Identification and Tissue Expression Profiles of Odorant Receptor Genes in the Green Peach Aphid Myzus persicae
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Insect Rearing and Sample Collection
2.2. RNA Extraction, cDNA Library Construction and Illumina Sequencing
2.3. Bioinformatics Analysis
2.4. Identification of M. persicae ORs by Bioinformatics
2.5. Phylogenetic Analysis
2.6. Verification of OR Sequences by Cloning and Sequencing
2.7. Genomic Structure Analysis of M. persicae ORs
2.8. Expression Profiles of M. persicae ORs in Different Tissues
3. Results
3.1. Identification of M. persicae ORs
3.2. Phylogenetic Analyses of MperORs
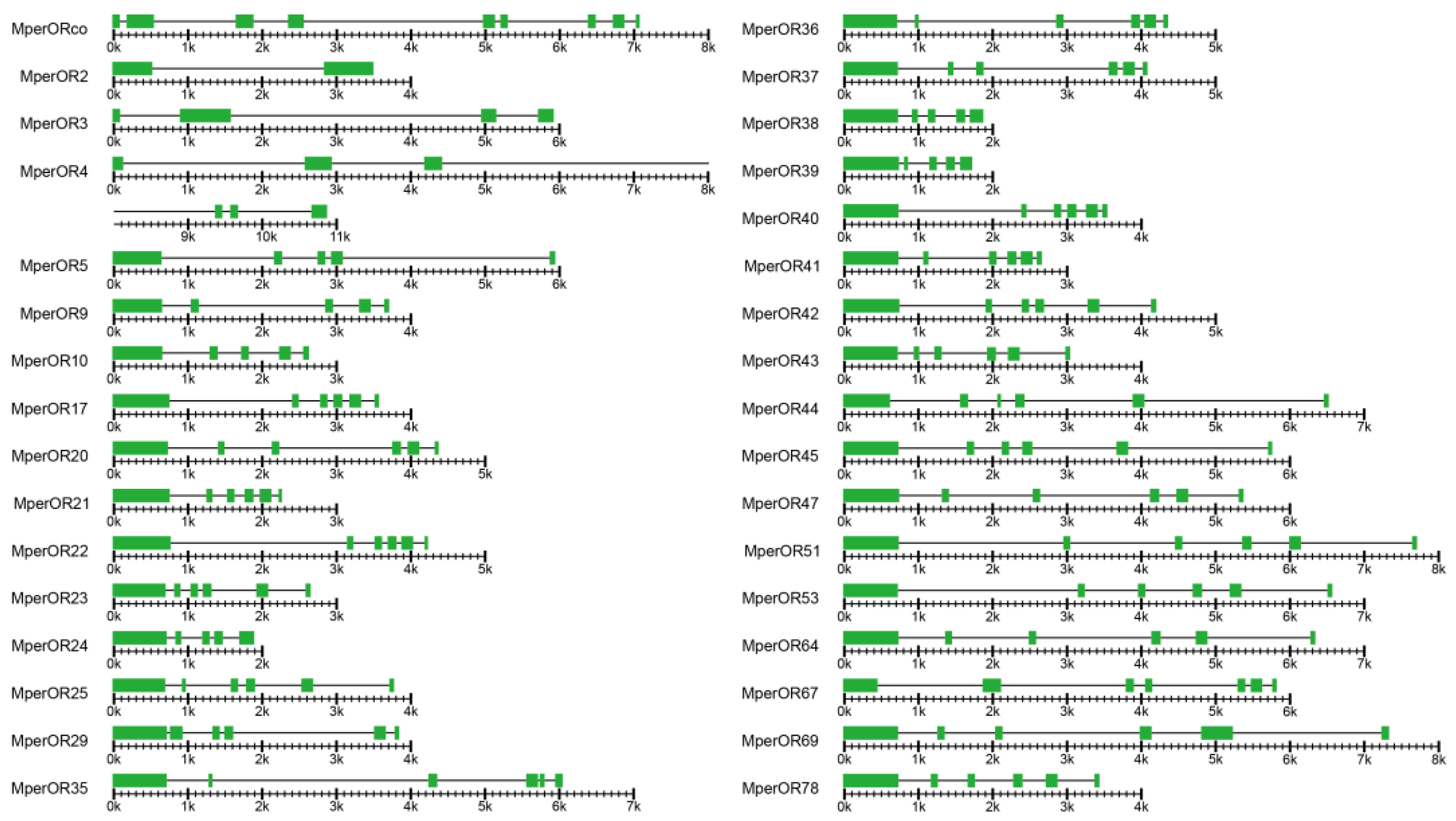
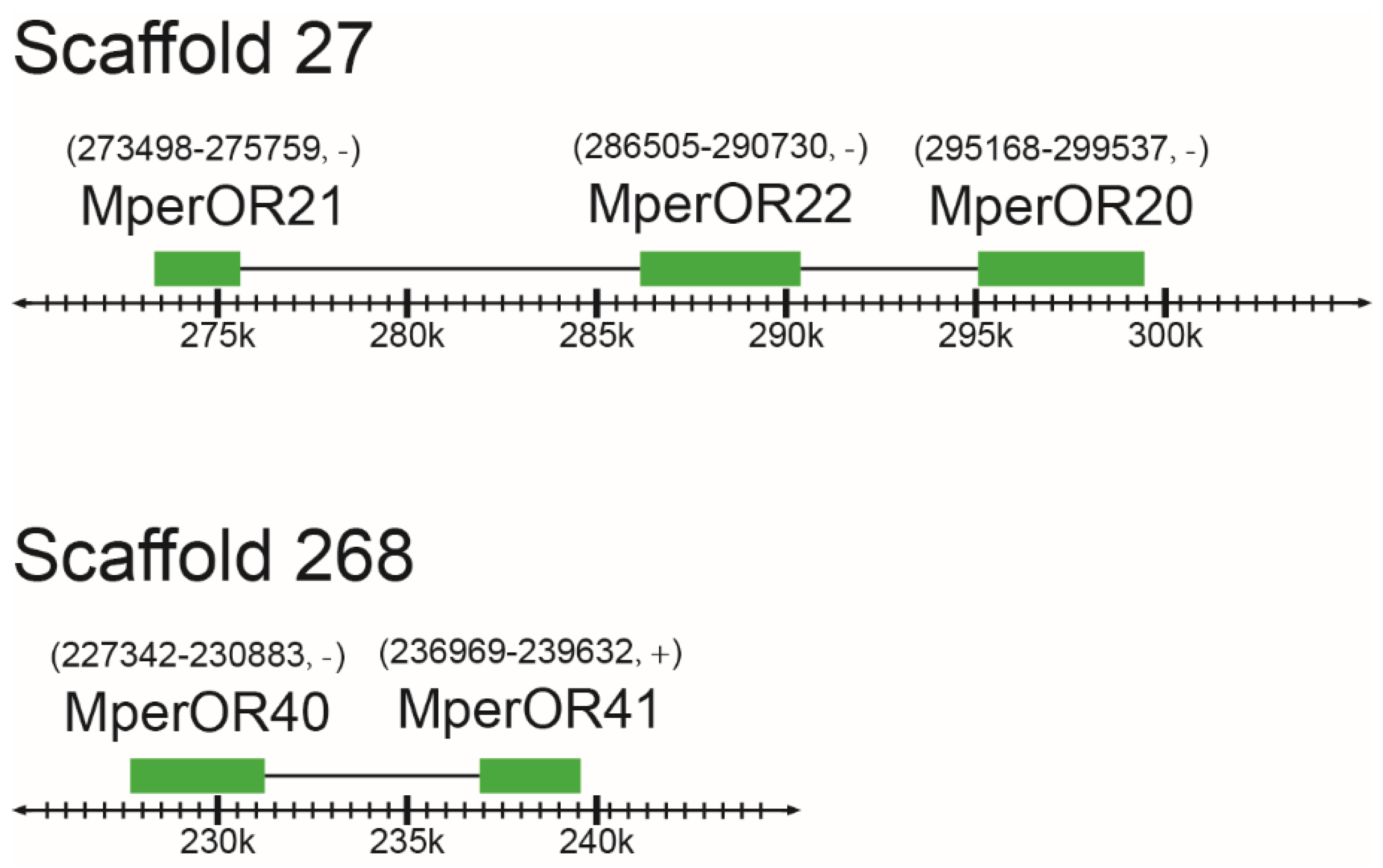
3.3. Genomic Structure of M. persicae ORs
3.4. Expression Profiles of M. perszicae ORs in Different Tissues
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Weber, G. Ecological genetics of host plant exploitation in the green peach aphid, Myzus persicae. Entomol. Exp. Appl. 1986, 40, 161–168. [Google Scholar] [CrossRef]
- Troncoso, A.J.; Vargas, R.R.; Tapia, D.H.; Olivares-Donoso, R.; Niemeyer, H.M. Host selection by the generalist aphid Myzus persicae (Hemiptera: Aphididae) and its subspecies specialized on tobacco, after being reared on the same host. Bull. Entomol. Res. 2005, 95, 23–28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Emden, H.F.; Eastop, V.F.; Hughes, D.P.; Way, M.J. The Ecology of Myzus persicae. Annu. Rev. Entomol. 1969, 14, 197–270. [Google Scholar] [CrossRef]
- Pickett, J.A.; Glinwood, R.T. Chemical Ecology. In Aphids as Crop Pests; VanEmden, H.F., Harrington, R., Eds.; CAB International Press: Wallingford, UK, 2007; pp. 235–260. [Google Scholar]
- Read, D.P.; Feeny, P.P.; Root, R.B. Habitat selection by the aphid parasite Diaeretiella rapae (Hymenoptera: Braconidae) and hyperparasite Charips brassicae (Hymenoptera: Cynipidae). Can. Entomol. 1970, 102, 1567–1578. [Google Scholar] [CrossRef]
