In Silico Analysis of Potential Off-Target Effects of a Next-Generation dsRNA Acaricide for Varroa Mites (Varroa destructor) and Lack of Effect on a Bee-Associated Arthropod
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
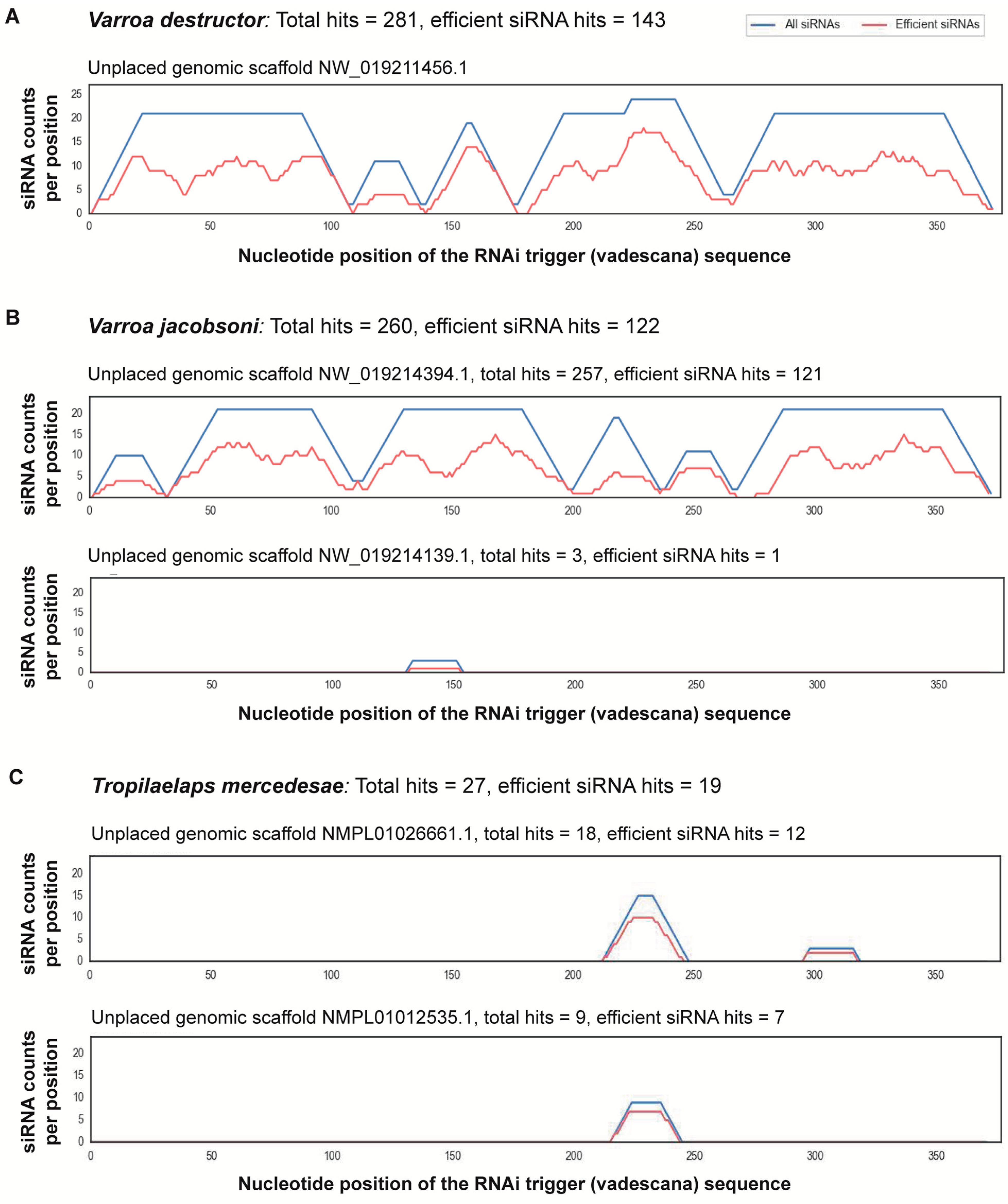
2.1. Predicting Off-Target Effects Using Bioinformatics
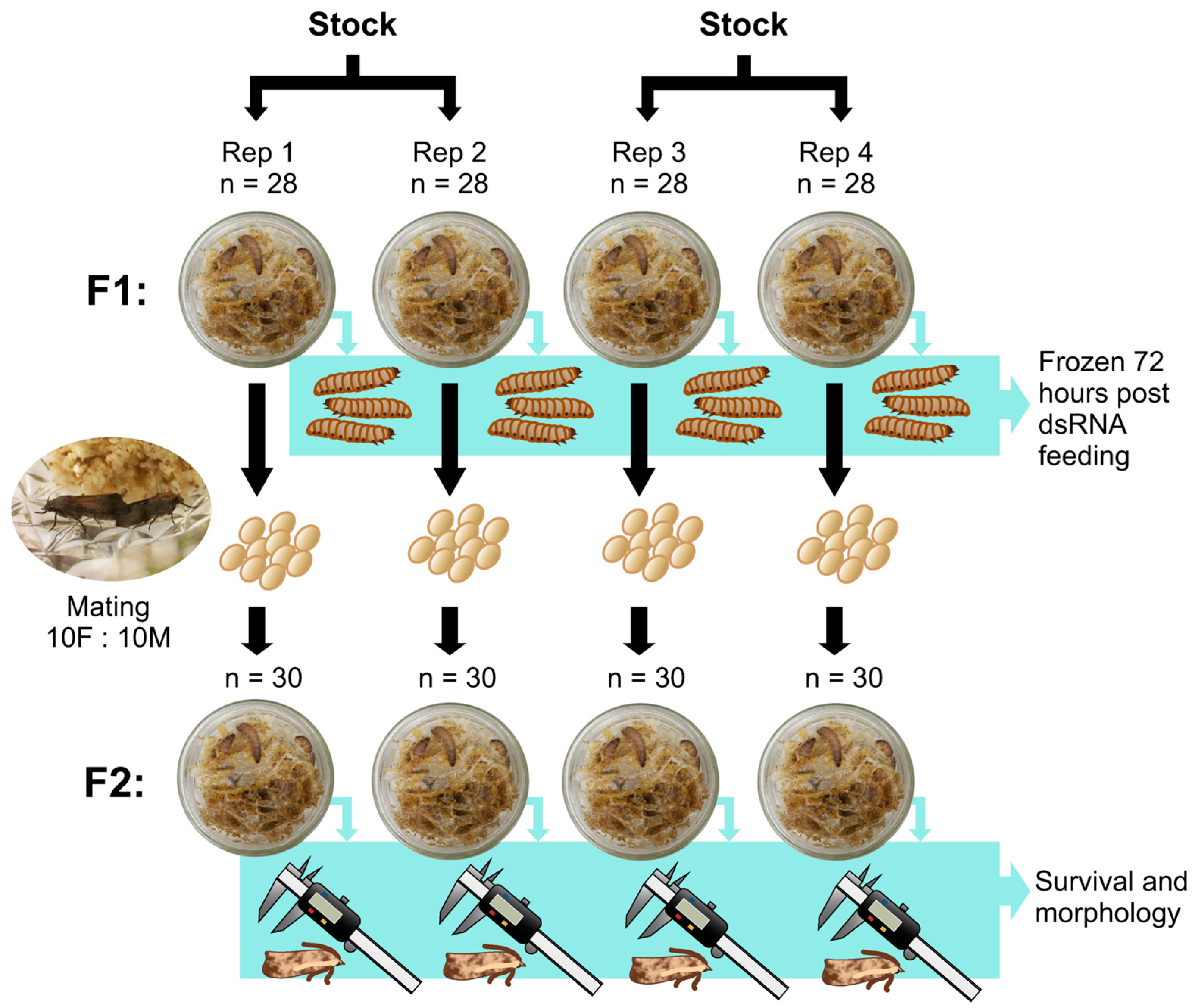
2.2. Insect Husbandry and dsRNA Feeding Treatments
2.2.1. dsRNA Setup/Dilution
2.2.2. Wax Moths
2.2.3. RNA Extraction, Reverse Transcription, and Quantitative PCR
3. Results
3.1. Predicting Off-Target Effects Using Bioinformatics
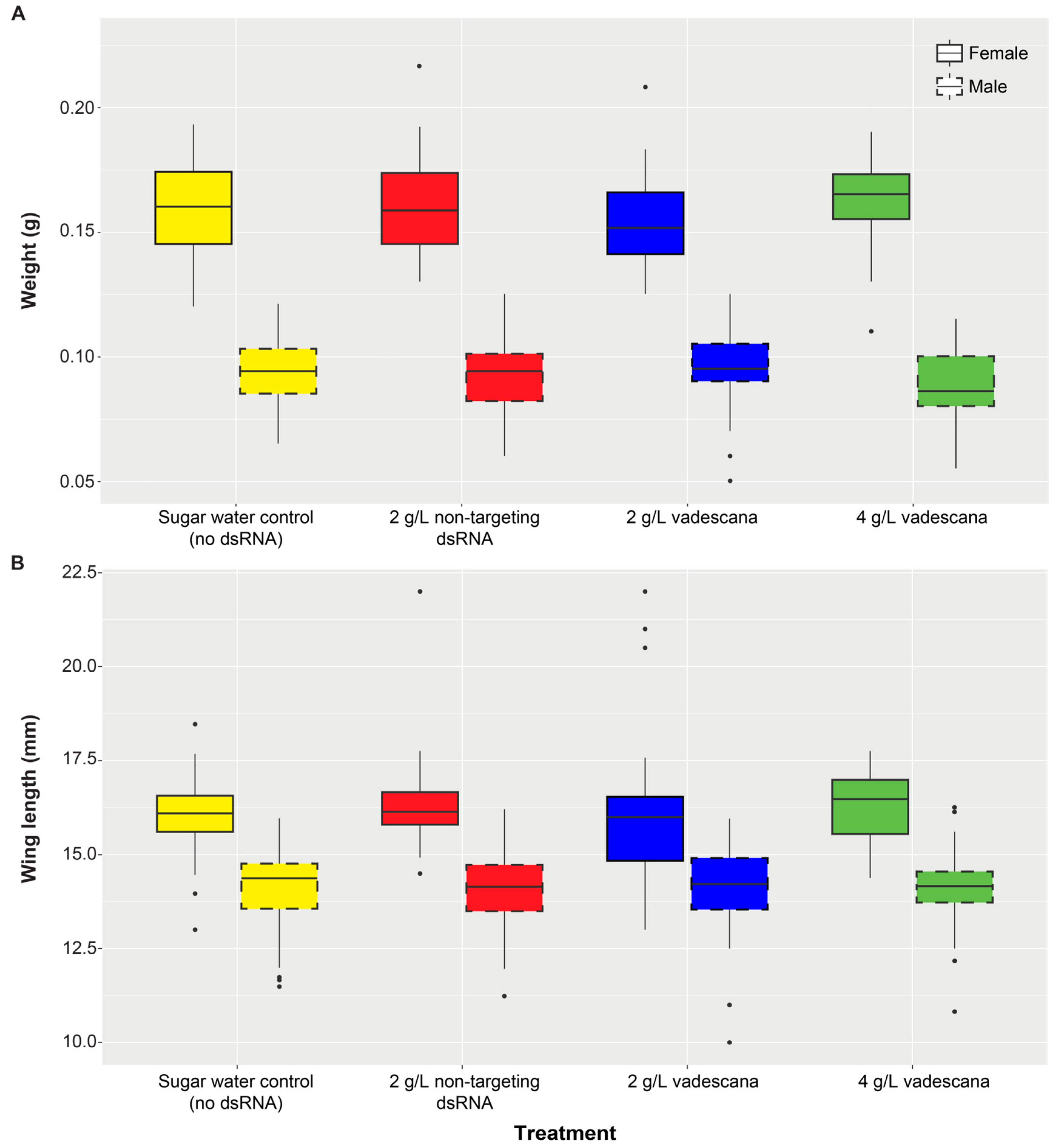
3.2. dsRNA Feeding Treatments
Wax Moth Survival, Morphology, and Calmodulin Expression
