Identification and Functional Analysis of V-ATPaseA and C Genes in Hyphantria cunea
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Insects
2.2. Cloning and Validation of cDNA Seqences of HcV-ATPases
2.3. Bioinformatics Analysis of HcV-ATPases
2.4. Expression Pattern of HcV-ATPases in Different Developmental Stages and Tissues
2.5. Functional Analysis of HcV-ATPases by RNAi
2.6. H&E Staining and Transmission Electron Microscopy of Midgut
2.7. Immunohistochemistry
2.8. Statistical Analysis
3. Results
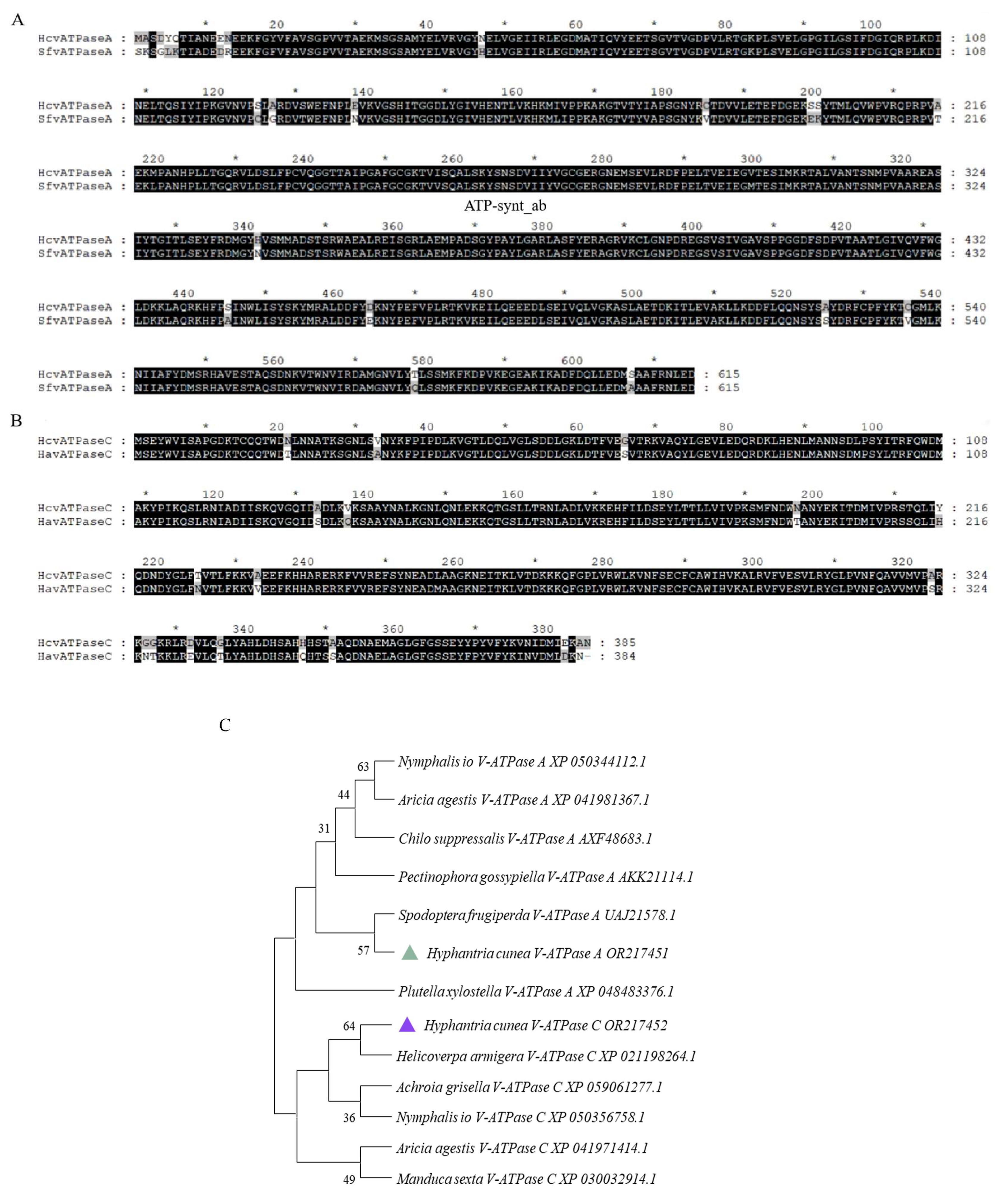
3.1. Identification and Characterization of HcV-ATPase A and C
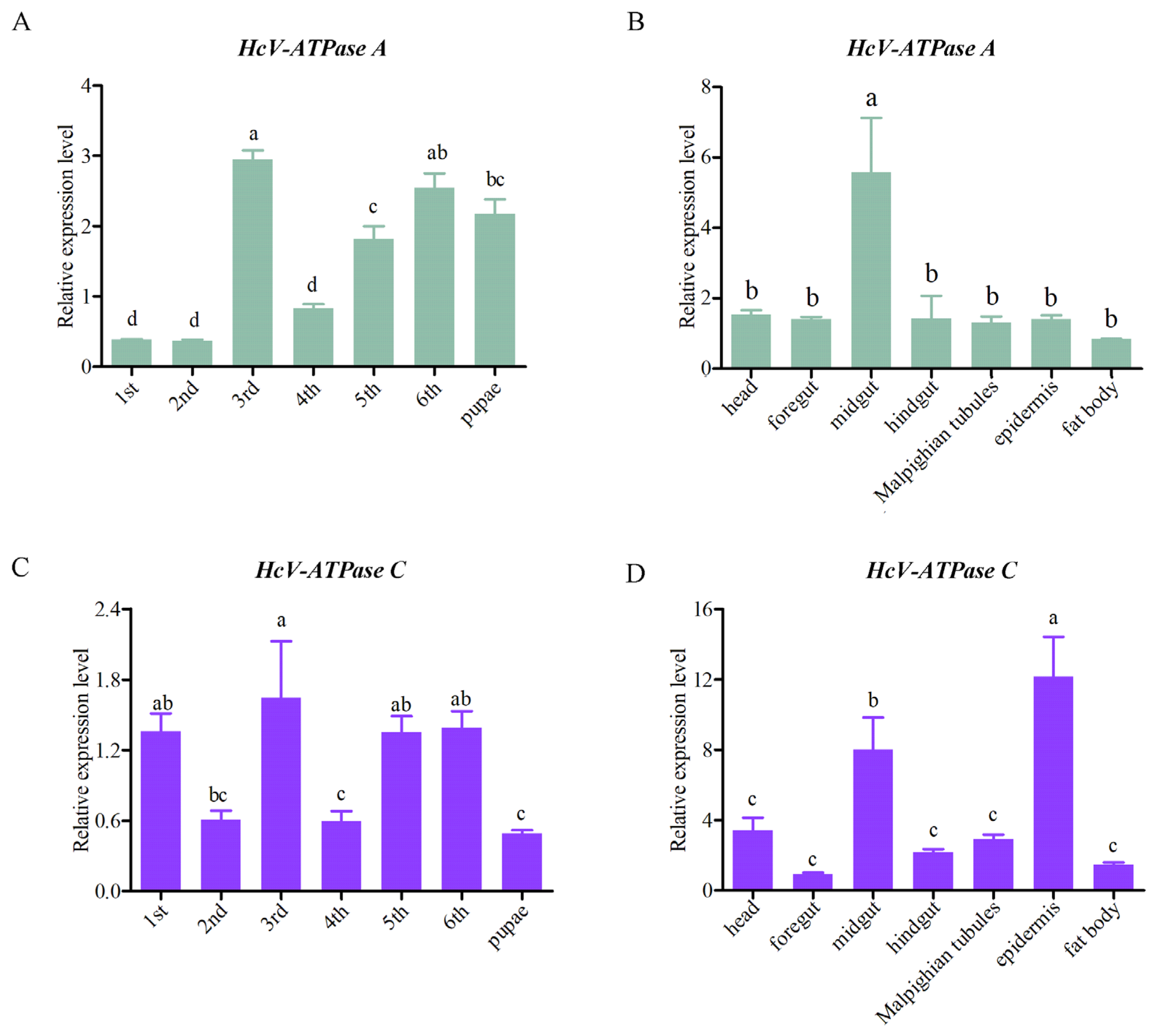
3.2. Expression Patterns of HcV-ATPase A and C
3.3. Effect on H. cunea Survival after HcV-ATPase A and C RNAi
3.4. Immunohistochemistry
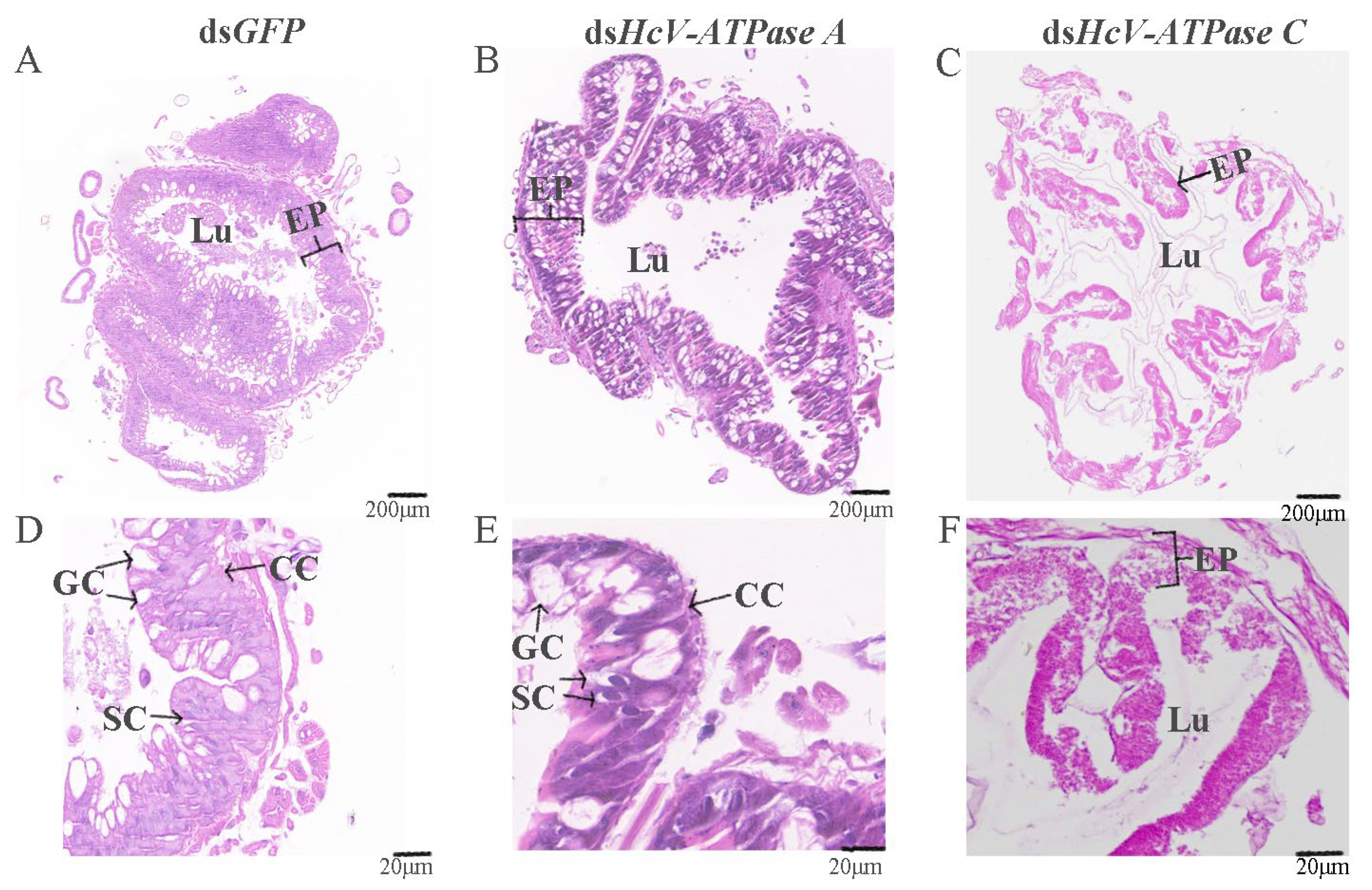
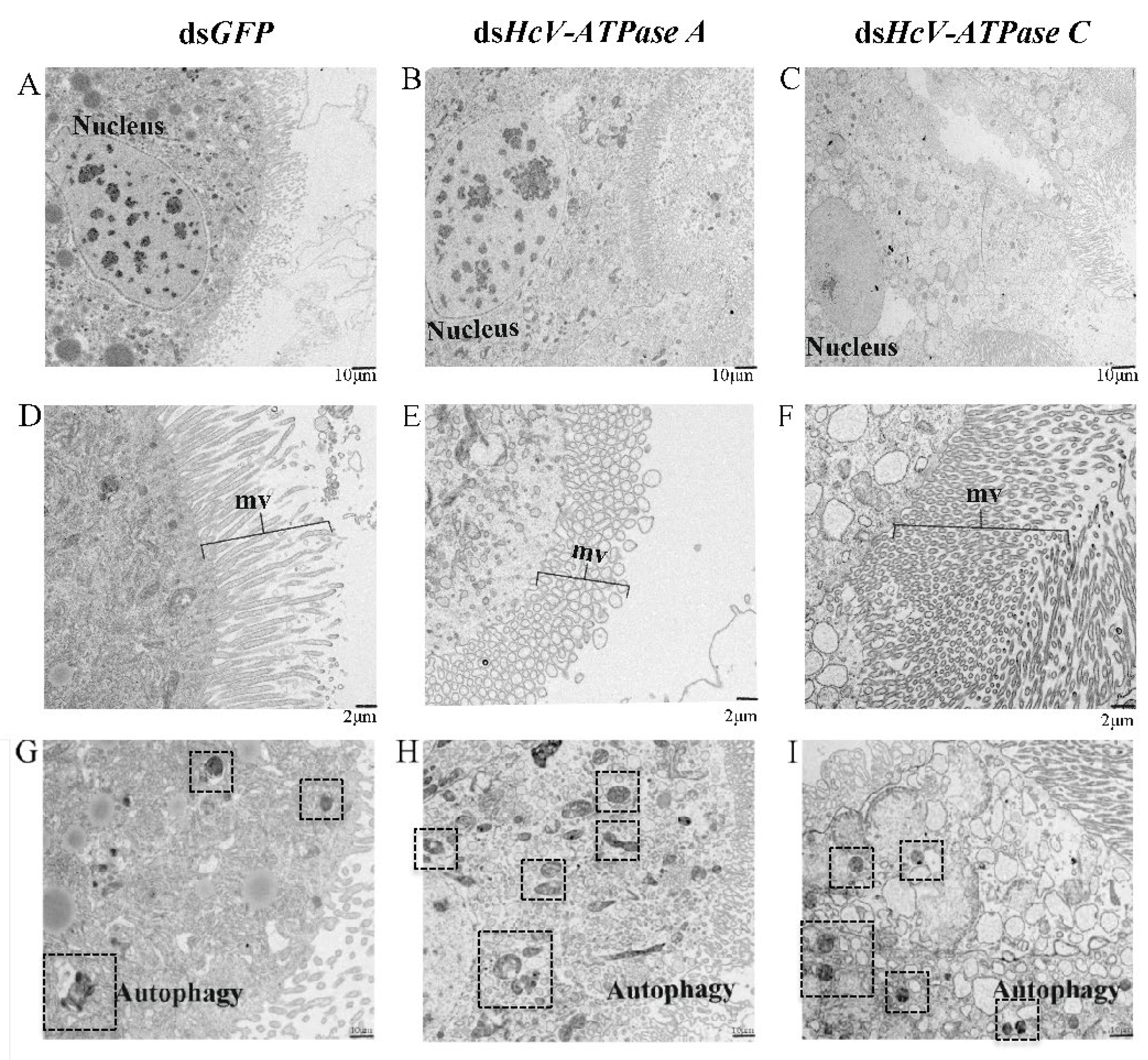
3.5. Effect of RNAi of HcV-ATPase A and C on H. cunea Midgut
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Appendix A
| Primer Name | Sequence (5′-3′) | Application | Product Size (bp) |
|---|---|---|---|
| HcV-ATPase AF | CCGGAATTCATGGCGTCGGATTATCAG | cDNA verification | 1845 |
| HcV-ATPase AR | CCAAGCTTCTTAGTCCTCGAGGTTGCG | ||
| HcV-ATPase CF | CCGGAATTCATGTCTGAATACTGGGTGA | 1158 | |
| HcV-ATPase CR | CGCGTCGACCATTAGTTCGCCTTCTCG | ||
| qHcV-ATPase AF | GTGATGTCAGCTGGGAATT | qRT-PCR | 156 |
| qHcV-ATPase AR | AGAAGGCGCGATGTATGT | ||
| qHcV-ATPase CF | CTGATGACCTTGGTAAGCTTG | 154 | |
| qHcV-ATPase CR | TATTTCGCCATGTCCCAT | ||
| actinF | CTACCTCACGCCATTCTC | 150 | |
| actinR | AGCTTCTCCTTGATGTCAC | ||
