MiR-10a, 27a, 34b/c, and 300 Polymorphisms are Associated with Ischemic Stroke Susceptibility and Post-Stroke Mortality
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Participants
2.2. Genotyping
2.3. Post-Stroke Mortality
2.4. Statistical Analysis
3. Results
3.1. Clinical Profiles of Ischemic Stroke Patients and Controls
3.2. Comparisons for the Four miRNA Polymorphisms among Patients with Ischemic Stroke, Ischemic Stroke Subtypes, and Controls
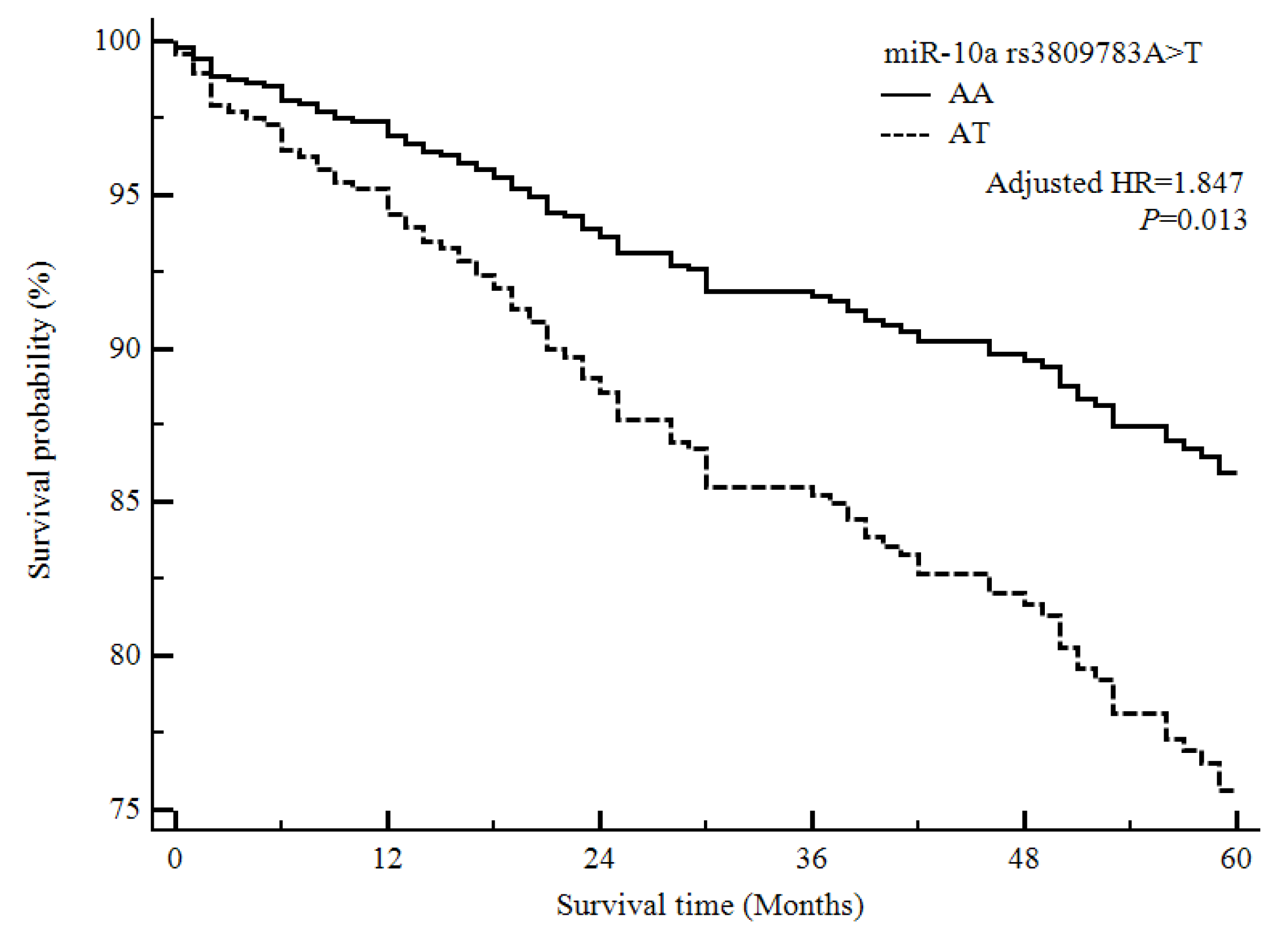
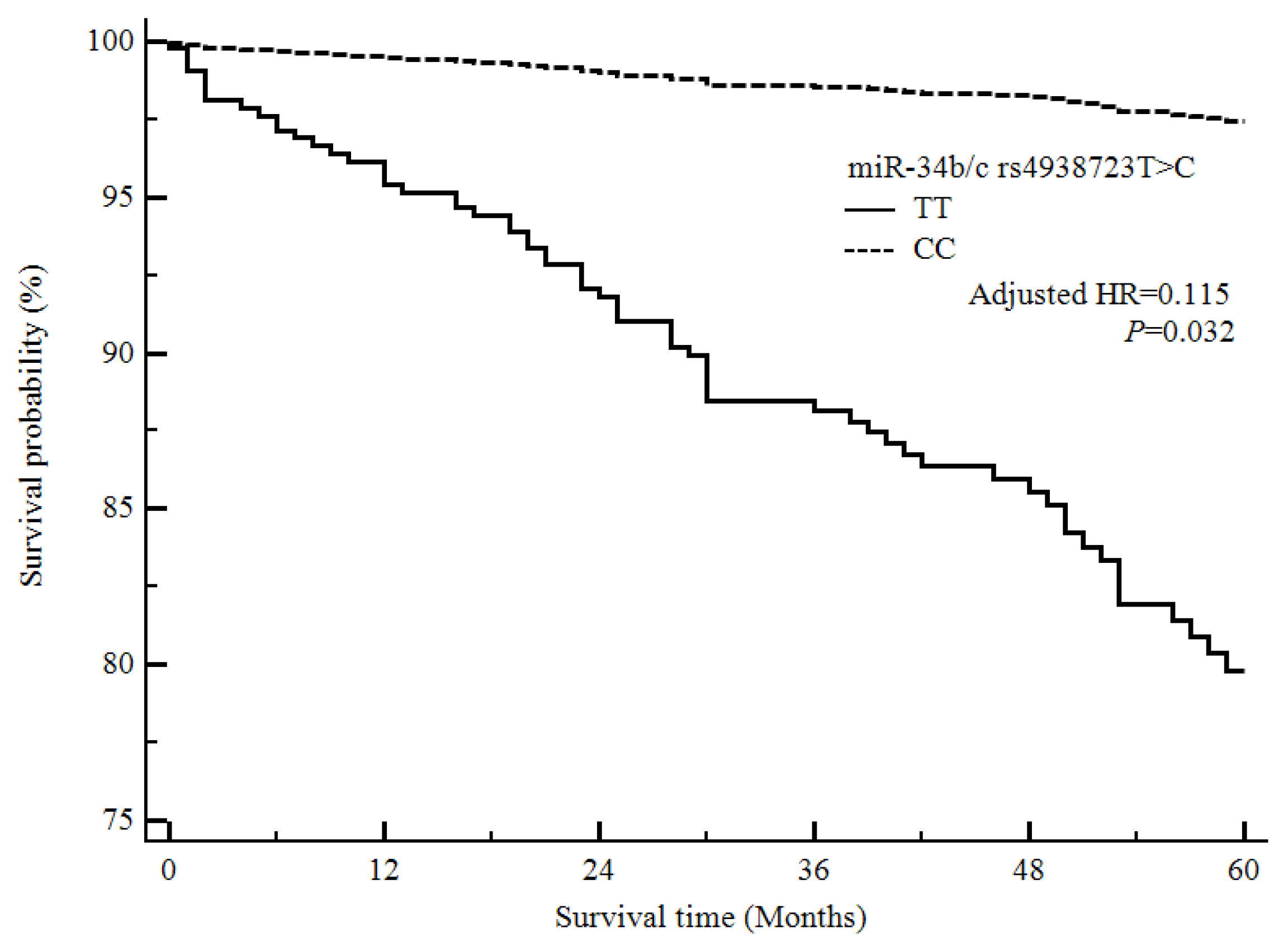
3.3. Analysis of the Four miRNA Polymorphisms, with Respect to Survival, in Ischemic Stroke Patients
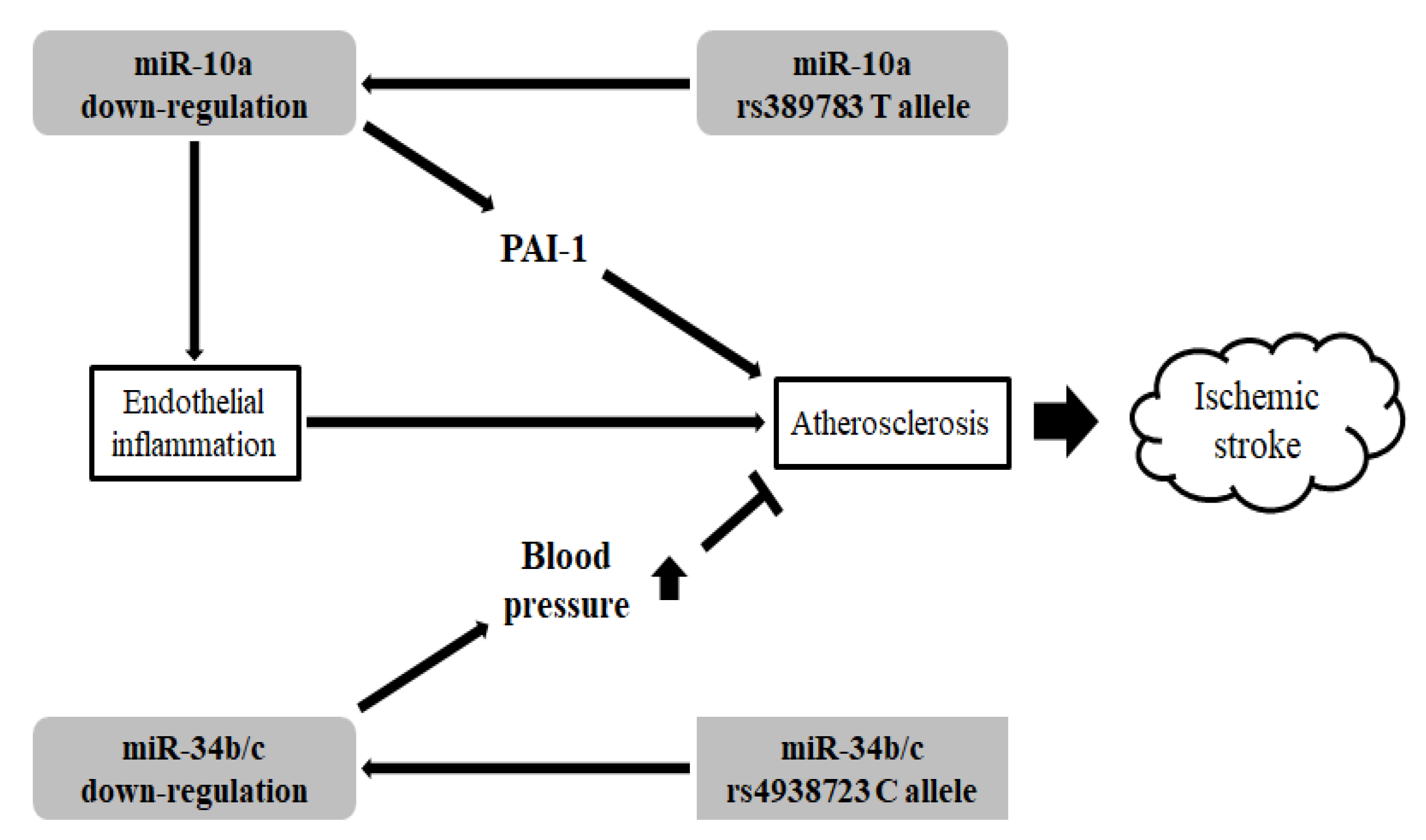
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Grysiewicz, R.A.; Thomas, K.; Pandey, D.K. Epidemiology of Ischemic and Hemorrhagic Stroke: Incidence, Prevalence, Mortality, and Risk Factors. Neurol. Clin. 2008, 26, 871–895. [Google Scholar] [CrossRef] [PubMed]
- Soto-Cámara, R.; González-Santos, J.; González-Bernal, J.; Martín-Santidrian, A.; Cubo, E.; Trejo-Gabriel-Galán, J. Factors Associated with Shortening of Prehospital Delay among Patients with Acute Ischemic Stroke. J. Clin. Med. 2019, 8, 1712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guzik, A.; Bushnell, C. Stroke Epidemiology and Risk Factor Management. Continuum (Minneap. Minn.) 2017, 23, 15–39. [Google Scholar] [CrossRef] [PubMed]
- Makris, K.; Haliassos, A.; Chondrogianni, M.; Tsivgoulis, G. Blood biomarkers in ischemic stroke: Potential role and challenges in clinical practice and research. Crit. Rev. Clin. Lab. Sci. 2018, 55, 294–328. [Google Scholar] [CrossRef]
- Monbailliu, T.; Goossens, J.; Hachimi-Idrissi, S. Blood protein biomarkers as diagnostic tool for ischemic stroke: A systematic review. Biomark. Med. 2017, 11, 503–512. [Google Scholar] [CrossRef]
- Vijayan, M.; Reddy, P.H. Peripheral biomarkers of stroke: Focus on circulatory microRNAs. Biochim. Biophys. Acta 2016, 1862, 1984–1993. [Google Scholar] [CrossRef]
- Dewdney, B.; Trollope, A.; Moxon, J.; Manapurathe, D.T.; Biros, E.; Golledge, J. Circulating MicroRNAs as Biomarkers for Acute Ischemic Stroke: A Systematic Review. J. Stroke Cerebrovasc. Dis. 2018, 27, 522–530. [Google Scholar] [CrossRef]
- Iorio, M.V.; Croce, C.M. MicroRNA dysregulation in cancer: Diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol. Med. 2012, 4, 143–159. [Google Scholar] [CrossRef]
- Dragomir, M.P.; Mafra, A.C.P.; Dias, S.M.G.; Vasilescu, C.; Calin, G.A. Using microRNA Networks to Understand Cancer. Int. J. Mol. Sci. 2018, 19, 1871. [Google Scholar] [CrossRef] [Green Version]
- Lee, Y.S.; Dutta, A. MicroRNAs in cancer. Annu. Rev. Pathol. 2009, 4, 199–227. [Google Scholar] [CrossRef]
- Sheinerman, K.S.; Toledo, J.B.; Tsivinsky, V.G.; Irwin, D.; Grossman, M.; Weintraub, D.; Hurtig, H.I.; Chen-Plotkin, A.; Wolk, D.A.; McCluskey, L.F.; et al. Circulating brain-enriched microRNAs as novel biomarkers for detection and differentiation of neurodegenerative diseases. Alzheimer’s Res. Ther. 2017, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Otto, T.; Candido, S.V.; Pilarz, M.S.; Sicinska, E.; Bronson, R.T.; Bowden, M.; Lachowicz, I.A.; Mulry, K.; Fassl, A.; Han, R.C.; et al. Cell cycle-targeting microRNAs promote differentiation by enforcing cell-cycle exit. Proc. Natl. Acad. Sci. USA 2017, 114, 10660–10665. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Njock, M.-S.; Cheng, H.S.; Dang, L.T.; Nazari-Jahantigh, M.; Lau, A.C.; Boudreau, E.; Roufaiel, M.; Cybulsky, M.I.; Schober, A.; Fish, J.E. Endothelial cells suppress monocyte activation through secretion of extracellular vesicles containing antiinflammatory microRNAs. Blood 2015, 125, 3202–3212. [Google Scholar] [CrossRef] [Green Version]
