Simple Summary
The brown planthopper, Nilaparvata lugens, is a destructive insect pest that poses a serious threat to rice production. Because the brown planthopper is a phloem-sap-feeding insect and relies on sugars as their main source of carbon, sugar transporters are particularly important. We identified 34 sugar transporters in N. lugens (NlSTs) and inferred their possible function based on their expression patterns. We identified a number of NlST genes that are expressed at higher levels in the gut than in other tissues, such as NlST2, 3, 4, 7, 20, 27, 28, and 31. This study provides a basis for further exploring the function of NlST genes in brown planthopper.
Abstract
Sugar transporters play important roles in controlling carbohydrate transport and are responsible for mediating the movement of sugars into cells in numerous organisms. In insects, sugar transporters not only play a role in sugar transport but may also act as receptors for virus entry and the accumulation of plant defense compounds. The brown planthopper, Nilaparvata lugens, inflicts damage on rice plants by feeding on their phloem sap, which is rich in sugars. In the present study, we identified 34 sugar transporters in N. lugens, which were classified into three subfamilies based on phylogenetic analysis. The motif numbers varied from seven to eleven, and motifs 2, 3, and 4 were identified in the functional domains of all 34 NlST proteins. Chromosome 1 was found to possess the highest number of NlST genes, harboring 15. The gut, salivary glands, fat body, and ovary were the different tissues enriched with NlST gene expression. The expression levels of NlST2, 3, 4, 7, 20, 27, 28, and 31 were higher in the gut than in the other tissues. When expressed in a Saccharomyces cerevisiae hexose transporter deletion mutant (strain EBY.VW4000), only ApST4 (previously characterized) and NlST4, 28, and 31 were found to transport glucose and fructose, resulting in functional rescue of the yeast mutant. These results provide valuable data for further studies on sugar transporters in N. lugens and lay a foundation for finding potential targets to control N. lugens.
1. Introduction
Sugar primarily provides energy and carbon skeletons for multicellular organisms [1,2,3]. Phloem-feeding insects mainly feed on phloem sap that contains high concentrations of sucrose (a disaccharide sugar comprising glucose and fructose) [4,5]. However, the ingested sucrose cannot be directly absorbed through the gut epithelium and must first be hydrolyzed into glucose and fructose [6]. The transportation of glucose, fructose, and trehalose (the primary circulating sugar in insect blood or hemolymph) across the gut epithelium is facilitated by sugar transporters (STs) [7,8]. For example, in Polypedilum vanderplanki, TRET1 (a trehalose transporter gene) is responsible for the release of trehalose from the fat body and the incorporation of trehalose into other tissues that require a carbon source, thereby regulating trehalose levels in the hemolymph [9,10]. The enhanced gut expression of ApST4 and the transport specificity of its product are consistent with ApST4 functioning as a gut glucose/fructose transporter in Acyrthosiphon pisum [11]. In addition, STs can remove monosaccharides (mainly glucose) from the gut lumen to cells, thereby regulating the high osmotic pressure caused by phloem sap entering the gut [12].
In addition to their basic transport functions, sugar transporters take the form of receptors on the surface of the cell membrane and may act as receptors for virus entry. These transporters are used for the transport of sugar molecules, which in turn makes them potential targets for viruses to enter host cells [13]. In the susceptible Bombyx mori race, there is constitutive expression of sugar transporter genes, causing the host to succumb to viral infection [14]. In Laodelphax striatellus, LsST6 can mediate viral entry into midgut epithelial cells and lead to successful transmission [15]. Additionally, sugar transporters have been demonstrated to be involved in plant–insect interactions. PaGTRs enable leaf beetles to accumulate plant defense compounds [16]. In a recent report, a cluster of Bombycidae-specific sugar transporter duplicates with complementary temporal expression synergistically facilitated the uptake of flavonoids, thus determining the development of the green cocoon [17]. There are eight families of sugar transporters present in plants [18], but the classification of sugar transporters in insects remain poorly understood.
Sequencing is a dedicated technology for genome analysis that enriches the information of genome sequences and provides insights into genome organization, genetic variation, and gene expression [19,20]. With the advent of whole-genome sequencing, more gene families have been identified in insects [21], including sugar transporters. A total of 19 sugar transporters (STs) were identified in the Acyrthosiphon pisum genome [11]. A genome-wide annotation of the silkworm B. mori revealed the existence of 100 putative sugar transporter (BmST) genes [14]. Moreover, 137 sugar transporters were identified based on an analysis of the genome and transcriptome of Bemisia tabaci MEAM1 [22].
The brown planthopper, Nilaparvata lugens Stål, is a monophagous rice herbivore and the most notorious pest of rice (Oryza sativa) [23,24]. It sucks the sap from the rice phloem using its stylet, causing direct damage. It is also a vector of viral diseases that cause secondary damage to rice [25,26]. The sugar in phloem sap is regarded as a major energy source in the brown planthopper [27]. Elucidating the mechanisms of sugar uptake in the gut and hemolymph is important for understanding the energy acquisition of plant-feeding insects and identifying new targets for controlling these pests. Several previous studies have reported on STs in the brown planthopper. In this species, NlHT1 (N. lugens hexose transporter 1) is likely to play an important role in glucose transport from the gut and in carbon nutrition in vivo [28]. In another study, 18 putative sugar transporter genes were identified from a brown planthopper EST (expressed sequence tags) database [29]. NlST6 is a facilitative glucose/fructose transporter that mediates sugar uptake from rice phloem sap in the N. lugens midgut [29]. Nlst6 knockdown significantly affected oviposition development and decreased the fat body and ovarian protein content in the brown planthoppers. Nlst6 plays an important role in N. lugens growth and fecundity, and it has potential as a novel target gene for the control of phloem-feeding pest insects [30].
Although several sugar transporters have been studied in N. lugens, genome-wide identification and systematic study of the STs family are still limited. In the current study, genome-wide identification and expression analyses of sugar transporters were conducted based on the genome and transcriptome of the brown planthopper. The transcriptomic data were used to mine for highly expressed NlST genes in the gut and to screen for candidate NlST genes responsible for transporting glucose and fructose. The overall goal was to increase our understanding of these STs and to identify those that might serve as targets for controlling the brown planthopper.
2. Materials and Methods
2.1. Brown Planthopper Rearing and Growth Conditions
The brown planthopper (Nilaparvata lugens Stål) biotype 1, which does not contain any N. lugens resistance gene, was originally obtained from the Center for Excellence in Molecular Plant Sciences in Shanghai, China. The N. lugens were reared on 1-month-old plants of the susceptible rice varieties Taichung Native 1 under controlled environmental conditions (26 °C ± 2 °C, 16 h light/8 h dark cycle) in Zhoukou, China. These insects were transferred to fresh rice seedlings every 14 days to ensure adequate nutrition.
2.2. De novo Identification of Sugar Transporters in Brown Planthopper
The current version of the genome of N. lugens (https://www.ncbi.nlm.nih.gov/datasets/taxonomy/108931/ (accessed on 9 September 2020)) was annotated through a homolog search and de novo predicting. The current version contains 18,160 protein models. Putative N. lugens sugar transporters matching the TIGRFAM sugar porter (SP) family motif (TIGR00879 [31]) were identified using the hmmsearch program, which is part of the HMMER package (version 3.0) [32]. All identified sugar porter family transporters had a hmmsearch sequence score of >237.80 (trusted cutoff) [11]. Subsequently, domain sequences with e-values greater than 1.2 × 10−22 were screened to construct species-specific sugar transport protein gene family domains of the brown planthopper. The family members retrieved were submitted to the SMART, CDD, and Pfam databases for further domain confirmation, and 34 sugar transporter gene family members were finally retained. Bioperl [33] was used to calculate the physicochemical property of NlSTs, including theoretical isoelectric point, instability index, aliphatic index, and grand average of hydropathicity.
2.3. Conservation Motif and Gene Structure Analysis of NlST Genes in N. lugens
The protein sequences of brown planthopper were downloaded from NCBI. For comparison, the protein sequences of sugar porter family transporters in whitefly (Bemisia tabaci) and pea aphid (Acyrthosiphon pisum) were used [11,22]. A phylogenetic tree was then constructed using the maximum likelihood (ML) method with 1000 guided replications in MEGA-CC 7.0 [34].
