Genetics 2017, 206, 2207–2223. [Google Scholar] [CrossRef]
- Bader, M.W.; Sanowar, S.; Daley, M.E.; Schneider, A.R.; Cho, U.S.; Xu, W.Q.; Klevit, R.E.; Le Moual, H.; Miller, S. Recognition of antimicrobial peptides by a bacterial sensor kinase. Cell 2005, 122, 461–472. [Google Scholar] [CrossRef] [PubMed]
- Moss, J.E.; Fisher, P.E.; Vick, B.; Groisman, E.A.; Zychilinsky, A. The regulatory protein PhoP controls susceptibility to the host inflammatory response in Shigella flexneri. Cell. Microbiol. 2000, 2, 443–452. [Google Scholar] [CrossRef]
- Oyston, P.C.F.; Dorrell, N.; Williams, K.; Li, S.R.; Green, M.; Titball, R.W.; Wren, B.W. The response regulator PhoP is important for survival under conditions of macrophage-induced stress and virulence in Yersinia pestis. Infect. Immun. 2000, 68, 3419–3425. [Google Scholar] [CrossRef]
- Venkatesh, B.; Babujee, L.; Liu, H.; Hedley, P.; Fujikawa, T.; Birch, P.; Toth, I.; Tsuyumu, S. The Erwinia chrysanthemi 3937 PhoQ sensor kinase regulates several virulence determinants. J. Bacteriol. 2006, 188, 3088–3098. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.W.; Jeong, K.S.; Han, S.W.; Lee, S.E.; Phee, B.K.; Hahn, T.R.; Ronald, P. The Xanthomonas oryzae pv. Oryzae PhoPQ two-component system is required for AvrXA21 activity, hrpG expression, and virulence. J. Bacteriol. 2008, 190, 2183–2197. [Google Scholar] [CrossRef] [PubMed]
- Sriariyanun, M.; Seo, Y.S.; Phetsom, J. Identification of PhoP regulon members in Xanthomonas oryzae pv. oryzae PXO99A. In 4th International Conference on Chemical, Biological And Environmental Engineering; IACSIT Press: Singapore, 2012. [Google Scholar]
- Perez, J.C.; Shin, D.; Zwir, I.; Latifi, T.; Hadley, T.J.; Groisman, E.A. Evolution of a bacterial regulon controlling virulence and Mg2+Homeostasis. PLoS Genet. 2009, 5, e1000428. [Google Scholar] [CrossRef] [PubMed]
- Zwir, I.; Latifi, T.; Perez, J.C.; Huang, H.; Groisman, E.A. The promoter architectural landscape of the Salmonella PhoP regulon. Mol. Microbiol. 2012, 84, 463–485. [Google Scholar] [CrossRef] [PubMed]
- Monsieurs, P.; De Keersmaecker, S.; Navarre, W.W.; Bader, M.W.; De Smet, F.; Mcclelland, M.; Fang, F.C.; De Moor, B.; Vanderleyden, J.; Marchal, K. Comparison of the PhoPQ regulon in Escherichia coli and Salmonella typhimurium. J. Mol. Evol. 2005, 60, 462–474. [Google Scholar] [CrossRef]
- Rigano, L.A.; Siciliano, F.; Enrique, R.; Sendin, L.; Filippone, P.; Torres, P.S.; Questa, J.; Dow, J.M.; Castagnaro, A.P.; Vojnov, A.A.; et al. Biofilm formation, epiphytic fitness, and canker development in Xanthomonas axonopodis pv. citri. Mol. Plant. Microbe Inter. 2007, 20, 1222–1230. [Google Scholar] [CrossRef]
- Shen, Y.W.; Chern, M.; Silva, F.G.; Ronald, P. Isolation of a Xanthomonas oryzae pv. oryzae flagellar operon region and molecular characterization of flhF. Mol. Plant. Microbe Inter. 2001, 14, 204–213. [Google Scholar] [CrossRef]
- Tang, J.L.; Liu, Y.N.; Barber, C.E.; Dow, J.M.; Wootton, J.C.; Daniels, M.J. Genetic and molecular analysis of a cluster of rpf genes involved in positive regulation of synthesis of extracellular enzymes and polysaccharide in Xanthomonas campestris pathovar campestris. Mol. Gen. Genet. 1991, 226, 409–417. [Google Scholar] [CrossRef]