- Zhao, L.; Wu, S.; Huang, E.; Gnatenko, D.; Bahou, W.F.; Zhu, W. Integrated micro/messenger RNA regulatory networks in essential thrombocytosis. PLoS ONE 2018, 13, e0191932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartel, D.P. MicroRNAs: Target Recognition and Regulatory Functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Williams, D.; Sur, S.; Wang, J.-Y.; Jo, H. Role of flow-sensitive microRNAs and long noncoding RNAs in vascular dysfunction and atherosclerosis. Vasc. Pharmacol. 2019, 114, 76–92. [Google Scholar] [CrossRef]
- Gaudet, A.D.; Fonken, L.K.; Watkins, L.R.; Nelson, R.J.; Popovich, P.G. MicroRNAs: Roles in Regulating Neuroinflammation. Neuroscientist 2018, 24, 221–245. [Google Scholar] [CrossRef] [Green Version]
- Tan, J.R.; Tan, K.S.; Yong, F.L.; Armugam, A.; Wang, C.W.; Jeyaseelan, K.; Wong, P.T.-H. MicroRNAs regulating cluster of differentiation 46 (CD46) in cardioembolic and non-cardioembolic stroke. PLoS ONE 2017, 12, e0172131. [Google Scholar] [CrossRef]
- Hermeking, H. The miR-34 family in cancer and apoptosis. Cell Death Differ. 2010, 17, 193–199. [Google Scholar] [CrossRef]
- Zhao, Y.; Pogue, A.I.; Lukiw, W.J. MicroRNA (miRNA) Signaling in the Human CNS in Sporadic Alzheimer’s Disease (AD)-Novel and Unique Pathological Features. Int. J. Mol. Sci. 2015, 16, 30105–30116. [Google Scholar] [CrossRef] [Green Version]
- Kabaria, S.; Choi, D.C.; Chaudhuri, A.D.; Mouradian, M.M.; Junn, E. Inhibition of miR-34b and miR-34c enhances α-synuclein expression in Parkinson’s disease. FEBS Lett. 2015, 589, 319–325. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Caputo, V.; Ciolfi, A.; Macrì, S.; Pizzuti, A. The emerging role of MicroRNA in schizophrenia. CNS Neurol. Disord. Drug Targets 2015, 14, 208–221. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, S.S.; Nygaard, A.-B.; Nielsen, M.-Y.; Jensen, K.; Christensen, T. miRNA Expression Profiles in Cerebrospinal Fluid and Blood of Patients with Acute Ischemic Stroke. Transl. Stroke Res. 2014, 5, 711–718. [Google Scholar] [CrossRef] [PubMed]
- Rink, C.; Khanna, S. MicroRNA in ischemic stroke etiology and pathology. Physiol. Genom. 2010, 43, 521–528. [Google Scholar] [CrossRef] [Green Version]
- Sessa, F.; Maglietta, F.; Bertozzi, G.; Salerno, M.; Di Mizio, G.; Messina, G.; Montana, A.; Ricci, P.; Pomara, C. Human Brain Injury and miRNAs: An Experimental Study. Int. J. Mol. Sci. 2019, 20, 1546. [Google Scholar] [CrossRef] [Green Version]
- Winter, J.; Jung, S.; Keller, S.; Gregory, R.I.; Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat. Cell Biol. 2009, 11, 228–234. [Google Scholar] [CrossRef]
- Xiao, S.; Ma, Y.; Zhu, H.; Sun, H.; Yin, Y.; Feng, G. miRNA functional synergistic network analysis of mice with ischemic stroke. Neurol. Sci. 2015, 36, 143–148. [Google Scholar] [CrossRef]
- Miao, W.; Bao, T.-H.; Han, J.-H.; Yin, M.; Zhang, J.; Yan, Y.; Zhu, Y.-H. Neuroprotection induced by post-conditioning following ischemia/reperfusion in mice is associated with altered microRNA expression. Mol. Med. Rep. 2016, 14, 2582–2588. [Google Scholar] [CrossRef] [Green Version]