To search for conservative motifs, the online tool MEME (http://meme-suite.org/ (accessed on 24 August 2023)) was used with optimization parameters set to a maximum of 10 conserved domains in each gene, with a minimum and maximum width of each motif of 15–38 amino acid sequences. The gene structure of the NlSTs was predicted using the online tool GSDS 2.0 (http://gsds.gao-lab.org/ (accessed on 20 April 2021)) [35]. Finally, TBtools v. 2.019 was used to visualize the results of the conserved motif analysis, genome annotation, motif information, and the constructed evolutionary tree.
2.4. Chromosomal Localization Analysis of NlSTs in N. lugens
The chromosomal distribution information of the NlSTs was obtained from the reference N. lugens database, and the distribution of NlSTs on the chromosomes was visualized using TBtools and plotted on the chromosome location images using MapChart [36].
2.5. Expression Analysis of NlSTs in N. lugens
The RNA-seq raw data from different developmental stages (eggs, first, second, third, fourth, fifth, and adult; SRP310488) and various tissues (ovary, salivary glands, antenna, gut, head, and integument; SRP375431) were derived from NCBI [37]. The expression level of NlST genes was calculated as reads per kilobase exon model per million mapped reads (RPKM). The expression data were hierarchically clustered with average linkage and displayed in the Tutools platform (http://www.cloudtutu.com) [38].
The primers for all genes involved in the experiment were designed for RT-qPCR using Primer3 software [39] (Table S1). The spatial gene expression of the NlSTs were investigated using RT-qPCR as follows: ovary, salivary glands, antenna, gut, head, and integument were dissected from 3 d females of N. lugens using a stereomicroscope. Total RNA was isolated from each type of tissue using RNAiso Plus (Takara, Dalian, Liaoning, China, Cat no. 9108). First-strand cDNA was obtained from all samples through reverse transcription using the PrimeScript RT Reagent Kit with gDNA Eraser (Takara, Dalian, Liaoning, China, Cat no. RR047A) according to the manufacturer’s instructions, followed by amplification with qPCR using the Bio-Rad CFX-96 Real-Time PCR system with the iTaq Universal SYBR Green Supermix Kit (Bio-Rad, CA, USA, Cat no. 1725121). As an endogenous control to normalize expression levels with average threshold cycle numbers, a partial fragment of the N. lugens actin gene was amplified with primers qNlActin-F and qNlActin-R (Table S1). A relative quantitative method (2−ΔΔCt) was applied to evaluate the variation in expression among samples [40].
2.6. NlST Expression Constructs and Yeast Transformation
Full-length coding sequences for the gut-expressed sugar transporters NlST2, 3, 4, 7, 20, 27, 28, and NlST31 were amplified from N. lugens cDNA using KOD Plus Neo polymerase (TOYOBO, Osaka, Japan, Cat no. KOD-201). The PCR primers used contained a 5′ BamHI site and a 3′ EcoRI site (primer sequences are shown in Table S1). The DNA was cloned into the pDRTXa vector using a ClonExpress II One Step Cloning Kit (Vazyme, Nanjing, China, Cat no. C112-01). NlST expression constructs were fully sequenced and used to transform Saccharomyces cerevisiae hexose transporter deletion mutant EBY.VW4000 [41] using the lithium acetate/PEG method [42]. Due to the deletion of at least 20 hexose transporters, EBY.VW4000 has very low hexose transport activity and is unable to grow on minimal medium plates containing hexose sugars as the sole carbon source [41]. Transformants were selected on synthetic complete (SC) media, pH 5.6 (0.67% yeast nitrogen base, 2% maltose, 1% agar supplemented with uracil drop-out mix), at 30 °C for 3–4 days. Positive transformants were replica-plated on SC media lacking maltose but containing 60 mM glucose, fructose, galactose, or mannose. The recovery of EBY.W4000 growth was assessed after 3 days at 30 °C.
3. Results
3.1. Identification and Related Information of the Sugar Transporter Gene Family in Nilaparvata lugens
A total of 34 genes encoding STs were identified through the sugar porter family motif analysis and homology search from the N. lugens genome and were named NlST1-34 based on their location on the chromosome. Table 1 provides details of the gene ID number, genomic length, cDNA length, protein length, molecular weight (MW), isoelectric point (pI), and other properties of the identified NlST family proteins. The proteins encoded by the 34 genes showed different physicochemical properties, with protein length varying from 450 (NlST25) to 824 (NlST22) amino acids, a molecular size between 49.317 kDa and 91.969 kDa, and theoretical isoelectric points between 4.97 and 9.41. Sequence analyses revealed that the number of transmembrane domains ranged from seven to twelve, with most containing twelve.

Table 1.
Structural and biochemical information of the sugar transporter gene family in Nilaparvata lugens. TMD: transmembrane domains; MW: molecular weight; PI: theoretical isoelectric point; II: instability index; AI: aliphatic index; GRAVY: grand average of hydropathicity.
3.2. Conserved Motifs and Exon–Intron Organization of Sugar Transporter Genes
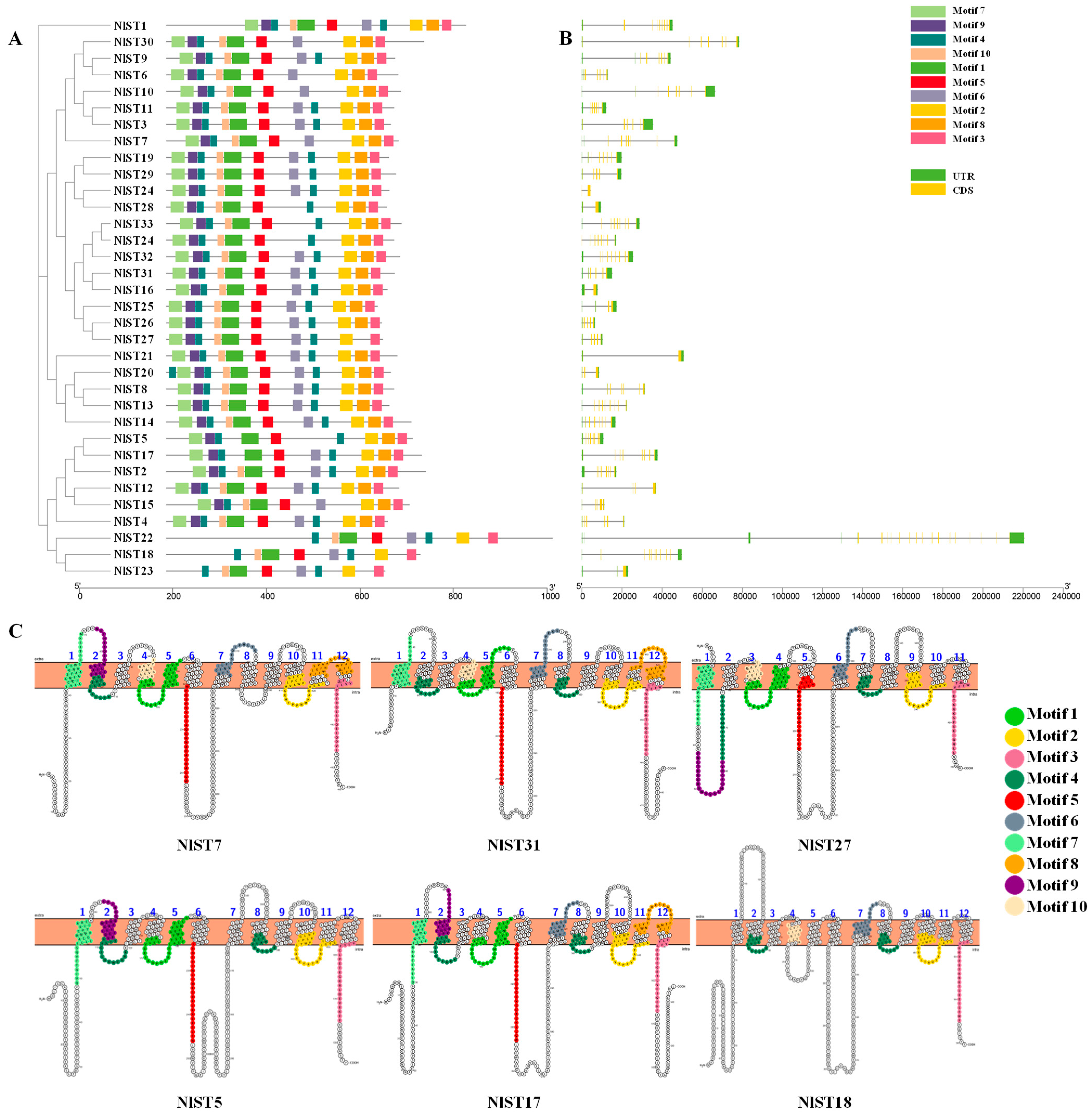
To gain further insight into the structural characteristics of the NlSTs in N. lugens, we employed the MEME server to identify ten conserved motifs (Figure S1). We observed variations in the number of motifs, ranging from seven to eleven. Notably, motifs 2, 3, and 4 were identified within the functional domains of all 34 NlST proteins, suggesting their significance for these proteins in N. lugens (Figure 1A). Interestingly, motif 4 was present twice in most NlST proteins, except NlST6, NlST7, NlST10, NlST15, and NlST30, indicating that it plays a vital role (Figure 1A). Most members had all 11 motifs, such as NlST31. Some members lacked motifs, for example, NlST27 lacked motif 8, and NlST5 lacked motifs 6 and 1. NlST22, NlST18, and NlST23 only had eight motifs, lacking motifs 7, 8, and 9. We also analyzed the distribution of conserved motifs on the transmembrane helices in NlST7, NlST31, NlST27, NlST5, NlST17, and NlST22 (Figure 1C).

