Machine Learning in Identifying Marker Genes for Congenital Heart Diseases of Different Cardiac Cell Types
Abstract
:1. Introduction
2. Materials and Methods
2.1. Data from the Single-Nucleus RNA Sequencing of Heart Tissues
2.2. Feature-Ranking Methods Used to Rank Features in Order of Importance
2.2.1. Categorical Boosting
2.2.2. Least Absolute Shrinkage and Selection Operator
2.2.3. Light Gradient Boosting Machine
2.2.4. Monte Carlo Feature Selection
2.2.5. Random Forest
2.2.6. eXtreme Gradient Boosting
2.3. Incremental Feature Selection
2.4. Synthetic Minority Oversampling Technique
2.5. Classification Algorithm
2.6. Performance Evaluation
2.7. Functional Enrichment Analysis
3. Results
3.1. Feature Ranking Results
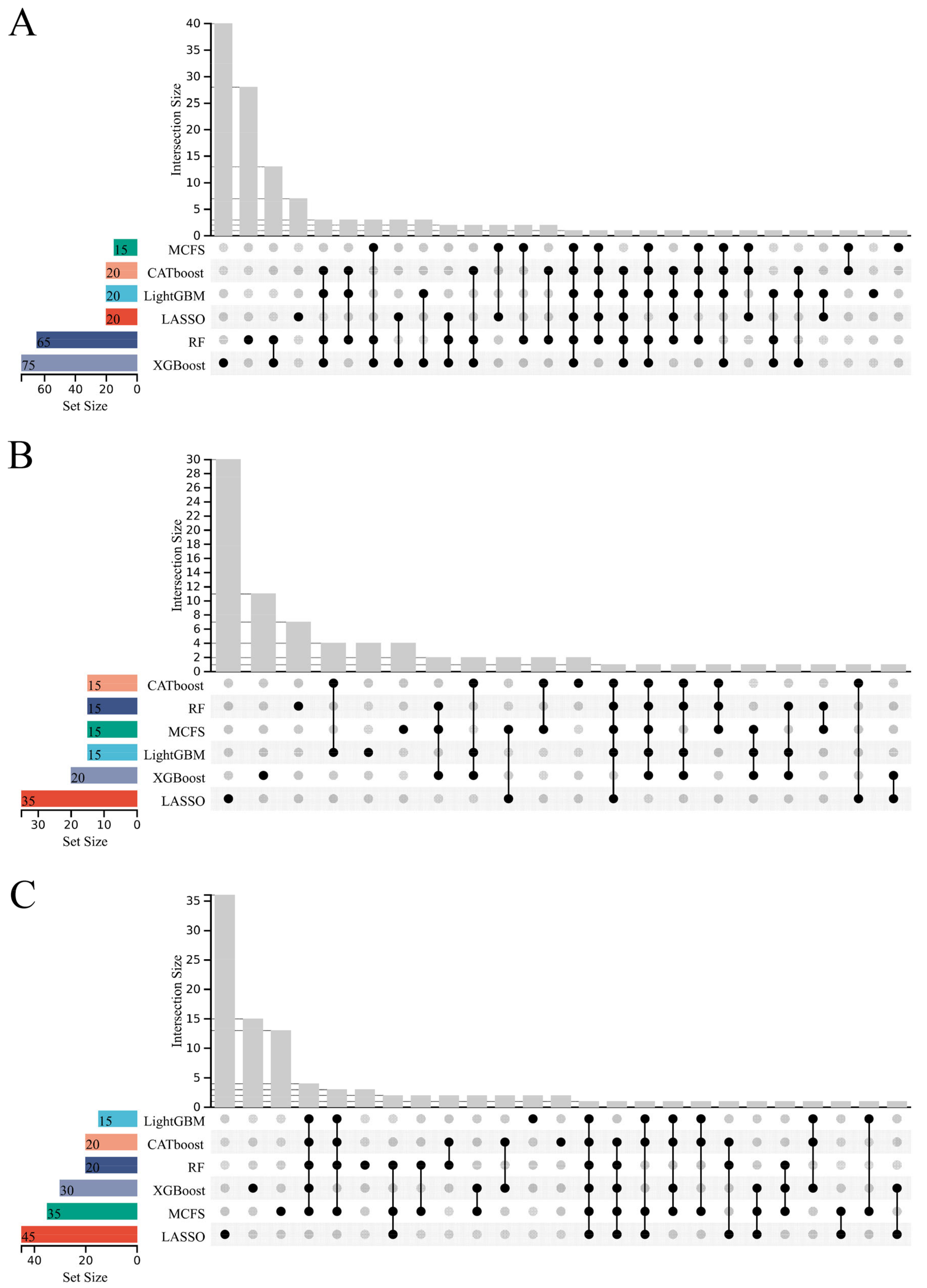
3.2. IFS Results and Feature Intersections for Finding Key Features Associated with Heart
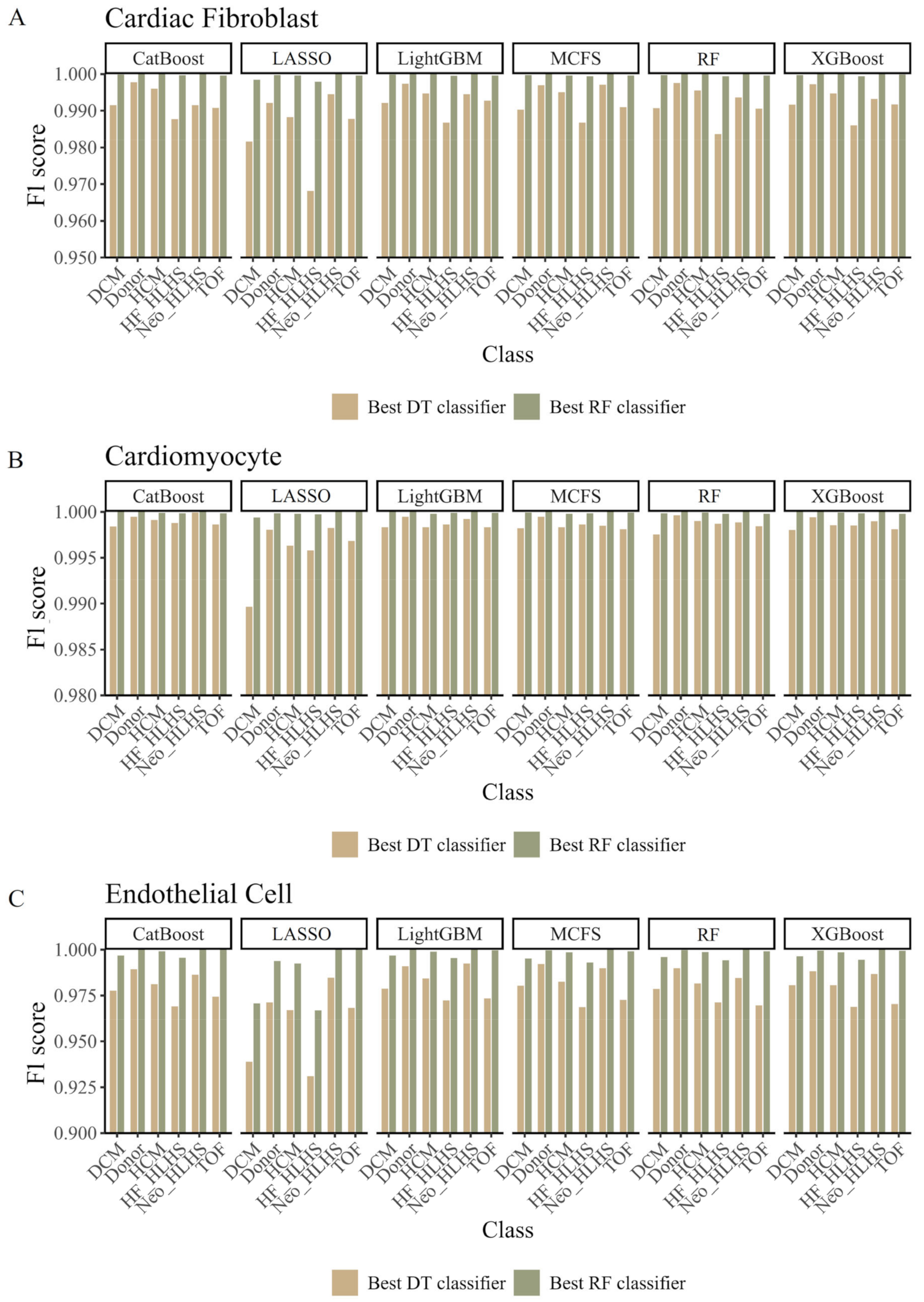
3.3. Establishing Classification Rules for Identifying Congenital Heart Diseases
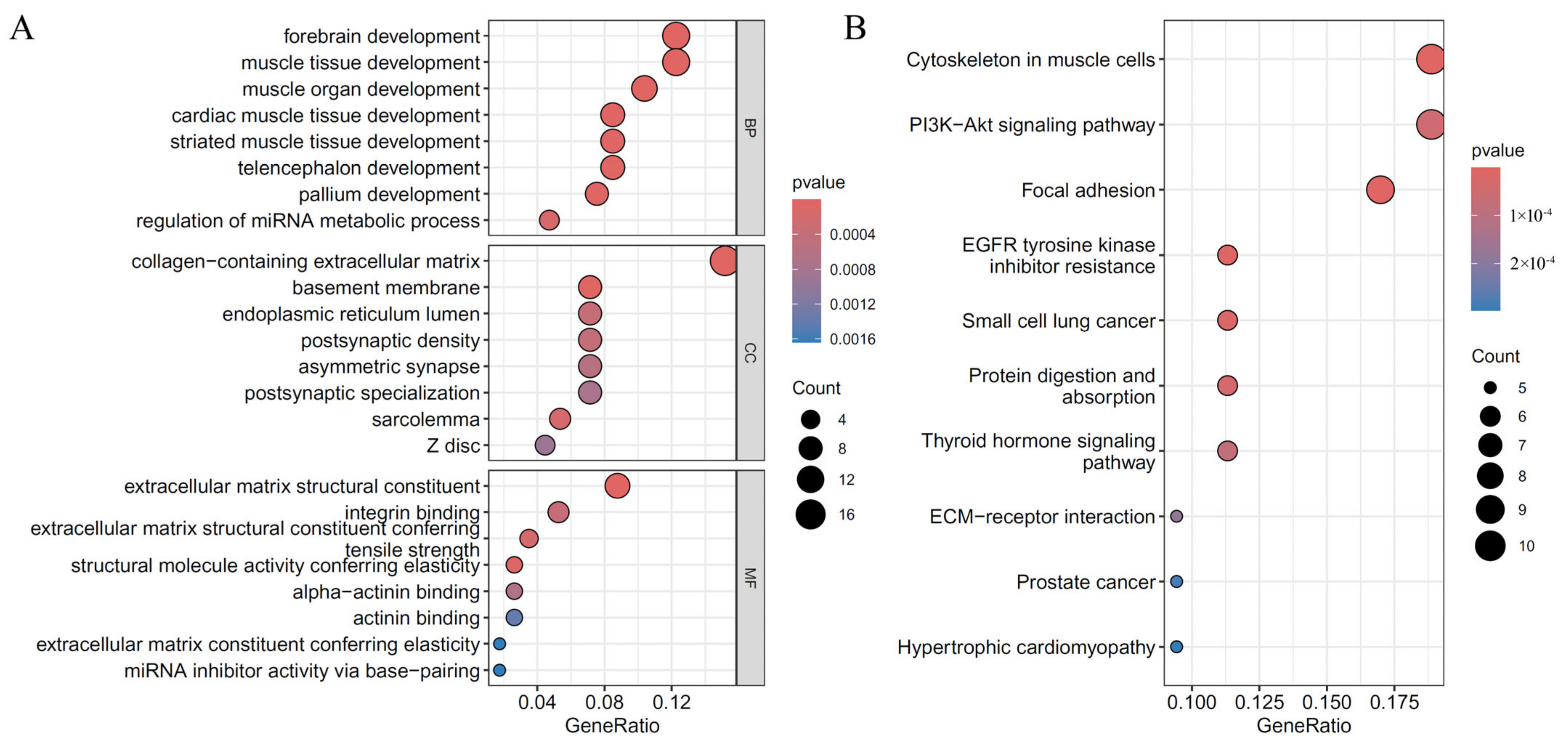
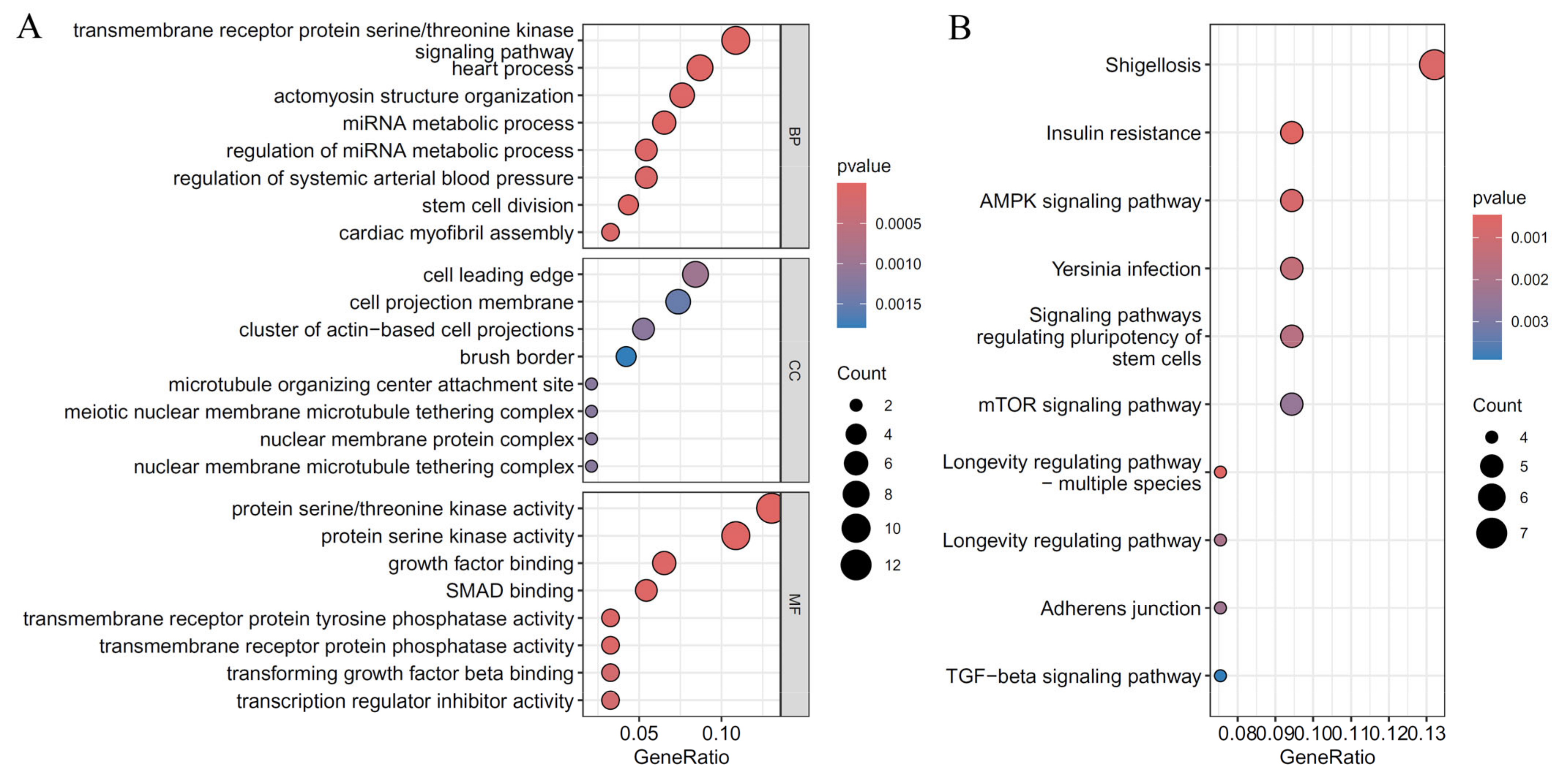
3.4. Enrichment Analysis for Essential Genes
4. Discussion
4.1. Optimized Features Selected by LASSO
4.2. Optimized Features Selected by LightGBM
4.3. Optimized Features Selected by CatBoost
4.4. Optimized Features Selected by MCFS
4.5. Optimized Features Selected by RF
4.6. Optimized Features Selected by XGBoost
4.7. Functional Analysis of the Key Features of CHD
4.8. Redundancy of Predicted Genes across Different Congenital Heart Disease Subtypes
4.9. Variants and Expression Profiling Congenital Heart Disease Subtyping
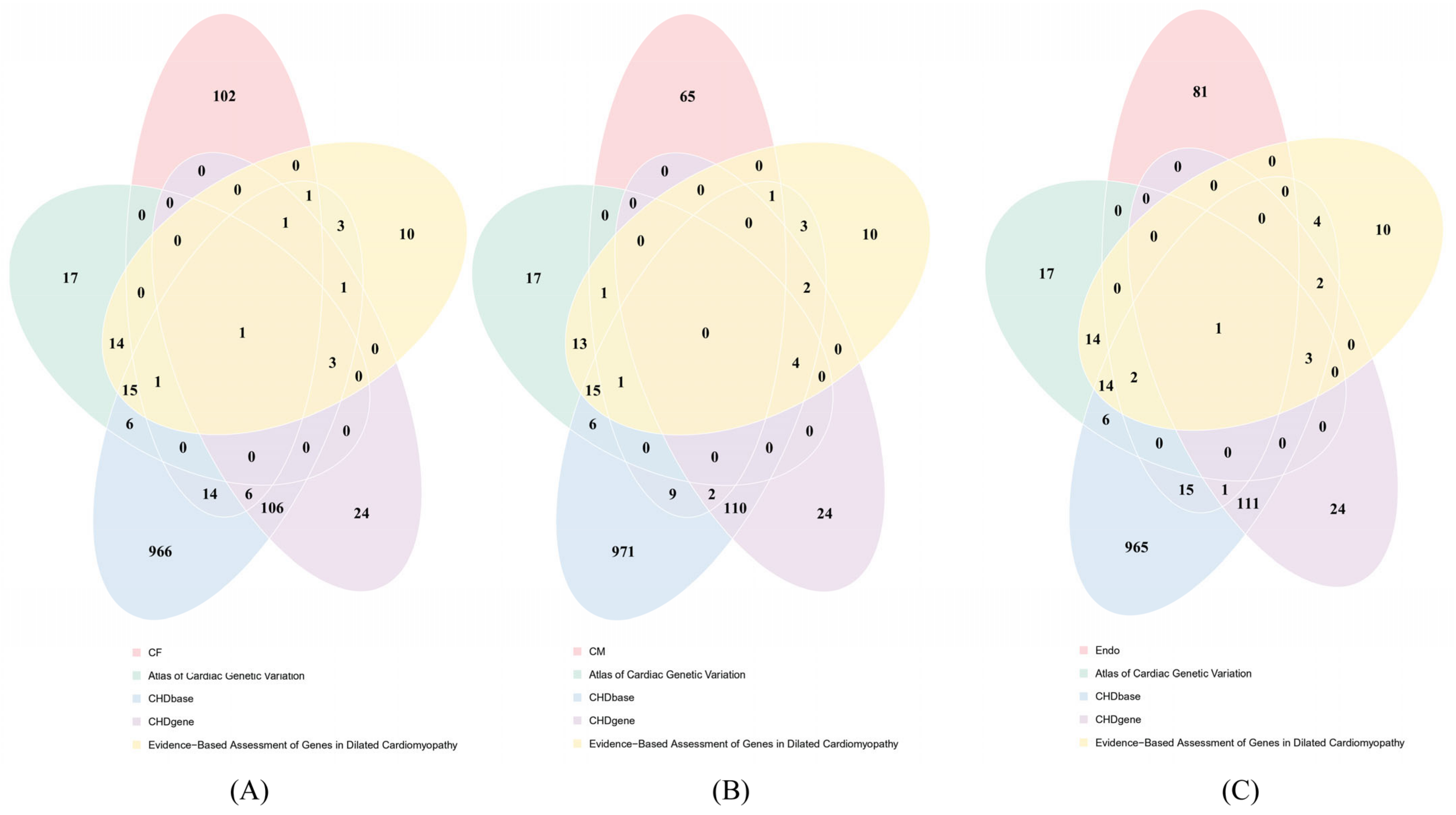
4.10. Comparison of the Public CHD-Related Genes
4.11. Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sun, R.; Liu, M.; Lu, L.; Zheng, Y.; Zhang, P. Congenital heart disease: Causes, diagnosis, symptoms, and treatments. Cell Biochem. Biophys. 2015, 72, 857–860. [Google Scholar] [CrossRef] [PubMed]
- Van der Bom, T.; Zomer, A.C.; Zwinderman, A.H.; Meijboom, F.J.; Bouma, B.J.; Mulder, B.J. The changing epidemiology of congenital heart disease. Nat. Rev. Cardiol. 2011, 8, 50–60. [Google Scholar] [CrossRef]
- Clark, B.C.; Berul, C.I. Arrhythmia diagnosis and management throughout life in congenital heart disease. Expert Rev. Cardiovasc. Ther. 2016, 14, 301–320. [Google Scholar] [CrossRef] [PubMed]
- GBD 2017 Congenital Heart Disease Collaborators. Global, regional, and national burden of congenital heart disease, 1990–2017: A systematic analysis for the global burden of disease study 2017. Lancet Child Adolesc. Health 2020, 4, 185–200. [Google Scholar] [CrossRef]
- Gilboa, S.M.; Devine, O.J.; Kucik, J.E.; Oster, M.E.; Riehle-Colarusso, T.; Nembhard, W.N.; Xu, P.; Correa, A.; Jenkins, K.; Marelli, A.J. Congenital heart defects in the united states: Estimating the magnitude of the affected population in 2010. Circulation 2016, 134, 101–109. [Google Scholar] [CrossRef]
- Heusch, G.; Libby, P.; Gersh, B.; Yellon, D.; Böhm, M.; Lopaschuk, G.; Opie, L. Cardiovascular remodelling in coronary artery disease and heart failure. Lancet 2014, 383, 1933–1943. [Google Scholar] [CrossRef] [PubMed]
- Aburawi, E.H.; Pesonen, E. Pathophysiology of coronary blood flow in congenital heart disease. Int. J. Cardiol. 2011, 151, 273–277. [Google Scholar] [CrossRef]
- Chung, I.-M.; Rajakumar, G. Genetics of congenital heart defects: The NKX2-5 gene, a key player. Genes 2016, 7, 6. [Google Scholar] [CrossRef]
- Reamon-Buettner, S.M.; Cho, S.-H.; Borlak, J. Mutations in the 3′-untranslated region of GATA4 as molecular hotspots for congenital heart disease (CHD). BMC Med. Genet. 2007, 8, 38. [Google Scholar] [CrossRef]
- Van Herck, J.L.; De Meyer, G.R.; Martinet, W.; Van Hove, C.E.; Foubert, K.; Theunis, M.H.; Apers, S.; Bult, H.; Vrints, C.J.; Herman, A.G. Impaired fibrillin-1 function promotes features of plaque instability in apolipoprotein e–deficient mice. Circulation 2009, 120, 2478–2487. [Google Scholar] [CrossRef]