- Mirzaei, H.; Momeni, F.; Saadatpour, L.; Sahebkar, A.; Goodarzi, M.; Masoudifar, A.; Kouhpayeh, S.; Salehi, H.; Mirzaei, H.R.; Jaafari, M.R. MicroRNA: Relevance to stroke diagnosis, prognosis, and therapy. J. Cell. Physiol. 2018, 233, 856–865. [Google Scholar] [CrossRef]
- Li, G.; Morris-Blanco, K.C.; Lopez, M.; Yang, T.; Zhao, H.; Vemuganti, R.; Luo, Y. Impact of microRNAs on ischemic stroke: From pre- to post-disease. Prog. Neurobiol. 2018, 59–78. [Google Scholar] [CrossRef]
- Vijayan, M.; Kumar, S.; Yin, X.; Zafer, D.; Chanana, V.; Cengiz, P.; Reddy, P.H. Identification of novel circulatory microRNA signatures linked to patients with ischemic stroke. Hum. Mol. Genet. 2018, 27, 2318–2329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khoshnam, S.E.; Winlow, W.; Farzaneh, M. The Interplay of MicroRNAs in the Inflammatory Mechanisms Following Ischemic Stroke. J. Neuropathol. Exp. Neurol. 2017, 76, 548–561. [Google Scholar] [CrossRef]
- Paschon, V.; Takada, S.H.; Ikebara, J.M.; Sousa, E.; Raeisossadati, R.; Ulrich, H.D.; Kihara, A.H. Interplay Between Exosomes, microRNAs and Toll-Like Receptors in Brain Disorders. Mol. Neurobiol. 2015, 53, 2016–2028. [Google Scholar] [CrossRef]
- Exin, H.; Eli, Y.; Echopp, M. Exosomes/miRNAs as mediating cell-based therapy of stroke. Front. Cell. Neurosci. 2014, 8, 377. [Google Scholar]
- Eyileten, C.; Wicik, Z.; De Rosa, S.; Mirowska-Guzel, D.M.; Soplinska, A.; Indolfi, C.; Kurkowska-Jastrzebska, I.; Członkowska, A.; Postula, M. MicroRNAs as Diagnostic and Prognostic Biomarkers in Ischemic Stroke—A Comprehensive Review and Bioinformatic Analysis. Cells 2018, 7, 249. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Jiang, C. MicroRNA Expression Profiles Identify Biomarker for Differentiating the Embolic Stroke from Thrombotic Stroke. BioMed Res. Int. 2018, 2018, 4514178. [Google Scholar] [CrossRef]
- Li, C.-X.; Weng, H.; Zheng, J.; Feng, Z.-H.; Ou, J.-L.; Liao, W.-J. Association Between MicroRNAs Polymorphisms and Risk of Ischemic Stroke: A Meta-Analysis in Chinese Individuals. Front. Aging Neurosci. 2018, 10, 82. [Google Scholar] [CrossRef] [Green Version]
- Huang, S.; Zhou, S.; Zhang, Y.; Lv, Z.; Li, S.; Xie, C.; Ke, Y.; Deng, P.; Geng, Y.; Zhang, Q.; et al. Association of the Genetic Polymorphisms in Pre-MicroRNAs with Risk of Ischemic Stroke in a Chinese Population. PLoS ONE 2015, 10, e0117007. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.; Xu, G.; Cai, B.; Zhang, H.; Zhu, W.; Liu, X. Genetic Variants in MicroRNAs Predict Recurrence of Ischemic Stroke. Mol. Neurobiol. 2016, 54, 2776–2780. [Google Scholar] [CrossRef]
- Zhou, G.; Lin, W.; Fang, P.; Lin, X.; Zhuge, L.; Hu, Z.; Jin, L. MiR-10a improves hepatic fibrosis by regulating the TGFbetal/Smads signal transduction pathway. Exp. Ther. Med. 2016, 12, 1719–1722. [Google Scholar] [CrossRef] [Green Version]
- Huang, R.; Ma, J.; Niu, B.; Li, J.; Chang, J.; Zhang, Y.; Liu, P.; Luan, X. MiR-34b Protects Against Focal Cerebral Ischemia-Reperfusion (I/R) Injury in Rat by Targeting Keap1. J. Stroke Cerebrovasc. Dis. 2019, 28, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Li, D.-B.; Liu, J.-L.; Wang, W.; Luo, X.-M.; Zhou, X.; Li, J.-P.; Cao, X.-L.; Long, X.-H.; Chen, J.-G.; Qin, C. Plasma Exosomal miRNA-122-5p and miR-300-3p as Potential Markers for Transient Ischaemic Attack in Rats. Front. Aging Neurosci. 2018, 10, 24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, C.; Zhu, H.-Y.; Bai, W.-D.; Su, L.; Liu, J.-Q.; Cai, W.-X.; Zhao, B.; Gao, J.-X.; Han, S.-C.; Li, J.; et al. MiR-10a and miR-181c regulate collagen type I generation in hypertrophic scars by targeting PAI-1 and uPA. FEBS Lett. 2014, 589, 380–389. [Google Scholar] [CrossRef] [PubMed]
