Abstract
Topological indices (molecular descriptors) are numerical values of a chemical structure and represented by a graph. Molecular descriptors are used in QSPR/QSAR modeling to determine a chemical structure’s physical, biological, and chemical properties. The cycle graphs are symmetric graphs for any number vertices. In this paper, recently defined neighborhood degree sum-based molecular descriptors and polynomials are studied. NM-polynomials and molecular descriptors of some cycle-related graphs, which consist of the wheel graph, gear graph, helm graph, flower graph, and friendship graph, are computed and compared.
1. Introduction
Chemical graph theory is a field of graph theory with molecular graphs obtained by representing chemical structures with graphs. Molecular descriptors or topological indices are numerically representative of a chemical structure and represented by a graph. They are used to predict the physical, biological, and chemical properties of new structures obtained from the molecule or molecular compound [1,2]. The first-known application was studied to find the physical properties of a chemical structure in 1947 [3].
Molecular descriptors depend on the vertex degree of the graph, the distance between the two vertices, the eigenvalues of the graph, etc. Finding the topological polynomials instead of calculating the molecular descriptors one by one makes it easier to provide information about the molecular graph. M-polynomials are polynomials of molecular descriptors based on the vertex degree of a G graph and were defined by Deutsch and Klavžar [4]. Using this definition, NM-polynomials are defined. These polynomials depend on the sum of adjacent vertex degrees [5,6].
The quality of a molecular descriptor is measured by its ability to successfully predict molecules. Kirmani et al. argued that the neighborhood second-modified Zagreb index is best to predict the index for the molar refractivity and polarizability properties of COVID-19 drugs [6]. Havare argued that the neighborhood harmonic index is the best-predicting index for the molar volume of cancer drugs [7]. NM-polynomials and these molecular descriptors for various chemical graphs were studied (see for detail [8,9,10,11]).
Cycle-related graphs are molecular graphs of many chemical structures in chemistry, such as cycloalkanes. It is also a graph representation of many networks [12]. Sowmya studied total-eccentricity polynomials [13]. Asif et al. computed the Mostar index of cycle-related chemical structures [14]. Natarajan et al. computed leap-Zagreb indices of cycle-related special graphs [15]. Basanagoud et al. studied the M-polynomials of cycle-related graphs [16]. Havare computed the Mostar index and edge-Mostar index of cycle-related graphs [17]. Javaraju et al. studied the reciprocal-leap indices of wheel graphs [18].
2. Materials and Methods
If the graph has and , then the numbers of these sets are defined as and , respectively. The degree of the vertex of the graph is denoted by . The neighborhood of a vertex in a graph G, ), is the set of all vertices adjacent to [19]. See Reference [19] for basic definitions and notations on graph theory. Let and .
The neighborhood M-polynomial is defined by Verma et al. [20] as
Table 1 shows descriptors based on the sum of neighborhood degrees.

Table 1.
Molecular descriptors and expressions.
The following operators are used in Table 1:
If and , then neighborhood degree sum-based molecular descriptors are obtained.
3. Results
In this section, NM-polynomials of the wheel, gear, helm, friendship, and flower graphs are obtained. Molecular descriptors based on the neighborhood degree sum using these polynomials are computed.
The wheel graph for is obtained from the cycle with orders and a central vertex . The orders of the wheel graph are and the size of it is [19].
Theorem 1.
Letbe the wheel graph withorders. The neighborhood M-polynomial ofis
Proof of Theorem 1.
From the definition of the wheel graph, it is divided as From Equation (1),
or
□
Corollary 1.
The molecular descriptors based on the neighborhood degree of thegraph are
|
Proof of Corollary 1.
Using the formulas in Table 1, the following equations can be written.
Using the above equations and Table 1, the following equations are obtained.
The Helm graph is obtained by joining a pendant edge attached to each vertex of of the wheel graph [24]. The orders of the helm graph are and the size of the helm graph is . □
Theorem 2.
Letbe the helm graph withorders. Then,
for.
Proof of Theorem 2.
From the definition of the helm graph, it is divided as From Equation (1),
or
□
Corollary 2.
The molecular descriptors based on the neighborhood degree of thegraph are
|
Proof of Corollary 2.
Using the formulas in Table 1, the following equations can be written.
□
The gear graph is obtained by a vertex added between each pair of adjacent vertices of of the wheel graph [24]. The orders of the gear graph are and the size of the helm graph is .
Theorem 3.
Letbe the helm graph withorders. Then,
for.
Proof of Theorem 3.
From the definition of the gear graph, it is divided as From Equation (1),
or
□
Corollary 3.
The molecular descriptors based on the neighborhood degree of thegraph are
|
Proof of Corollary 3.
Using the formulas in Table 1, the following equations can be written.
□
The flower graph is obtained from a helm by combining the pendant vertices of the helm graph with the central vertex of the helm graph with one edge [24]. The orders of the flower graph are and the size of the flower graph is .