| dsHcV-ATPase AF | TAATACGACTCACTATAGGGAGAAGATGCCGGCCAAC | RNAi | 527 |
| dsHcV-ATPase AR | TAATACGACTCACTATAGGCGTCCAGCTCGCTCGTAG | ||
| dsHcV-ATPase CF | TAATACGACTCACTATAGGCCTACAATGCACTCAAGGGC | 502 | |
| dsHcV-ATPase CR | TAATACGACTCACTATAGGGATCCAGGCGCAGAAACAC | ||
| dsGFPF | TAATACGACTCACTATAGGCCACAAGTTCAGCGTGT | 500 | |
| dsGFPR | TAATACGACTCACTATAGGAGTTCACCTTGATGCCG |
References
- Yang, F.; Sendi, J.J.; Johns, R.C.; Takeda, M. Haplotype diversity of mtCOI in the fall webworm Hyphantria cunea (Lepidoptera: Arctiidae) in introduced regions in China, Iran, Japan, Korea, and its homeland, the United States. Appl. Entomol. Zoolog. 2017, 52, 401–406. [Google Scholar] [CrossRef]
- Ning, J.; Lu, P.; Fan, J.; Ren, L.; Zhao, L. American fall webworm in China: A new case of global biological invasions. Innovation 2022, 3, 100201. [Google Scholar] [CrossRef] [PubMed]
- Edosa, T.T.; Jo, Y.H.; Keshavarz, M.; Anh, Y.S.; Noh, M.Y.; Han, Y.S. Current status of the management of fall webworm, Hyphantria cunea: Towards the integrated pest management development. J. Appl. Entomol. 2019, 143, 1–10. [Google Scholar] [CrossRef]
- Zhu, K.Y.; Palli, S.R. Mechanisms, applications, and challenges of insect RNA interference. Annu. Rev. Entomol. 2020, 65, 293–311. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.Q.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Graham, L.A.; Finnigan, G.C.; Kane, P.M. Some assembly required: Contributions of Tom Stevens’ lab to the V-ATPase field. Traffic 2018, 19, 385–390. [Google Scholar] [CrossRef]