- Mathers, T.C.; Chen, Y.; Kaithakottil, G.; Legeai, F.; Mugford, S.T.; Baa-Puyoulet, P.; Bretaudeau, A.; Clavijo, B.; Colella, S.; Collin, O.; et al. Rapid transcriptional plasticity of duplicated gene clusters enables a clonally reproducing aphid to colonise diverse plant species. Genome Biol. 2017, 18, 1–20. [Google Scholar] [CrossRef] [Green Version]
- Gadenne, C.; Barrozo, R.B.; Anton, S. Plasticity in insect olfaction: To smell or not to smell? Annu. Rev. Entomol. 2016, 61, 317–333. [Google Scholar] [CrossRef]
- Haverkamp, A.; Hansson, B.S.; Knaden, M. Combinatorial codes and labeled lines: How insects use olfactory cues to find and judge food, mates, and oviposition sites in complex environments. Front. Physiol. 2018, 9, 49. [Google Scholar] [CrossRef]
- Touhara, K.; Vosshall, L.B. Sensing odorants and pheromones with chemosensory receptors. Annu. Rev. Physiol. 2009, 71, 307–332. [Google Scholar] [CrossRef]
- Dahanukar, A.; Hallem, E.A.; Carlson, J.R. Insect chemoreception. Curr. Opin. Neurobiol. 2005, 15, 423–430. [Google Scholar] [CrossRef]
- Benton, R.; Sachse, S.; Michnick, S.W.; Vosshall, L.B. Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS Biol. 2006, 4, e20. [Google Scholar] [CrossRef] [Green Version]
- Lundin, C.; Käll, L.; Kreher, S.A.; Kapp, K.; Sonnhammer, E.L.; Carlson, J.R.; von Heijne, G.; Nilsson, I. Membrane topology of the Drosophila OR83b odorant receptor. FEBS Lett. 2007, 581, 5601–5604. [Google Scholar] [CrossRef] [Green Version]
- Smart, R.; Kiely, A.; Beale, M.; Vargas, E.; Carraher, C.; Kralicek, A.V.; Christie, D.L.; Chen, C.; Newcomb, R.D.; Warr, C.G. Drosophila odorant receptors are novel seven transmembrane domain proteins that can signal independently of heterotrimeric G proteins. Insect Biochem. Mol. Biol. 2008, 38, 770–780. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Pellegrino, M.; Nakagawa, T.; Nakagawa, T.; Vosshall, L.B.; Touhara, K. Insect olfactory receptors are heteromeric ligand-gated ion channels. Nature 2008, 452, 1002–1006. [Google Scholar] [CrossRef] [PubMed]
- Wicher, D.; Schäfer, R.; Bauernfeind, R.; Stensmyr, M.C.; Heller, R.; Heinemann, S.H.; Hansson, B.S. Drosophila odorant receptors are both ligand-gated and cyclic-nucleotide-activated cation channels. Nature 2008, 452, 1007–1011. [Google Scholar] [CrossRef] [PubMed]