Figure 1.
Analysis of motifs, exon–intron structure, and transmembrane helices (TMs) of NlST genes in Nilaparvata lugens. (A) Phylogenetic tree and motifs of NlSTs. Different colors represent different motifs. (B) Gene structures of NlSTs. Yellow boxes, green boxes, and lines represent exons, untranslated region (UTR), and introns, respectively. The lengths of the boxes and lines were scaled according to the gene length. (C) Lineup of conserved motifs on the TMs. The transmembrane structure is marked with blue numbers. The conserved motif is indicated in different colors.
To better comprehend the structure of NlSTs, the exon and intron boundaries were analyzed. Exon–intron borders are very often conserved over long evolutionary distances [43], and structure determines function [44], so they play significant roles in the evolution of gene families. The results revealed that different NlSTs contained different exon numbers, ranging from 2 to 20 (Figure 1B). The gene NlST22 contained the greatest number of exons (20), whereas NlST16 and NlST21 only contained two.
3.3. Phylogenic Analysis and Classification
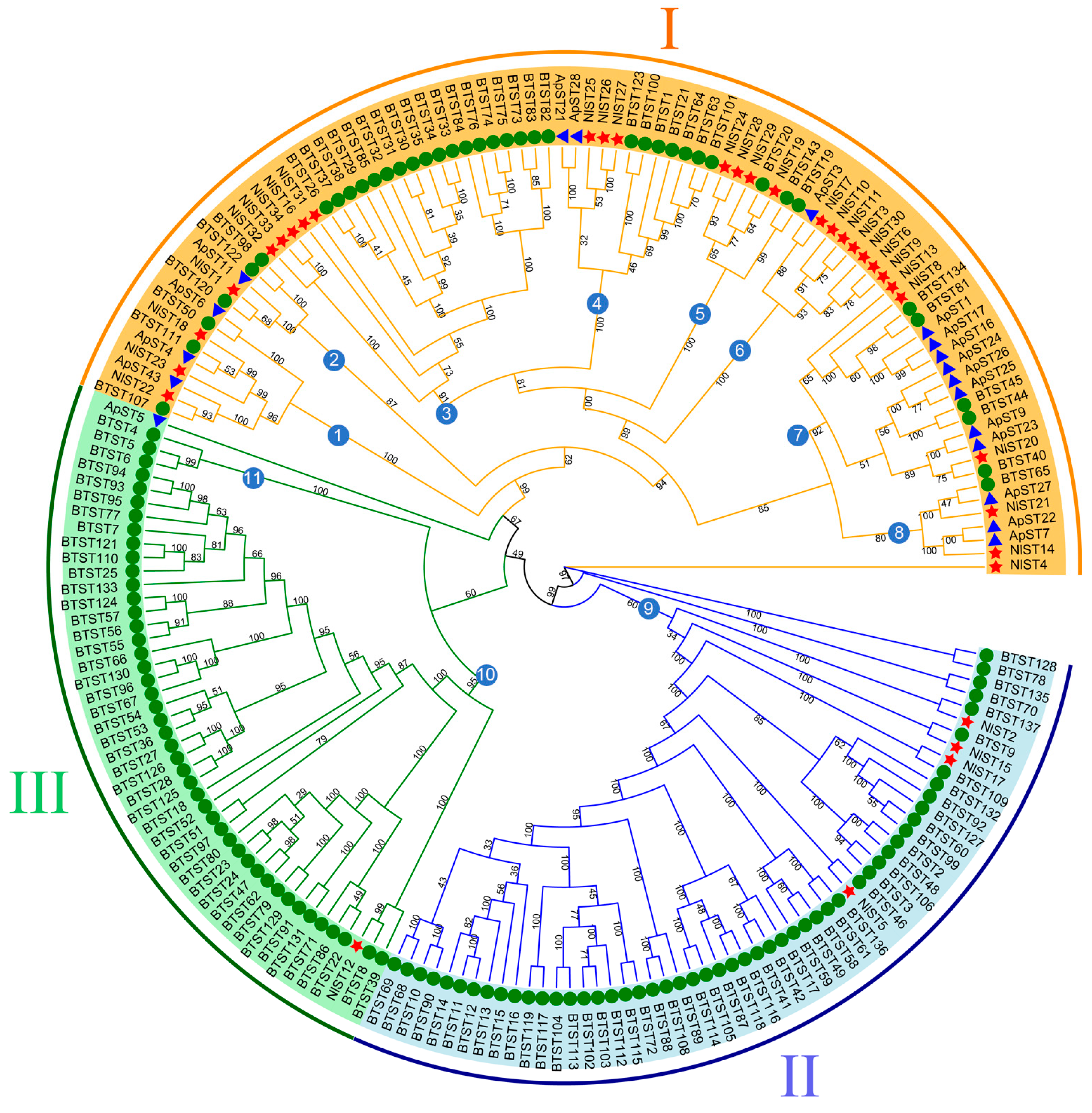
To characterize the evolutionary relationships of the ST gene family, a maximum likelihood tree was created (Figure 2). A phylogenetic tree was constructed by iqtree2 for the 34 NlST proteins, 137 B. tabaci sugar transporter proteins (BtSTs), and 18 A. pisum sugar transporter proteins (ApSTs) after performing multiple sequence alignments. Their protein sequence data are shown as Table S2. According to the topology of the ML phylogenetic tree, the sugar transporter genes of N. lugens, whitefly (B. tabaci), and pea aphid (A. pisum) could be divided into three major groups and eleven subgroups. Three major groups, namely, I, II, and III, contain 29, 4, and 1 NlST proteins, respectively. Group I marked in yellow is the largest subfamily, which contains 29 NlST proteins and all 18 of the ApST proteins. Most homologous genes from the same species are clustered in different subgroups.
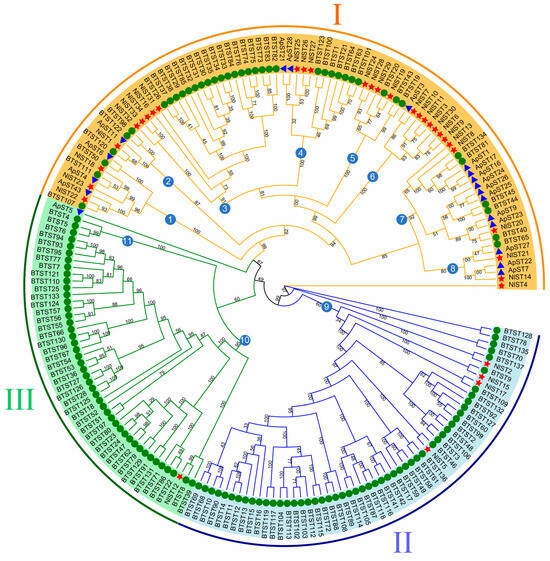
Figure 2.
Phylogenetic tree of brown planthopper (N. lugens), whitefly (Bemisia tabaci), and pea aphid (Acyrthosiphon pisum) sugar transporter proteins. The picture represents the phylogenetic tree as constructed via the maximum likelihood method, where the numbers on the branch represent the bootstrap values. The red star, green circle, and blue triangle represent brown planthopper, whitefly, and pea aphid ST proteins, respectively. Major groups are marked in different colors, with each group marked outside the circle as I, II, and III. Subgroups are indicated by the numbers in the blue circles.
