Evaluation of Pre-Analytical and Analytical Methods for Detecting SARS-CoV-2 in Municipal Wastewater Samples in Northern Italy
Abstract
:1. Introduction
2. Materials and Methods
2.1. Generation of Wastewater Samples Stock
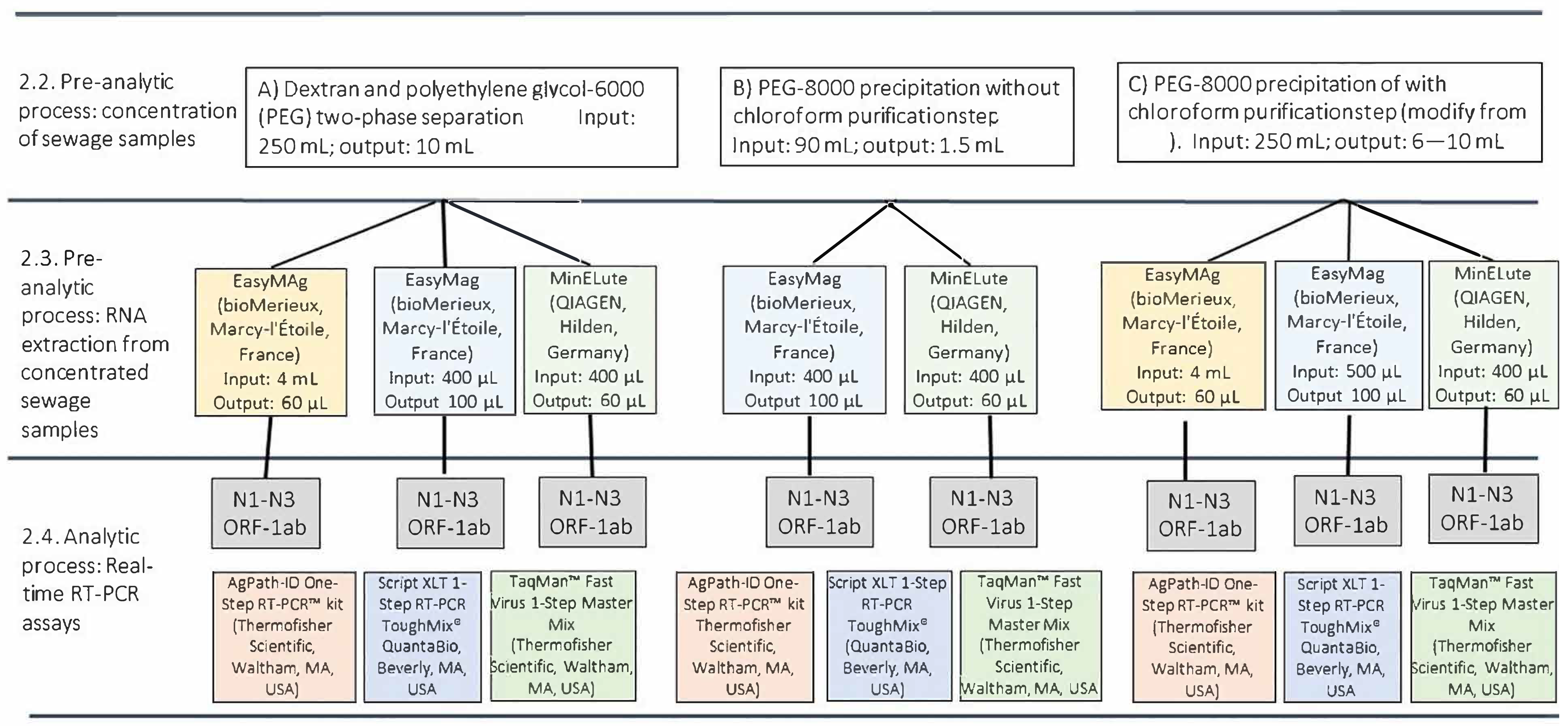
2.2. Pre-Analytical Process: Concentration of Sewage Samples
- (1)
- Dextran and polyethylene glycol-6000 (PEG) two-phase separation according to the 2003 WHO Guidelines for Environmental Surveillance of Poliovirus protocol [14], omitting the chloroform treatment to preserve the integrity of the SARS-CoV-2 envelope, as described, firstly, by La Rosa, G. et al. [15]. Briefly, 250 mL of wastewater sample was centrifuged for 30 min at 4500× g to pellet the wastewater solids, retaining the pellet for further processing. The clarified wastewater was mixed with dextran and PEG-6000 (19.8 mL of 22% dextran, 143.5 mL 29% PEG 6000, 17.5 mL 5 N NaCl); after a constant agitation for 30 min using a horizontal shaker, the mixture was left to stand overnight at 4 °C in a separation funnel. The bottom layer and the interphase were then collected drop-wise; this concentrate was added to the wastewater solids [14].