- Kakinuma, Y.; Ohsumi, Y.; Anraku, Y. Properties of H+-translocating adenosine triphosphatase in vacuolar membranes of SAccharomyces cerevisiae. J. Biol. Chem. 1981, 256, 10859–10863. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Guo, M.J.; Yang, C.X.; Liu, Z.Q.; Chen, S.M.; Lu, J.; Qiu, B.L.; Zhang, Y.J.; Zhou, X.G.; Pan, H.P. RNA interference-mediated silencing of vATPase subunits A and E affect survival and development of the 28-spotted ladybeetle, Henosepilachna vigintioctopunctata. Insect Sci. 2021, 28, 1664–1676. [Google Scholar] [CrossRef]
- Breton, S.; Brown, D. Regulation of Luminal Acidification by the V-ATPase. Physiology 2013, 28, 318–329. [Google Scholar] [CrossRef]
- Zhang, X.; Fan, Z.; Zhang, R.; Kong, X.; Liu, F.; Fang, J.; Zhang, S.; Zhang, Z. Bacteria-mediated RNAi for managing fall webworm, Hyphantria cunea: Screening target genes and analyzing lethal effect. Pest. Manag. Sci. 2023, 79, 1566–1577. [Google Scholar] [CrossRef]
- Liu, X.J.; Liang, X.Y.; Guo, J.; Shi, X.K.; Merzendorfer, H.; Zhu, K.Y.; Zhang, J.Z. V-ATPase subunit a is required for survival and midgut development of Locusta migratoria. Insect Mol. Biol. 2022, 31, 60–72. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Miyata, K.; Ozawa, S.; Hasegawa, K. Systemic RNAi of V-ATPase subunit B causes molting defect and developmental abnormalities in Periplaneta fuliginosa. Insect Sci. 2019, 26, 721–731. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Xia, Y. Vacuolar ATPase subunit H is essential for the survival and moulting of Locusta migratoria manilensis. Insect Mol. Biol. 2012, 21, 405–413. [Google Scholar] [CrossRef] [PubMed]