- Francis, S.M.S.; Davidson, M.R.; Tan, M.E.; Wright, C.M.; Clarke, B.E.; Duhig, E.E.; Bowman, R.V.; Hayward, N.K.; Fong, K.M.; Yang, I.A. MicroRNA-34c is associated with emphysema severity and modulates SERPINE1 expression. BMC Genom. 2014, 15, 88. [Google Scholar]
- Fernandes, T.; Hashimoto, N.Y.; Magalhães, F.C.; Fernandes, F.B.; Casarini, D.E.; Carmona, A.K.; Krieger, J.E.; Phillips, M.I.; Oliveira, E.M. Aerobic exercise training-induced left ventricular hypertrophy involves regulatory MicroRNAs, decreased angiotensin-converting enzyme-angiotensin ii, and synergistic regulation of angiotensin-converting enzyme 2-angiotensin (1-7). Hypertension 2011, 58, 182–189. [Google Scholar] [CrossRef] [Green Version]
- Gu, Q.; Wang, B.; Zhang, X.-F.; Ma, Y.-P.; Liu, J.-D.; Wang, X.-Z. Contribution of renin-angiotensin system to exercise-induced attenuation of aortic remodeling and improvement of endothelial function in spontaneously hypertensive rats. Cardiovasc. Pathol. 2014, 23, 298–305. [Google Scholar] [CrossRef]
- Tasdemir, S.; Erdem, H.B.; Sahin, I.; Ozel, L.; Ozdemir, G.; Eroz, R.; Tatar, A. Correlation with Platelet Parameters and Genetic Markers of Thrombophilia Panel (Factor II g.20210G>A, Factor V Leiden, MTHFR (C677T, A1298C), PAI-1, beta-Fibrinogen, Factor XIIIA (V34L), Glycoprotein IIIa (L33P)) in Ischemic Strokes. NeuroMolecular Med. 2016, 18, 170–176. [Google Scholar] [CrossRef]
- Selim, M.; Savitz, S.I.; Linfante, I.; Caplan, L.R.; Schlaug, G. Effect of pre-stroke use of ACE inhibitors on ischemic stroke severity. BMC Neurol. 2005, 5, 10. [Google Scholar] [CrossRef] [Green Version]
- Zonneveld, T.P.; Richard, E.; Vergouwen, M.D.; Nederkoorn, P.J.; de Haan, R.; Roos, Y.B.; Kruyt, N.D. Blood pressure-lowering treatment for preventing recurrent stroke, major vascular events, and dementia in patients with a history of stroke or transient ischaemic attack. Cochrane Database Syst. Rev. 2018, 7, CD007858. [Google Scholar] [CrossRef] [Green Version]
- Wei, L.K.; Au, A.; Menon, S.; Gan, S.H.; Griffiths, L.R. Clinical Relevance of MTHFR, eNOS, ACE, and ApoE Gene Polymorphisms and Serum Vitamin Profile among Malay Patients with Ischemic Stroke. J. Stroke Cerebrovasc. Dis. 2015, 24, 2017–2025. [Google Scholar] [CrossRef]
- Zhang, Z.; Ran, Y.; Shaw, T.S.; Peng, Y. MicroRNAs 10a and 10b Regulate the Expression of Human Platelet Glycoprotein Ibalpha for Normal Megakaryopoiesis. Int. J. Mol. Sci. 2016, 17, 1873. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, L.; Chen, B.; Li, S.; Wei, X.; Liu, T.; Huang, Y.; Lin, X. Plasma miR-10a: A Potential Biomarker for Coronary Artery Disease. Dis. Markers 2016, 2016, 3841927. [Google Scholar] [CrossRef] [Green Version]
- Yasmeen, S.; Kaur, S.; Mirza, A.H.; Brodin, B.; Pociot, F.; Kruuse, C. miRNA-27a-3p and miRNA-222-3p as Novel Modulators of Phosphodiesterase 3a (PDE3A) in Cerebral Microvascular Endothelial Cells. Mol. Neurobiol. 2019, 56, 5304–5314. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Bao, J.; Kim, M.; Yuan, S.; Tang, C.; Zheng, H.; Mastick, G.S.; Xu, C.; Yan, W. Two miRNA clusters, miR-34b/c and miR-449, are essential for normal brain development, motile ciliogenesis, and spermatogenesis. Proc. Natl. Acad. Sci. USA 2014, 111, E2851–E2857. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Wang, X.-Q.; Zhang, L.; Lv, X.-D.; Su, X.; Tian, S.; Liu, C.-M.; Ma, X.; Xia, H.-F. A SNP in pri-miR-10a is associated with recurrent spontaneous abortion in a Han-Chinese population. Oncotarget 2016, 7, 8208–8222. [Google Scholar] [CrossRef] [Green Version]