3.4. Chromosomal Localization
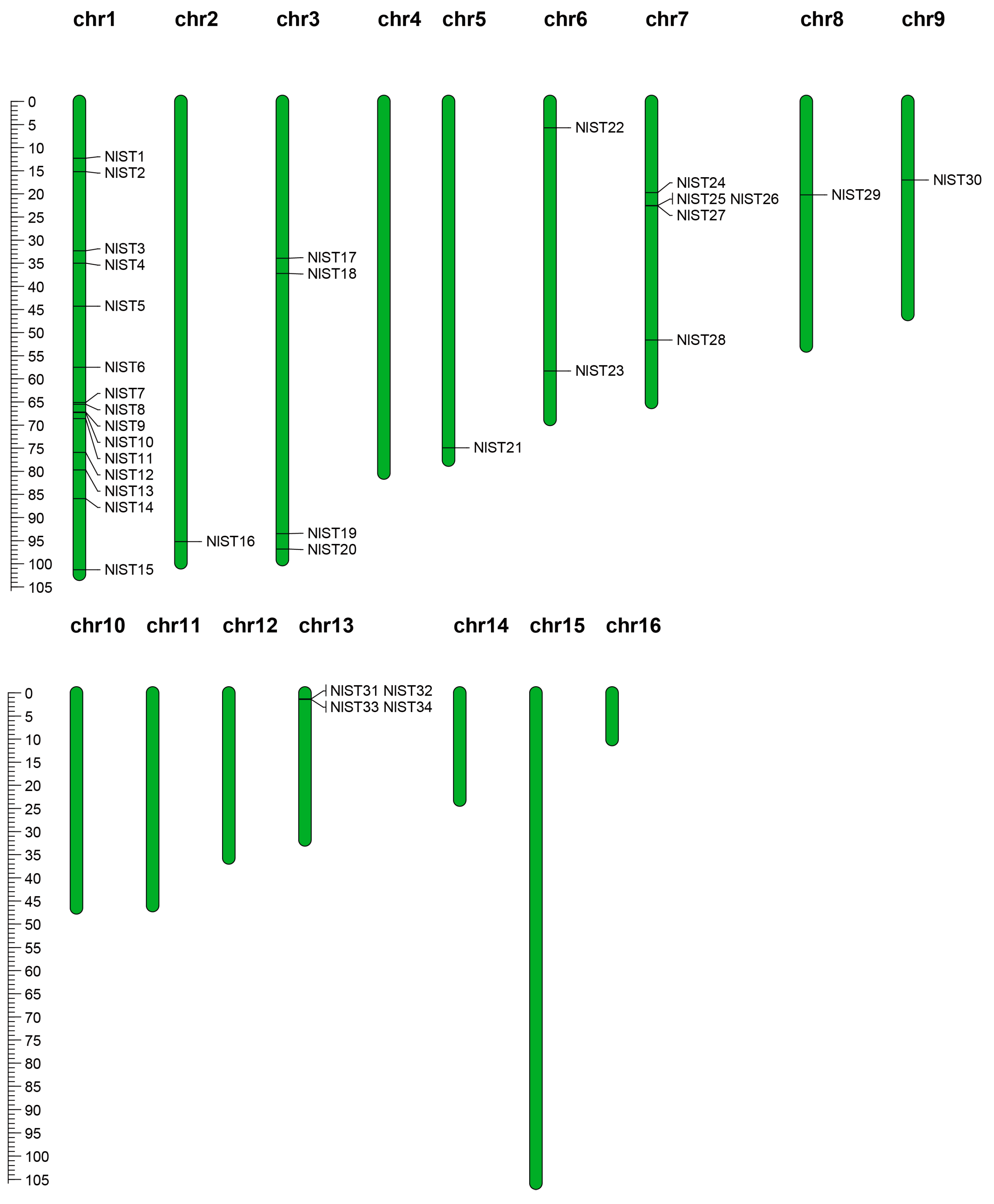
The NlST genes are not distributed on all sixteen chromosomes and were found only on nine N. lugens chromosomes, presenting an unbalanced distribution (Figure 3). The number of NlST genes on the nine chromosomes ranged from one to eight, with the largest number of FtTH genes, which was eight on the first chromosome. Fifteen NlST genes were concentrated on chromosome 1, while a range from one to five genes was found on eight chromosomes (Table 1). Subsequently, gene cluster expansion events of NlSTs in the N. lugens genome were analyzed. In terms of the sequence similarity analysis and phylogenetic relationship, we identified three groups (e.g., NlST9 and NlST10 on chromosome 1; NlST25, NlST26, and NlST27 on chromosome 7; NlST31, NlST32, NlST33, and NlST34 on chromosome 13) of NlST genes. For example, the NlST31, NlST32, NlST33, and NlST34 protein sequences shared 63.25% similarity, whereas those of NlST25, NlST26, and NlST27 showed 59.08% similarity.

Figure 3.
Chromosomal distribution of NlST genes in the N. lugens genome. The chromosomal position of each NlST gene was mapped according to the N. lugens genome. The chromosome number is indicated at the top of each chromosome. The scale bar on the left represents the length (Mb) of the N. lugens chromosomes.
3.5. Expression Profiles of N. lugens Sugar Transporter Genes
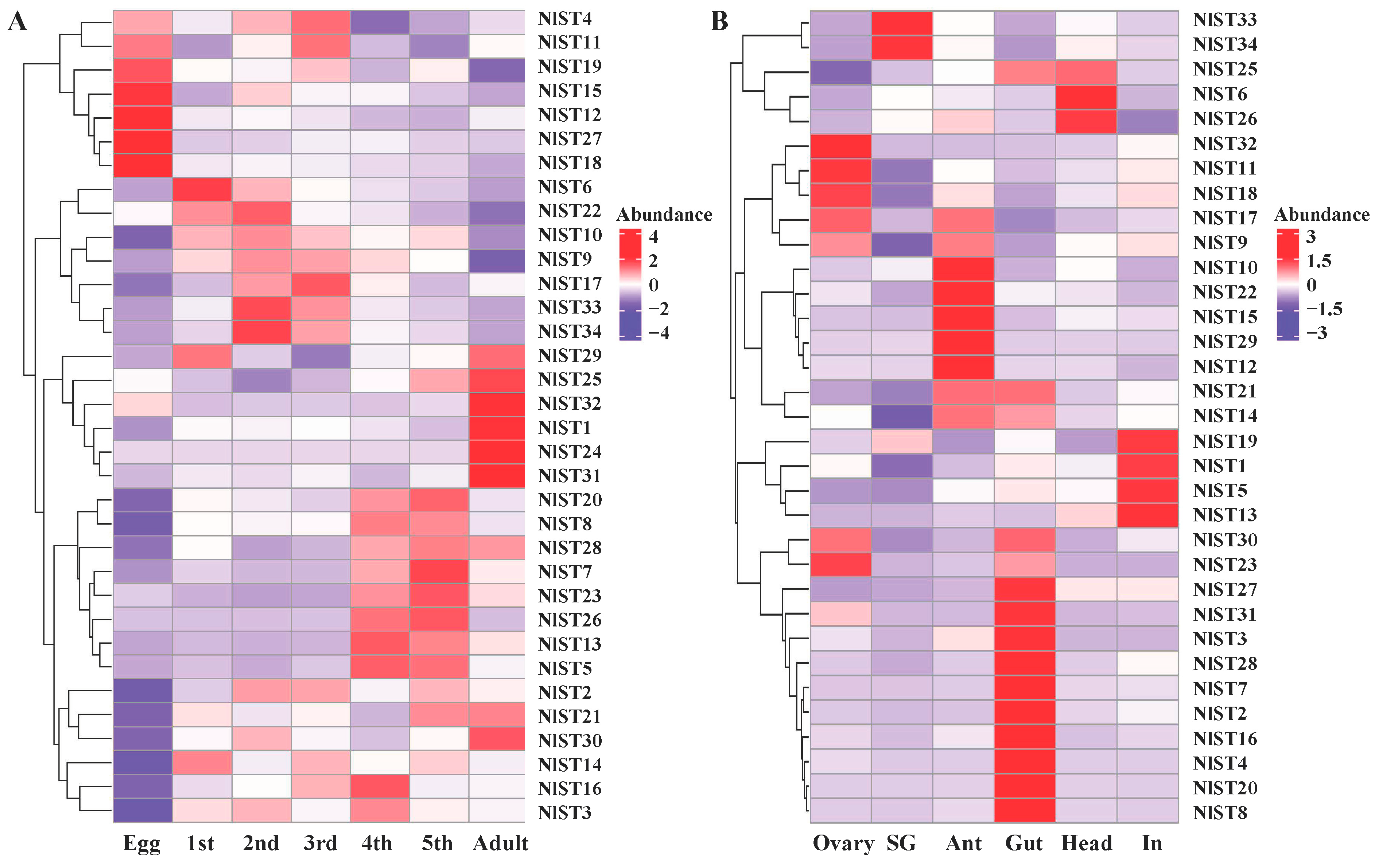
The expression patterns of NlSTs were investigated using RNAseq datasets obtained from different developmental stages of N. lugens. Expression analysis revealed that the NlSTs have a wide expression profile during all life stages, suggesting that they play important roles in development. The transcript expression of most NlSTs was the lowest in eggs, while the expression of NlST4, NlST11, NlST19, NlST15, NlST12, NlST27, and NlST18 was high in eggs but low in other stages (Figure 4A). The expression levels of NlST29, 25, 32, 1, 24, and 31 were higher in the adult than other stages (Figure 4A).
