1. Introduction
Genomic selection (GS) uses genome-wide marker information to predict the genomic estimated breeding values (GEBV) of individuals [
1]. In GS, a population with known phenotype and genotype information (training population) is used to predict the GEBV of individuals having only genotype data (test or validation population) [
2]. GEBV can be then used as a basis for selecting which lines to advance and/or be used as parents in the breeding program. The utility of GEBV for choosing parents for the next generation of crossing, together with the introgression of favorable alleles, can be used to fast-forward genetic gain [
3]. The performance of GS depends on predictive ability, defined as the correlation between GEBV and observed phenotype [
4,
5]. The predictive power of GS is a factor of genetic relatedness between the training and test populations, prediction models used, number of markers, population structure, including secondary correlated traits in the model, among others [
6,
7,
8,
9,
10].
In recent years, recommender algorithms have been developed to streamline the selection process in electronic commerce (e-commerce) based on the customer’s preferences through generating a list of recommended items [
11,
12]. Item-based collaborative filtering (IBCF) is a popular recommender approach for suggesting which items to buy [
13]. In this process, relationships between the different items are first identified, and this information is subsequently used to compute recommendations, where a database of preferences for users is ultimately established [
11]. In an IBCF recommender system, a set of items that has been rated (i.e., selected or bought) by a specific user
ui is first evaluated and compared to the target item
Y; after which, the most similar items to
Y are selected. The similarity coefficients between the most similar items are then computed, and once the most similar items are identified, prediction for the
ui and the target item
Y is calculated through a simple weighted average of the similar items [
13]. The IBCF recommender system has been recently used in the context of genomic selection (GS) to predict performance of wheat lines in multi-trait, multi-environment trials [
13,
14,
15]. In an IBCF-GS approach, phenotypic data for each line
i for every trait–environment combination
j is first built. Afterward, raw phenotypic data from each line is standardized using the mean and standard deviation for each
j. Using the correlation (similarity) coefficients between each
j, scaled predictive phenotype for each
i on a trait–environment combination, and the adjusted trait values, predicted values for the missing phenotypes are calculated [
16].
A key to the successful evaluation and implementation of GS is an improved predictive ability for target traits [
13]. Hence, to make GS more relevant in the breeding program, it is necessary to explore alternative models which could potentially result in improved predictive ability. As more empirical evidence is required to get a clearer picture of the performance of the IBCF recommender approach [
16], it would be essential to assess how this system would be useful in the context of GS in a winter wheat breeding program. Therefore, the objectives of this study were to (1) evaluate the potential of using an IBCF recommender system for GS of different traits in winter wheat; (2) predict grain yield using secondary spectral traits from high-throughput phenotyping using an IBCF approach; and (3) compare the predictive ability of IBCF with a univariate genomic BLUP (GBLUP) prediction model.
4. Discussion
The potential of a multi-trait, multi-environment IBCF recommender system for GS was assessed across different traits for winter wheat lines evaluated in US Pacific Northwest growing conditions. In the current study, we demonstrated the successful implementation of this approach in the context of a winter wheat breeding program, particularly in using genomic marker information for trait adjustments and improving the predictive ability of GS for yield, agronomic, and disease resistance traits.
Predicting a trait for a single year using combined information from multiple years would be the most effective strategy when predicting across years using the IBCF approach. It was observed that the correlation between years played an important role in achieving optimal cross-validation accuracies. Predicting performance of single years for GY in LND (average correlation
r between environments of 0.21) resulted in higher mean predictive ability compared to PUL (mean
r of 0.08) (0.16 vs. 0.03). The correlation between environments could therefore be regarded as the major driver for increased predictive ability across years. Likewise, PH, which had higher mean correlations between environments (
r = 0.11), had the highest predictive ability overall (0.29) compared to GY and HD. These observations were consistent with studies [
13,
14,
15] which previously demonstrated that for predicting complex traits, an IBCF approach would be most efficient if the correlation between environments were moderate to high. A lack of correlation between years would therefore be unfavorable for implementing a recommender approach for GS as low or negative correlations between environments could result in low predictive ability [
2]. Further, the presence of genotype-by-environment (G x E) interactions, which could result in ‘weak’ correlations among years, especially when one year is different from the others, could impose challenges and limitations in the implementation of recommender systems for GS in breeding programs [
13]. The variability in terms of weather patterns, for example, could have resulted in the differences across the environments. The 2015 growing season was considered to be having one of the warmest Spring temperatures in PUL at 22 °C (72 °F) (Loyd and Hoogenboom [
29]
https://weather.wsu.edu/index.php?page=AWN_Spring_2015_Weather_Review; retrieved 7 May 2020), whereas the 2018 season one of the wettest, with >10 consecutive days with precipitation in both LND and PUL environments (Washington State University AgWeatherNet;
http://weather.wsu.edu/; retrieved 7 May 2020; [
30]). Selecting stable lines with minimal G x E effects to ensure better correlation of phenotypes across years would therefore be crucial in the success of an IBCF recommender system for predicting future performance of lines in breeding programs. Recently, analyses of G x E interactions for this population of US Pacific Northwest winter wheat lines was conducted and stable lines based on AMMI and Finlay-Wilkinson regression coefficients were identified [
20]. Furthermore, genetic mapping for yield and yield stability identified loci controlling both sets of traits demonstrating the potential of simultaneously selecting stable and higher-yielding lines [
20].
