TMT-Based Proteomic Analysis of Continuous Cropping Response in Codonopsis tangshen Oliv.
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals and Instrumentation
2.2. Plant Materials and Sample Collection
2.3. Measurement of Physiological and Biochemical Properties
2.4. Protein Extraction and Digestion
2.5. Labelling and Peptide Fractionation
2.6. Liquid Chromatography–Mass Spectrometry (LC-MS/MS) Analysis and Database Search
2.7. Protein Quantification and Bioinformatics Analysis
2.8. Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR) Analysis
3. Results
3.1. Physiological and Biochemical Properties
3.2. Mass Spectrometry Identification
3.3. Gene Ontology (GO) Analysis of DEPs in Response to Continuous Cropping
3.4. KEGG Pathway Analysis of Responsive DEPs
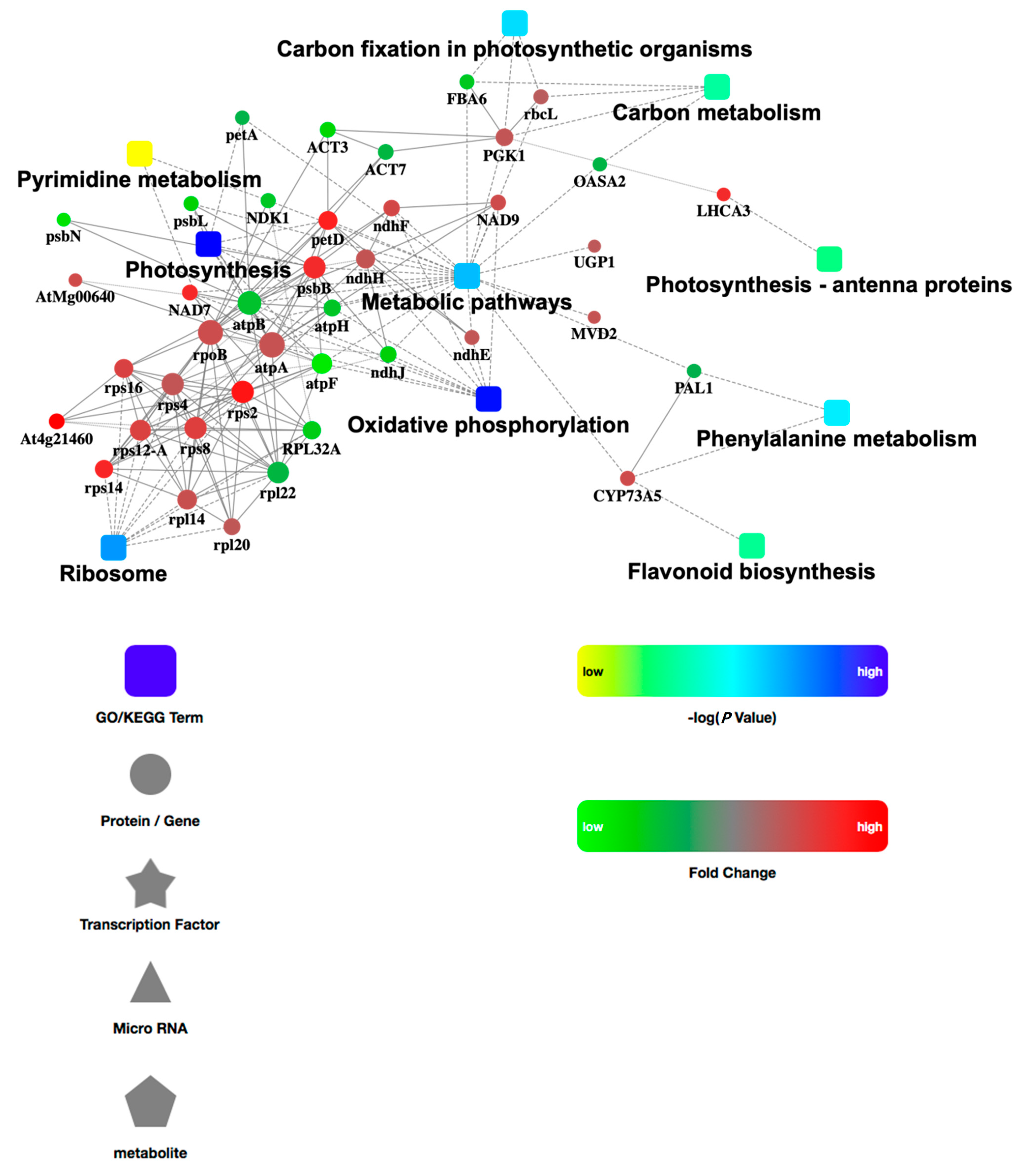
3.5. PPI Network Analysis
3.6. Validation of the DEPs by qRT-PCR
4. Discussion
4.1. Physiological and Biochemical Response in Continuously Cropped C. tangshen
4.2. Proteomic Patterns of C. tangshen in Response to Continuous Cropping
4.3. DEPs Related to Photosynthesis Pathway
4.4. DEPs Associated with Oxidative Phosphorylation
4.5. DEPs Related to Ribosome Activity
4.6. DEPs Related to Secondary Metabolites
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lin, L.C.; Tsai, T.H.; Chao, L.K. Chemical constituents comparison of Codonopsis tangshen, Codonopsis pilosula var. modesta and Codonopsis pilosula. Nat. Prod. Res. 2013, 27, 1812–1815. [Google Scholar] [CrossRef] [PubMed]
- He, J.Y.; Zhu, S.; Komatsu, K.; Yukihiro, G.; Cai, S.Q. Genetic polymorphism of medicinally-used Codonopsis species in internal transcribed spacer sequence of nuclear ribosomal DNA and its application to authenticate Codonopsis Radix. J. Nat. Med.-Tokyo 2013, 68, 112–124. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.M.; Luo, M.; Yao, X.D.; Yu, L. Purification, structural characterization, and antioxidant activity of the COP-W1 polysaccharide from Codonopsis tangshen Oliv. Carbohyd. Polym. 2020, 236, 116020. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhou, N.; Tan, J. Preliminary analysis of the quality difference of Codonopsis pilosula from different producing areas. Asia-Pac. Tradit. Med. 2017, 13, 37–39. [Google Scholar]
- He, Y.S.; Zhang, M.D.; Zhou, W.X.; Ai, L.Q.; You, J.W.; Liu, H.H.; You, J.M.; Wang, H.; Wassie, W.; Wang, M.; et al. Transcriptome analysis reveals novel insights into the continuous cropping induced response in Codonopsis tangshen, a medicinal herb. Plant Physiol. Bioch. 2019, 141, 279–290. [Google Scholar] [CrossRef]