- Stengl, M.; Funk, N.W. The role of the coreceptor Orco in insect olfactory transduction. J. Comp. Physiol. A 2013, 199, 897–909. [Google Scholar] [CrossRef] [PubMed]
- Vosshall, L.B.; Amrein, H.; Morozov, P.; Rzhetsky, A.; Axel, R. A spatial map of olfactory receptor expression in the Drosophila antenna. Cell 1999, 96, 725–736. [Google Scholar] [CrossRef] [Green Version]
- Gao, Q.; Chess, A. Identification of Candidate Drosophila Olfactory Receptors from Genomic DNA Sequence. Genomics 1999, 60, 31–39. [Google Scholar] [CrossRef]
- Clyne, P.J.; Warr, C.G.; Freeman, M.R.; Lessing, D.; Kim, J.; Carlson, J.R. A novel family of divergent seven-transmembrane proteins: Candidate odorant receptors in Drosophila. Neuron 1999, 22, 327–338. [Google Scholar] [CrossRef] [Green Version]
- Richards, S.; Gibbs, R.A.; Weinstock, G.M.; Brown, S.J.; Denell, R.; Beeman, R.W.; Gibbs, R.; Bucher, G.; Friedrich, M.; Grimmelikhuijzen, C.J. The genome of the model beetle and pest Tribolium castaneum. Nature 2008, 452, 949–955. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Yang, P.; Chen, D.; Jiang, F.; Li, Y.; Wang, X.; Kang, L. Identification and functional analysis of olfactory receptor family reveal unusual characteristics of the olfactory system in the Migratory locust. Cell. Mol. Life Sci. 2015, 72, 4429–4443. [Google Scholar] [CrossRef] [Green Version]
- Matthews, B.B.; Dos Santos, G.; Crosby, M.A.; Emmert, D.B., St.; Pierre, S.E.; Gramates, L.S.; Zhou, P.; Schroeder, A.J.; Falls, K.; Strelets, V. Gene model annotations for Drosophila melanogaster: Impact of high-throughput data. G3 Genes Genomes Genet. 2015, 5, 1721–1736. [Google Scholar] [CrossRef] [Green Version]