- Rybak, M.; Minsavage, G.V.; Stall, R.E.; Jones, J.B. Identification of Xanthomonas citri ssp. citri host specificity genes in a heterologous expression host. Mol. Plant. Pathol. 2009, 10, 249–262. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Southey-Pillig, C.J.; Davies, D.G.; Sauer, K. Characterization of temporal protein production in Pseudomonas aeruginosa biofilms. J. Bacteriol. 2005, 187, 8114–8126. [Google Scholar] [CrossRef]
- Branda, S.S.; Vik, A.; Friedman, L.; Kolter, R. Biofilms: The matrix revisited. Trends Microbiol. 2005, 13, 20–26. [Google Scholar] [CrossRef] [PubMed]
- Gottig, N.; Garavaglia, B.S.; Daurelio, L.D.; Valentine, A.; Gehring, C.; Orellano, E.G.; Ottado, J. Xanthomonas axonopodis pv. citri uses a plant natriuretic peptide-like protein to modify host homeostasis. Proc. Natl. Acad. Sci. USA. 2008, 5, 18631–18636. [Google Scholar] [CrossRef] [PubMed]
- Gehring, C.A.; Irving, H.R. Natriuretic peptides—A class of heterologous molecules in plants. Inter. J. Biochem. Cell Biol. 2003, 35, 1318–1322. [Google Scholar] [CrossRef]
- Ludidi, N.; Morse, M.; Sayed, M.; Wherrett, T.; Shabala, S.; Gehring, C. A recombinant plant natriuretic peptide causes rapid and spatially differentiated K+, Na+ and H+ flux changes in Arabidopsis thaliana roots. Plant. Cell Physiol. 2004, 45, 1093–1098. [Google Scholar] [CrossRef] [PubMed]
- Bijlsma, J.J.E.; Groisman, E.A. The PhoP/PhoQ system controls the intramacrophage type three secretion system of Salmonella enterica. Mol. Microbiol. 2005, 57, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Adams, P.; Fowler, R.; Kinsella, N.; Howell, G.; Farris, M.; Coote, P.; O’Connor, C.D. Proteomic detection of PhoPQ- and acid-mediated repression of Salmonella motility. Proteomics 2001, 1, 597–607. [Google Scholar] [CrossRef]
- Matilla, M.A.; Krell, T. The effect of bacterial chemotaxis on host infection and pathogenicity. FEMS Microbiol. Rev. 2017, 42, fux052. [Google Scholar] [CrossRef]
- Kim, T.J.; Young, B.M.; Young, G.M. Effect of flagellar mutations on Yersinia enterocolitica biofilm formation. Appl. Environ. Microbiol. 2008, 74, 5466–5474. [Google Scholar] [CrossRef]
- Liang, Y.L.; Gao, H.C.; Chen, J.R.; Dong, Y.Y.; Wu, L.; He, Z.L.; Liu, X.D.; Qiu, G.Z.; Zhou, J.Z. Pellicle formation in Shewanella oneidensis. BMC Microbiol. 2010, 10, 291. [Google Scholar] [CrossRef]
- Dow, J.M.; Crossman, L.; Findlay, K.; He, Y.Q.; Feng, J.X.; Tang, J.L. Biofilm dispersal in Xanthomonas campestris is controlled by cell–cell signaling and is required for full virulence to plants. Proc. Natl. Acad. Sci. USA 2003, 100, 10995–11000. [Google Scholar] [CrossRef] [PubMed]
- Malamud, F.; Torres, P.S.; Roeschlin, R.; Rigano, L.A.; Enrique, R.; Bonomi, H.R.; Castagnaro, A.P.; Marano, M.R.; Vojnov, A.A. The Xanthomonas axonopodis pv. citri flagellum is required for mature biofilm and canker development. Microbiology 2011, 157, 819–829. [Google Scholar] [CrossRef]