- Tan, G.; Liu, Y.J.; Peng, S.G.; Yin, H.Q.; Meng, D.L.; Tao, J.M.; Gu, Y.B.; Li, J.; Yang, S.; Xiao, N.W.; et al. Soil potentials to resist continuous cropping obstacle: Three field cases. Environ. Res. 2021, 200, 111319. [Google Scholar] [CrossRef]
- Liu, Z.X.; Liu, J.J.; Yu, Z.H.; Yao, Q.; Li, Y.S.; Liang, A.Z.; Zhang, W.; Mi, G.; Jin, J.; Liu, X.B.; et al. Long-term continuous cropping of soybean is comparable to crop rotation in mediating microbial abundance, diversity and community composition. Soil. Till. Res. 2020, 197, 104503. [Google Scholar] [CrossRef]
- Zhang, X.H.; Lang, D.Y.; Zhang, E.H.; Wang, Z.S. Effect of Autotoxicity and Soil Microbes in Continuous Cropping Soil on Angelica sinensis Seedling Growth and Rhizosphere Soil Microbial Population. Chin. Herb. Med. 2015, 7, 88–93. [Google Scholar]
- Feng, F.J.; Yang, C.Y.; Li, M.J.; Zhan, S.Y.; Liu, H.Y.; Chen, A.G.; Wang, J.M.; Zhang, Z.Y.; Li, G. Key molecular events involved in root exudates-mediated replanted disease of Rehmannia glutinosa. Plant Physiol. Bioch. 2022, 172, 136–150. [Google Scholar] [CrossRef]
- Li, M.J.; Yang, Y.H.; Feng, F.J.; Zhang, B.; Chen, S.Q.; Yang, C.Y.; Li, G.; Wang, F.Q.; Zhang, J.Y.A.; Chen, A.G.; et al. Differential proteomic analysis of replanted Rehmannia glutinosa roots by iTRAQ reveals molecular mechanisms for formation of replant disease. BMC Plant Biol. 2017, 17, 116. [Google Scholar] [CrossRef] [Green Version]
- Tan, Y.; Cui, Y.S.; Li, H.Y.; Kuang, A.X.; Li, X.R.; Wei, Y.L.; Ji, X.L. Rhizospheric soil and root endogenous fungal diversity and composition in response to continuous Panax notoginseng cropping practices. Microbiol. Res. 2017, 194, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Li, J.R.; Chen, X.Z.; Zhan, R.T.; He, R. Transcriptome profiling reveals metabolic alteration in Andrographis paniculata in response to continuous cropping. Ind. Crop. Prod. 2019, 137, 585–596. [Google Scholar] [CrossRef]
- Ross, A.B.; Langer, J.D.; Jovanovic, M. Proteome Turnover in the Spotlight: Approaches, Applications, and Perspectives. Mol. Cell. Proteomics. 2021, 20, 100016. [Google Scholar] [CrossRef] [PubMed]
- Li, L.H.; Li, D.; Hu, T.; Fu, J. Antioxidant Enzyme Activity and Gene Expression in Response to Lead Stress in Perennial Ryegrass. J. Am. Soc. Hortic. Sci. Am. Soc. Hortic. Sci. 2012, 137, 80–85. [Google Scholar] [CrossRef] [Green Version]