- Liao, Y.; Chen, K.-H.; Dong, X.-M.; Fang, Y.; Li, W.-G.; Huang, G.-Y.; Song, W. A role of pre-mir-10a coding region variant in host susceptibility to coxsackie virus-induced myocarditis. Eur. Rev. Med. Pharmacol. Sci. 2015, 19, 3500–3507. [Google Scholar]
- Xu, C.; Yang, C.; Zhang, A.; Xu, Y.; Li, X.; Liu, Z.; Liu, S.; Sun, N.; Zhang, K. The interaction of miR-34b/c polymorphisms and negative life events increases susceptibility to major depressive disorder in Han Chinese population. Neurosci. Lett. 2017, 651, 65–71. [Google Scholar] [CrossRef]
- Yang, B.; Song, B.; Yan, G.; Hao, H. rs11671784 G/A and rs895819 A/G Polymorphisms Inversely Affect Gastric Cancer Susceptibility and miR-27a Expression in a Chinese Population. Med. Sci. Monit. 2014, 20, 2318–2326. [Google Scholar] [CrossRef] [Green Version]
- Adams, H.P.; Bendixen, B.H.; Kappelle, L.J.; Biller, J.; Love, B.B.; Gordon, D.L.; Marsh, E.E. Classification of subtype of acute ischemic stroke. Definitions for use in a multicenter clinical trial. TOAST. Trial of Org 10172 in Acute Stroke Treatment. Stroke 1993, 24, 35–41. [Google Scholar] [CrossRef] [Green Version]
- Ryu, C.S.; Bae, J.; Kim, I.J.; Kim, J.; Oh, S.H.; Kim, O.J.; Kim, N.K. MPG and NPRL3 Polymorphisms are Associated with Ischemic Stroke Susceptibility and Post-Stroke Mortality. Diagnostics 2020, 10, 947. [Google Scholar] [CrossRef]
- Kim, I.J.; Kim, S.H.; Cha, D.H.; Lim, S.W.; Moon, J.Y.; Kim, J.O.; Ryu, C.S.; Park, H.S.; Sung, J.H.; Kim, N.K. Association of COX2 −765G>C promoter polymorphism and coronary artery disease in Korean population. Genes Genom. 2019, 41, 1055–1062. [Google Scholar] [CrossRef] [PubMed]
- Maitrias, P.; Meuth, V.M.-L.; Nader, J.; Reix, T.; Caus, T.; Metzinger, L. The Involvement of miRNA in Carotid-Related Stroke. Arter. Thromb. Vasc. Biol. 2017, 37, 1608–1617. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rokavec, M.; Li, H.; Jiang, L.; Hermeking, H. The p53/miR-34 axis in development and disease. J. Mol. Cell Biol. 2014, 6, 214–230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Ling, C.C.; Li, L.; Qin, Y.; Qi, J.; Liu, X.; You, B.; Shi, Y.; Zhang, J.; Jiang, Q.; et al. MicroRNA-10a/10b represses a novel target gene mib1 to regulate angiogenesis. Cardiovasc. Res. 2016, 110, 140–150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, H.; Fan, L.; Zhan, R.; Wu, S.; Niu, W. Expression of microRNA-10a, microRNA-342-3p and their predicted target gene TIAM1 in extranodal NK/T-cell lymphoma, nasal type. Oncol. Lett. 2016, 11, 345–351. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goyal, R.; Goyal, D.; Leitzke, A.; Gheorghe, C.P.; Longo, L.D. Brain Renin-Angiotensin System: Fetal Epigenetic Programming by Maternal Protein Restriction During Pregnancy. Reprod. Sci. 2009, 17, 227–238. [Google Scholar] [CrossRef]
- Jickling, G.C.; Ander, B.P.; Zhan, X.; Noblett, D.; Stamova, B.; Liu, D. microRNA Expression in Peripheral Blood Cells following Acute Ischemic Stroke and Their Predicted Gene Targets. PLoS ONE 2014, 9, e99283. [Google Scholar] [CrossRef]