Using secondary correlated traits that are more heritable than the target trait for GS has been shown to improve predictive ability [
17,
19,
31,
32]. Predicting yield using spectral traits collected from high-throughput phenotyping using the IBCF recommender system resulted in a 50% higher predictive ability compared with using single and multi-trait partial least square regression models [
17] (0.24 vs. 0.16). Our results were comparable to Juliana et al. [
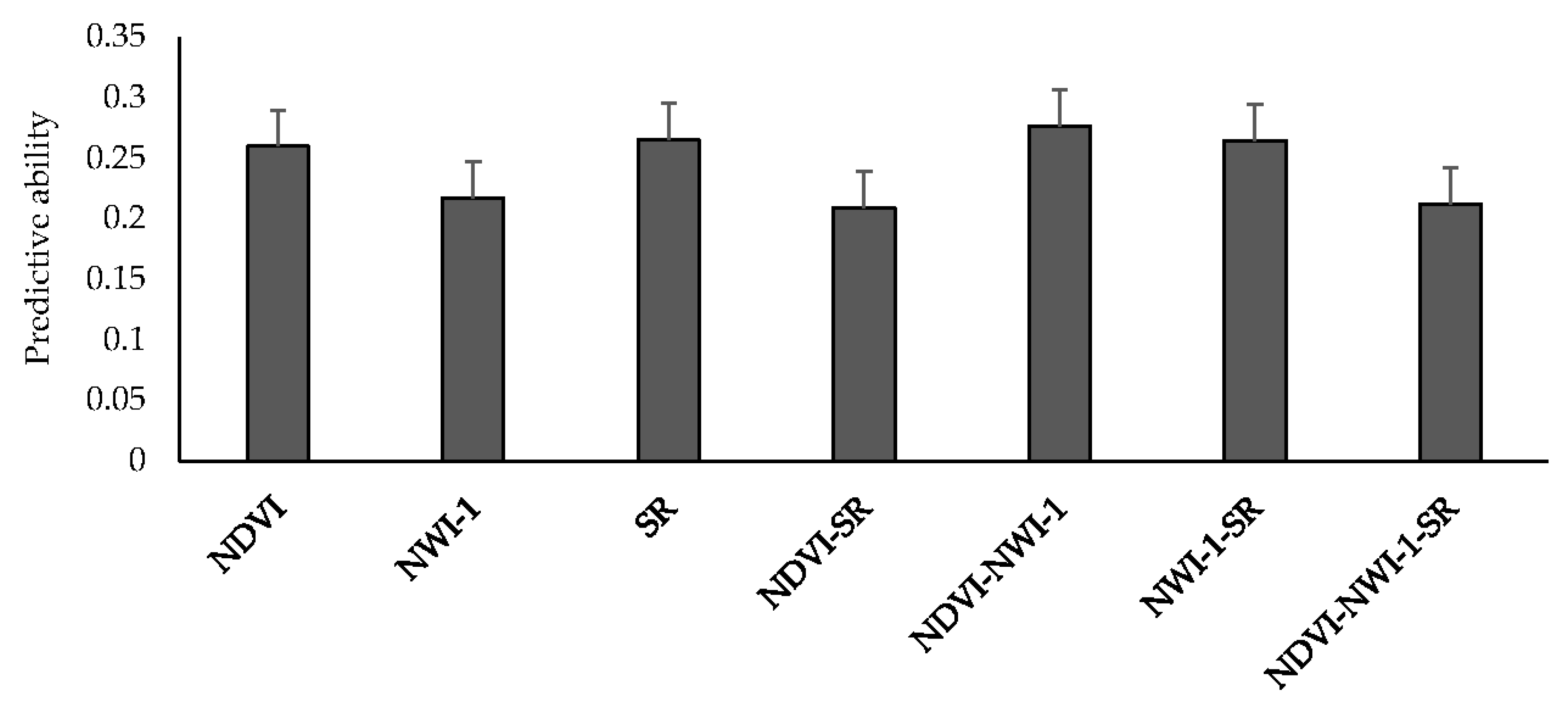
15] who observed an average predictive ability of 0.22 in spring wheat lines with a narrow range of heading dates when green NVDI was used to predict GY in an IBCF approach. In the current study, using two traits instead of three resulted in the highest predictive ability in which predicting GY using NDVI-NWI-1 and NWI-1-SR resulted in a mean predictive ability of 0.28 and 0.26, respectively. Collinearity which could affect model predictive ability could result from including too many traits as predictors [
33]; hence, using only a few highly heritable traits with high correlation with the target trait might be the ideal strategy to achieve better predictive ability in this GS scenario.