- Ma, G.J.; Zhang, M.D.; Xu, J.L.; Zhou, W.X.; Cao, L.W. Transcriptomic analysis of short-term heat stress response in Pinellia ternata provided novel insights into the improved thermotolerance by spermidine and melatonin. Ecotoxicol. Environ. Saf. 2020, 202, 110877. [Google Scholar] [CrossRef]
- Jiang, X.; Wassie, M.; Zhou, W.; Wang, H.; You, J.; Ma, G.; Zhang, M. Transcriptomic Analysis Provides Novel Insights into the Heat Stress-Induced Response in Codonopsis tangshen. Life 2023, 13, 168. [Google Scholar] [CrossRef]
- Liu, X.; Qiu, H.Z.; Zhang, W.M.; Zhang, C.H.; Wang, Y.F.; Ma, X. Sink-source relationship of potato plants and its role involved in the reduction of tuber yield in continuous cropping system. Yingyong Shengtai Xuebao 2017, 28, 1571–1582. [Google Scholar]
- Gu, L.; Niu, M.M.; Zheng, H.Y.; Wang, J.M.; Wu, L.K.; Li, Z.F.; Zhang, Z.Y. Effect of continuous cropping of Rehmannia on its morphological and physiological characteristics. Zhong Yao Cai 2013, 36, 691–695. [Google Scholar]
- Zhao, G.; Hong, Y.; Li, L.; Zhang, H.; Xu, R.; Hao, Y. Selection and characterization of plant-derived alkaloids with strong antialgal inhibition: Growth inhibition selectivity and inhibitory mechanism. Harmful Algae 2022, 117, 102272. [Google Scholar] [CrossRef]
- Mao, R.J.; He, Z.G. Pinellia ternata (Thunb.) Breit: A review of its germplasm resources, genetic diversity and active components. J. Ethnopharmacol. 2020, 263, 113252. [Google Scholar] [CrossRef]
- Mierke, C.T. Cellular Mechanics and Biophysics, Structure and Function of Basic Cellular Components Regulating Cell Mechanics: Translation and Post-translational Modifications in Protein Biosynthesis; Springer: Cham, Switzerland, 2020; pp. 595–665. [Google Scholar]
- Chen, L.; Kashina, A. Post-translational Modifications of the Protein Termini. Front. Cell Dev. Biol. 2021, 9, 719590. [Google Scholar] [CrossRef]
- Zancani, M.; Braidot, E.; Filippi, A.; Lippe, G. Structural and functional properties of plant mitochondrial F-ATP synthase. Mitochondrion 2020, 53, 178–193. [Google Scholar] [CrossRef]
- Allen, J.F. Photosynthesis of ATP—Electrons, Proton Pumps, Rotors, and Poise. Cell 2002, 110, 273–276. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pagliano, C.; Saracco, G.; Barber, J. Structural, functional and auxiliary proteins of Photosystem II. Photosynth. Res. 2013, 116, 167–188. [Google Scholar] [CrossRef] [PubMed]