- Hu, X.; Zan, X.; Xie, Z.; Li, Y.; Lin, S.; Li, H.; You, C. Association Between Plasminogen Activator Inhibitor-1 Genetic Polymorphisms and Stroke Susceptibility. Mol. Neurobiol. 2017, 54, 328–341. [Google Scholar] [CrossRef]
- Sabino, A.D.P.; Ribeiro, D.D.; Domingueti, C.P.; Dos Santos, M.S.; Gadelha, T.; Dusse, L.M.S.; Carvalho, M.D.G.; Fernandes, A.P. Plasminogen activator inhibitor-1 4G/5G promoter polymorphism and PAI-1 plasma levels in young patients with ischemic stroke. Mol. Biol. Rep. 2011, 38, 5355–5360. [Google Scholar] [CrossRef]
- Raafat, I.R.; Amer, R.A.; Abozeid, A.A.; Elsharaby, R.M.; Shafik, N.M. Micro RNA 146a gene variant/TNF-α/IL-6/IL-1 beta; A cross-link axis inbetween oxidative stress, endothelial dysfunction and neuro-inflammation in acute ischemic stroke and chronic schizophrenic patients. Arch. Biochem. Biophys. 2019, 679, 108193. [Google Scholar]
- Li, J.; Li, K.; Chen, X. Inflammation-regulatory microRNAs: Valuable targets for intracranial atherosclerosis. J. Neurosci. Res. 2019, 97, 1242–1252. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.K.; Vaughan, D.E. PAI-1 in tissue fibrosis. J. Cell. Physiol. 2012, 227, 493–507. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, S.; Wei, X.; He, J.; Tian, X.; Yuan, S.; Sun, L. Plasminogen activator inhibitor-1 in cancer research. Biomed. Pharmacother. 2018, 105, 83–94. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Cheng, J.; Guo, X.; Mo, J.; Gao, B.; Zhou, H.; Wu, Y.; Li, Z. The roles of PAI-1 gene polymorphisms in atherosclerotic diseases: A systematic review and meta-analysis involving 149,908 subjects. Gene 2018, 673, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Rabieian, R.; Boshtam, M.; Zareei, M.; Kouhpayeh, S.; Masoudifar, A.; Mirzaei, H. Plasminogen Activator Inhibitor Type-1 as a Regulator of Fibrosis. J. Cell. Biochem. 2018, 119, 17–27. [Google Scholar] [CrossRef]
- Fogo, A.B.; Vaughan, U.E.; Fogo, M.A.B. Compound interest: ACE and PAI-1 polymorphisms and risk of thrombosis and fibrosis. Kidney Int. 1998, 54, 1765–1766. [Google Scholar] [CrossRef] [Green Version]
- Malueka, R.G.; Dwianingsih, E.K.; Sutarni, S.; Bawono, R.G.; Bayuangga, H.F.; Gofir, A.; Setyopranoto, I. The D allele of the angiotensin-converting enzyme (ACE) insertion/deletion (I/D) polymorphism is associated with worse functional outcome of ischaemic stroke. Int. J. Neurosci. 2017, 128, 697–704. [Google Scholar] [CrossRef]
- Jeon, Y.J.; Kim, O.-J.; Kim, S.Y.; Oh, S.H.; Oh, D.; Shin, B.S.; Kim, N.K. Association of the miR-146a, miR-149, miR-196a2, and miR-499 Polymorphisms with Ischemic Stroke and Silent Brain Infarction Risk. Arter. Thromb. Vasc. Biol. 2013, 33, 420–430. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.; Choi, G.H.; Ko, K.H.; Kim, J.O.; Oh, S.H.; Park, Y.S.; Kim, O.J.; Kim, N.K. Association of the Single Nucleotide Polymorphisms in microRNAs 130b, 200b, and 495 with Ischemic Stroke Susceptibility and Post-Stroke Mortality. PLoS ONE 2016, 11, e0162519. [Google Scholar] [CrossRef]
- Zhu, R.; Wang, Q.W.; Zhao, J.; Liu, X.; He, Z.T. miR-149 and miR-499 gene polymorphism and the incident of ischemic stroke in the Asian population: From a case-control study to meta-analysis. Clin. Neurol. Neurosurg. 2020, 193, 105789. [Google Scholar] [CrossRef]