An increased correlation between GY and spectral indices was also related to improved predictive ability, similar in the case of predicting across years. Including spectral traits as fixed effects in ridge regression prediction models, nevertheless, might still be more advantageous in improving predictive ability compared to using a recommender approach for GS. Previously, a mean predictive ability of 0.43 for GY was observed using the same population of Pacific Northwest winter wheat lines when NDVI, NWI-1, and SR were included as fixed effects in an RRBLUP prediction model [
19]. Canopy temperature and NDVI resulted in a 70% increase in predictive ability when predicting GY using multivariate models [
34]. In other studies, incorporating markers for major growth and developmental genes such as
Vrn and
Ppd as fixed effects in the model improved predictive ability for GY, where combining multiple genes in the model resulted in optimal values [
35,
36]. Using GWAS-derived markers as fixed effects was also observed to have positive effects on the predictive ability for yield, stability, and related traits [
37,
38]. Predictive ability for GY using secondary spectral traits under an IBCF approach is ultimately a factor of the strength of correlation between traits across different environments.
Incorporating genomic information from molecular marker data under the IBCF recommender approach improved predictive ability for yield and agronomic traits compared to the other GS scenarios evaluated in the current study. By default, the IBCF algorithm does not allow for the direct incorporation of genomic data and therefore it is necessary to perform trait adjustments by markers using regression models first to obtain breeding values. Although we observed no significant differences among the various Bayesian regression models used for trait adjustments, using Bayesian LASSO resulted in the highest mean predictive ability across the datasets, indicating that this model could be preferentially used to achieve optimal predictive ability under an IBCF approach. Negative prediction accuracies were nevertheless still observed for the LND18 and PUL15 datasets, even when traits were already adjusted for marker effects. One disadvantage of IBCF is that poor predictions could result when genomic relationship matrices that do not represent the phenotypic correlations between genotypes are used as proxy values in performing predictions [
14]. This could be the reason why negative prediction accuracies were observed for these datasets. In the present study, using an Illumina SNP chip for genotyping lines resulted in a moderate to high predictive ability across different environments for the evaluated traits. In the case of using genotyping-by-sequencing (GBS) markers, where there is an abundance of missing data for GS, imputation of missing markers can be carried out when there is a high correlation between the column of SNPs [
14], consequently allowing for the effective implementation of an IBCF approach using GBS-derived SNP markers.
Among the advantages of an IBCF recommender approach in the context of GS is that its implementation is fast, straightforward, and does not require a lot of computing power [
14,
16]. However, in the current study, one drawback observed for the IBCF approach is its execution time. Compared to a univariate GBLUP model implemented in iPat, the time to execute an IBCF analysis where trait values were adjusted for marker effects using Bayesian models was longer (mean of ~64 min vs. ~13 min. for nine environments, 456 lines, and 15,229 SNP markers across three traits). Using a recommender system for moderate to large datasets would nonetheless still be a desirable alternative to the traditional GS models used in the breeding program in terms of achieving competitive prediction accuracies. When compared to other recommender approach such as matrix factorization, IBCF was also previously observed to be a better method for predicting GY, PH, and HD in wheat [
14]. Finally, the IBCF.MTME is not a model-based approach, and hence it cannot estimate genetic variances or variance components [
16], but nonetheless could still be used as an alternative approach for predicting traits for GS in plant breeding programs. In the future, the predictive power of other approaches for phenotypic trait adjustments such as generalized linear models via penalized maximum likelihood and frequentist LASSO (‘glmnet’ package in R; [
39,
40]) and non-local priors [
41,
42], and multivariate GBLUP [
43] and compound symmetry GBLUP models should also be evaluated and compared with IBCF recommender system in the context of multi-environment, multi-trait GS. Exploring alternative strategies could facilitate the improvement of predictive ability for recommender approaches and other prediction models for GS in plant breeding programs [
16].