- Mao, J.; Zhang, P.; Liu, C.; Zeng, F. Co-silence of the coatomer beta and V-ATPase A genes by siRNA feeding reduces larval survival rate and weight gain of cotton bollworm, Helicoverpa armigera. Pestic. Biochem. Physiol. 2015, 118, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Rahmani, S.; Bandani, A.R. A gene silencing of V-ATPase subunit A interferes with survival and development of the tomato leafminer, Tuta absoluta. Arch. Insect Biochem. Physiol. 2021, 106, 9. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.Z.; Yang, B.; Zhang, A.H.; Ding, D.R.; Wang, G.R. Plant-Mediated RNAi for Controlling Apolygus lucorum. Front. Plant Sci. 2019, 10, 64. [Google Scholar] [CrossRef] [PubMed]
- Vélez, A.M.; Jurzenski, J.; Matz, N.; Zhou, X.G.; Wang, H.C.; Ellis, M.; Siegfried, B.D. Developing an in vivo toxicity assay for RNAi risk assessment in honey bees, Apis mellifera L. Chemosphere 2016, 144, 1083–1090. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Liu, X.; Cooper, A.M.; Silver, K.; Merzendorfer, H.; Zhu, K.Y.; Zhang, J. Vacuolar (H+)-ATPase subunit c is essential for the survival and systemic RNA interference response in Locusta migratoria. Pest. Manag. Sci. 2022, 78, 1555–1566. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Wang, X.; Xie, K.; Li, Y.; Zhao, P. Gene cloning and expression characteristics of vacuolar-type ATPase subunit B in Bombyx mori. Chin. J. Biotechnol. 2016, 32, 487–496. [Google Scholar] [CrossRef]