- Sarti, C.; Gallagher, J. The metabolic syndrome: Prevalence, CHD risk, and treatment. J. Diabetes Its Complicat. 2006, 20, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Van der Ven, J.P.G.; van den Bosch, E.; Bogers, A.; Helbing, W.A. Current outcomes and treatment of Tetralogy of Fallot. F1000Research 2019, 8, F1000 Faculty Rev-1530. [Google Scholar] [CrossRef] [PubMed]
- Wise-Faberowski, L.; Asija, R.; McElhinney, D.B. Tetralogy of Fallot: Everything you wanted to know but were afraid to ask. Paediatr. Anaesth. 2019, 29, 475–482. [Google Scholar] [CrossRef] [PubMed]
- Hill, M.C.; Kadow, Z.A.; Long, H.; Morikawa, Y.; Martin, T.J.; Birks, E.J.; Campbell, K.S.; Nerbonne, J.; Lavine, K.; Wadhwa, L.; et al. Integrated multi-omic characterization of congenital heart disease. Nature 2022, 608, 181–191. [Google Scholar] [CrossRef]
- Somerville, J.; Becú, L. Congenital heart disease associated with hypertrophic cardiomyopathy. Br. Heart J. 1978, 40, 1034–1039. [Google Scholar] [CrossRef]
- Japp, A.G.; Gulati, A.; Cook, S.A.; Cowie, M.R.; Prasad, S.K. The diagnosis and evaluation of dilated cardiomyopathy. J. Am. Coll. Cardiol. 2016, 67, 2996–3010. [Google Scholar] [CrossRef]
- Metcalf, M.K.; Rychik, J. Outcomes in hypoplastic left heart syndrome. Pediatr. Clin. N. Am. 2020, 67, 945–962. [Google Scholar] [CrossRef] [PubMed]
- Trivedi, D.V.; Adhikari, A.S.; Sarkar, S.S.; Ruppel, K.M.; Spudich, J.A. Hypertrophic cardiomyopathy and the myosin mesa: Viewing an old disease in a new light. Biophys. Rev. 2018, 10, 27–48. [Google Scholar] [CrossRef] [PubMed]
- McNally, E.M.; Mestroni, L. Dilated cardiomyopathy: Genetic determinants and mechanisms. Circ. Res. 2017, 121, 731–748. [Google Scholar] [CrossRef] [PubMed]
- Hosseinpour, A.R.; González-Calle, A.; Adsuar-Gómez, A.; Ho, S.Y. The predicament of surgical correction of tetralogy of fallot. Pediatr. Cardiol. 2021, 42, 1252–1257. [Google Scholar] [CrossRef]
- Orphanou, N.; Papatheodorou, E.; Anastasakis, A. Dilated cardiomyopathy in the era of precision medicine: Latest concepts and developments. Heart Fail. Rev. 2022, 27, 1173–1191. [Google Scholar] [CrossRef]
- Norwood, W.I., Jr.; Jacobs, M.L.; Murphy, J.D. Fontan procedure for hypoplastic left heart syndrome. Ann. Thorac. Surg. 1992, 54, 1025–1030. [Google Scholar] [CrossRef] [PubMed]
- Feinstein, J.A.; Benson, D.W.; Dubin, A.M.; Cohen, M.S.; Maxey, D.M.; Mahle, W.T.; Pahl, E.; Villafañe, J.; Bhatt, A.B.; Peng, L.F.; et al. Hypoplastic left heart syndrome: Current considerations and expectations. J. Am. Coll. Cardiol. 2012, 59, S1–S42. [Google Scholar] [CrossRef] [PubMed]
- Spirito, P.; Seidman, C.E.; McKenna, W.J.; Maron, B.J. The management of hypertrophic cardiomyopathy. New Engl. J. Med. 1997, 336, 775–785. [Google Scholar] [CrossRef] [PubMed]
- Miranda, A.M.A.; Janbandhu, V.; Maatz, H.; Kanemaru, K.; Cranley, J.; Teichmann, S.A.; Hübner, N.; Schneider, M.D.; Harvey, R.P.; Noseda, M. Single-Cell transcriptomics for the assessment of cardiac disease. Nat. Rev. Cardiol. 2023, 20, 289–308. [Google Scholar] [CrossRef]
- Molenaar, B.; Timmer, L.T.; Droog, M.; Perini, I.; Versteeg, D.; Kooijman, L.; Monshouwer-Kloots, J.; de Ruiter, H.; Gladka, M.M.; van Rooij, E. Single-Cell transcriptomics following ischemic injury identifies a role for B2M in cardiac repair. Commun. Biol. 2021, 4, 146. [Google Scholar] [CrossRef] [PubMed]
- Nicin, L.; Schroeter, S.M.; Glaser, S.F.; Schulze-Brüning, R.; Pham, M.-D.; Hille, S.S.; Yekelchyk, M.; Kattih, B.; Abplanalp, W.T.; Tombor, L.; et al. A human cell atlas of the pressure-induced hypertrophic heart. Nat. Cardiovasc. Res. 2022, 1, 174–185. [Google Scholar] [CrossRef]
- Alimadadi, A.; Munroe, P.B.; Joe, B.; Cheng, X. Meta-Analysis of dilated cardiomyopathy using cardiac RNA-Seq transcriptomic datasets. Genes 2020, 11, 60. [Google Scholar] [CrossRef]
- Burrell, L.M.; Harrap, S.B.; Velkoska, E.; Patel, S.K. The ACE2 gene: Its potential as a functional candidate for cardiovascular disease. Clin. Sci. 2013, 124, 65–76. [Google Scholar] [CrossRef]
- Jiang, Z.; Zhou, M. Neuregulin signaling and heart failure. Curr. Heart Fail. Rep. 2010, 7, 42–47. [Google Scholar] [CrossRef]