- Kumar Verma, R.; Samal, B.; Chatterjee, S. Xanthomonas oryzae pv. oryzae chemotaxis components and chemoreceptor Mcp2 are involved in the sensing of constituents of xylem sap and contribute to the regulation of virulence-associated functions and entry into rice. Mol. Plant. Pathol. 2018, 19, 2397–2415. [Google Scholar] [CrossRef]
- Dunger, G.; Arabolaza, A.L.; Gottig, N.; Orellano, E.G.; Ottado, J. Participation of Xanthomonas axonopodis pv. citri hrp cluster in citrus canker and nonhost plant responses. Plant. Pathology. 2005, 54, 781–788. [Google Scholar] [CrossRef]
- Baptista, J.C.; Machado, M.A.; Homem, R.A.; Torres, P.S.; Vojnov, A.A.; do Amaral, A.M. Mutation in the xpsD gene of Xanthomonas axonopodis pv. citri affects cellulose degradation and virulence. Genet. Mol. Biol. 2010, 33, 146–153. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Wang, N. High-throughput screening and analysis of genes of Xanthomonas citri subsp. citri involved in citrus canker symptom development. Mol. Plant. Microbe Inter. 2012, 25, 69–84. [Google Scholar] [CrossRef]
- Matsumoto, H.; Jitareerat, P.; Baba, Y.; Tsuyumu, S. Comparative study of regulatory mechanisms for pectinase production by Erwinia carotovora subsp carotovora and Erwinia chrysanthemi. Mol. Plant. Microbe Inter. 2003, 16, 226–237. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Calva., E.; Oropeza, R. Two-component signal transduction systems, environmental signals, and virulence. Microbial. Ecol. 2006, 51, 166–176. [Google Scholar] [CrossRef]
- Alfano, J.R.; Collmer, A. Bacterial pathogens in plants: Life up against the wall. Plant. Cell. 1996, 8, 1683–1698. [Google Scholar] [CrossRef]
- Segura, A.; Moreno, M.; Madueno, F.; Molina, A.; Garcia-Olmedo, F. Snakin-1, a peptide from potato that is active against plant pathogens. Mol. Plant. Microbe Inter. 1999, 12, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Osbourn, A.E. Preformed antimicrobial compounds and plant defense against fungal attack. Plant. Cell. 1996, 8, 1821–1831. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.H.; Xu, C.G.; Yang, Y.B.; Xing, X.X.; Liu, X.; Qu, Q.W.; Ding, W.Y.; Bello-Onaghise, G.; Li, Y.H. Histidine metabolism and IGPD play a key role in cefquinome inhibiting biofilm formation of Staphylococcus xylosus. Front. Microbiol. 2018, 9, 665. [Google Scholar] [CrossRef] [PubMed]
- Matas, I.M.; Lambertsen, L.; Rodrguez-Moreno, L.; Ramos, C. Identification of novel virulence genes and metabolic pathways required for full fitness of Pseudomonas savastanoi pv. savastanoi in olive (Olea europaea) knots. New Phytologist. 2012, 196, 1182–1196. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.Y.; Yu, X.Y.; Pan, X.Y.; Wu, J.; Duan, Y.B.; Wang, J.X.; Zhou, M.G. A thiadiazole reduces the virulence of Xanthomonas oryzae pv. oryzae by inhibiting the histidine utilization pathway and quorum sensing. Mol. Plant Pathol. 2018, 19, 16–128. [Google Scholar] [CrossRef] [PubMed]
- Christie, P.J. Type IV secretion: Intercellular transfer of macromolecules by systems ancestrally related to conjugation machines. Mol. Microbiol. 2001, 40, 294–305. [Google Scholar] [CrossRef]
- Zhao, Z.M.; Sagulenko, E.; Ding, Z.Y.; Christie, P.J. Activities of virE1 and the VirE1 secretion chaperone in export of the multifunctional VirE2 effector via an Agrobacterium type IV secretion pathway. J. Bacteriol. 2001, 183, 3855–3865. [Google Scholar] [CrossRef]