Theorem 4.
Letbe the flower graph withorders. Then,
for.
Proof of Theorem 4.
From the definition of , it is divided as From Equation (1),
or
□
Corollary 4.
The molecular descriptors based on the neighborhood degree of thegraph are
|
Proof of Corollary 4.
Using the formulas in Table 1, the following equations can be written.
□
The friendship graph is obtained by deleting the alternate edges of the of [24]. The orders of the friendship graph are and the size of the friendship graph is .
Theorem 5.
Letbe the friendship graph withorders. Then,
for.
Proof of Theorem 5.
From the definition of , it is divided as From Equation (1),
or
□
Corollary 5.
The molecular descriptors based on the neighborhood degree of thegraph are
|
Proof of Corollary 5.
Using the formulas in Table 1, the following equations can be written.
□
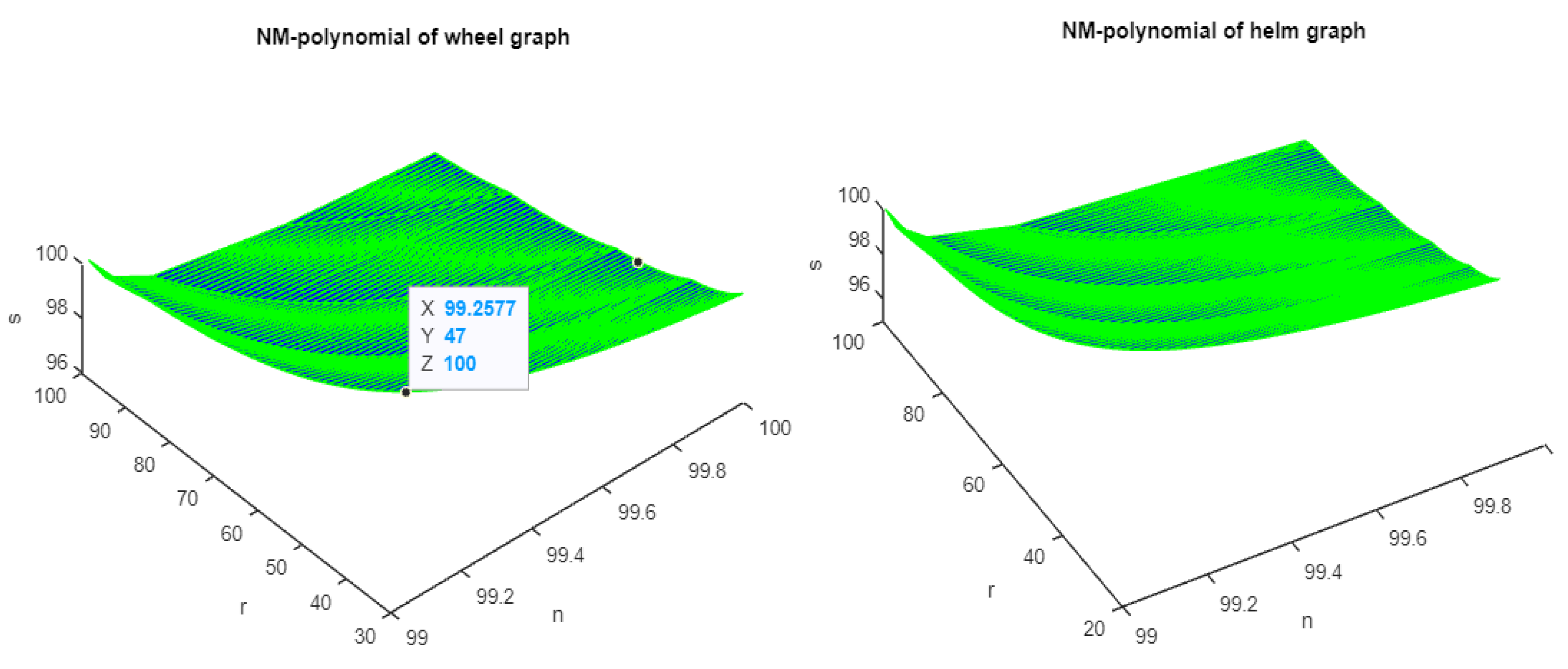
Figure 1, Figure 2 and Figure 3 show the 3D plots of the NM-polynomial of the wheel, helm, gear, flower, and friendship graphs.
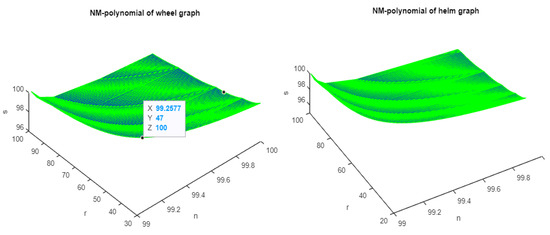
Figure 1.