- Gu, J.-N.; Yang, C.-X.; Ding, Y.-Y.; Qiao, Q.; Di, R.-M.; Sun, Y.-M.; Wang, J.; Yang, L.; Xu, Y.-J.; Yang, Y.-Q. Identification of BMP10 as a novel gene contributing to dilated cardiomyopathy. Diagnostics 2023, 13, 242. [Google Scholar] [CrossRef] [PubMed]
- Togănel, R.; Muntean, I.; Duicu, C.; Făgărăşan, A.; Gozar, L.; Bănescu, C. The role of eNOS and AGT gene polymorphisms in secondary pulmonary arterial hypertension in romanian children with congenital heart disease. Rev. Romana Med. Lab. 2013, 21, 267–274. [Google Scholar] [CrossRef]
- Zhang, L.; Wang, Q.; Xie, X. Identification of biomarkers related to immune cell infiltration with gene coexpression network in myocardial infarction. Dis. Markers 2021, 2021, 2227067. [Google Scholar] [CrossRef] [PubMed]
- Samad, T.; Wu, S.M. Single cell RNA sequencing approaches to cardiac development and congenital heart disease. Semin. Cell Dev. Biol. 2021, 118, 129–135. [Google Scholar] [CrossRef]
- Doyle, M.J.; Lohr, J.L.; Chapman, C.S.; Koyano-Nakagawa, N.; Garry, M.G.; Garry, D.J. Human induced pluripotent stem cell-derived cardiomyocytes as a model for heart development and congenital heart disease. Stem Cell Rev. Rep. 2015, 11, 710–727. [Google Scholar] [CrossRef] [PubMed]
- Kathiriya, I.S.; Rao, K.S.; Iacono, G.; Devine, W.P.; Blair, A.P.; Hota, S.K.; Lai, M.H.; Garay, B.I.; Thomas, R.; Gong, H.Z.; et al. Modeling human TBX5 haploinsufficiency predicts regulatory networks for congenital heart disease. Dev. Cell 2021, 56, 292–309.e9. [Google Scholar] [CrossRef]
- Sugimoto, M.; Kuwata, S.; Kurishima, C.; Kim, J.H.; Iwamoto, Y.; Senzaki, H. Cardiac biomarkers in children with congenital heart disease. World J. Pediatr. 2015, 11, 309–315. [Google Scholar] [CrossRef]
- Emoto, T.; Lu, J.; Sivasubramaniyam, T.; Maan, H.; Khan, A.B.; Abow, A.A.; Schroer, S.A.; Hyduk, S.J.; Althagafi, M.G.; McKee, T.D.; et al. Colony stimulating factor-1 producing endothelial cells and mesenchymal stromal cells maintain monocytes within a perivascular bone marrow niche. Immunity 2022, 55, 862–878.e8. [Google Scholar] [CrossRef]
- Dorogush, A.V.; Ershov, V.; Gulin, A. Catboost: Gradient boosting with categorical features support. arXiv 2018, arXiv:1810.11363. [Google Scholar]
- Geurts, P.; Ernst, D.; Wehenkel, L. Extremely randomized trees. Mach. Learn. 2006, 63, 3–42. [Google Scholar] [CrossRef]
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.-Y. LightGBM: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst. 2017, 30, 3146–3154. [Google Scholar]
- Draminski, M.; Rada-Iglesias, A.; Enroth, S.; Wadelius, C.; Koronacki, J.; Komorowski, J. Monte Carlo feature selection for supervised classification. Bioinformatics 2008, 24, 110–117. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; Association for Computing Machinery: New York, NY, USA, 2016; pp. 785–794. [Google Scholar]
- Liu, H.; Setiono, R. Incremental feature selection. Appl. Intell. 1998, 9, 217–230. [Google Scholar] [CrossRef]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. Smote: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Safavian, S.R.; Landgrebe, D. A survey of decision tree classifier methodology. IEEE Trans. Syst. Man Cybern. 1991, 21, 660–674. [Google Scholar] [CrossRef]
- Ren, J.; Chen, L.; Guo, W.; Feng, K.; Huang, T.; Cai, Y.-D. Patterns of gene expression profiles associated with colorectal cancer in colorectal mucosa by using machine learning methods. Comb. Chem. High Throughput Screen. 2024, 27, 2921–2934. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Gao, Q.; Zhou, X.; Chen, L.; Guo, W.; Feng, K.; Huang, T.; Cai, Y.-D. Identification of key gene expression associated with quality of life after recovery from COVID-19. Med. Biol. Eng. Comput. 2024, 62, 1031–1048. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Zhou, X.; Huang, K.; Chen, L.; Guo, W.; Feng, K.; Huang, T.; Cai, Y.-D. Identification of key genes associated with persistent immune changes and secondary immune activation responses induced by influenza vaccination after COVID-19 recovery by machine learning methods. Comput. Biol. Med. 2024, 169, 107883. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; Fu, M.; Li, J.; Chen, L.; Feng, K.; Huang, T.; Cai, Y.-D. Analysis and prediction of protein stability based on interaction network, gene ontology, and KEGG pathway enrichment scores. BBA-Proteins Proteom. 2023, 1871, 140889. [Google Scholar] [CrossRef]