- Zou, D.; Liu, C.; Zhang, Q.; Li, X.; Qin, G.; Huang, Q.; Meng, Y.; Chen, L.; Wei, J. Association between polymorphisms in microRNAs and ischemic stroke in an Asian population: Evidence based on 6083 cases and 7248 controls. Clin. Interv. Aging 2018, 13, 1709–1726. [Google Scholar] [CrossRef] [Green Version]
- Jiang, H.; Toscano, J.F.; Song, S.S.; Schlick, K.H.; Dumitrascu, O.M.; Pan, J.; Lyden, P.D.; Saver, J.L.; Gonzalez, N. Differential expression of circulating exosomal microRNAs in refractory intracranial atherosclerosis associated with antiangiogenesis. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Liao, C.; Jian, C.; Liang, L.; Liu, J.; Tang, Y.; Wei, Y.-S. Evaluation of miR-34b/c polymorphisms to the risk of ischemic stroke. J. Hypertens. 2020, 38, 1481–1487. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Li, H.; Du, Y.; Shi, Q.; Zhao, L. Downregulation of microRNA-34b is responsible for the elevation of blood pressure in spontaneously hypertensive rats. Mol. Med. Rep. 2017, 15, 1031–1036. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Characteristics | Controls (n = 403) | Stroke Patients (n = 530) | p-Value | LAD Patients (n = 184) | p-Value | SVD Patients (n = 151) | p-Value | CE Patients (n = 56) | p-Value |
|---|---|---|---|---|---|---|---|---|---|
| Age (years, mean ± SD) | 62.70 ± 11.55 | 63.14 ± 11.63 | 0.566 | 63.80 ± 11.16 | 0.281 | 60.88 ± 11.61 | 0.100 | 67.13 ± 12.65 | 0.008 |
| Male (%) | 172 (42.7) | 245 (46.2) | 0.281 | 82 (44.6) | 0.669 | 76 (41.3) | 0.107 | 24 (42.9) | 0.980 |
| Smoking (%) | 137 (34.0) | 208 (39.2) | 0.083 | 71 (38.6) | 0.241 | 56 (30.4) | 0.464 | 18 (32.1) | 0.784 |
| Hypertension (%) | 162 (40.2) | 342 (64.5) | <0.0001 | 117 (63.6) | <0.0001 | 94 (51.1) | <0.0001 | 32 (57.1) | 0.016 |
| Hyperlipidemia (%) | 81 (20.1) | 161 (30.4) | 0.0004 | 63 (34.2) | 0.0002 | 47 (25.5) | 0.006 | 12 (21.4) | 0.817 |
| Diabetes mellitus (%) | 52 (12.9) | 148 (27.9) | <0.0001 | 48 (26.1) | 0.0001 | 45 (24.5) | <0.0001 | 14 (25.0) | 0.016 |
| MetS (%) | 60 (14.9) | 212 (40.0) | <0.0001 | 85 (46.2) | <0.0001 | 60 (32.6) | <0.0001 | 19 (33.9) | 0.0004 |
| Genotypes | Controls (n = 402) | Stroke Patients (n = 530) | AOR (95% CI) | p-Value | LAD (n = 184) | AOR (95% CI) | p-Value | SVD (n = 151) | AOR (95% CI) | p-Value | CE (n = 56) | AOR (95% CI) | p-Value |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| miR-10a rs3809783 A>T | |||||||||||||
| AA | 340 (84.6) | 435 (82.1) | 1.000 (reference) | 152 (82.6) | 1.000 (reference) | 121 (80.1) | 1.000 (reference) | 48 (85.7) | 1.000 (reference) | ||||
| AT | 58 (14.4) | 89 (16.8) | 1.126 (0.770–1.649) | 0.540 | 30 (16.3) | 1.107 (0.679–1.804) | 0.685 | 26 (17.2) | 1.256 (0.731–2.157) | 0.409 | 8 (14.3) | 1.008 (0.447–2.274) | 0.984 |
| TT | 4 (1.0) | 6 (1.1) | 1.172 (0.307–4.481) | 0.816 | 2 (1.1) | 0.999 (0.176–5.663) | 0.999 | 4 (2.6) | 3.103 (0.678–14.204) | 0.145 | 0 (0.0) | NA | 0.994 |
| Dominant (AA vs AT + TT) | 1.129 (0.780–1.636) | 0.520 | 1.103 (0.686–1.774) | 0.686 | 1.370 (0.817–2.297) | 0.232 | 0.949 (0.422–2.135) | 0.900 | |||||
| Recessive (AA + AT vs TT) | 1.153 (0.302–4.409) | 0.835 | 1.045 (0.186–5.877) | 0.961 | 2.962 (0.649–13.522) | 0.161 | NA | 0.994 | |||||
| HWE-P | 0.393 | 0.549 | |||||||||||
| miR-27a rs895819 T>C | |||||||||||||
| TT | 203 (50.5) | 254 (47.9) | 1.000 (reference) | 88 (47.8) | 1.000 (reference) | 73 (48.3) | 1.000 (reference) | 34 (60.7) | 1.000 (reference) | ||||