The 3D plots of NM polynomial of wheel and helm graphs.
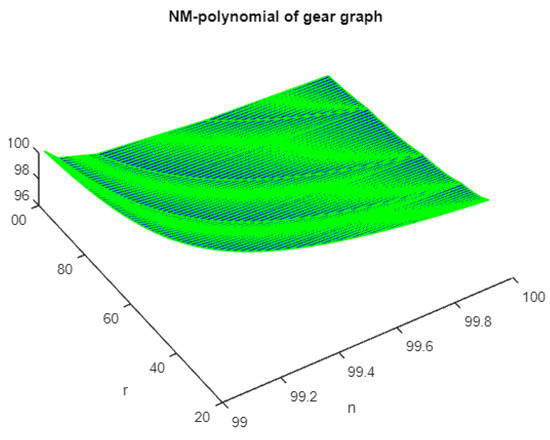
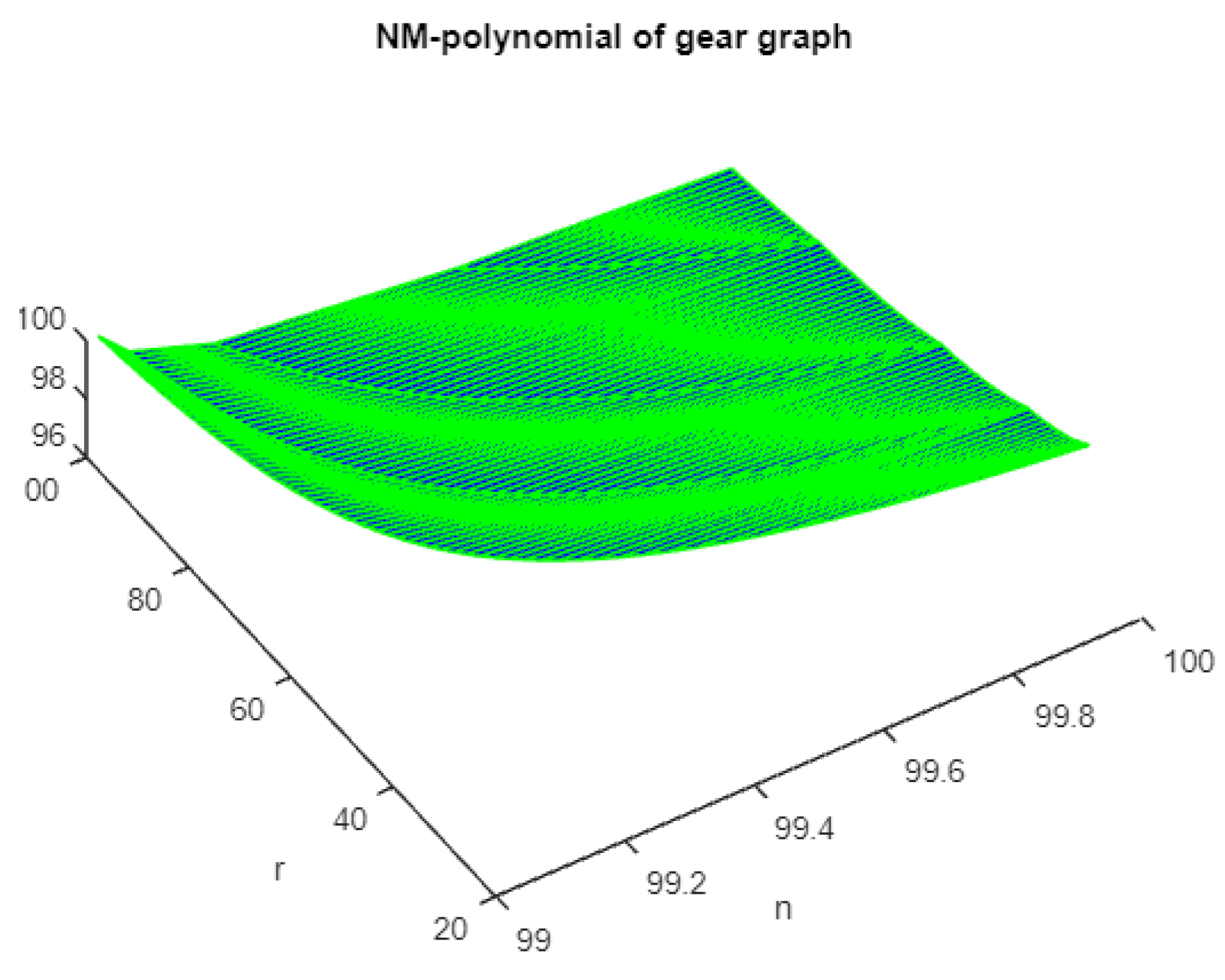
Figure 2.
The 3D plot of NM polynomial of gear graph.