- Tanaka, K.; Uda, Y.; Ono, Y.; Nakagawa, T.; Suwa, M.; Yamaoka, R.; Touhara, K. Highly selective tuning of a silkworm olfactory receptor to a key mulberry leaf volatile. Curr. Biol. 2009, 19, 881–890. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, X.K.; Sun, L.; Liu, H.W.; Liu, D.F.; Ding, Y.X.; Li, L.M.; Zhang, Y.J.; Guo, Y.Y. Identification and expression analysis of an olfactory receptor gene family in green plant bug Apolygus lucorum (Meyer-Dur). Sci. Rep. 2016, 6, 37870. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, S.-N.; Peng, Y.; Lu, Z.-Y.; Dhiloo, K.H.; Gu, S.-H.; Li, R.-J.; Zhou, J.-J.; Zhang, Y.-J.; Guo, Y.-Y. Identification and Expression Analysis of Putative Chemosensory Receptor Genes in Microplitis mediator by Antennal Transcriptome Screening. Int. J. Biol. Sci. 2015, 11, 737–751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grosse-Wilde, E.; Kuebler, L.S.; Bucks, S.; Vogel, H.; Wicher, D.; Hansson, B.S. Antennal transcriptome of Manduca sexta. Proc. Natl. Acad. Sci. USA 2011, 108, 7449–7454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smadja, C.; Shi, P.; Butlin, R.K.; Robertson, H.M. Large Gene Family Expansions and Adaptive Evolution for Odorant and Gustatory Receptors in the Pea Aphid, Acyrthosiphon pisum. Mol. Biol. Evol. 2009, 26, 2073–2086. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.D.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anderson, I.; Brass, A. Searching DNA databases for similarities to DNA sequences: When is a match significant? Bioinformatics 1998, 14, 349–356. [Google Scholar] [CrossRef]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997v2. Available online: https://arxiv.org/abs/1303.3997v2 (accessed on 26 May 2013).
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L.L. Predicting transmembrane protein topology with a hidden markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef] [Green Version]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. ProtTest 3: Fast selection of best-fit models of protein evolution. Bioinformatics 2011, 27, 1164–1165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiao, Y.; Sun, L.; Ma, X.Y.; Dong, K.; Liu, H.W.; Wang, Q.; Guo, Y.Y.; Liu, Z.W.; Zhang, Y.J. Identification and characterization of the distinct expression profiles of candidate chemosensory membrane proteins in the antennal transcriptome of Adelphocoris lineolatus (Goeze). Insect Mol. Biol. 2017, 26, 74–91. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCt Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Ramakers, C.; Ruijter, J.M.; Deprez, R.H.; Moorman, A.F. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 2003, 339, 62–66. [Google Scholar] [CrossRef]
- Modrek, B.; Lee, C. A genomic view of alternative splicing. Nat. Genet. 2002, 30, 13–19. [Google Scholar] [CrossRef]
- Cao, D.; Liu, Y.; Walker, W.B.; Li, J.; Wang, G. Molecular characterization of the Aphis gossypii olfactory receptor gene families. PLoS ONE 2014, 9, e101187. [Google Scholar] [CrossRef]
- Wang, Q.; Zhou, J.J.; Liu, J.T.; Huang, G.Z.; Xu, W.Y.; Zhang, Q.; Chen, J.L.; Zhang, Y.J.; Li, X.C.; Gu, S.H. Integrative transcriptomic and genomic analysis of odorant binding proteins and chemosensory proteins in aphids. Insect Mol. Biol. 2019, 28, 1–22. [Google Scholar] [CrossRef]
- Kwon, H.W.; Lu, T.; Rutzler, M.; Zwiebel, L.J. Olfactory responses in a gustatory organ of the malaria vector mosquito Anopheles gambiae. Proc. Natl. Acad. Sci. USA 2006, 103, 13526–13531. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edwards, L.J.; Siddall, J.B.; Dunham, L.L.; Uden, P.; Kislow, C.J. Trans-β-farnesene, Alarm Pheromone of the Green Peach Aphid, Myzus persicae (Sulzer). Nature 1973, 241, 126–127. [Google Scholar] [CrossRef]
- Zhang, R.; Wang, B.; Grossi, G.; Falabella, P.; Liu, Y.; Yan, S.; Lu, J.; Xi, J.; Wang, G. Molecular Basis of Alarm Pheromone Detection in Aphids. Curr. Biol. 2017, 27, 55–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gu, S.H.; Wu, K.M.; Guo, Y.Y.; Field, L.M.; Pickett, J.A.; Zhang, Y.J.; Zhou, J.J. Identification and Expression Profiling of Odorant Binding Proteins and Chemosensory Proteins between Two Wingless Morphs and a Winged Morph of the Cotton Aphid Aphis gossypii Glover. PLoS ONE 2013, 8, e73524. [Google Scholar] [CrossRef] [Green Version]


| Gene Name | ORF (bp) | Amino Acids | Scaffold * | Antennae RPKM | Head RPKM | Legs RPKM | Body RPKM | Homology Search with A. Pisum | ||
|---|---|---|---|---|---|---|---|---|---|---|