- Huang, F.; Ma, Q.; Ren, J.; Li, J.; Wang, F.; Huang, T.; Cai, Y.-D. Identification of smoking associated transcriptome aberration in blood with machine learning methods. BioMed Res. Int. 2023, 2023, 5333361. [Google Scholar] [CrossRef]
- Ren, J.; Zhang, Y.; Guo, W.; Feng, K.; Yuan, Y.; Huang, T.; Cai, Y.-D. Identification of genes associated with the impairment of olfactory and gustatory functions in COVID-19 via machine-learning methods. Life 2023, 13, 798. [Google Scholar] [CrossRef]
- Powers, D. Evaluation: From precision, recall and F-measure to ROC., informedness, markedness & correlation. J. Mach. Learn. Technol. 2011, 2, 37–63. [Google Scholar]
- Chen, L.; Chen, Y. RMTLysPTM: Recognizing multiple types of lysine PTM sites by deep analysis on sequences. Brief. Bioinform. 2024, 25, bbad450. [Google Scholar] [CrossRef]
- Chen, L.; Qu, R.; Liu, X. Improved multi-label classifiers for predicting protein subcellular localization. Math. Biosci. Eng. 2024, 21, 214–236. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Zhang, C.; Xu, J. PredictEFC: A fast and efficient multi-label classifier for predicting enzyme family classes. BMC Bioinform. 2024, 25, 50. [Google Scholar] [CrossRef]
- Chen, L.; Zhao, X. PCDA-HNMP: Predicting circRNA-disease association using heterogeneous network and meta-path. Math. Biosci. Eng. 2023, 20, 20553–20575. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Li, L. Prediction of drug pathway-based disease classes using multiple properties of drugs. Curr. Bioinform. 2024, 19, 859–872. [Google Scholar] [CrossRef]
- Matthews, B. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. BBA-Protein Struct. 1975, 405, 442–451. [Google Scholar] [CrossRef]
- Gorodkin, J. Comparing two K-Category assignments by a K-Category correlation coefficient. Comput. Biol. Chem. 2004, 28, 367–374. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L.; et al. ClusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef]
- Hildebrandt, L.; Dieterlen, M.T.; Klaeske, K.; Haunschild, J.; Saeed, D.; Eifert, S.; Borger, M.A.; Jawad, K. Myostatin/AKT/FOXO signaling is altered in human non-ischemic dilated cardiomyopathy. Life 2022, 12, 1418. [Google Scholar] [CrossRef]
- Li, W.; Zhu, L.; Huang, H.; He, Y.; Lv, J.; Li, W.; Chen, L.; He, W. Identification of susceptible genes for complex chronic diseases based on disease risk functional snps and interaction networks. J. Biomed. Inform. 2017, 74, 137–144. [Google Scholar] [CrossRef]
- Durbin, M.D.; O’Kane, J.; Lorentz, S.; Firulli, A.B.; Ware, S.M. SHROOM3 is downstream of the planar cell polarity pathway and loss-of-function results in congenital heart defects. Dev. Biol. 2020, 464, 124–136. [Google Scholar] [CrossRef]
- Zheng, M.; Zhao, L.; Yang, X. Expression profiles of long noncoding rna and mrna in epicardial adipose tissue in patients with heart failure. BioMed Res. Int. 2019, 2019, 3945475. [Google Scholar] [CrossRef]
- Jefferies, J.L.; Towbin, J.A. Dilated cardiomyopathy. Lancet 2010, 375, 752–762. [Google Scholar] [CrossRef]
- Holm, H.; Gudbjartsson, D.F.; Arnar, D.O.; Thorleifsson, G.; Thorgeirsson, G.; Stefansdottir, H.; Gudjonsson, S.A.; Jonasdottir, A.; Mathiesen, E.B.; Njølstad, I. Several common variants modulate heart rate, PR interval and QRS duration. Nat. Genet. 2010, 42, 117–122. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.; Xu, J.; Wang, F.; Wang, J.; Zhao, H.; Feng, D. LncRNA XIST promotes myocardial infarction by regulating FOS through targeting miR-101a-3p. Aging 2020, 12, 7232. [Google Scholar] [CrossRef]
- Baharlooie, M.; Peymani, M.; Nasr-Esfahani, M.H.; Ghaedi, K. Network construction revealed that gestational diabetes mellitus may lead to congenital heart disease via potential lncRNAs-miRNAs regulating PPARγ. Human Gene 2022, 33, 201062. [Google Scholar] [CrossRef]