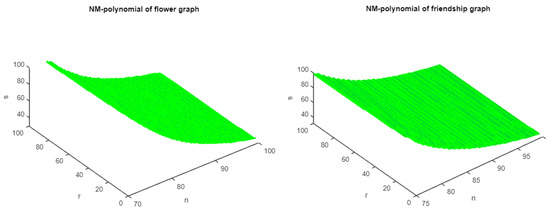
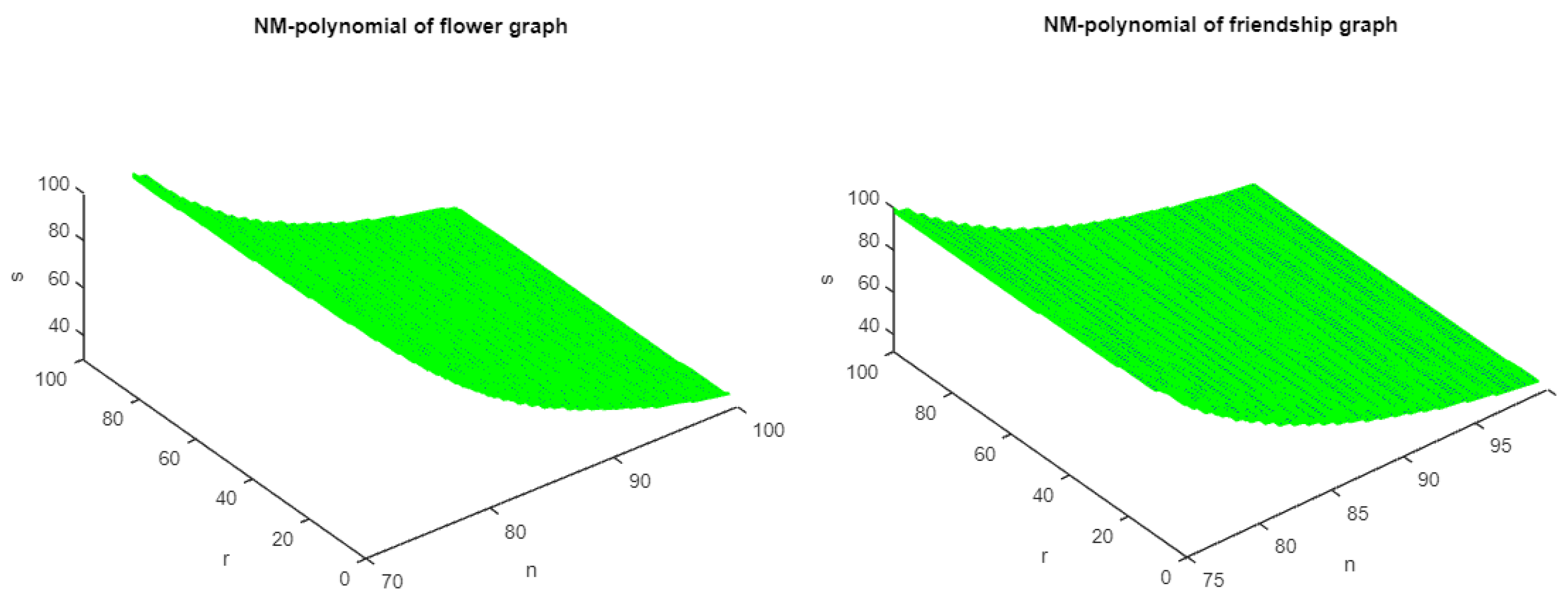
Figure 3.
The 3D plots of NM polynomial of flower and friendship graphs.