| Most Similar | %Identity | TM Domains | ||||||||
| MperORco | 1494 | 498 | 173 (307,518–314,584) | 16.04 | 1.32 | 0.15 | 1.05 | ApisOR1 | 96 | 7 |
| MperOR2 | 1203 | 401 | 200 (329,352–332,846) | 4.19 | 0.14 | 0.07 | 0.14 | ApisOR2 | 93 | 5 |
| MperOR3 | 1206 | 402 | 27 (538,815–544,733) | 2.76 | 0.14 | 0.04 | 0.08 | ApisOR3 | 94 | 8 |
| MperOR4 | 1167 | 389 | 2 (1,669,537–1,680,404) | 0.66 | 0.25 | 0.08 | 0.12 | ApisOR4 | 94 | 6 |
| MperOR5 | 1104 | 368 | 185 (390,783–396,720) | 1.28 | 0.08 | 0.00 | 0.02 | ApisOR5 | 87 | 8 |
| MperOR9 | 1107 | 369 | 52 (795,957–799,660) | 1.23 | 0.03 | 0.03 | 0.03 | ApisOR9 | 82 | 6 |
| MperOR10 | 1122 | 374 | 448 (232,245–234,867) | 1.23 | 0.03 | 0.03 | 0.03 | ApisOR10 | 68 | 7 |
| MperOR17 | 1296 | 432 | 54 (655,522–659,088) | 1.36 | 4.15 | 1.44 | 5.60 | ApisOR17 | 84 | 6 |
| MperOR20 | 1266 | 422 | 27 (295,168–299,537) | 3.45 | 0.14 | 0.62 | 0.32 | ApisOR20 | 89 | 6 |
| MperOR21 | 1272 | 424 | 27 (273,498–275,759) | 1.39 | 0.20 | 0.05 | 0.05 | ApisOR21 | 84 | 7 |
| MperOR22 | 1290 | 430 | 27 (286,505–290,732) | 0.09 | 0.00 | 0.00 | 0.00 | ApisOR22 | 80 | 6 |
| MperOR23 | 1248 | 416 | 128 (606,788–609,435) | 0.42 | 0.00 | 0.00 | 0.02 | ApisOR23 | 90 | 7 |
| MperOR24 | 1224 | 408 | 128 (619,838–621,712) | 1.49 | 5.02 | 1.90 | 4.38 | ApisOR23 | 43 | 4 |
| MperOR25 | 1203 | 401 | 1011 (10,792–14,561) | 1.30 | 0.01 | 0.01 | 0.05 | ApisOR25 | 80 | 5 |
| MperOR29 | 1242 | 414 | 495 (178,577–182,414) | 0.21 | 0.06 | 0.00 | 0.01 | ApisOR29 | 72 | 6 |
| MperOR35 | 1221 | 407 | 139 (21,194–26,450) | 1.32 | 0.01 | 0.01 | 0.08 | ApisOR35 | 59 | 6 |
| MperOR36 | 1221 | 407 | 0 (1,027,879–1,031,452) | 0.19 | 0.02 | 0.00 | 0.00 | ApisOR36 | 75 | 6 |
| MperOR37 | 1251 | 417 | 533 (71,681–75,764) | 1.95 | 0.01 | 0.05 | 0.04 | ApisOR37 | 88 | 7 |
| MperOR38 | 1221 | 407 | 16 (1,217,873–1,219,749) | 0.26 | 0.03 | 0.07 | 0.05 | ApisOR38 | 84 | 6 |
| MperOR39 | 1188 | 396 | 257 (222,543–224,270) | 0.12 | 0.00 | 0.02 | 0.03 | ApisOR39 | 93 | 5 |
| MperOR40 | 1266 | 422 | 268 (227,342–230,883) | 0.53 | 0.00 | 0.01 | 0.00 | ApisOR40 | 86 | 6 |