- Zhao, X.; Gou, X.; Liu, W.; Ma, E.; Moussian, B.; Li, S.; Zhu, K.; Zhang, J. The wing-specific cuticular protein LmACP7 is essential for normal wing morphogenesis in the migratory locust. Insect Biochem. Mol. Biol. 2019, 112, 103206. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Collins, P.J. A new resistance to pyrethroids in Tribolium castaneum (herbst). Pestic. Sci. 2006, 28, 101–115. [Google Scholar] [CrossRef]
- Jagadeesan, R.; Collins, P.J.; Daglish, G.J.; Ebert, P.R.; Schlipalius, D.I. Phosphine Resistance in the Rust Red Flour Beetle, Tribolium castaneum (Coleoptera: Tenebrionidae): Inheritance, Gene Interactions and Fitness Costs. PLoS ONE 2012, 7, e31582. [Google Scholar] [CrossRef]
- Opit, G.P.; Phillips, T.W.; Aikins, M.J.; Hasan, M.M. Phosphine resistance in Tribolium castaneum and Rhyzopertha dominica from stored wheat in Oklahoma. J. Econ. Entomol. 2012, 105, 1107–1114. [Google Scholar] [CrossRef] [PubMed]
- Forgac, M. Vacuolar ATPases: Rotary proton pumps in physiology and pathophysiology. Nat. Rev. Mol. Cell Biol. 2007, 8, 917–929. [Google Scholar] [CrossRef] [PubMed]
- Nishi, T.; Forgac, M. The vacuolar (H+)-ATPases -: Nature’s most versatile proton pumps. Nat. Rev. Mol. Cell Biol. 2002, 3, 94–103. [Google Scholar] [CrossRef]
- Leng, X.H.; Manolson, M.F.; Liu, Q.; Forgac, M. Site-directed mutagenesis of the 100-kDa subunit (Vph1p) of the yeast vacuolar (H+)-ATPase. J. Biol. Chem. 1996, 271, 22487–22493. [Google Scholar] [CrossRef]
- Toyomura, T.; Oka, T.; Yamaguchi, C.; Wada, Y.; Futai, M. Three subunit a isoforms of mouse vacuolar H+-ATPase -: Preferential expression of the a3 isoform during osteoclast differentiation. J. Biol. Chem. 2000, 275, 8760–8765. [Google Scholar] [CrossRef] [PubMed]