- Zou, Z.; Li, M.Y.; Jia, R.Z.; Zhao, H.; He, P.P.; Zhang, Y.L.; Guo, A.P. Genes encoding light-harvesting chlorophyll a/b-binding proteins in papaya (Carica papaya L.) and insight into lineage-specific evolution in Brassicaceae. Gene 2020, 748, 144685. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.H.; Li, X.P.; Razeghifard, R.; Anderson, J.M.; Niyogi, K.K.; Pogson, B.J.; Chow, W.S. The multiple roles of light-harvesting chlorophyll a/b-protein complexes define structure and optimize function of Arabidopsis chloroplasts: A study using two chlorophyll b-less mutants. BBA-Bioenerg. 2009, 1787, 973–984. [Google Scholar] [CrossRef] [Green Version]
- Stec, B. Structural mechanism of RuBisCO activation by carbamylation of the active site lysine. Proc. Natl. Acad. Sci. USA 2012, 109, 18785–18790. [Google Scholar] [CrossRef] [Green Version]
- Maheshwari, C.; Coe, R.; Karki, S.; Covshoff, S.; Tapia, R.; Tyagi, A.; Hibberd, J.M.; Furbank, R.T.; Quick, W.P.; Lin, H.C. Targeted knockdown of ribulose-1, 5-bisphosphate carboxylase-oxygenase in rice mesophyll cells. J. Plant Physiol. 2021, 260, 153395. [Google Scholar] [CrossRef]
- Quiles, M.J.; López, N.I. Photoinhibition of photosystems I and II induced by exposure to high light intensity during oat plant growth: Effects on the chloroplast NADH dehydrogenase complex. Plant Sci. 2004, 166, 815–823. [Google Scholar] [CrossRef]
- Reyes-Galindo, M.; Suarez, R.; Esparza-Perusquía, M.; de Lira-Sánchez, J.; Pardo, J.P.; Martínez, F.; Flores-Herrera, O. Mitochondrial respirasome works as a single unit and the cross-talk between complexes I, III2 and IV stimulates NADH dehydrogenase activity. BBA-Bioenerg. 2019, 1860, 618–627. [Google Scholar] [CrossRef]
- Bouché, N.; Fait, A.; Bouchez, D.; Moller, S.; Fromm, H. Mitochondrial succinic-semialdehyde dehydrogenase of the ??-aminobutyrate shunt is required to restrict level of reactive oxygen intermediate in plants. Proc. Natl. Acad. Sci. USA 2003, 100, 6843–6848. [Google Scholar] [CrossRef] [Green Version]
- Sweetlove, L.; Heazlewood, J.; Herald, V.; Day, D.; Leaver, C.; Millar, A. The impact of oxidative stress on Arabidopsis mitochondria. Plant J. 2003, 32, 891–904. [Google Scholar] [CrossRef] [PubMed]
- Byrne, M.E. A role for the ribosome in development. Trends Plant. Sci. 2009, 14, 512–519. [Google Scholar] [CrossRef] [PubMed]