- Zhu, S.; Deng, M.; Law, A.; Poon, E.; Lo, J.; Liang, R.; Kwong, A.; Ng, K.; Tse, H.; Chan, G. Dmd—Animal models & preclinical treatment: P. 204 patient-derived induced pluripotent stem cells differentiated cardiomyocytes as platform for disease modelling for X-linked dilated cardiomyopathy. Neuromuscul. Disord. 2020, 30, S106. [Google Scholar]
- Boisson, M.; Cordier, A.G.; Martinovic, J.; Receveur, A.; Mouka, A.; Diot, R.; Egoroff, C.; Esnault, G.; Drévillon, L.; Benachi, A. Copy number variations analysis in a cohort of 47 fetuses and newborns with congenital diaphragmatic hernia. Prenat. Diagn. 2022, 42, 1627–1635. [Google Scholar] [CrossRef]
- Ingelman-Sundberg, M.; Pirmohamed, M. Precision medicine in cardiovascular therapeutics: Evaluating the role of pharmacogenetic analysis prior to drug treatment. J. Intern. Med. 2024, 295, 583–598. [Google Scholar] [CrossRef]
- Tsuchihashi, T.; Maeda, J.; Shin, C.H.; Ivey, K.N.; Black, B.L.; Olson, E.N.; Yamagishi, H.; Srivastava, D. Hand2 function in second heart field progenitors is essential for cardiogenesis. Dev. Biol. 2011, 351, 62–69. [Google Scholar] [CrossRef]
- Peng, Y.; Shan, J.; Qi, X.; Xue, H.; Rong, C.; Yao, D.; Guo, Z.; Zheng, S. Effects of catecholamine-β-adrenoceptor-camp system on severe patients with heart failure. Chin. Med. J. 2003, 116, 1459–1463. [Google Scholar]
- Reuter, M.S.; Chaturvedi, R.R.; Liston, E.; Manshaei, R.; Aul, R.B.; Bowdin, S.; Cohn, I.; Curtis, M.; Dhir, P.; Hayeems, R.Z.; et al. The cardiac genome clinic: Implementing genome sequencing in pediatric heart disease. Genet. Med. 2020, 22, 1015–1024. [Google Scholar] [CrossRef]
- Wada, K.; Misaka, T.; Yokokawa, T.; Kimishima, Y.; Kaneshiro, T.; Oikawa, M.; Yoshihisa, A.; Takeishi, Y. Blood-Based epigenetic markers of FKBP5 gene methylation in patients with dilated cardiomyopathy. J. Am. Heart Assoc. 2021, 10, e021101. [Google Scholar] [CrossRef]
- Fatkin, D.; Ohanian, M.; Brown, K.J. A novel role for FKBP5 in atrial cardiomyopathy. Circ. Res. 2023, 133, 45–47. [Google Scholar] [CrossRef]
- Wei, Y.; Cao, H.; Peng, Y.-Y.; Zhang, B. Alterated gene expression in dilated cardiomyopathy after left ventricular assist device support by bioinformatics analysis. Front. Cardiovasc. Med. 2023, 10, 1013057. [Google Scholar] [CrossRef]
- Gerull, B.; Gramlich, M.; Atherton, J.; McNabb, M.; Trombitás, K.; Sasse-Klaassen, S.; Seidman, J.; Seidman, C.; Granzier, H.; Labeit, S. Mutations of TTN, encoding the giant muscle filament titin, cause familial dilated cardiomyopathy. Nat. Genet. 2002, 30, 201–204. [Google Scholar] [CrossRef] [PubMed]
- Gill, K.; Sasaki, J.; Jayakar, P.; Sosa, L.; Welch, E. Chromosomal microarray detects genetic risks of neurodevelopmental disorders in newborns with congenital heart disease. Cardiol. Young 2021, 31, 1275–1282. [Google Scholar] [CrossRef] [PubMed]
- Mika, D.; Bobin, P.; Lindner, M.; Boet, A.; Hodzic, A.; Lefebvre, F.; Lechène, P.; Sadoune, M.; Samuel, J.-L.; Algalarrondo, V.; et al. Synergic PDE3 and PDE4 control intracellular cAMP and cardiac excitation-contraction coupling in a porcine model. J. Mol. Cell. Cardiol. 2019, 133, 57–66. [Google Scholar] [CrossRef]
- Radhakrishna, U.; Vishweswaraiah, S.; Veerappa, A.M.; Zafra, R.; Albayrak, S.; Sitharam, P.H.; Saiyed, N.M.; Mishra, N.K.; Guda, C.; Bahado-Singh, R. Newborn blood DNA epigenetic variations and signaling pathway genes associated with Tetralogy of Fallot (TOF). PLoS ONE 2018, 13, e0203893. [Google Scholar] [CrossRef]
- Gambetta, K.; Al-Ahdab, M.K.; Ilbawi, M.N.; Hassaniya, N.; Gupta, M. Transcription repression and blocks in cell cycle progression in hypoplastic left heart syndrome. Am. J. Physiol.-Heart Circ. Physiol. 2008, 294, H2268–H2275. [Google Scholar] [CrossRef]
- Shoja-Taheri, F.; George, A.; Agarwal, U.; Platt, M.O.; Gibson, G.; Davis, M.E. Using statistical modeling to understand and predict pediatric stem cell function. Circ. Genom. Precis. Med. 2019, 12, e002403. [Google Scholar] [CrossRef]
- Chen, Y.; Liu, X.; Chen, L.; Chen, W.; Zhang, Y.; Chen, J.; Wu, X.; Zhao, Y.; Wu, X.; Sun, G. The long noncoding RNA XIST protects cardiomyocyte hypertrophy by targeting miR-330-3p. Biochem. Biophys. Res. Commun. 2018, 505, 807–815. [Google Scholar] [CrossRef]