| MperOR41 | 1248 | 416 | 268 (236,969–239,632) | 0.61 | 0.04 | 0.18 | 0.28 | ApisOR41 | 84 | 6 |
| MperOR42 | 1290 | 430 | 129 (610,966–615,169) | 0.12 | 2.27 | 0.21 | 0.65 | ApisOR42 | 75 | 7 |
| MperOR43 | 1254 | 418 | 138 (86,064–89,109) | 1.32 | 0.15 | 0.01 | 0.08 | ApisOR43 | 92 | 8 |
| MperOR44 | 1140 | 380 | 115 (575,915–582,437) | 0.10 | 0.05 | 0.00 | 0.18 | ApisOR44 | 59 | 8 |
| MperOR45 | 1296 | 432 | 13 (1,040,637–1,046,400) | 0.50 | 0.22 | 0.05 | 0.64 | ApisOR45 | 58 | 5 |
| MperOR47 | 1302 | 434 | 109 (556,358–561,734) | 0.08 | 0.00 | 0.05 | 0.13 | ApisOR47 | 73 | 7 |
| MperOR51 | 1293 | 431 | 1192 (10,158–17,868) | 0.53 | 0.06 | 0.00 | 0.03 | ApisOR70 | 70 | 6 |
| MperOR53 | 1281 | 427 | 700 (104,988–111,553) | 0.84 | 0.04 | 0.00 | 0.02 | ApisOR53 | 70 | 6 |
| MperOR64 | 1290 | 430 | 380 (259,518–265,853) | 0.53 | 0.06 | 0.00 | 0.03 | ApisOR64 | 71 | 5 |
| MperOR67 | 1239 | 413 | 354 (145,937–151,752) | 5.87 | 0.06 | 0.05 | 0.09 | ApisOR67 | 81 | 6 |
| MperOR69 | 1626 | 542 | 304 (223,678–231,005) | 0.36 | 0.04 | 0.06 | 0.02 | ApisOR69 | 59 | 5 |
| MperOR78 | 1290 | 430 | 140 (453,957–457,387) | 0.36 | 0.05 | 0.03 | 0.08 | ApisOR78 | 62 | 6 |
| Gene Name | Genomic DNA Size (bp) and Strand (+/−) | No. of Intron | Total Length of Introns (bp) | Average Intron Size (bp) | No. of Exon | Total Length of Exons (bp) | Average Exon Size (bp) |
|---|---|---|---|---|---|---|---|
| MperORco | 7066(+) | 8 | 5572 | 697 | 9 | 1494 | 166 |
| MperOR2 | 3494(−) | 1 | 2291 | 2291 | 2 | 1203 | 602 |
| MperOR3 | 5918(+) | 3 | 4712 | 1571 | 4 | 1206 | 302 |
| MperOR4 | 10,867(+) | 5 | 9700 | 1940 | 6 | 1167 | 195 |
| MperOR5 | 5937(+) | 4 | 4833 | 1208 | 5 | 1104 | 221 |
| MperOR9 | 3703(+) | 4 | 2596 | 649 | 5 | 1107 | 221 |
| MperOR10 | 2622(+) | 4 | 1500 | 375 | 5 | 1122 | 224 |
| MperOR17 | 3566(−) | 5 | 2270 | 454 | 6 | 1296 | 216 |
| MperOR20 | 4369(−) | 5 | 3103 | 621 | 6 | 1266 | 211 |
| MperOR21 | 2261(−) | 5 | 989 | 198 | 6 | 1272 | 212 |
| MperOR22 | 4227(−) | 5 | 2937 | 587 | 6 | 1290 | 215 |
| MperOR23 | 2647(−) | 5 | 1399 | 280 | 6 | 1248 | 208 |