4. Discussion and Conclusions
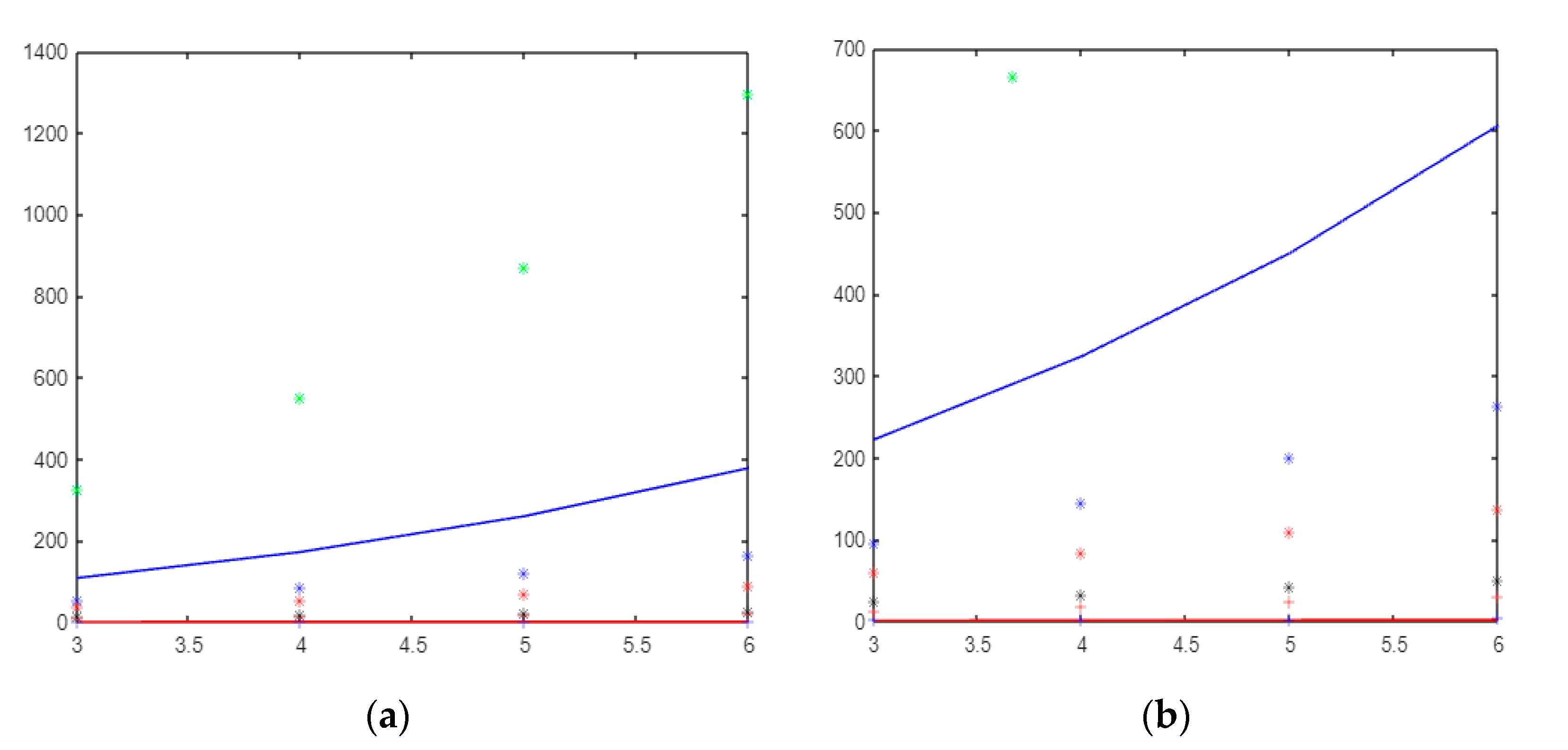
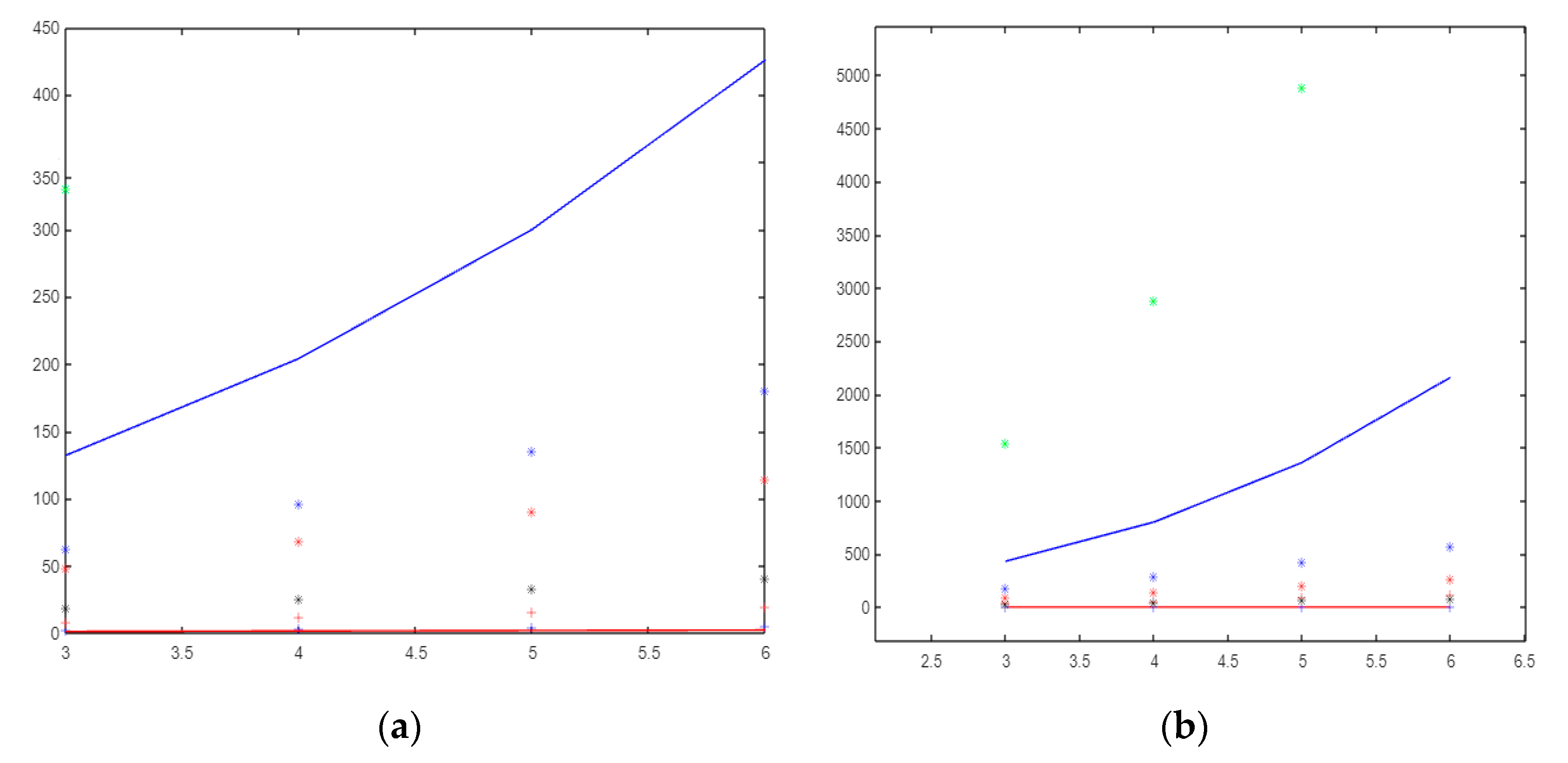
This paper shows that the gear, helm, and friendship graphs have the same orders and size, but NM-polynomials are different from each other. In Figure 4, Figure 5 and Figure 6, green * = , blue − = , blue * = , red * = , black * = , red + = , red − =, and blue + = .
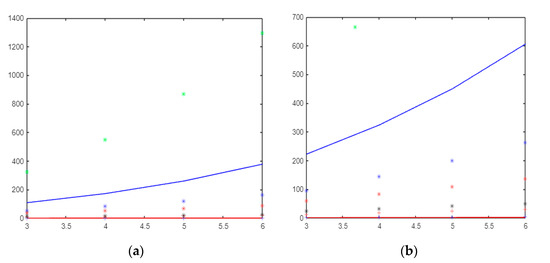
Figure 4.
(a)The molecular descriptors based on the neighborhood sum degree of and (b) The molecular descriptors based on the neighborhood sum degree of graph.
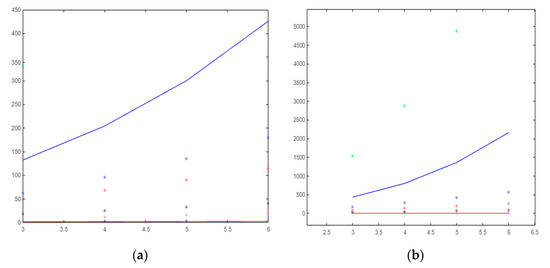
Figure 5.
(a) The molecular descriptors based on the neighborhood sum degree of and (b) The molecular descriptors based on the neighborhood sum degree of graph.
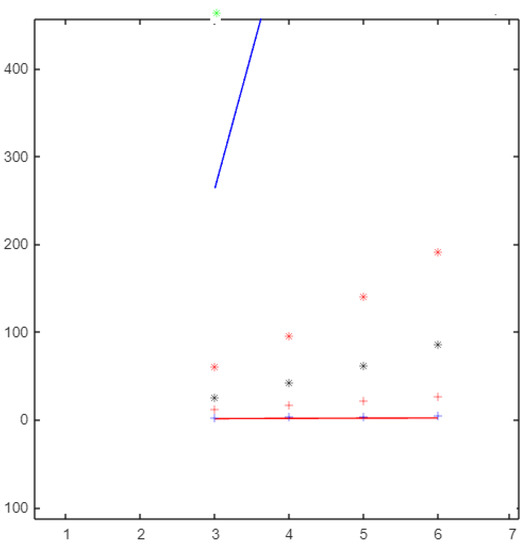
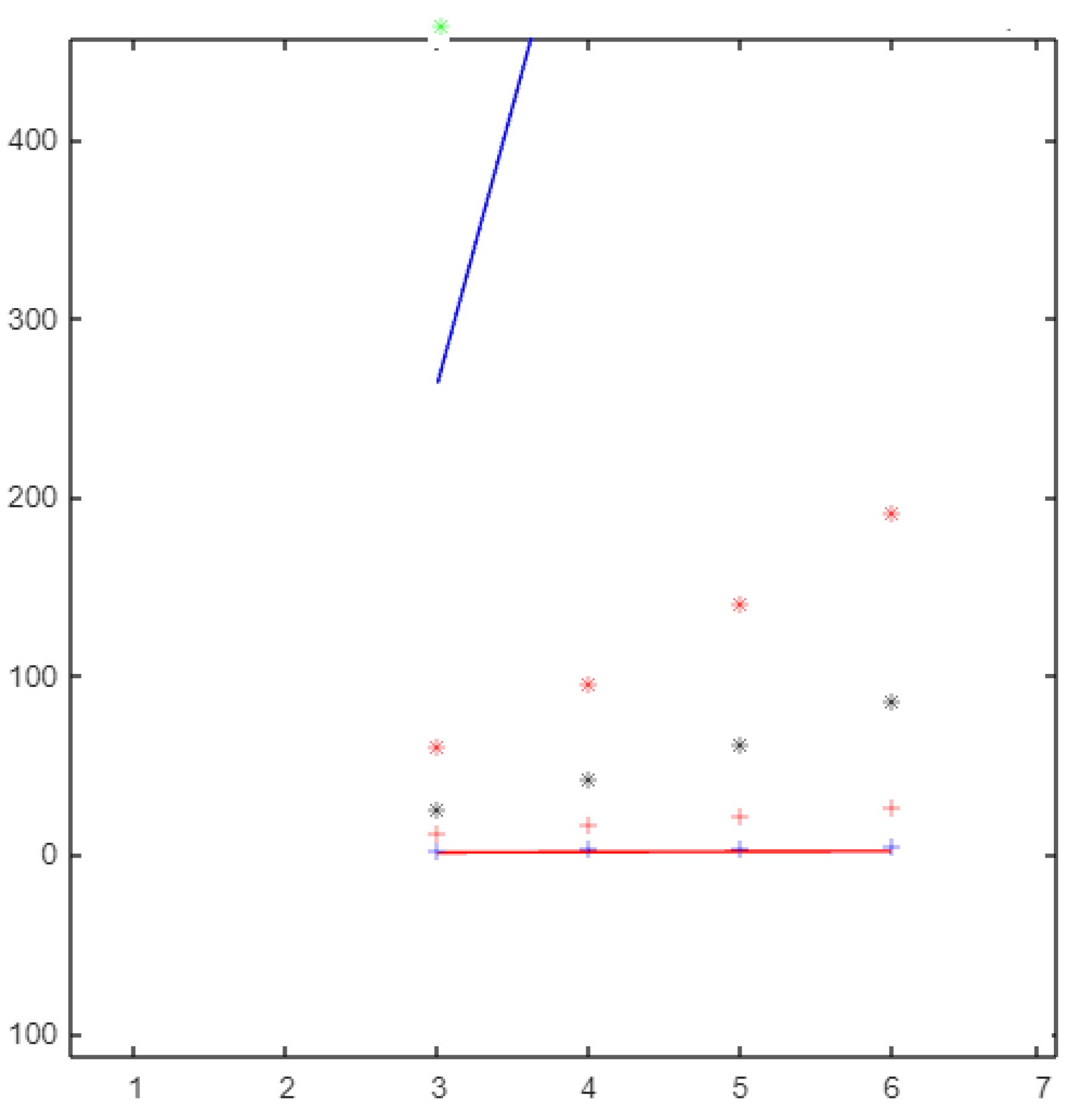