- Bell, K.S.; Sebaihia, M.; Pritchard, L.; Holden, M.T.G.; Hyman, L.J.; Holeva, M.C.; Thomson, N.R.; Bentley, S.D.; Churcher, L.J.C.; Mungall, K.; et al. Genome sequence of the enterobacterial phytopathogen Erwinia carotovora subsp. atroseptica and characterization of virulence factors. Proc. Natl. Acad. Sci. USA 2004, 101, 11105–11110. [Google Scholar] [CrossRef]







| Strain and Plasmid | Relevant Characteristics | Origin |
|---|---|---|
| X. citri subsp. citri | ||
| XHG3 | Wild-type strain | This study |
| ΔphoP | phoP knock-out mutant of XHG3 | This study |
| R-phoP | complementation strain of ΔphoP | This study |
| E. coli | ||
| K12 CC118 | gyrA,recA,λ pir | Lab collection |
| DH5α | deoR,recA,endA,hsdR,supE,thi,gyrA,relA | Lab collection |
| Plasmids | ||
| pKNG101 | SmR,SacB,mobRK2,oriR6K (pir-minus) | Lab collection |
| pKNG-ΔphoP | PhoP knock-out fragment ligated on pKNG101 | This study |
| pBBR1- phoP | PhoP complementation fragment ligated on pBBR1MCS4 | This study |
| Gene | Locus tag | log2.Fold_change. | q-value |
|---|---|---|---|
| hrpF | XAC0394 | −1.77 | 6.33 × 10−14 |
| hpaB | XAC0396 | −3.20 | 0 |
| hrpE | XAC0397 | −5.16 | 7.95 × 10−4 |
| hrpD6 | XAC0398 | −1.95 | 8.94 × 10−7 |
| hrpD5 | XAC0399 | −2.82 | 1.82 × 10−5 |
| hpaA | XAC0400 | −2.46 | 2.68 × 10−3 |
| hrcS | XAC0401 | −3.86 | 1.11 × 10−17 |
| hrcR | XAC0403 | −4.39 | 2.06 × 10−16 |
| hrcQ | XAC0404 | −4.26 | 1.14 × 10−12 |
| hrcV | XAC0405 | −2.03 | 7.26 × 10−12 |
| hrcU | XAC0406 | −2.99 | 8.22 × 10−28 |
| hrpB1 | XAC0407 | −4.21 | 3.45 × 10−12 |
| hrpB2 | XAC0408 | −4.02 | 6.59 × 10−20 |
| hrcJ | XAC0409 | −4.48 | 1.95 × 10−12 |
| hrpB4 | XAC0410 | −3.03 | 6.46 × 10−7 |
| hrpB5 | XAC0411 | −4.05 | 6.51 × 10−9 |
| hrcN | XAC0412 | −3.08 | 2.02 × 10−4 |
| hrpB7 | XAC0413 | −3.76 | 2.94 × 10−6 |
| hrcT | XAC0414 | −2.85 | 5.01 × 10−7 |
| hrcC | XAC0415 | −1.69 | 3.18 × 10−183 |
| hpa1 | XAC0416 | −-6.06 | 6.33 × 10−14 |
| Effector family | Locus tag | log2.Fold_change. | q-value |
|---|---|---|---|
| AvrBs2 | XAC0076 | −1.82 | 5.36 × 10−8 |
| AvrBs3(PthA1) | XACa002 | - | - |
| AvrBs3(PthA2) | XACa0039 | - | - |
| AvrBs3(PthA3) | XACb0015 | - | - |
| AvrBs3(PthA4) | XACb0065 | - | - |
| HrpW (PopW) | XAC2922 | −2.52 | 2.41 × 10−55 |
| XopAD (Skwp, RSc3401) | XAC4213 | −2.13 | 2.92 × 10−44 |
| XopAE (HpaF/G/PopC) | XAC0393 | N | N |
| XopAI (HopO1 (HopPto, HopPtoS), HopAI1 (HolPtoAI)) | XAC3230 | −1.99 | 2.05 × 10−26 |
| XopAK (HopAK1 (HopPtoK, HolPtoAB) C terminal domain | XAC3666 | −2.05 | 2.38 × 10−7 |
| XopE1 (AvrXacE1, HopX, AvrPPhE) | XAC0286 | −1.79 | 1.61 × 10−17 |
| XopE2 (AvrXacE3, AvrXccE1) | XACb0011 | N | N |
| XopE3 (AvrXacE2, HopX, AvrPPhE) | XAC3224 | −1.11 | 2.24 × 10−3 |
| XopF2 | XAC2785 | N | N |
| XopI | XAC0754 | −2.35 | 1.60 × 10−7 |
| XopK | XAC3085 | −3.20 | 1.40 × 10−32 |
| XopL | XAC3090 | N | N |
| XopN (HopAU1) | XAC2786 | −2.28 | 6.35 × 10−23 |
| XopP | XAC1208 | −1.24 | 8.87 × 10−4 |
| XopQ (HopQ1) | XAC4333 | −3.84 | 5.58 × 10−19 |
| XopR | XAC0277 | −2.61 | 4.79 × 10−25 |
| XopV | XAC0601 | N | N |