- (2)
- (3)
- PEG-8000 precipitation of 250 mL of sewage, modified from Wu, F. et al. [16], as follows:
2.3. Pre-Analytical Process: RNA Extraction from Concentrated Sewage Samples
2.4. Analytical Process: Real-Time RT-PCR Assays
2.5. Pre-Analytical and Analytical Workflows
2.6. Evaluation of SARS-CoV-2 Recovery Efficiency
SARS-CoV-2 copies/µL seeded
2.7. Evaluation of SARS-CoV-2 RT-PCR Assays Efficiency
2.8. Data Analysis
3. Results
3.1. Performance of the Concentration Methods
3.2. SARS-CoV-2 Recovery Efficiency
3.3. Real-Time RT-PCR Efficiency
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Vandenberg, O.; Martiny, D.; Rochas, O.; van Belkum, A.; Kozlakidis, Z. Considerations for diagnostic COVID-19 tests. Nat. Rev. Microbiol. 2021, 19, 171–183. [Google Scholar] [CrossRef] [PubMed]
- Durant, T.J.S.; Peaper, D.R.; Ferguson, D.; Schulz, W.L. Impact of COVID-19 Pandemic on Laboratory Utilization. J. Appl. Lab. Med. 2020, 5, 1194–1205. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Bivins, A.; Bertsch, P.M.; Bibby, K.; Choi, P.M.; Farkas, K.; Gyawali, P.; Hamilton, K.A.; Haramoto, E.; Kitajima, M.; et al. Surveillance of SARS-CoV-2 RNA in wastewater: Methods optimisation and quality control are crucial for generating reliable public health information. Curr. Opin. Environ. Sci. Health 2020, 17, 82–93. [Google Scholar] [CrossRef]
- Michael-Kordatou, I.; Karaolia, P.; Fatta-Kassinos, D. Sewage analysis as a tool for the COVID-19 pandemic response and management: The urgent need for optimised protocols for SARS-CoV-2 detection and quantification. J. Environ. Chem. Eng. 2020, 8, 104306. [Google Scholar] [CrossRef]
- Medema, G.; Been, F.; Heijnen, L.; Petterson, S. Implementation of environmental surveillance for SARS-CoV-2 virus to support public health decisions: Opportunities and challenges. Curr. Opin. Environ. Sci. Health 2020, 17, 49–71. [Google Scholar] [CrossRef]
- Peccia, J.; Zulli, A.; Brackney, D.E.; Grubaugh, N.D.; Kaplan, E.H.; Casanovas-Massana, A.; Ko, A.I.; Malik, A.A.; Wang, D.; Wang, M.; et al. Measurement of SARS-CoV-2 RNA in wastewater tracks community infection dynamics. Nat. Biotechnol. 2020, 38, 1164–1167. [Google Scholar] [CrossRef]
- Larsen, D.A.; Wigginton, K.R. Tracking COVID-19 with wastewater. Nat. Biotechnol. 2020, 38, 1151–1153. [Google Scholar] [CrossRef]
- Agrawal, S.; Orschler, L.; Lackner, S. Long-term monitoring of SARS-CoV-2 RNA in wastewater of the Frankfurt metropolitan area in Southern Germany. Sci. Rep. 2021, 11, 5372. [Google Scholar] [CrossRef]
- Alygizakis, N.; Markou, A.N.; Rousis, N.I.; Galani, A.; Avgeris, M.; Adamopoulos, P.G.; Scorilas, A.; Lianidou, E.S.; Paraskevis, D.; Tsiodras, S.; et al. Analytical methodologies for the detection of SARS-CoV-2 in wastewater: Protocols and future perspectives. Trends Anal. Chem. 2021, 134, 116125. [Google Scholar] [CrossRef]
- Philo, S.E.; Keim, E.K.; Swanstrom, R.; Ong, A.Q.; Burnor, E.A.; Kossik, A.L.; Harrison, J.C.; Demeke, B.A.; Zhou, N.A.; Beck, N.K.; et al. A comparison of SARS-CoV-2 wastewater concentration methods for environmental surveillance. Sci. Total Environ. 2021, 760, 144215. [Google Scholar] [CrossRef] [PubMed]
- Word Health Organization (WHO). Contributions of the Polio Network to COVID-19 Response. Available online: https://apps.who.int/iris/bitstream/handle/10665/336261/9789240011533-eng.pdf?sequence=1&isAllowed=y (accessed on 5 March 2022).