- Horiguchi, G.; Van Lijsebettens, M.; Candela, H.; Micol, J.L.; Tsukaya, H. Ribosomes and translation in plant developmental control. Plant Sci. 2012, 191–192, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Schippers, J.; Mueller-Roeber, B. Ribosomal composition and control of leaf development. Plant Sci. 2010, 179, 307–315. [Google Scholar] [CrossRef]
- Motoyama, K.; Unno, H.; Hattori, A.; Takaoka, T.; Ishikita, H.; Kawaide, H.; Yoshimura, T.; Hemmi, H. A Single Amino Acid Mutation Converts (R)-5-Diphosphomevalonate Decarboxylase into a Kinase*. J. Biol. Chem. 2017, 292, 2457–2469. [Google Scholar] [CrossRef] [Green Version]
- Li, Z.H.; Wang, Q.; Ruan, X.; Pan, C.D.; Jiang, D.A. Phenolics and Plant Allelopathy. Molecules 2010, 15, 8933–8952. [Google Scholar] [CrossRef] [Green Version]
- Chin, D.C.; Senthil, K.R.; Suen, C.S.; Chien, C.Y.; Wang, M.J.; Hsu, C.H.; Xuhan, X.; Lai, Z.X.; Yeh, K.W. Plant Cytosolic Ascorbate Peroxidase with Dual Catalytic Activity Modulates Abiotic Stress Tolerances. Iscience 2019, 16, 31–49. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Liu, H.; Wang, Y.; Teng, R.M.; Liu, J.; Lin, S.; Zhuang, J. Cytosolic ascorbate peroxidase 1 modulates ascorbic acid metabolism through cooperating with nitrogen regulatory protein P-II in tea plant under nitrogen deficiency stress. Genomics 2020, 112, 3497–3503. [Google Scholar] [CrossRef]
- Zhang, Q.; Ma, C.; Xue, X.; Xu, M.; Li, J.; Wu, J.X. Overexpression of a Cytosolic Ascorbate Peroxidase Gene, OsAPX2, Increases Salt Tolerance in Transgenic Alfalfa. J. Integr. Agr. 2014, 13, 2500–2507. [Google Scholar] [CrossRef] [Green Version]
- Li, G.; Wu, Y.F.; Li, M.J.; Wang, F.Q.; Li, Z.F.; Yuan, F.Y.; Zhang, Z.Y. Over-immunity mediated abnormal deposition of lignin arrests the normal enlargement of the root tubers of Rehmannia glutinosa under consecutive monoculture stress. Plant Physiol. Bioch. 2021, 165, 36–46. [Google Scholar]
- Denness, L.; McKenna, J.; Segonzac, C.; Wormit, A.; Madhou, P.; Bennett, M.; Mansfield, J.; Zipfel, C.; Hamann, T. Cell Wall Damage-Induced Lignin Biosynthesis Is Regulated by a Reactive Oxygen Species- and Jasmonic Acid-Dependent Process in Arabidopsis. Plant Physiol. 2011, 156, 1364–1374. [Google Scholar] [CrossRef] [PubMed] [Green Version]





| Accession Number | Pathway and Annotation | Mean Ratio | MW [kDa] | pI | Score Mascot |
|---|---|---|---|---|---|
| Photosynthesis pathway | |||||
| A0A1U7AF74 | Cytochrome b6-f complex subunit 4 | 0.322 | 18.9 | 6.04 | 58 |
| A0A2D1PBS6 | Photosystem II 47 kDa protein | 0.368 | 56.1 | 6.90 | 665 |