- Tang, B.-Y.; Ge, J.; Wu, Y.; Wen, J.; Tang, X.-H. The role of ADAM17 in inflammation-related atherosclerosis. J. Cardiovasc. Transl. Res. 2022, 15, 1283–1296. [Google Scholar] [CrossRef]
- Noack, C.; Iyer, L.M.; Liaw, N.Y.; Schoger, E.; Khadjeh, S.; Wagner, E.; Woelfer, M.; Zafiriou, M.-P.; Milting, H.; Sossalla, S.; et al. KLF15-Wnt-Dependent cardiac reprogramming up-regulates SHISA3 in the mammalian heart. J. Am. Coll. Cardiol. 2019, 74, 1804–1819. [Google Scholar] [CrossRef]
- Audain, E.; Wilsdon, A.; Breckpot, J.; Izarzugaza, J.M.; Fitzgerald, T.W.; Kahlert, A.-K.; Sifrim, A.; Wünnemann, F.; Perez-Riverol, Y.; Abdul-Khaliq, H.; et al. Integrative analysis of genomic variants reveals new associations of candidate haploinsufficient genes with congenital heart disease. PLoS Genet. 2021, 17, e1009679. [Google Scholar]
- Zheng, Y.; Lang, Y.; Qi, Z.; Qi, B.; Gao, W.; Hu, X.; Li, T. Macrophage-Related genes biomarkers in left ventricular remodeling induced by heart failure. Rev. Cardiovasc. Med. 2022, 23, 109. [Google Scholar] [CrossRef] [PubMed]
- Winsvold, B.S.; Kitsos, I.; Thomas, L.F.; Skogholt, A.H.; Gabrielsen, M.E.; Zwart, J.A.; Nilsen, K.B. Genome-Wide association study of 2,093 cases with idiopathic polyneuropathy and 445,256 controls identifies first susceptibility loci. Front. Neurol. 2021, 12, 789093. [Google Scholar] [CrossRef]
- Butler, C.L.; Hickey, M.J.; Jiang, N.; Zheng, Y.; Gjertson, D.; Zhang, Q.; Rao, P.; Fishbein, G.A.; Cadeiras, M.; Deng, M.C.; et al. Discovery of non-HLA antibodies associated with cardiac allograft rejection and development and validation of a non-HLA antigen multiplex panel: From bench to bedside. Am. J. Transplant. 2020, 20, 2768–2780. [Google Scholar] [CrossRef] [PubMed]
- McCormick, D. Investigating MicroRNAs during Cardiac Development in the Chick. Ph.D. Thesis, University of East Anglia, Norwich, UK, 2014. [Google Scholar]
- Sicko, R.J.; Browne, M.L.; Rigler, S.L.; Druschel, C.M.; Liu, G.; Fan, R.; Romitti, P.A.; Caggana, M.; Kay, D.M.; Brody, L.C.; et al. Genetic variants in isolated ebstein anomaly implicated in myocardial development pathways. PLoS ONE 2016, 11, e0165174. [Google Scholar] [CrossRef] [PubMed]
- Perry, D.J.; Mullen, C.R.; Carvajal, H.G.; Brar, A.K.; Eghtesady, P. Familial screening for left-sided congenital heart disease: What is the evidence? What is the cost? Diseases 2017, 5, 29. [Google Scholar] [CrossRef] [PubMed]
- Schneider, M.; Al-Shareffi, E.; Haltiwanger, R.S. Biological functions of fucose in mammals. Glycobiology 2017, 27, 601–618. [Google Scholar] [CrossRef] [PubMed]
- Navas-Acien, A.; Domingo-Relloso, A.; Subedi, P.; Riffo-Campos, A.L.; Xia, R.; Gomez, L.; Haack, K.; Goldsmith, J.; Howard, B.V.; Best, L.G.; et al. Blood DNA methylation and incident coronary heart disease: Evidence from the strong heart study. JAMA Cardiol. 2021, 6, 1237–1246. [Google Scholar] [CrossRef] [PubMed]
- Kalayinia, S.; Maleki, M.; Mahdavi, M.; Mahdieh, N. Whole-Exome sequencing reveals a novel mutation of flna gene in an iranian family with nonsyndromic Tetralogy of Fallot. Lab. Med. 2021, 52, 614–618. [Google Scholar] [CrossRef] [PubMed]
- Lim, T.B.; Foo, S.Y.R.; Chen, C.K. The role of epigenetics in congenital heart disease. Genes 2021, 12, 390. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Bai, X.; Yan, W. Lnc-MALAT1, as a biomarker of neonatal BPD, exacerbates the pathogenesis of BPD by targeting miR-206. Am. J. Transl. Res. 2021, 13, 462. [Google Scholar] [PubMed]
- Dueñas, A.; Expósito, A.; Aranega, A.; Franco, D. The role of non-coding RNA in congenital heart diseases. J. Cardiovasc. Dev. Dis. 2019, 6, 15. [Google Scholar] [CrossRef]
- Yin, X.-Y.; Chen, H.-X.; Chen, Z.; Yang, Q.; Han, J.; He, G.-W. Genetic variants of ISL1 gene promoter identified from congenital tetralogy of fallot patients alter cellular function forming disease basis. Biomolecules 2023, 13, 358. [Google Scholar] [CrossRef]