- Regione Lombardia. Regione Lombardia, Delibera N.4127—Accordo di Collaborazione con il Ministero Della Salute-Centro Nazionale per la Prevenzione e il Controllo delle Malattie e Convenzioni con l’Istituto Superiore di Sanità e l’ATS Milano Città Metropolitana per la Realizzazione del Progetto CCM 2020—Area Progettuale “Epidemiologia delle Acque Reflue: Implementazione del Sistema di Sorveglianza per L’identificazione Precoce di Agenti Patogeni, con Particolare Riferimento al SARS-CoV2”. Available online: https://www.dati.lombardia.it/api/views/q639-trxu/rows.xml?accessType=DOWNLOAD (accessed on 5 March 2022).
- Word Health Organization (WHO). WHO Guidelines for Environmental Surveillance of Poliovirus Protocol—WHO/V&B/03.03. Available online: https://polioeradication.org/wp-content/uploads/2016/07/WHO_V-B_03.03_eng.pdf (accessed on 5 March 2022).
- la Rosa, G.; Iaconelli, M.; Mancini, P.; Ferraro, G.B.; Veneri, C.; Bonadonna, L.; Lucentini, L.; Suffredini, E. First detection of SARS-CoV-2 in untreated wastewaters in Italy. Sci. Total Environ. 2020, 736, 139652. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Xiao, A.; Zhang, J.; Moniz, K.; Endo, N.; Armas, F.; Bushman, M.; Chai, P.R.; Duvallet, C.; Erickson, T.B.; et al. Wastewater surveillance of SARS-CoV-2 across 40 U.S. states from February to June 2020. Water Res. 2021, 202, 117400. [Google Scholar] [CrossRef] [PubMed]
- Castiglioni, S.; Schiarea, S.; Pellegrinelli, L.; Primache, V.; Galli, C.; Bubba, L.; Mancinelli, F.; Marinelli, M.; Cereda, D.; Ammoni, E.; et al. SARS-CoV-2 RNA in urban wastewater samples to monitor the COVID-19 pandemic in Lombardy, Italy (March-June 2020). Sci. Total Environ. 2022, 806 Pt 4, 150816. [Google Scholar] [CrossRef]
- Haramoto, E.; Kitajima, M.; Hata, A.; Torrey, J.R.; Masago, Y.; Sano, D.; Katayama, H. A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Res. 2018, 135, 168–186. [Google Scholar] [CrossRef]
- Bisseux, M.; Colombet, J.; Mirand, A.; Roque-Afonso, A.-M.; Abravanel, F.; Izopet, J.; Archimbaud, C.; Peigue-Lafeuille, H.; Debroas, D.; Bailly, J.-L.; et al. Monitoring human enteric viruses in wastewater and relevance to infections encountered in the clinical setting: A one-year experiment in central France, 2014 to 2015. Eurosurveillance 2018, 23, 17-00237. [Google Scholar] [CrossRef] [Green Version]
- Pellegrinelli, L.; Bubba, L.; Primache, V.; Pariani, E.; Battistone, A.; Delogu, R.; Fiore, S.; Binda, S. Surveillance of poliomyelitis in Northern Italy: Results of acute flaccid paralysis surveillance and environmental surveillance, 2012–2015. Hum. Vaccines Immunother. 2016, 13, 332–338. [Google Scholar] [CrossRef]
- Hellmér, M.; Paxéus, N.; Magnius, L.O.; Enache, L.; Arnholm, B.; Johansson, A.M.; Bergström, T.; Norder, H. Detection of Pathogenic Viruses in Sewage Provided Early Warnings of Hepatitis A Virus and Norovirus Outbreaks. Appl. Environ. Microbiol. 2014, 80, 6771–6781. [Google Scholar] [CrossRef] [Green Version]