| A0A291F2T6 | Photosystem II reaction center protein L | 2.211 | 4.5 | 4.50 | 112 |
| A0A1L6BTC9 | ATP synthase subunit alpha | 0.574 | 54.9 | 6.34 | 671 |
| A0A291F242 | ATP synthase subunit alpha | 0.571 | 54.9 | 6.04 | 600 |
| B9U4M6 | ATP synthase subunit beta | 0.104 | 50.5 | 5.36 | 1120 |
| B9U4I8 | ATP synthase subunit beta | 0.479 | 50.7 | 6.06 | 1198 |
| H6VP95 | ATP synthase subunit beta | 0.369 | 47.1 | 5.83 | 1173 |
| K9UVR2 | Chlorophyll a-b binding protein | 0.368 | 21.3 | 5.55 | 461 |
| A0A288W750 | ATPase subunit 4 | 0.524 | 21.9 | 9.69 | 26 |
| A0A075DZC7 | Protein PsbN | 2.621 | 4.7 | 4.28 | 31 |
| Carbon fixation in photosynthetic organisms | |||||
| T1WIC8 | Ribulose bisphosphate carboxylase large chain | 1.950 | 26.9 | 6.01 | 842 |
| A0A023Q1G8 | Ribulose bisphosphate carboxylase large chain | 0.488 | 41.1 | 7.61 | 2141 |
| A0A023Q3H2 | Ribulose bisphosphate carboxylase large chain | 0.629 | 41.7 | 7.21 | 2563 |
| U6BNI3 | Ribulose bisphosphate carboxylase large chain | 0.203 | 37.9 | 8.03 | 2216 |
| C7AQR5 | Ribulose bisphosphate carboxylase large chain | 0.615 | 23.2 | 7.80 | 553 |
| A0A023Q2I2 | Ribulose bisphosphate carboxylase large chain | 0.420 | 28.6 | 9.13 | 1361 |
| A0A023Q1M8 | Ribulose bisphosphate carboxylase large chain | 0.175 | 36.2 | 8.53 | 2133 |
| C7AQT3 | Ribulose bisphosphate carboxylase large chain | 2.386 | 50.1 | 7.02 | 1668 |
| H6VPA6 | Ribulose bisphosphate carboxylase large chain | 0.580 | 48.6 | 6.80 | 1569 |
| Q37184 | Ribulose bisphosphate carboxylase large chain | 0.639 | 52.5 | 6.70 | 2548 |
| C3S8M7 | Ribulose bisphosphate carboxylase large chain | 0.538 | 50.7 | 6.68 | 1821 |
| G8D4X5 | Ribulose bisphosphate carboxylase large chain | 1.511 | 22.4 | 6.05 | 607 |
| C3S8S0 | Ribulose bisphosphate carboxylase large chain | 0.374 | 50.2 | 6.71 | 1209 |
| A2VAL3 | Ribulose bisphosphate carboxylase large chain | 0.426 | 51.7 | 6.95 | 736 |
| A0A1L6BRS7 | Ribulose bisphosphate carboxylase large chain | 0.628 | 52.6 | 6.70 | 1522 |
| A6XBE1 | Ribulose-1,5-bisphosphate carboxylase/oxygenase | 0.319 | 20.3 | 7.27 | 1105 |
| A0A1D6XPB7 | Ribulose-1,5-bisphosphate carboxylase/oxygenase | 0.632 | 12.0 | 7.34 | 230 |
| Ribosome | |||||
| A0A240FG30 | 30S ribosomal protein S14 | 0.344 | 11.8 | 11.53 | 28 |
| A0A220D819 | 50S ribosomal protein L20 | 0.610 | 15.0 | 11.27 | 26 |
| A0A2D1PBT7 | Chloroplast 30S ribosomal protein S4 | 0.594 | 23.3 | 10.51 | 131 |
| A0A1Z2QRI8 | Ribosomal protein L14 | 0.556 | 13.5 | 9.19 | 176 |