- Kolur, V.; Vastrad, B.; Vastrad, C.; Kotturshetti, S.; Tengli, A. Identification of candidate biomarkers and therapeutic agents for heart failure by bioinformatics analysis. BMC Cardiovasc. Disord. 2021, 21, 329. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Chen, J.; Wang, H.; Song, X.; Sun, Z.; Zhang, R.; Zhong, J.; Gu, X.; Wang, M.; Peng, C.; et al. Marginal zinc deficiency alters the heart proteome of rats. Food Funct. 2023, 14, 4117–4128. [Google Scholar] [CrossRef] [PubMed]
- Jain, P.N.; Robertson, M.; Lasa, J.J.; Shekerdemian, L.; Guffey, D.; Zhang, Y.; Lingappan, K.; Checchia, P.; Coarfa, C. Altered metabolic and inflammatory transcriptomics after cardiac surgery in neonates with congenital heart disease. Sci. Rep. 2021, 11, 4965. [Google Scholar] [CrossRef] [PubMed]
- Polyakova, E.; van Gils, J.M.; Stöger, J.L.; Kiès, P.; Egorova, A.D.; Koopmann, T.T.; van Dijk, T.; DeRuiter, M.C.; Barge-Schaapveld, D.Q.; Jongbloed, M.R. New genetic variant in the MYH7 gene associated with hypoplastic right heart syndrome and hypertrophic cardiomyopathy in the same family. Circ. Genom. Precis. Med. 2023, 16, e004184. [Google Scholar] [CrossRef] [PubMed]
- Zhong, H.; Wang, T.; Lian, G.; Xu, C.; Wang, H.; Xie, L. TRPM7 regulates angiotensin ii-induced sinoatrial node fibrosis in sick sinus syndrome rats by mediating smad signaling. Heart Vessel. 2018, 33, 1094–1105. [Google Scholar] [CrossRef] [PubMed]
- Yin, X.; Liu, J.; Feng, R.; Xu, M.; Liu, J. Novel PIK3R1 mutation of short syndrome: A case report with a 6-month follow up. J. Diabetes Investig. 2021, 12, 1919–1922. [Google Scholar] [CrossRef] [PubMed]
- Luyckx, I.; Walton, I.S.; Boeckx, N.; Van Schil, K.; Pang, C.; De Praeter, M.; Lord, H.; Watson, C.M.; Bonthron, D.T.; Van Laer, L.; et al. Homozygous SMAD6 variants in two unrelated patients with craniosynostosis and radioulnar synostosis. J. Med. Genet. 2024, 61, 363–368. [Google Scholar] [CrossRef]
- Chen, T.; Li, S.J.; Chen, B.; Huang, Q.; Kong, X.Y.; Shen, C.; Gu, H.T.; Wang, X.W. Akt3 is a target of miR-29c-3p and serves an important function in the pathogenesis of congenital heart disease. Int. J. Mol. Med. 2019, 43, 980–992. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Luo, P.; Zhang, Y. Identification of differentially expressed long non-coding rnas associated with dilated cardiomyopathy using integrated bioinformatics approaches. Drug Discov. Ther. 2020, 14, 181–186. [Google Scholar] [CrossRef]
- Gabriel, A.F.; Costa, M.C.; Enguita, F.J. Interactions among regulatory non-coding RNAs involved in cardiovascular diseases. Adv. Exp. Med. Biol. 2020, 1229, 79–104. [Google Scholar]
- Gu, J.; Liu, C.; Huo, Z. Mirna and cardiac hypertrophy. Sci. Insights 2019, 28, 17–24. [Google Scholar] [CrossRef]
- Walsh, R.; Thomson, K.L.; Ware, J.S.; Funke, B.H.; Woodley, J.; McGuire, K.J.; Mazzarotto, F.; Blair, E.; Seller, A.; Taylor, J.C.; et al. Reassessment of mendelian gene pathogenicity using 7,855 cardiomyopathy cases and 60,706 reference samples. Genet. Med. 2017, 19, 192–203. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.-Z.; Li, W.; Shen, H.; Wang, R.W.; Chen, W.; Zhang, Y.; Zeng, Q.; Wang, H.; Yuan, M.; Zeng, Z. Chdbase: A comprehensive knowledgebase for congenital heart disease-related genes and clinical manifestations. Genom. Proteom. Bioinform. 2023, 21, 216–227. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.; Alankarage, D.; Cuny, H.; Ip, E.K.; Almog, M.; Lu, J.; Das, D.; Enriquez, A.; Szot, J.O.; Humphreys, D.T.; et al. Chdgene: A curated database for congenital heart disease genes. Circ. Genom. Precis. Med. 2022, 15, e003539. [Google Scholar] [CrossRef]
- Jordan, E.; Peterson, L.; Ai, T.; Asatryan, B.; Bronicki, L.; Brown, E.; Celeghin, R.; Edwards, M.; Fan, J.; Ingles, J.; et al. Evidence-Based assessment of genes in dilated cardiomyopathy. Circulation 2021, 144, 7–19. [Google Scholar] [CrossRef]




| Cell Type | Feature-Ranking Algorithm | Classification Algorithm | Number of Features | ACC | MCC | Macro F1 | Weighted F1 |
|---|---|---|---|---|---|---|---|
| CF | CatBoost | RF | 65 | 0.9998 | 0.9997 | 0.9998 | 0.9998 |
| DT | 150 | 0.9929 | 0.9909 | 0.9924 | 0.9929 | ||