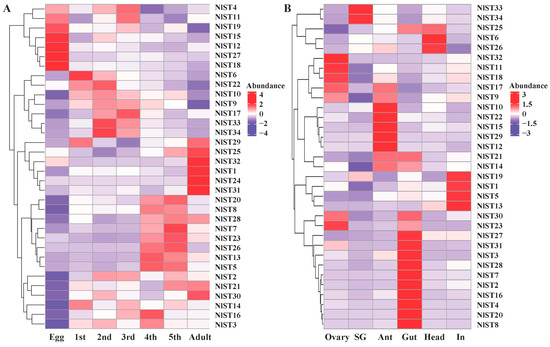
Figure 4.
Gene expression of NlSTs across different developmental stages (A) and various tissues (B). Heat map and hierarchical cluster display differential expression profile of NlST genes across different developmental stages and tissues. Different tissues including ovary, salivary glands (SG), antenna (Ant), gut, head, and integument (In) were dissected from 3 d females for RNA-seq analysis. The expression levels are presented in the heatmap using fold-change values transformed to log2. Bright red, dark purple, and white indicate higher, lower, and equal expression values, respectively.
The expression levels of the NlST genes were examined in the ovary, salivary glands, antenna, gut, head, and integument using transcriptome data obtained in a previous study [37]. The different NlSTs showed variable levels of expression in the tested tissues. The results revealed that the expression level of NlST2, 3, 4, 7, 8, 16, 20, 27, 28, and 31 was higher in the gut than other selected tissues (Figure 4B). NlST33 and NlST34 were highly expressed in the salivary glands; NlST1, 3, and 19 were highly expressed in the fat body; and NlST23, 18, 11, and 32 were highly expressed in the ovary. NlST24 showed a low level of expression in all tissues, and a distinctive expression pattern could not be determined (Figure 4B).
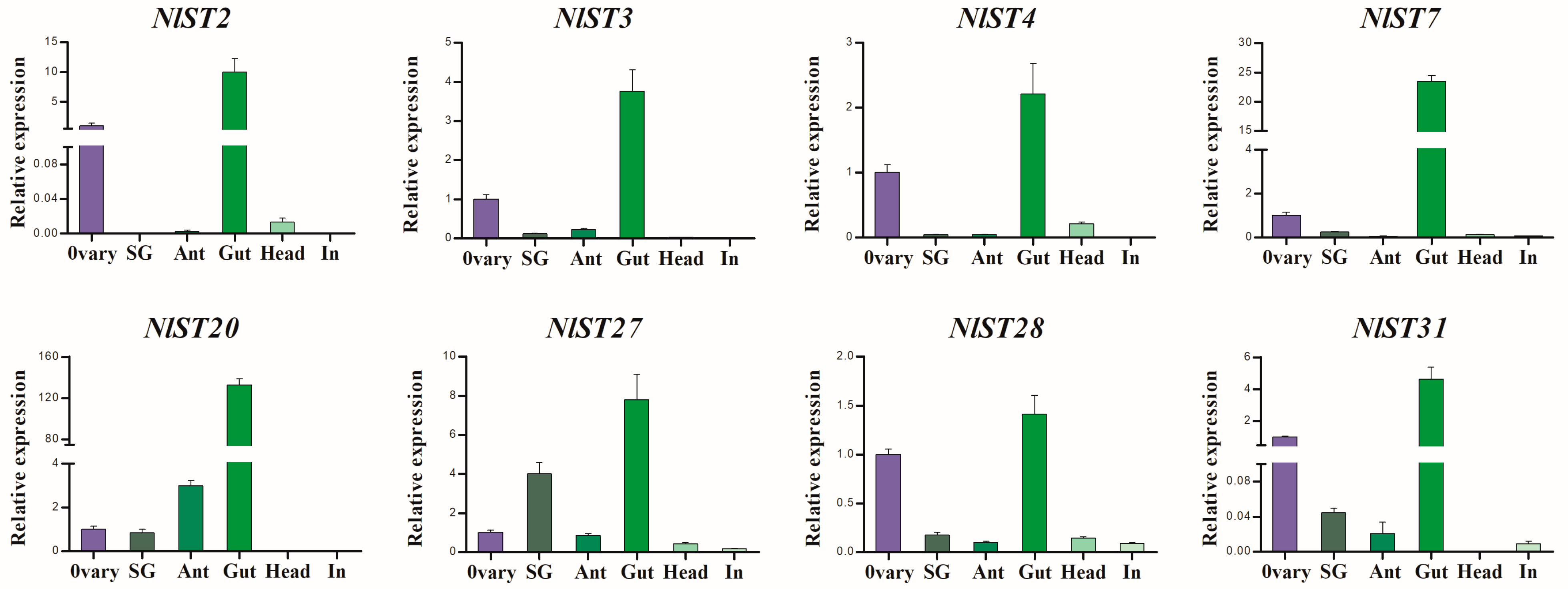
RT-qPCR was used to further determine the expression pattern of NlSTs in N. lugens. As shown in Figure 5, NlST2, 3, 4, 7, 20, 27, 28, and 31 had significantly higher expression levels in the gut than in other tissues. Interestingly, the expression of NlST2, 3, 4, 7, 28, and 31 genes in the ovary was second only to that in the gut, which indicates these genes may have a dual function.

Figure 5.
The expression of several NlSTs in different tissues of N. lugens by RT-qPCR. Different tissues including ovary, salivary glands (SG), antenna (Ant), gut, head, and integument (In) were dissected from 3 d females.
3.6. N. lugens Gut-Expressed Sugar Transporters Transport Hexose Sugars
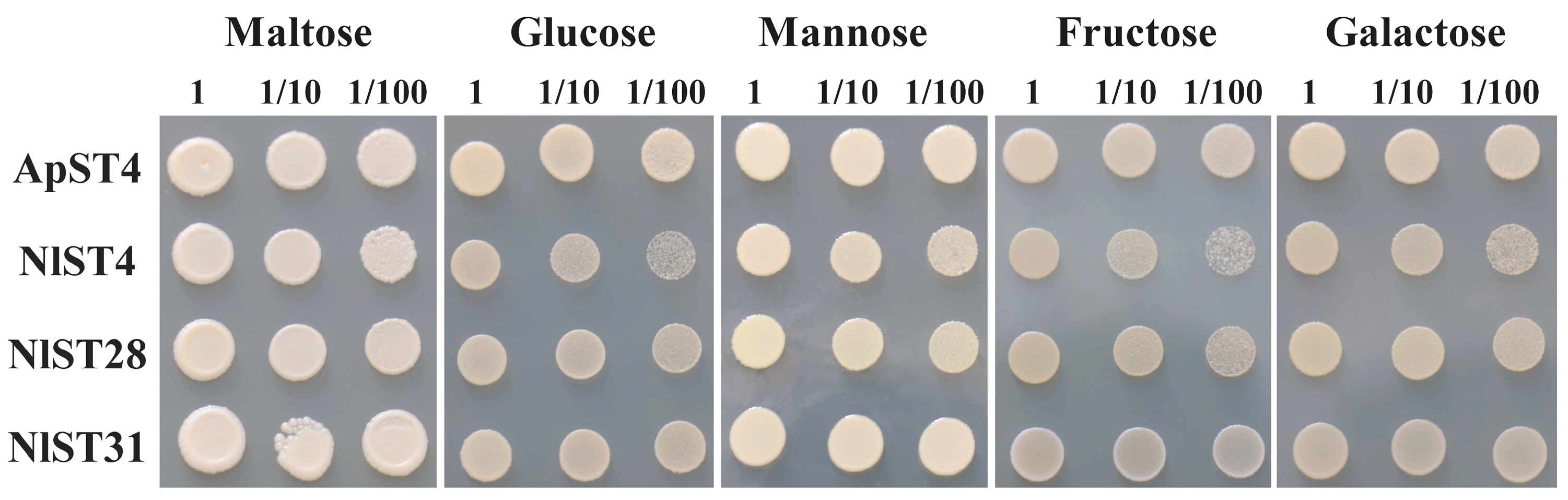
The utilization of ingested sugar as a nutrient resource and the maintenance of osmotic balance depend on the transit of sugar transporters through the gut epithelium and into cells throughout the insect [45]. All eight N. lugens gut-expressed sugar transporters (NlST2, 3, 4, 7, 20, 27, 28, and 31) were screened for hexose transport activity through functional complementation of the S. cerevisiae hexose-uptake-deficient strain EBY.VW4000 [41]. Yeast cells transformed with either ApST4 (positive control) or NlST expression constructs produced recombinant transporter proteins. As shown in Figure 6, ApST4, NlST4, NlST28, and NlST31 were able to restore EBY.VW4000 growth on minimal media containing glucose, fructose, galactose, or mannose as the sole carbon source. These results showed that the transporters NlST4, NlST28, and NlST31 are functional hexose transporters that mediate the efficient transport of hexose sugars across the yeast plasma membrane.