- Toei, M.; Saum, R.; Forgac, M. Regulation and isoform function of the V-ATPases. Biochemistry 2010, 49, 4715–4723. [Google Scholar] [CrossRef]
- Cooper, A.M.; Silver, K.; Zhang, J.Z.; Park, Y.; Zhu, K.Y. Molecular mechanisms influencing efficiency of RNA interference in insects. Pest. Manag. Sci. 2019, 75, 18–28. [Google Scholar] [CrossRef]
- Wang, J.; Zhang, X.; Deng, S.; Ma, E.; Zhang, J.; Xing, S. Molecular characterization and RNA interference analysis of the DEAD-box gene family in Locusta migratoria. Gene 2020, 728, 144297. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Zhao, D.; Zhang, Y.; Guo, W.; Wang, W.; Zhao, K.; Gao, Y.; Wang, X. Identification and characterization of chitin deacetylase2 from the American white moth, Hyphantria cunea (Drury). Gene 2018, 670, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Baum, J.A.; Bogaert, T.; Clinton, W.; Heck, G.R.; Feldmann, P.; Ilagan, O.; Johnson, S.; Plaetinck, G.; Munyikwa, T.; Pleau, M.; et al. Control of coleopteran insect pests through RNA interference. Nat. Biotechnol. 2007, 25, 1322–1326. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.X.; Singh, N.D.; Li, L.B.; Zhang, X.L.; Daniell, H. Engineered chloroplast dsRNA silences cytochrome p450 monooxygenase, V-ATPase and chitin synthase genes in the insect gut and disrupts Helicoverpa armigera larval development and pupation. Plant Biotechnol. J. 2015, 13, 435–446. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Cooper, A.M.W.; Zhang, J.; Zhu, K.Y. Biosynthesis, modifications and degradation of chitin in the formation and turnover of peritrophic matrix in insects. J. Insect Physiol. 