| MperOR24 | 1874(−) | 4 | 650 | 163 | 5 | 1224 | 245 |
| MperOR25 | 3769(+) | 5 | 2566 | 513 | 6 | 1203 | 201 |
| MperOR29 | 3837(+) | 5 | 2595 | 519 | 6 | 1242 | 207 |
| MperOR35 | 5256(−) | 5 | 4035 | 807 | 6 | 1221 | 204 |
| MperOR36 | 3573(−) | 5 | 2352 | 470 | 6 | 1221 | 204 |
| MperOR37 | 4083(+) | 5 | 2832 | 566 | 6 | 1251 | 209 |
| MperOR38 | 1876(+) | 4 | 655 | 164 | 5 | 1221 | 244 |
| MperOR39 | 1727(+) | 4 | 539 | 135 | 5 | 1188 | 238 |
| MperOR40 | 3541(−) | 5 | 2275 | 455 | 6 | 1266 | 211 |
| MperOR41 | 2663(+) | 5 | 1415 | 283 | 6 | 1248 | 208 |
| MperOR42 | 4203(+) | 5 | 2913 | 583 | 6 | 1290 | 215 |
| MperOR43 | 3045(+) | 5 | 1791 | 358 | 6 | 1254 | 209 |
| MperOR44 | 6522(−) | 5 | 5382 | 1076 | 6 | 1140 | 190 |
| MperOR45 | 5763(+) | 5 | 4467 | 893 | 6 | 1296 | 216 |
| MperOR47 | 5376(+) | 5 | 4074 | 815 | 6 | 1302 | 217 |
| MperOR51 | 7710(−) | 5 | 6417 | 1283 | 6 | 1293 | 216 |
| MperOR53 | 6565(+) | 5 | 5284 | 1057 | 6 | 1281 | 214 |
| MperOR64 | 6335(+) | 5 | 5045 | 1009 | 6 | 1290 | 215 |
| MperOR67 | 5815(−) | 6 | 4576 | 763 | 7 | 1239 | 177 |
| MperOR69 | 7327(+) | 5 | 5701 | 1140 | 6 | 1626 | 271 |
| MperOR78 | 3430(+) | 5 | 2140 | 428 | 6 | 1290 | 215 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, J.; Xie, J.; Khashaveh, A.; Zhou, J.; Zhang, Y.; Dong, H.; Cong, B.; Gu, S. Identification and Tissue Expression Profiles of Odorant Receptor Genes in the Green Peach Aphid Myzus persicae. Insects 2022, 13, 398. https://doi.org/10.3390/insects13050398
Liu J, Xie J, Khashaveh A, Zhou J, Zhang Y, Dong H, Cong B, Gu S. Identification and Tissue Expression Profiles of Odorant Receptor Genes in the Green Peach Aphid Myzus persicae. Insects. 2022; 13(5):398. https://doi.org/10.3390/insects13050398
Chicago/Turabian StyleLiu, Jingtao, Jiaoxin Xie, Adel Khashaveh, Jingjiang Zhou, Yongjun Zhang, Hui Dong, Bin Cong, and Shaohua Gu. 2022. "Identification and Tissue Expression Profiles of Odorant Receptor Genes in the Green Peach Aphid Myzus persicae" Insects 13, no. 5: 398. https://doi.org/10.3390/insects13050398