Figure 6.
Functional identification of Saccharomyces cerevisiae hexose transport mutant EBY.VW4000 expressing ApST4 and NlSTs. Positive control cells (+) were transformed with ApST4. Yeast cell suspensions containing 10 μL of 1, 1/10, and 1/100 OD600 units were plated on minimal media containing 60 mM of the indicated sugar as the sole carbon source.
4. Discussion
The whole-genome sequencing of N. lugens combined with a substantial amount of transcriptome sequencing data provided a wealth of invaluable information to explore the evolutionary mechanism, structure, and function of the NlST genes. In the current study, 34 sugar transporter genes were identified from the genome of N. lugens. The 34 NlSTs were located on nine chromosomes, with fifteen concentrated on chromosome 1, showing an unbalanced distribution. In B. tabaci, some BTSTs formed four different gene clusters that were located on four different scaffolds [22]. A similar phenomenon was observed in sugar transporter genes in B. mori, and chromosome 27 had a maximum of 22 BmST genes distributed [14].
According to the evolutionary relationship inferred by phylogenetic analysis, these genes were classified into three main clades, consistent among brown planthopper (N. lugens), whitefly (B. tabaci), and pea aphid (A. pisum) [11,22]. According to the phylogenetic tree analysis, although the majority of NlST, BtST, and ApST orthologs can be divided into 11 distinct subgroups, most homologous genes from the same species are clustered in different subgroups. In addition, previous studies have found that homologous genes in the same branch can exhibit the same or similar biological functions [46]. Up to now, the biological functions of most NlSTs have remained unclear. However, the characterized function of STs in other species, such as A. pisum, can help to predict the gene functions in N. lugens via ortholog analysis in the same subfamily. For example, ApST3, the most abundantly expressed sugar transporter gene in A. pisum, is mainly involved in gut sugar transport [45]. The N. lugens ST genes NlST7, NlST3, NlST30, and NlST6, which are orthologous to ApST3 and grouped in subgroup 6, are highly expressed in the gut and might be essential sugar transporters. Silencing BTST40 and BTST44 can significantly increase the mortality rate of B. tabaci at days 2 and 4. NlST20, BTST40, and BTST44 are grouped in subgroup 7, which indicates NlST20 may be used as a potential RNAi target for the bio-control of N. lugens.
We found that the complex expression of the NlST gene at different developmental stages may be due to differences in nutrient requirements and physiological behavior. Seven NlST genes were highly expressed in eggs. This indicates that they may play vital roles in egg development, because N. lugens does not feed while in the egg. Similar results have been found in Bemisia tabaci [22]. In B. mori, the expression levels of St genes remained at low levels in diapause eggs, whereas high gene expressions of trehalose transporter 1 (Tret1), St4, and St3 were detected in developing eggs [47]. In Harmonia axyridisa, a lack of HaGlut4 can impair ovarian development and oocyte maturation and result in decreased fecundity [48]. Our results also found that some NlST genes were highly expressed in the gut, which is consistent with previous findings that NlST genes may be involved in carbohydrate incorporation from the gut cavity into the hemolymph. For instance, NlST16 was shown to facilitate glucose transport along gradients in N. lugens [49]. Simultaneous knockdown of the five sugar homeostasis genes in the potato psyllid gut yielded high mortality [50].
In addition, the insect gut is the first major barrier limiting virus acquisition; L. striatellus sugar transporters 6 (LsST6) mediates viral entry into midgut epithelial cells and leads to successful transmission by the insect vector [15]. Most of the NlST genes are expressed at lower levels in the salivary glands, except NlST33 and NlST34, suggesting that these two may have a specific role in the pathway of sugar metabolism. In Anopheles stephensi, AsST in the salivary glands is strongly downregulated in response to blood feeding compared to sugar feeding [51]. In addition, we found that the NlSTs were differentially or highly expressed in antennae and ovary, suggesting that these genes may be involved in olfactory detection and breeding. However, we cannot exclude the possibility that these genes are involved in other physiological functions.
The hexose-deficient yeast EBY.VW4000 has been used in many studies to verify whether sugar transporters have hexose transport activity [52,53]. In this study, we used this method to determine whether some N. lugens sugar transporters have glucose and fructose transport activities, such as NlST4, 28, and 31. They are highly expressed in gut tissue and may be involved in moving glucose and fructose from high concentrations in the gut lumen to low concentrations in the hemolymph, such as ApST3 [45] and ApST4 [11]. Sugar transporters belong to the major facilitator superfamily (MFS) [45]. MFS transporters function as either facilitative transporters or secondary active transporters. Facilitator transporters facilitate passive solute transport across membranes by moving the solute along its concentration gradient without expending energy [54]. ApST4 is a facilitative hexose transporter, which is concentration dependent [11]. However, it is not clear whether NlST4, 28, and 31 in N. lugens depend on concentration. Therefore, further studies will include functional complementation assays and uptake experiments in EBY.VW4000 yeast cells.
RNAi played a major role in advances in insect biology [55]. Injection of dsRNA [56], feeding dsRNA [57,58], or spraying dsRNA [59] induces efficient RNAi in N. lugens. So, more precise targets in N. lugens need to be identified and characterized. In our study, NlST4, 28, and 31 were expressed highly in the gut and may be the potential targets of RNAi. Our findings will help to improve the design of effective resistance management strategies to control N. lugens in agriculture.
Supplementary Materials
The following supporting information can be downloaded at https://www.mdpi.com/article/10.3390/insects15070509/s1. Figure S1: Sequence logos for 10 motifs of sugar transporter domains using the MEME program. Table S1: List of primers used in this study. Table S2: Amino acid sequences of the NlSTs, BTSTs, and ApSTs.
Author Contributions
Conceptualization, X.S.; methodology, X.S. and X.Z.; software, X.S. and X.Z.; validation, X.S., X.Z. and M.Z.; formal analysis, X.S., Y.Z., X.L. and M.Y.; investigation, X.Y., S.W., L.G. and L.W.; resources, X.S. and H.C.; data curation, K.X.; writing—original draft preparation, X.S. and X.Z.; writing—review and editing, Y.Z., X.L. and K.X.; visualization, X.S. and M.Z.; supervision, K.X.; project administration, X.S.; funding acquisition, X.S., Y.Z., X.L. and X.Z. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Natural Science Foundation of China, grant numbers 32001921 and 32102487, the Foundation of Henan Science and Technology Committee, grant numbers 232102111094 and 242102110238, and the Department of Science and Technology Planning Project of Henan Province, grant number 242102110299.
Data Availability Statement
The genome and transcriptome data of Nilaparvata lugens are available from NCBI under the BioProject numbers PRJNA913551 and PRJNA901050, respectively. The data presented in this study are available on request from the corresponding author.