Figure 6.
The molecular descriptors based on the neighborhood sum degree of graph.
If it is compared, topological indices depend on the sum of the neighboring degrees of the wheel graph; the index has a very large value and creates a very fast curve, while has the lowest value. Although the index is close to the index, it has larger values and creates a faster curve (see Figure 4a).
Figure 4b shows the molecular descriptors based on the neighborhood sum degree of the graph. Figure 4b shows that is the fastest curve-forming index. The index that produces values closest to zero is the mM index. If we order the indices from that with the largest value to that with the smallest value, the list would be as follows: , , , , ,, , and .
For indices dependent on the sum of the adjacent vertex degrees of the gear graph, is the most curvilinear-growing, followed by . The most linear-growing is . If they are sorted in order from the index with the lowest value to the index with the highest value, the list would be as follows: , , , , , , , and .
If we list the indices for the flower graph, as the number increases, the index with the smallest value would be , , , , , , , and .
From Figure 6, if the topological index values for are ordered from smallest to the largest, the list would be as follows: ,, , , , , , and .
For the NM-polynomials of the cycle-related graphs and the topological indices based on them, the index with the fastest increasing and the slowest increasing values is . These results can be used to predict the properties of new cycle-related molecules in QSPR/QSPR studies.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The author declares no conflict of interest.