2019, 114, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Vitavska, O.; Merzendorfer, H.; Wieczorek, H. The V-ATPase subunit C binds to polymeric F-actin as well as to monomeric G-actin and induces cross-linking of actin filaments. J. Biol. Chem. 2005, 280, 1070–1076. [Google Scholar] [CrossRef]
- Hiesinger, P.R.; Fayyazuddin, A.; Mehta, S.Q.; Rosenmund, T.; Schulze, K.L.; Zhai, R.G.; Verstreken, P.; Cao, Y.; Zhou, Y.; Kunz, J.; et al. The V-ATPase Vo subunit a1 is required for a late step in synaptic vesicle exocytosis in Drosophila. Cell 2005, 121, 607–620. [Google Scholar] [CrossRef]
- Wang, D.; Epstein, D.; Khalaf, O.; Srinivasan, S.; Williamson, W.R.; Fayyazuddin, A.; Quiocho, F.A.; Hiesinger, P.R. Ca2+-Calmodulin regulates SNARE assembly and spontaneous neurotransmitter release via v-ATPase subunit Voa1. J. Cell Biol. 2014, 205, 21–31. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, X.; Zhao, D.; Wang, Q.; Liu, Y.; Lu, X.; Guo, W. Identification and Functional Analysis of V-ATPaseA and C Genes in Hyphantria cunea. Insects 2024, 15, 515. https://doi.org/10.3390/insects15070515
Wang X, Zhao D, Wang Q, Liu Y, Lu X, Guo W. Identification and Functional Analysis of V-ATPaseA and C Genes in Hyphantria cunea. Insects. 2024; 15(7):515. https://doi.org/10.3390/insects15070515
Chicago/Turabian StyleWang, Xiaojie, Dan Zhao, Qian Wang, Yanan Liu, Xiujun Lu, and Wei Guo. 2024. "Identification and Functional Analysis of V-ATPaseA and C Genes in Hyphantria cunea" Insects 15, no. 7: 515. https://doi.org/10.3390/insects15070515