Acknowledgments
We acknowledge the generosity of Eckhard Boles for the contribution of S. cerevisiae strain EBY.VW4000.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Thompson, S.N. Trehalose—The Insect ‘Blood’ Sugar. Adv. Insect Physiol. 2003, 31, 205–285. [Google Scholar]
- Liu, N.; Wei, Z.; Min, X.; Yang, L.; Zhang, Y.; Li, J.; Yang, Y. Genome-Wide Identification and Expression Analysis of the SWEET Gene Family in Annual Alfalfa (Medicago polymorpha). Plants 2023, 12, 1948. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Chen, Y.; Wang, S.; Xue, H.; Su, Y.; Yang, J.; Li, X. Identification of candidate genes involved in the sugar metabolism and accumulation during pear fruit post-harvest ripening of ‘Red Clapp’s Favorite’ (Pyrus communis L.) by transcriptome analysis. Hereditas 2018, 155, 11. [Google Scholar] [CrossRef] [PubMed]
- Douglas, A.E.; Price, D.R.G.; Minto, L.B.; Jones, E.; Pescod, K.V.; François, C.L.M.J.; Pritchard, J.; Boonham, N. Sweet problems: Insect traits defining the limits to dietary sugar utilisation by the pea aphid, Acyrthosiphon pisum. J. Exp. Biol. 2006, 209, 1395–1403. [Google Scholar] [CrossRef] [PubMed]
- Sattar, S.; Thompson, G.A. Small RNA Regulators of Plant-Hemipteran Interactions: Micromanagers with Versatile Roles. Front. Plant Sci. 2016, 7, 1241. [Google Scholar] [CrossRef]
- Anand, R.; Divya, D.; Mazumdar-Leighton, S.; Bentur, J.S.; Nair, S. Expression Analysis Reveals Differentially Expressed Genes in BPH and WBPH Associated with Resistance in Rice RILs Derived from a Cross between RP2068 and TN1. Int. J. Mol. Sci. 2023, 24, 13982. [Google Scholar] [CrossRef] [PubMed]
- Mueckler, M. Facilitative glucose transporters. Eur. J. Biochem. 1994, 219, 713–725. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Lei, G.; Chen, Y.; You, M.; You, S. PxTret1-like Affects the Temperature Adaptability of a Cosmopolitan Pest by Altering Trehalose Tissue Distribution. Int. J. Mol. Sci. 2022, 23, 9019. [Google Scholar] [CrossRef] [PubMed]
- Kanamori, Y.; Saito, A.; Hagiwara-Komoda, Y.; Tanaka, D.; Mitsumasu, K.; Kikuta, S.; Watanabe, M.; Cornette, R.; Kikawada, T.; Okuda, T. The trehalose transporter 1 gene sequence is conserved in insects and encodes proteins with different kinetic properties involved in trehalose import into peripheral tissues. Insect Biochem. Mol. Biol. 2010, 40, 30–37. [Google Scholar] [CrossRef]
- Kikawada, T.; Saito, A.; Kanamori, Y.; Nakahara, Y.; Iwata, K.-i.; Tanaka, D.; Watanabe, M.; Okuda, T. Trehalose transporter 1, a facilitated and high-capacity trehalose transporter, allows exogenous trehalose uptake into cells. Proc. Natl. Acad. Sci. USA 2007, 104, 11585–11590. [Google Scholar] [CrossRef]
- Price, D.R.; Gatehouse, J.A. Genome-wide annotation and functional identification of aphid GLUT-like sugar transporters. BMC Genom. 2014, 15, 647. [Google Scholar] [CrossRef] [PubMed]
- Tzin, V.; Yang, X.; Jing, X.; Zhang, K.; Jander, G.; Douglas, A.E. RNA interference against gut osmoregulatory genes in phloem-feeding insects. J. Insect Physiol. 2015, 79, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.T.; Leu, J.H.; Huang, P.Y.; Chen, L.L. A putative cell surface receptor for white spot syndrome virus is a member of a transporter superfamily. PLoS ONE 2012, 7, e33216. [Google Scholar] [CrossRef]
- Govindaraj, L.; Gupta, T.; Esvaran, V.G.; Awasthi, A.K.; Ponnuvel, K.M. Genome-wide identification, characterization of sugar transporter genes in the silkworm Bombyx mori and role in Bombyx mori nucleopolyhedrovirus (BmNPV) infection. Gene 2016, 579, 162–171. [Google Scholar] [CrossRef] [PubMed]
- Carr, J.; Qin, F.; Liu, W.; Wu, N.; Zhang, L.; Zhang, Z.; Zhou, X.; Wang, X. Invasion of midgut epithelial cells by a persistently transmitted virus is mediated by sugar transporter 6 in its insect vector. PLoS Pathog. 2018, 14, e1007201. [Google Scholar] [CrossRef]
- Yang, Z.L.; Nour-Eldin, H.H.; Hanniger, S.; Reichelt, M.; Crocoll, C.; Seitz, F.; Vogel, H.; Beran, F. Sugar transporters enable a leaf beetle to accumulate plant defense compounds. Nat. Commun. 2021, 12, 2658. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Luo, J.; An, E.; Lu, B.; Wei, Y.; Chen, X.; Lu, K.; Liang, S.; Hu, H.; Han, M.; et al. Deciphering the Genetic Basis of Silkworm Cocoon Colors Provides New Insights into Biological Coloration and Phenotypic Diversification. Mol. Biol. Evol. 2023, 40, msad017. [Google Scholar] [CrossRef] [PubMed]
- Sun, N.; Liu, Y.; Xu, T.; Zhou, X.; Xu, H.; Zhang, H.; Zhan, R.; Wang, L. Genome-wide analysis of sugar transporter genes in maize (Zea mays L.): Identification, characterization and their expression profiles during kernel development. PeerJ 2023, 11, e16423. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Shang, L.; Zhu, Q.H.; Fan, L.; Guo, L. Twenty years of plant genome sequencing: Achievements and challenges. Trends Plant Sci. 2022, 27, 391–401. [Google Scholar] [CrossRef]
- Li, F.; Zhao, X.; Li, M.; He, K.; Huang, C.; Zhou, Y.; Li, Z.; Walters, J.R. Insect genomes: Progress and challenges. Insect Mol. Biol. 2019, 28, 739–758. [Google Scholar] [CrossRef]
- Kumar, V.; Garg, S.; Gupta, L.; Gupta, K.; Diagne, C.T.; Misse, D.; Pompon, J.; Kumar, S.; Saxena, V. Delineating the Role of Aedes aegypti ABC Transporter Gene Family during Mosquito Development and Arboviral Infection via Transcriptome Analyses. Pathogens 2021, 10, 1127. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Xia, J.; Pan, H.; Gong, C.; Xie, W.; Guo, Z.; Zheng, H.; Yang, X.; Yang, F.; Wu, Q.; et al. Genome-Wide Characterization and Expression Profiling of Sugar Transporter Family in the Whitefly, Bemisia tabaci (Gennadius) (Hemiptera: Aleyrodidae). Front. Physiol. 2017, 8, 322. [Google Scholar] [CrossRef] [PubMed]
- Jing, S.; Zhao, Y.; Du, B.; Chen, R.; Zhu, L.; He, G. Genomics of interaction between the brown planthopper and rice. Curr. Opin. Insect Sci. 2017, 19, 82–87. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Zhu, L.; He, G. Genetic and molecular understanding of host rice resistance and Nilaparvata lugens adaptation. Curr. Opin. Insect Sci. 2021, 45, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Sōgawa, K. THE RICE BROWN PLANTHOPPER: Feeding Physiology and Host Plant Interactions. Annu. Rev. Entomol. 1982, 27, 49–73. [Google Scholar] [CrossRef]
- Hibino, H. Biology and epidemiology of rice viruses. Annu. Rev. Phytopathol. 1996, 34, 249–274. [Google Scholar] [CrossRef] [PubMed]
- Douglas, A.E. Phloem-sap feeding by animals: Problems and solutions. J. Exp. Bot. 2006, 57, 747–754. [Google Scholar] [CrossRef] [PubMed]
- Price, D.R.G.; Wilkinson, H.S.; Gatehouse, J.A. Functional expression and characterisation of a gut facilitative glucose transporter, NlHT1, from the phloem-feeding insect Nilaparvata lugens (rice brown planthopper). Insect Biochem. Mol. Biol. 2007, 37, 1138–1148. [Google Scholar] [CrossRef] [PubMed]
- Kikuta, S.; Kikawada, T.; Hagiwara-Komoda, Y.; Nakashima, N.; Noda, H. Sugar transporter genes of the brown planthopper, Nilaparvata lugens: A facilitated glucose/fructose transporter. Insect Biochem. Mol. Biol. 2010, 40, 805–813. [Google Scholar] [CrossRef]