- Battistone, A.; Buttinelli, G.; Fiore, S.; Amato, C.; Bonomo, P.; Patti, A.M.; Vulcano, A.; Barbi, M.; Binda, S.; Pellegrinelli, L.; et al. Sporadic Isolation of Sabin-Like Polioviruses and High-Level Detection of Non-Polio Enteroviruses during Sewage Surveillance in Seven Italian Cities, after Several Years of Inactivated Poliovirus Vaccination. Appl. Environ. Microbiol. 2014, 80, 4491–4501. [Google Scholar] [CrossRef] [Green Version]
- Pellegrinelli, L.; Galli, C.; Binda, S.; Primache, V.; Tagliacarne, C.; Pizza, F.; Mazzini, R.; Pariani, E.; Romanò, L. Molecular Characterization and Phylogenetic Analysis of Enteroviruses and Hepatitis A Viruses in Sewage Samples, Northern Italy, 2016. Food Environ. Virol. 2019, 11, 393–399. [Google Scholar] [CrossRef]
- Ahmed, W.; Angel, N.; Edson, J.; Bibby, K.; Bivins, A.; O’Brien, J.W.; Choi, P.M.; Kitajima, M.; Simpson, S.L.; Li, J.; et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 2020, 728, 138764. [Google Scholar] [CrossRef] [PubMed]
- Randazzo, W.; Truchado, P.; Cuevas-Ferrando, E.; Simón, P.; Allende, A.; Sánchez, G. SARS-CoV-2 RNA in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 2020, 181, 115942. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Malla, B.; Thakali, O.; Kitajima, M. First Environmental Surveillance for the Presence of SARS-CoV-2 RNA in Wastewater and River Water in Japan. Sci. Total Environ. 2020, 737, 140405. [Google Scholar] [CrossRef] [PubMed]
- Sherchan, S.P.; Shahin, S.; Ward, L.M.; Tandukar, S.; Aw, T.G.; Schmitz, B.; Ahmed, W.; Kitajima, M. First detection of SARS-CoV-2 RNA in wastewater in North America: A study in Louisiana, USA. Sci. Total Environ. 2020, 743, 140621. [Google Scholar] [CrossRef] [PubMed]
- Westhaus, S.; Weber, F.-A.; Schiwy, S.; Linnemann, V.; Brinkmann, M.; Widera, M.; Greve, C.; Janke, A.; Hollert, H.; Wintgens, T.; et al. Detection of SARS-CoV-2 in raw and treated wastewater in Germany—Suitability for COVID-19 surveillance and potential transmission risks. Sci. Total Environ. 2021, 751, 141750. [Google Scholar] [CrossRef]
- Torii, S.; Oishi, W.; Zhu, Y.; Thakali, O.; Malla, B.; Yu, Z.; Zhao, B.; Arakawa, C.; Kitajima, M.; Hata, A.; et al. Comparison of five polyethylene glycol precipitation procedures for the RT-qPCR based recovery of murine hepatitis virus, bacteriophage phi6, and pepper mild mottle virus as a surrogate for SARS-CoV-2 from wastewater. Sci. Total Environ. 2021, 807, 150722. [Google Scholar] [CrossRef]
- Ahmed, W.; Bertsch, P.M.; Bivins, A.; Bibby, K.; Farkas, K.; Gathercole, A.; Haramoto, E.; Gyawali, P.; Korajkic, A.; McMinn, B.R.; et al. Comparison of virus concentration methods for the RT-qPCR-based recovery of murine hepatitis virus, a surrogate for SARS-CoV-2 from untreated wastewater. Sci. Total Environ. 2020, 739, 139960. [Google Scholar] [CrossRef]