| XopX (HolPsyAE) | XAC0543 | −2.79 | 7.16 × 10−51 |
| XopZ (HopAS, AWR) | XAC2009XAC2990 | −2.35N | 6.46 × 10−19N |
| Gene | Locus tag | log2.Fold_change. | q-value |
|---|---|---|---|
| virB4 | XAC2614 | 2.34 | 1.57 × 10−258 |
| virB1 | XAC2617 | 2.90 | 8.15 × 10−188 |
| virB11 | XAC2618 | 2.18 | 5.55 × 10−99 |
| virB10 | XAC2619 | 2.19 | 6.58 × 10−139 |
| virB8 | XAC2621 | 1.86 | 2.10 × 10−93 |
| virD4 | XAC2623 | 1.94 | 1.22 × 10−115 |
| XAC0096 | XAC0096 | 2.99 | 6.06 × 10−134 |
| XAC1918 | XAC1918 | 1.62 | 1.14 × 10−12 |
| XAC2609 | XAC2609 | 1.77 | 2.55 × 10−135 |
| XAC2610 | XAC2610 | 1.69 | 3.98 × 10−128 |
| phlA | XAC2885 | 2.04 | 1.60 × 10−42 |
| XAC0323 | XAC0323 | 2.12 | 2.22 × 10−26 |
| HI | XAC0466 | 2.22 | 8.25 × 10−106 |
| Cw1L | XAC3634 | 2.1 | 9.38 × 10−43 |
| Gene_id | log2.Fold_change | P-value | Gene Description |
|---|---|---|---|
| XAC2447 | −2.7359 | 1.47 × 10−4 | Chemotaxis protein CheW/cheW |
| XAC2448 | −2.908 | 2.51 × 10−16 | Methyl-accepting chemotaxis protein I/tsr |
| XAC3213 | −1.4002 | 4.098 × 10−3 | Methyl-accepting chemotaxis protein I/GN=tsr |
| XAC2866 | −1.6455 | 0 | Methyl-accepting chemotaxis citrate transducer/tcp |
| XAC1894 | −1.6016 | 9.78 × 10−5 | Methyl-accepting chemotaxis protein I/tsr |
| XAC1899 | −1.257 | 2.948 × 10−3 | Methyl-accepting chemotaxis protein I/tsr |
| XAC1892 | −1.9605 | 2.22 × 10−5 | Methyl-accepting chemotaxis protein I/tsr |
| XAC1895 | 1.37 | 2.06 × 10−8 | Methyl-accepting chemotaxis serine transducer/ |
| XAC1930 | −2.3919 | 9.80 × 10−19 | Chemotaxis protein CheA/cheA |
| XAC1931 | −2.147 | 6.31 × 10−9 | Protein phosphatase CheZ/cheZ |
| XAC1932 | −2.0968 | 3.54 × 10−8 | Chemotaxis protein CheY/cheY |
| XAC1933 | −2.6854 | 7.24 × 10−12 | RNA polymerase sigma factor for flagellar operon/fliA |
| XAC1934 | −2.3675 | 3.02 × 10−10 | Flagellum site-determining protein YlxH/ylxH |
| XAC1935 | −4.4217 | 6.82 × 10−24 | Flagellar biosynthesis protein FlhF/flhF |
| XAC1936 | −2.9695 | 6.93 × 10−18 | Flagellar biosynthesis protein FlhA/flhA |
| XAC1937 | −3.0715 | 2.47 × 10−5 | Flagellar biosynthetic protein FlhB/flhB |
| XAC1938 | −1.8277 | 1.47 × 10−5 | Uncharacterized signaling protein PA1727/PA1727//0 |
| XAC1940 | −2.2302 | 9.73 × 10−10 | Uncharacterized signaling protein CC_0091/GN=CC_0091 |
| XAC1941 | −2.2199 | 1.924 × 10−3 | Flagellar biosynthetic protein FliR/fliR |
| XAC1942 | −2.7865 | 3.83 × 10−4 | -//- |
| XAC1944 | −1.7605 | 3.91 × 10−9 | Flagellar biosynthetic protein FliP/fliP |
| XAC1945 | −3.5139 | 1.58 × 10−6 | -//- |
| XAC1946 | −3.9225 | 0.853 × 10−3 | Flagellar motor switch protein FliN/fliN |
| XAC1947 | −3.492 | 1.35 × 10−8 | Flagellar motor switch protein FliM/fliM |
| XAC1948 | −2.9248 | 1.76 × 10−8 | Flagellar protein FliL/GN=fliL |
| XAC1949 | −2.264 | 1.97 × 10−26 | -//- |
| XAC1950 | −1.583 | 8.74 × 10−4 | -//- |
| XAC1951 | −2.256 | 1.43 × 10−17 | Flagellum-specific ATP synthase/fliI |
| XAC1952 | −2.2921 | 7.37 × 10−9 | -//- |
| XAC1953 | −1.5637 | 1.43 × 10−8 | Flagellar motor switch protein FliG/fliG |
| XAC1954 | −2.7658 | 7.29 × 10−20 | Flagellar M-ring protein/fliF |
| XAC1955 | −4.3033 | 2.79 × 10−6 | Flagellar hook-basal body complex protein FliE/fliE |