| LASSO | RF | 305 | 0.9989 | 0.9986 | 0.9991 | 0.9989 | |
| DT | 245 | 0.9842 | 0.9797 | 0.9853 | 0.9842 | ||
| LightGBM | RF | 130 | 0.9998 | 0.9997 | 0.9998 | 0.9998 | |
| DT | 370 | 0.9931 | 0.9911 | 0.9929 | 0.9931 | ||
| MCFS | RF | 335 | 0.9996 | 0.9995 | 0.9996 | 0.9996 | |
| DT | 860 | 0.9924 | 0.9902 | 0.9927 | 0.9924 | ||
| RF | RF | 495 | 0.9997 | 0.9996 | 0.9997 | 0.9997 | |
| DT | 220 | 0.9920 | 0.9897 | 0.9918 | 0.9920 | ||
| XGBoost | RF | 255 | 0.9997 | 0.9996 | 0.9998 | 0.9997 | |
| DT | 535 | 0.9926 | 0.9905 | 0.9923 | 0.9926 | ||
| CM | CatBoost | RF | 130 | 0.9999 | 0.9999 | 0.9999 | 0.9999 |
| DT | 65 | 0.9991 | 0.9988 | 0.9990 | 0.9991 | ||
| LASSO | RF | 100 | 0.9998 | 0.9997 | 0.9998 | 0.9998 | |
| DT | 940 | 0.9965 | 0.9955 | 0.9958 | 0.9966 | ||
| LightGBM | RF | 200 | 0.9999 | 0.9999 | 0.9999 | 0.9999 | |
| DT | 60 | 0.9988 | 0.9985 | 0.9987 | 0.9988 | ||
| MCFS | RF | 310 | 0.9999 | 0.9999 | 0.9999 | 0.9999 | |
| DT | 105 | 0.9987 | 0.9983 | 0.9985 | 0.9987 | ||
| RF | RF | 980 | 0.9999 | 0.9998 | 0.9999 | 0.9999 | |
| DT | 270 | 0.9989 | 0.9986 | 0.9986 | 0.9990 | ||
| XGBoost | RF | 255 | 0.9999 | 0.9999 | 0.9999 | 0.9999 | |
| DT | 50 | 0.9988 | 0.9984 | 0.9986 | 0.9988 | ||
| Endo | CatBoost | RF | 190 | 0.9976 | 0.9970 | 0.9983 | 0.9976 |
| DT | 195 | 0.9791 | 0.9731 | 0.9795 | 0.9791 | ||
| LASSO | RF | 155 | 0.9812 | 0.9758 | 0.9871 | 0.9812 | |
| DT | 335 | 0.9524 | 0.9388 | 0.9600 | 0.9524 | ||
| LightGBM | RF | 165 | 0.9976 | 0.9970 | 0.9983 | 0.9976 | |
| DT | 50 | 0.9812 | 0.9758 | 0.9818 | 0.9812 | ||
| MCFS | RF | 425 | 0.9965 | 0.9955 | 0.9974 | 0.9965 | |
| DT | 40 | 0.9807 | 0.9752 | 0.9808 | 0.9808 | ||
| RF | RF | 250 | 0.9971 | 0.9962 | 0.9978 | 0.9971 | |
| DT | 105 | 0.9797 | 0.9739 | 0.9790 | 0.9797 | ||
| XGBoost | RF | 215 | 0.9971 | 0.9963 | 0.9979 | 0.9971 | |
| DT | 65 | 0.9794 | 0.9735 | 0.9791 | 0.9794 |
| Cell Type | ACC | MCC | Macro F1 | Weighted F1 |
|---|---|---|---|---|
| CF | 0.9976 | 0.9969 | 0.9978 | 0.9976 |
| CM | 0.9996 | 0.9995 | 0.9996 | 0.9996 |
| Endo | 0.9796 | 0.9740 | 0.9850 | 0.9796 |
| Cell Type | Feature-Ranking Algorithm | Classification Algorithm | Number of Features | ACC | MCC | Macro F1 | Weighted F1 |
|---|---|---|---|---|---|---|---|
| CF | CatBoost | RF | 20 | 0.9990 | 0.9987 | 0.9991 | 0.9990 |
| LASSO | RF | 20 | 0.9929 | 0.9909 | 0.9942 | 0.9929 | |
| LightGBM | RF | 20 | 0.9992 | 0.9990 | 0.9991 | 0.9992 | |
| MCFS | RF | 15 | 0.9927 | 0.9906 | 0.9940 | 0.9927 | |
| RF | RF | 65 | 0.9990 | 0.9988 | 0.9992 | 0.9991 | |
| XGBoost | RF | 75 | 0.9991 | 0.9989 | 0.9992 | 0.9991 | |
| CM | CatBoost | RF | 15 | 0.9995 | 0.9994 | 0.9995 | 0.9995 |
| LASSO | RF | 35 | 0.9992 | 0.9989 | 0.9991 | 0.9992 | |
| LightGBM | RF | 15 | 0.9996 | 0.9994 | 0.9996 | 0.9996 | |
| MCFS | RF | 15 | 0.9993 | 0.9990 | 0.9992 | 0.9993 | |
| RF | RF | 15 | 0.9992 | 0.9990 | 0.9991 | 0.9992 | |
| XGBoost | RF | 20 | 0.9993 | 0.9991 | 0.9993 | 0.9993 | |
| Endo | CatBoost | RF | 20 | 0.9921 | 0.9898 | 0.9927 | 0.9921 |
| LASSO | RF | 45 | 0.9723 | 0.9645 | 0.9805 | 0.9724 | |
| LightGBM | RF | 15 | 0.9903 | 0.9875 | 0.9914 | 0.9903 | |
| MCFS | RF | 35 | 0.9917 | 0.9894 | 0.9934 | 0.9917 | |
| RF | RF | 20 | 0.9904 | 0.9876 | 0.9915 | 0.9904 | |
| XGBoost | RF | 30 | 0.9900 | 0.9872 | 0.9915 | 0.9900 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, Q.; Zhang, Y.-H.; Guo, W.; Feng, K.; Huang, T.; Cai, Y.-D. Machine Learning in Identifying Marker Genes for Congenital Heart Diseases of Different Cardiac Cell Types. Life 2024, 14, 1032. https://doi.org/10.3390/life14081032
Ma Q, Zhang Y-H, Guo W, Feng K, Huang T, Cai Y-D. Machine Learning in Identifying Marker Genes for Congenital Heart Diseases of Different Cardiac Cell Types. Life. 2024; 14(8):1032. https://doi.org/10.3390/life14081032
Chicago/Turabian StyleMa, Qinglan, Yu-Hang Zhang, Wei Guo, Kaiyan Feng, Tao Huang, and Yu-Dong Cai. 2024. "Machine Learning in Identifying Marker Genes for Congenital Heart Diseases of Different Cardiac Cell Types" Life 14, no. 8: 1032. https://doi.org/10.3390/life14081032