- Lázaro-Perona, F.; Rodriguez-Antolín, C.; Alguacil-Guillén, M.; Gutiérrez-Arroyo, A.; Mingorance, J.; García-Rodriguez, J.; SARS-CoV-2 Working Group. Evaluation of two automated low-cost RNA extraction protocols for SARS-CoV-2 detection. PLoS ONE 2021, 16, e0246302. [Google Scholar] [CrossRef]
- Vogels, C.B.F.; Brito, A.F.; Wyllie, A.L.; Fauver, J.R.; Ott, I.M.; Kalinich, C.C.; Petrone, M.E.; Casanovas-Massana, A.; Muenker, M.C.; Moore, A.J.; et al. Analytical sensitivity and efficiency comparisons of SARS-CoV-2 RT-qPCR primer-probe sets. Nat. Microbiol. 2020, 5, 1299–1305. [Google Scholar] [CrossRef]
- Zhou, N.A.; Tharpe, C.; Meschke, J.S.; Ferguson, C.M. Survey of rapid development of environmental surveillance methods for SARS-CoV-2 detection in wastewater. Sci. Total Environ. 2021, 769, 144852. [Google Scholar] [CrossRef]

| Spiked Sample by SARS-CoV-2 Viral Load: 4.7 × 107 Copies/mL | Unknown Sample | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Volume of Sewage to be Concentrated | Methods of Concentration | Volume of Concentrating Sewage | RNA Extraction Kit | Extraction Input Elution Volume | RT-qPCR Kit | RT-PCR Instrument | Tournaround Time | Target | Positive Replicates | Mean Ct Value | SD Ct Value | ΔCt (Recoverded vs. Seeded) | Mean Recovery Efficiency (%) | Positive Replicates | Mean Ct Value | SD Ct Value |
| 250 mL | (A) Dextran and PEG-6000 two-phase separation | nearly 10 mL | QIAamp MinElute Virus Spin Kit (QIAGEN) | 400 μL 60 μL | AGPATH-ID ONE-STEP RT-PCR (Thermo Fisher) | Applied Biosystems 7500 Real-Time PCR System (Thermo Fisher) | 16-h | N1 | 100% | 29.2 | 0.62 | 7.2 | 2% | 17% | 37.27 | / |
| N3 | 100% | 30.4 | 0.85 | und | und | / | ||||||||||
| ORF | 100% | 36.5 | 0.79 | und | und | / | ||||||||||
| NucliSens EasyMag (bioMerieux) | 400 μL 100 μL | AGPATH-ID ONE-STEP RT-PCR (Thermo Fisher) | 16-h | N1 | 100% | 26.2 | 0.38 | 4.2 | 12% | 67% | 37.9 | 0.45 | ||||
| N3 | 100% | 25.6 | 0.41 | und | und | / | ||||||||||
| ORF | 100% | 28.9 | 0.23 | und | und | 0.02 | ||||||||||
| NucliSens EasyMag (bioMerieux) | 4 mL 60 μL | AGPATH-ID ONE-STEP RT-PCR (Thermo Fisher) | 16-h | N1 | 100% | 27.6 | 0.25 | 5.6 | 5% | 33% | 38.2 | 0.02 | ||||
| N3 | 100% | 27 | 0.22 | 67% | 37.6 | 0.2 | ||||||||||
| ORF | 100% | 30.4 | 0.17 | und | und | / | ||||||||||
| 80 mL | (B) PEG-8000 precipitation without chloroform purification step | 1.5 mL | QIAamp MinElute Virus Spin Kit (QIAGEN) | 400 μL 60 μL | AGPATH-ID ONE-STEP RT-PCR (Thermo Fisher) | Applied Biosystems 7500 Real-Time PCR System (Thermo Fisher) | 8-h | N1 | 100% | 20.1 | 0.13 | −1.9 | 76% | 100% | 33.1 | 0.4 |
| N3 | 100% | 20.5 | 0.20 | 100% | 33.4 | 0.34 | ||||||||||
| ORF | 100% | 24.6 | 0.30 | 100% | 37.5 | 0.75 | ||||||||||