References
- Gutman, I. A property of the Simple Topological Index. MATCH Commun. Math. Comput. Chem. 1990, 25, 131–140. [Google Scholar]
- Xiao, C.; Chen, H.; Raigorodskii, A.M. A connection between the Kekulé structures of pentagonal chains and the Hosoya index of caterpillar trees. Disc. Appl. Math. 2017, 232, 230–234. [Google Scholar] [CrossRef]
- Wiener, H. Structural Determination of Paraffin Boiling Points. J. Am. Chem. Soc. 1947, 69, 17–20. [Google Scholar] [CrossRef] [PubMed]
- Deutsch, E.; Klavzar, S. M-Polynomial and Degree-Based Topological Indices. Iran. J. Math. Chem. 2015, 6, 93–102. [Google Scholar]
- Kirmani, S.A.K.; Ali, P.; Azam, F. Topological indices and QSPR/QSAR analysis of some antiviral drugs being investigated for the treatment of COVID-19 patients. Int. J. Quantum Chem. 2021, 121, e26594. [Google Scholar] [CrossRef]
- Mondal, S.; Siddiqui, M.K.; De, N.; Pal, A. Neighborhood M-Polynomial of Crystallographic Structures. Biointerface Res. Appl. Chem. 2021, 11, 9372–9381. [Google Scholar]
- Havare, Ö.Ç. Topological Indices and QSPR modeling of Some Novel Drugs Used in the Cancer Treatment. Int. J. Quantum Chem. 2021, 121, e26813. [Google Scholar] [CrossRef]
- Mondal, S.; Imran, M.; De, N.; Pal, A. Neighborhood M-polynomial of titanium compounds. Arab. J. Chem. 2021, 14, 103244. [Google Scholar] [CrossRef]
- Mondal, S.; De, N.; Pal, A. Neighborhood degree sum-based molecular descriptors of fractal and Cayley tree dendrimers. Eur. Phys. J. Plus 2021, 136, 303. [Google Scholar] [CrossRef]
- Haoer, R.S. Topological indices of metal-organic networks via neighborhood M-polynomial. J. Discret. Math. Sci. Cryptogr. 2021, 24, 369–390. [Google Scholar] [CrossRef]
- Réti, T.; Ali, A.; Varga, P.; Bitay, E. Some properties of the neighborhood first Zagreb index. Discret. Math. Lett. 2019, 2, 10–17. [Google Scholar]
- Ponomarenko, E.I.; Raigorodskii, A.M. New upper bounds for the independence numbers of graphs with vertices in {−1, 0, 1} n and their applications to problems of the chromatic numbers of distance graphs. Math. Notes 2014, 96, 140–148. [Google Scholar] [CrossRef]
- Sowmya, S. On Topological Indices of Cycle Related Graph. Adv. Math. Sci. J. 2020, 9, 4221–4230. [Google Scholar] [CrossRef]
- Asif, F.; Kashif, A.; Zafar, S.; Ojiema, M.O. Mostar Index of Cycle-Related Structures. J. Chem. 2022, 2022, 9411947. [Google Scholar] [CrossRef]
- Natarajan, C.; Selvamuthukumaran, S.; Farahani, M.R. On leap Zagreb indices of some cycle related graphs. TWMS J. Appl. Eng. Math. 2022, 12, 954–968. [Google Scholar]
- Basavanagoud, B.; Barangi, A.P.; Jakkannavar, P. M-polynomial of some graph operations and cycle related graphs. Iran. J. Math. Chem. 2019, 10, 127–150. [Google Scholar]
- Havare, O.C. Mostar Index (Mo) and edge Mo index for some cycle related graphs. Rom. J. Math. Comput. Sci. 2020, 10, 53–66. [Google Scholar]
- Javaraju, S.; Alsinai, A.; Alwardi, A.; Ahmed, H.; Soner, N.D. Reciprocal leap indices of some wheel related graphs. J. Prime Res. Math. 2021, 17, 101–110. [Google Scholar]
- Chartrand, G.; Lesniak, L. Graphs and Digraphs; The CRS Press: Baltimore, MD, USA, 2005. [Google Scholar]
- Verma, A.; Mondal, S.; De, N.; Pal, A. Topological properties of bismuth Tri-iodide using neighborhood M-polynomial. Int. J. Math. Trends Technol. 2019, 67, 83–90. [Google Scholar]
- Ghorbani, M.; Hosseinzadeh, M.A. The third version of Zagreb index. Disc. Math. Algorithms Appl. 2013, 5, 1350039. [Google Scholar] [CrossRef]
- Mondal, S.; De, N.; Pal, A. On some new neighbourhood degree based index. Acta Chem. Iasi. 2019, 27, 31–46. [Google Scholar] [CrossRef]
- Mondal, S.; Dey, A.; De, N.; Pal, A. QSPR analysis of some novel neighbourhood degree-based topological descriptors. Complex Intell. Syst. 2021, 7, 977–996. [Google Scholar] [CrossRef]
- Gallian, J.A. A Dynamic Survey of Graph Labeling. Electron. J. Comb. 2018, 1, #DS6. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).