| XAC1969 | −1.3085 | 4.57 × 10−35 | RNA polymerase sigma-54 factor/rpoN |
| XAC1975 | −1.699 | 1.03 × 10−45 | A-type flagellin/fliC |
| XAC1976 | −2.816 | 7.13 × 10−19 | Flagellar hook-associated protein/flgL |
| XAC1977 | −3.0454 | 1.74 × 10−23 | Flagellar hook-associated protein/flgK |
| XAC1978 | −3.1714 | 9.31 × 10−16 | Peptidoglycan hydrolase FlgJ/flgJ |
| XAC1979 | −3.7131 | 1.32 × 10−14 | Flagellar P-ring protein/flgI |
| XAC1980 | −2.9579 | 1.89 × 10−8 | Flagellar L-ring protein/flgH |
| XAC1981 | −3.7208 | 9.54 × 10−24 | Flagellar basal-body rod protein FlgG/flgG |
| XAC1982 | −2.1116 | 1.26 × 10−8 | Flagellar basal-body rod protein FlgF/flgF |
| XAC1983 | −2.751 | 2.42 × 10−22 | Flagellar hook protein FlgE/flgE |
| XAC1984 | −3.135 | 3.25 × 10−11 | Basal-body rod modification protein FlgD/flgD |
| XAC1985 | −3.1884 | 2.24 × 10−4 | Flagellar basal-body rod protein FlgC/flgC |
| XAC1986 | −3.7208 | 1.21 × 10−12 | Flagellar basal body rod protein FlgB/flgB |
| XAC1988 | −2.8726 | 1.54 × 10−6 | -//- |
| Gene_id | log2.Fold_change | P-value | Gene Description |
|---|---|---|---|
| XAC1637 | −1.4138 | 4.91 × 10−7 | HutH/Histidine ammonia-lyase |
| XAC1635 | −1.8604 | 9.51 × 10−8 | HutU/Urocanate hydratase |
| XAC1834 | −2.8393 | 3.05 × 10−17 | HisF/Imidazole glycerol phosphate synthase subunit |
| XAC1833 | −2.906 | 2.31 × 10−20 | HisA/1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino] imidazole-4-carboxamide isomerase |
| XAC1835 | −2.9827 | 3.98 × 10−21 | HisIE/Histidine biosynthesis bifunctional protein |
| XAC1832 | −3.343 | 3.05 × 10−13 | HisH/Imidazole glycerol phosphate synthase subunit |
| XAC1831 | −3.3447 | 5.44 × 10−70 | HisB/Histidine biosynthesis bifunctional protein |
| XAC1829 | −3.4986 | 8.67 × 10−60 | HisD/Histidinol dehydrogenase |
| XAC1830 | −3.506 | 3.44 × 10−40 | HisC/Histidinol-phosphate aminotransferase |
| XAC1828 | −3.7577 | 7.18 × 10−43 | HisG/ATP phosphoribosyltransferase |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wei, C.; Ding, T.; Chang, C.; Yu, C.; Li, X.; Liu, Q. Global Regulator PhoP is Necessary for Motility, Biofilm Formation, Exoenzyme Production, and Virulence of Xanthomonas citri Subsp. citri on Citrus Plants. Genes 2019, 10, 340. https://doi.org/10.3390/genes10050340
Wei C, Ding T, Chang C, Yu C, Li X, Liu Q. Global Regulator PhoP is Necessary for Motility, Biofilm Formation, Exoenzyme Production, and Virulence of Xanthomonas citri Subsp. citri on Citrus Plants. Genes. 2019; 10(5):340. https://doi.org/10.3390/genes10050340
Chicago/Turabian StyleWei, Chudan, Tian Ding, Changqing Chang, Chengpeng Yu, Xingwei Li, and Qiongguang Liu. 2019. "Global Regulator PhoP is Necessary for Motility, Biofilm Formation, Exoenzyme Production, and Virulence of Xanthomonas citri Subsp. citri on Citrus Plants" Genes 10, no. 5: 340. https://doi.org/10.3390/genes10050340
APA StyleWei, C., Ding, T., Chang, C., Yu, C., Li, X., & Liu, Q. (2019). Global Regulator PhoP is Necessary for Motility, Biofilm Formation, Exoenzyme Production, and Virulence of Xanthomonas citri Subsp. citri on Citrus Plants. Genes, 10(5), 340. https://doi.org/10.3390/genes10050340