| NucliSens EasyMag (bioMerieux) | 500 μL 100 μL | AGPATH-ID ONE-STEP RT-PCR (Thermo Fisher) | 8-h | N1 | 100% | 21.4 | 0.38 | 0.6 | 31.4% | 100% | 34.1 | 0.22 | ||||
| N3 | 100% | 19.7 | 0.31 | 100% | 32.6 | 0.19 | ||||||||||
| ORF | 100% | 24.6 | 0.43 | 100% | 37.2 | 0.34 | ||||||||||
| 250 mL | (C) PEG-8000 precipitation with chloroform purification step | 6–10 mL | QIAamp MinElute Virus Spin Kit (QIAGEN) | 400 μL 60 μL | QScript XLT 1-Step RT-PCR ToughMix (QuantaBio) | CFX96 BioRad real-time PCR System (Biorad) | 8-h | N1 | 100% | 25.6 | 1.27 | 3.6 | 18% | 83% | 37.7 | 1.69 |
| N3 | 100% | 26.9 | 1.05 | 67% | 36.8 | 0.46 | ||||||||||
| ORF | 100% | 32.8 | 1.08 | und | und | / | ||||||||||
| NucliSens EasyMag (bioMerieux) | 400 μL 100 μL | QScript XLT 1-Step RT-PCR ToughMix (QuantaBio) | 8-h | N1 | 100% | 29.2 | 0.33 | 7.2 | 2% | 50% | 38.7 | 0.3 | ||||
| N3 | 100% | 29.5 | 0.43 | 67% | 37.4 | 0.01 | ||||||||||
| ORF | 100% | 32.5 | 0.57 | und | und | / | ||||||||||
| NucliSens EasyMag (bioMerieux) | 4 mL 60 μL | QScript XLT 1-Step RT-PCR ToughMix (QuantaBio) | 8-h | N1 | 100% | 29.3 | 0.5 | 7.3 | 1% | 17% | 38.03 | / | ||||
| N3 | 100% | 26.1 | 0.38 | 67% | 37.9 | 0.46 | ||||||||||
| ORF | 100% | 33.1 | 1.41 | und | und | / | ||||||||||
| Target | Standard Curve | R2 | Efficiency | |
|---|---|---|---|---|
| AgPath-ID One-Step RT-PCR™ kit | N1 | y = −3.3863x + 37.009 | 0.9984 | 98.4% |
| N3 | y = −3.3677x + 38.426 | 0.9982 | 98.2% | |
| QScript XLT 1-Step RT-PCR ToughMix® | N1 | y = −3.279x + 39.076 | 0.9988 | 98.8% |
| N3 | y = −3.3073x + 40.08 | 0.9995 | 99.5% | |
| TaqMan™ Fast Virus 1-Step Master Mix | N1 | y = −3.7356x + 41.786 | 0.9985 | 85.2% |
| N3 | y = −3.5652x + 38.426 | 0.9971 | 90.8% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pellegrinelli, L.; Castiglioni, S.; Cocuzza, C.E.; Bertasi, B.; Primache, V.; Schiarea, S.; Salmoiraghi, G.; Franzetti, A.; Musumeci, R.; Tilola, M.; et al. Evaluation of Pre-Analytical and Analytical Methods for Detecting SARS-CoV-2 in Municipal Wastewater Samples in Northern Italy. Water 2022, 14, 833. https://doi.org/10.3390/w14050833
Pellegrinelli L, Castiglioni S, Cocuzza CE, Bertasi B, Primache V, Schiarea S, Salmoiraghi G, Franzetti A, Musumeci R, Tilola M, et al. Evaluation of Pre-Analytical and Analytical Methods for Detecting SARS-CoV-2 in Municipal Wastewater Samples in Northern Italy. Water. 2022; 14(5):833. https://doi.org/10.3390/w14050833
Chicago/Turabian StylePellegrinelli, Laura, Sara Castiglioni, Clementina E. Cocuzza, Barbara Bertasi, Valeria Primache, Silvia Schiarea, Giulia Salmoiraghi, Andrea Franzetti, Rosario Musumeci, Michela Tilola, and et al. 2022. "Evaluation of Pre-Analytical and Analytical Methods for Detecting SARS-CoV-2 in Municipal Wastewater Samples in Northern Italy" Water 14, no. 5: 833. https://doi.org/10.3390/w14050833