| A0A291F5L8 | Ribosomal protein L22 | 1.687 | 32.5 | 9.54 | 75 |
| K9UUF3 | Ribosomal protein RPL32e | 2.096 | 10.6 | 10.29 | 26 |
| A0A1L6BUB7 | Ribosomal protein S12 | 0.469 | 14.1 | 11.46 | 45 |
| A0A2D1PBQ4 | Ribosomal protein S16 | 0.475 | 10.5 | 10.11 | 24 |
| A0A2D1PBU9 | Ribosomal protein S2 | 0.287 | 26.9 | 9.69 | 45 |
| A0A291F0G4 | Ribosomal protein S8 | 0.451 | 15.9 | 10.87 | 94 |
| Oxidative phosphorylation | |||||
| A0A2D1PBY9 | NAD(P)H-quinone oxidoreductase chain 5 | 0.504 | 84.9 | 9.10 | 21 |
| A0A2D1PBT2 | NAD(P)H-quinone oxidoreductase chain J | 2.160 | 18.8 | 6.79 | 37 |
| A0A240FG85 | NAD(P)H-quinone oxidoreductase subunit 4L | 0.593 | 11.2 | 9.72 | 33 |
| A0A088PYE4 | NAD(P)H-quinone oxidoreductase subunit H | 0.583 | 45.5 | 5.87 | 48 |
| A0A288W7F2 | NADH dehydrogenase subunit 7 | 0.354 | 44.2 | 7.05 | 69 |
| A0A0K1ZFQ7 | NADH dehydrogenase subunit 9 | 0.531 | 22.5 | 7.46 | 0 |
| A0A240FG45 | Cytochrome f | 1.622 | 35.1 | 8.97 | 822 |
| A0A1L6BSB5 | ATP synthase CF0 subunit I | 2.819 | 21.0 | 6.02 | 0 |
| H6VP90 | ATP synthase subunit beta (Fragment) | 2.091 | 47.8 | 5.19 | 1609 |
| A0A1Z2QTG2 | ATP synthase subunit beta | 2.267 | 53.7 | 5.58 | 1479 |
| B3SU35 | ATP synthase subunit beta, chloroplastic | 1.870 | 53.7 | 5.72 | 1461 |
| A0A220D850 | ATP synthase subunit c, chloroplastic | 1.969 | 11.0 | 4.64 | 31 |
| B9U4M3 | ATP synthase subunit beta (Fragment) | 1.907 | 49.5 | 5.08 | 1707 |
| A0A0K1Z6V1 | ATP synthase subunit alpha | 4.268 | 54.9 | 5.91 | 560 |
| Secondary metabolites | |||||
| V9PEM0 | Diphosphomevalonate decarboxylase | 0.578 | 46.0 | 6.38 | 19 |
| J7EQD9 | Phenylalanine ammonia-lyase | 1.557 | 77.1 | 6.10 | 264 |
| Q2WFK7 | Cytosolic ascorbate peroxidase | 1.620 | 27.9 | 6.00 | 242 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, X.; Zhou, W.; Wang, H.; You, J.; Liu, W.; Zhang, M. TMT-Based Proteomic Analysis of Continuous Cropping Response in Codonopsis tangshen Oliv. Life 2023, 13, 765. https://doi.org/10.3390/life13030765
Jiang X, Zhou W, Wang H, You J, Liu W, Zhang M. TMT-Based Proteomic Analysis of Continuous Cropping Response in Codonopsis tangshen Oliv. Life. 2023; 13(3):765. https://doi.org/10.3390/life13030765
Chicago/Turabian StyleJiang, Xiaogang, Wuxian Zhou, Hua Wang, Jinwen You, Wenlu Liu, and Meide Zhang. 2023. "TMT-Based Proteomic Analysis of Continuous Cropping Response in Codonopsis tangshen Oliv." Life 13, no. 3: 765. https://doi.org/10.3390/life13030765