| TC | 158 (39.3) | 223 (42.1) | 1.145 (0.858–1.529) | 0.358 | 80 (43.5) | 0.963 (0.658–1.408) | 0.844 | 63 (41.7) | 1.135 (0.746–1.726) | 0.555 | 17 (30.4) | 0.653 (0.348–1.225) | 0.184 |
| CC | 41 (10.2) | 53 (10.0) | 1.090 (0.678–1.753) | 0.723 | 16 (8.7) | 1.206 (0.670–2.172) | 0.532 | 15 (9.9) | 1.396 (0.698–2.793) | 0.345 | 5 (8.9) | 0.835 (0.301–2.316) | 0.729 |
| Dominant (TT vs TC + CC) | 1.139 (0.866–1.499) | 0.352 | 1.002 (0.703–1.429) | 0.991 | 1.183 (0.796–1.759) | 0.406 | 0.693 (0.387–1.240) | 0.217 | |||||
| Recessive (TT + TC vs CC) | 1.028 (0.654–1.616) | 0.906 | 1.216 (0.699–2.115) | 0.489 | 1.322 (0.685–2.549) | 0.406 | 0.992 (0.367–2.680) | 0.987 | |||||
| HWE-P | 0.217 | 0.693 | |||||||||||
| miR-34b/c rs4938723 T>C | |||||||||||||
| TT | 204 (50.7) | 295 (55.7) | 1.000 (reference) | 100 (54.3) | 1.000 (reference) | 87 (57.6) | 1.000 (reference) | 27 (48.2) | 1.000 (reference) | ||||
| TC | 161 (40.0) | 197 (37.2) | 0.839 (0.627–1.123) | 0.238 | 72 (39.1) | 1.055 (0.728–1.527) | 0.778 | 51 (33.8) | 0.732 (0.476–1.125) | 0.155 | 27 (48.2) | 1.269 (0.706–2.281) | 0.426 |
| CC | 37 (9.2) | 38 (7.2) | 0.703 (0.419–1.179) | 0.182 | 12 (6.5) | 0.852 (0.440–1.651) | 0.636 | 13 (8.6) | 0.883 (0.431–1.811) | 0.734 | 2 (3.6) | 0.374 (0.083–1.684) | 0.200 |
| Dominant (TT vs TC + CC) | 0.816 (0.620–1.074) | 0.147 | 1.008 (0.708–1.435) | 0.964 | 0.766 (0.515–1.141) | 0.190 | 1.080 (0.609-1.914) | 0.793 | |||||
| Recessive (TT + TC vs CC) | 0.762 (0.461–1.261) | 0.291 | 0.816 (0.432–1.541) | 0.531 | 1.022 (0.510–2.049) | 0.952 | 0.310 (0.071–1.359) | 0.120 | |||||
| HWE-P | 0.522 | 0.518 | |||||||||||
| miR-300 rs12894467 T>C | |||||||||||||
| TT | 231 (57.5) | 311 (58.7) | 1.000 (reference) | 114 (62.0) | 1.000 (reference) | 92 (60.9) | 1.000 (reference) | 23 (41.1) | 1.000 (reference) | ||||
| TC | 145 (36.1) | 195 (36.8) | 1.035 (0.774–1.384) | 0.818 | 61 (33.2) | 1.119 (0.771–1.625) | 0.554 | 52 (34.4) | 0.922 (0.605–1.404) | 0.705 | 30 (53.6) | 2.069 (1.141–3.753) | 0.017 |
| CC | 26 (6.5) | 24 (4.5) | 0.729 (0.397–1.341) | 0.309 | 9 (4.9) | 0.820 (0.375–1.791) | 0.618 | 7 (4.6) | 0.733 (0.298–1.800) | 0.498 | 3 (5.4) | 1.175 (0.317–4.359) | 0.809 |
| Dominant (TT vs TC + CC) | 0.986 (0.747–1.303) | 0.923 | 1.070 (0.749–1.531) | 0.709 | 0.898 (0.600–1.344) | 0.600 | 1.931 (1.078–3.459) | 0.027 | |||||
| Recessive (TT + TC vs CC) | 0.720 (0.397–1.306) | 0.280 | 0.782 (0.365–1.676) | 0.527 | 0.758 (0.313–1.838) | 0.540 | 0.823 (0.235–2.879) | 0.760 | |||||
| HWE-P | 0.615 | 0.343 | |||||||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ryu, C.S.; Oh, S.H.; Lee, K.O.; Park, H.S.; An, H.J.; Lee, J.Y.; Ko, E.J.; Park, H.W.; Kim, O.J.; Kim, N.K. MiR-10a, 27a, 34b/c, and 300 Polymorphisms are Associated with Ischemic Stroke Susceptibility and Post-Stroke Mortality. Life 2020, 10, 309. https://doi.org/10.3390/life10120309
Ryu CS, Oh SH, Lee KO, Park HS, An HJ, Lee JY, Ko EJ, Park HW, Kim OJ, Kim NK. MiR-10a, 27a, 34b/c, and 300 Polymorphisms are Associated with Ischemic Stroke Susceptibility and Post-Stroke Mortality. Life. 2020; 10(12):309. https://doi.org/10.3390/life10120309
Chicago/Turabian StyleRyu, Chang Soo, Seung Hun Oh, Kee Ook Lee, Han Sung Park, Hui Jeong An, Jeong Yong Lee, Eun Ju Ko, Hyeon Woo Park, Ok Joon Kim, and Nam Keun Kim. 2020. "MiR-10a, 27a, 34b/c, and 300 Polymorphisms are Associated with Ischemic Stroke Susceptibility and Post-Stroke Mortality" Life 10, no. 12: 309. https://doi.org/10.3390/life10120309