- Ge, L.-Q.; Jiang, Y.-P.; Xia, T.; Song, Q.-S.; Stanley, D.; Kuai, P.; Lu, X.-L.; Yang, G.-Q.; Wu, J.-C. Silencing a sugar transporter gene reduces growth and fecundity in the brown planthopper, Nilaparvata lugens (Stål) (Hemiptera: Delphacidae). Sci. Rep. 2015, 5, 12194. [Google Scholar] [CrossRef]
- Haft, D.; Loftus, B.; Richardson, D.; Yang, F.; Eisen, J.; Paulsen, I.; White, O. TIGRFAMs: A protein family resource for the functional identification of proteins. Nucleic Acids Res. 2001, 29, 41–43. [Google Scholar] [CrossRef] [PubMed]
- Johnson, L.; Eddy, S.; Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinform. 2010, 11, 431. [Google Scholar] [CrossRef]
- Stajich, J.E.; Block, D.; Boulez, K.; Brenner, S.E.; Chervitz, S.A.; Dagdigian, C.; Fuellen, G.; Gilbert, J.G.; Korf, I.; Lapp, H.; et al. The Bioperl toolkit: Perl modules for the life sciences. Genome Res. 2002, 12, 1611–1618. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Peterson, D.; Tamura, K. MEGA-CC: Computing core of molecular evolutionary genetics analysis program for automated and iterative data analysis. Bioinformatics 2012, 28, 2685–2686. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.; Guo, A.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Voorrips, R. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.J.; Zhang, J.L.; Xu, H.J. A genome-wide identification and analysis of the homeobox genes in the brown planthopper, Nilaparvata lugens (Hemiptera: Delphacidae). Arch. Insect Biochem. Physiol. 2021, 108, e21833. [Google Scholar] [CrossRef]
- Li, F.; Cao, L.; Bähre, H.; Kim, S.K.; Schroeder, K.; Jonas, K.; Koonce, K.; Mekonnen, S.A.; Mohanty, S.; Bai, F.; et al. Patatin-like phospholipase CapV in Escherichia coli-morphological and physiological effects of one amino acid substitution. NPJ Biofilms Microbiomes 2022, 8, 39. [Google Scholar] [CrossRef] [PubMed]
- Kaur, R.; Ahlawat, S.; Choudhary, V.; Kumari, A.; Chhabra, P.; Arora, R.; Sharma, R.; Vijh, R.K. Comparative expression profiling of cytokine genes in Theileria annulata-infected and healthy cattle. Trop. Anim. Health Prod. 2022, 54, 383. [Google Scholar] [CrossRef]
- Gupta, K.; Dhawan, R.; Kajla, M.; Misra, T.; Kumar, S.; Gupta, L. The evolutionary divergence of STAT transcription factor in different Anopheles species. Gene 2017, 596, 89–97. [Google Scholar] [CrossRef]
- Wieczorke, R.K.S.; Weierstall, T.; Freidel, K.; Hollenberg, C.P.; Boles, E. Concurrent knock-out of at least 20 transporter genes is required to block uptake of hexoses in Saccharomyces cerevisiae. FEBS Lett. 1999, 464, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Gietz, R.; Schiestl, R. Quick and easy yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2007, 2, 35–37. [Google Scholar] [CrossRef]
- Poverennaya, I.V.; Potapova, N.A.; Spirin, S.A. Is there any intron sliding in mammals? BMC Evol. Biol. 2020, 20, 164. [Google Scholar] [CrossRef]
- Han, S.; Han, X.; Qi, C.; Guo, F.; Yin, J.; Liu, Y.; Zhu, Y. Genome-Wide Identification of DUF668 Gene Family and Expression Analysis under F. solani, Chilling, and Waterlogging Stresses in Zingiber officinale. Int. J. Mol. Sci. 2024, 25, 929. [Google Scholar] [CrossRef] [PubMed]
- Price, D.R.G.; Tibbles, K.; Shigenobu, S.; Smertenko, A.; Russell, C.W.; Douglas, A.E.; Fitches, E.; Gatehouse, A.M.R.; Gatehouse, J.A. Sugar transporters of the major facilitator superfamily in aphids; from gene prediction to functional characterization. Insect Mol. Biol. 2010, 19, 97–112. [Google Scholar] [CrossRef]
- Liu, Y.; Wu, S.; Lan, K.; Wang, Q.; Ye, T.; Jin, H.; Hu, T.; Xie, T.; Wei, Q.; Yin, X. An Investigation of the JAZ Family and the CwMYC2-like Protein to Reveal Their Regulation Roles in the MeJA-Induced Biosynthesis of β-Elemene in Curcuma wenyujin. Int. J. Mol. Sci. 2023, 24, 15004. [Google Scholar] [CrossRef]
- Gu, S.; Lin, P.; Chang, C. Expressions of sugar transporter genes during Bombyx mori embryonic development. J. Exp. Zool. Part A Ecol. Integr. Physiol. 2023, 339, 788–798. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, S.S.; Wang, S.; Wang, S.G.; Tang, B.; Liu, F. Involvement of glucose transporter 4 in ovarian development and reproductive maturation of Harmonia axyridis (Coleoptera: Coccinellidae). Insect Sci. 2022, 29, 691–703. [Google Scholar] [CrossRef]
- Kikuta, S.; Nakamura, Y.; Hattori, M.; Sato, R.; Kikawada, T.; Noda, H. Herbivory-induced glucose transporter gene expression in the brown planthopper, Nilaparvata lugens. Insect Biochem. Mol. Biol. 2015, 64, 60–67. [Google Scholar] [CrossRef]
- Arad, N.; Paredes-Montero, J.R.; Mondal, M.H.; Ponvert, N.; Brown, J.K. RNA interference-mediated knockdown of genes involved in sugar transport and metabolism disrupts psyllid Bactericera cockerelli (Order: Hemiptera) gut physiology and results in high mortality. Front Insect. Sci. 2023, 3, 1283334. [Google Scholar] [CrossRef]
- Dixit, R.; Rawat, M.; Kumar, S.; Pandey, K.C.; Adak, T.; Sharma, A. Salivary gland transcriptome analysis in response to sugar feeding in malaria vector Anopheles stephensi. J. Insect Physiol. 2011, 57, 1399–1406. [Google Scholar] [CrossRef] [PubMed]
- Rashid, A.; Ruan, H.; Wang, Y. The Gene FvTST1 From Strawberry Modulates Endogenous Sugars Enhancing Plant Growth and Fruit Ripening. Front. Plant Sci. 2021, 12, 774582. [Google Scholar] [CrossRef]
- Zhou, Y.; Li, K.; Wen, S.; Yang, D.; Gao, J.; Wang, Z.; Zhu, P.; Bie, Z.; Cheng, J. Phloem unloading in cultivated melon fruits follows an apoplasmic pathway during enlargement and ripening. Hortic. Res. 2023, 10, uhad123. [Google Scholar] [CrossRef] [PubMed]
- Varela, M.F.; Stephen, J.; Bharti, D.; Lekshmi, M.; Kumar, S. Inhibition of Multidrug Efflux Pumps Belonging to the Major Facilitator Superfamily in Bacterial Pathogens. Biomedicines 2023, 11, 1448. [Google Scholar] [CrossRef] [PubMed]
- Palli, S.R. RNAi turns 25:contributions and challenges in insect science. Front. Insect Sci. 2023, 3, 1209478. [Google Scholar] [CrossRef]
- Wang, B.; Mao, Z.; Chen, Y.; Ying, J.; Wang, H.; Sun, Z.; Li, J.; Zhang, C.; Zhuo, J. Identification and Functional Analysis of the fruitless Gene in a Hemimetabolous Insect, Nilaparvata lugens. Insects 2024, 15, 262. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, D.; Yao, Q.; Zhang, J.; Dong, X.; Tian, H.; Chen, J.; Zhang, W. Feeding-based RNA interference of a trehalose phosphate synthase gene in the brown planthopper, Nilaparvata lugens. Insect Mol. Biol. 2010, 19, 777–786. [Google Scholar] [CrossRef] [PubMed]
- Zha, W.; Peng, X.; Chen, R.; Du, B.; Zhu, L.; He, G. Knockdown of midgut genes by dsRNA-transgenic plant-mediated RNA interference in the hemipteran insect Nilaparvata lugens. PLoS ONE 2011, 6, e20504. [Google Scholar] [CrossRef]
- Lyu, Z.; Chen, J.; Lyu, J.; Guo, P.; Liu, J.; Liu, J.; Zhang, W. Spraying double-stranded RNA targets UDP-N-acetylglucosamine pyrophosphorylase in the control of Nilaparvata lugens. Int. J. Biol. Macromol. 2024, 271, 132455. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).