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Murphy, J.T.; Breeze, T.D.; Willcox, B.; Kavanagh, S.; Stout, J.C. Globalisation and pollinators: Pollinator declines are an economic threat to global food systems. People Nat. 2022, 4, 773–785. [Google Scholar] [CrossRef]
- Beaurepaire, A.L.; Moro, A.; Mondet, F.; Le Conte, Y.; Neumann, P.; Locke, B. Population genetics of ectoparasitic mites suggest arms race with honeybee hosts. Sci. Rep. 2019, 9, 11355. [Google Scholar] [CrossRef] [PubMed]
- Leonard, S.P.; Powell, J.E.; Perutka, J.; Geng, P.; Heckmann, L.C.; Horak, R.D.; Davies, B.W.; Ellington, A.D.; Barrick, J.E.; Moran, N.A. Engineered symbionts activate honey bee immunity and limit pathogens. Science 2020, 367, 573–576. [Google Scholar] [CrossRef] [PubMed]
- Lester, P.J.; Felden, A.; Baty, J.W.; Bulgarella, M.; Haywood, J.; Mortensen, A.N.; Remnant, E.J.; Smeele, Z.E. Viral communities in the parasite Varroa destructor and in colonies of their honey bee host (Apis mellifera) in New Zealand. Sci. Rep. 2022, 12, 8809. [Google Scholar] [CrossRef] [PubMed]
- Wilfert, L.; Long, G.; Leggett, H.C.; Schmid-Hempel, P.; Butlin, R.; Martin, S.J.M.; Boots, M. Deformed wing virus is a recent global epidemic in honeybees driven by Varroa mites. Science 2016, 351, 594–597. [Google Scholar] [CrossRef]
- Martin, S.J.; Highfield, A.C.; Brettell, L.; Villalobos, E.M.; Budge, G.E.; Powell, M.; Nikaido, S.; Schroeder, D.C. Global honey bee viral landscape altered by a parasitic mite. Science 2012, 336, 1304–1306. [Google Scholar] [CrossRef]
- Traynor, K.S.; Mondet, F.; de Miranda, J.R.; Techer, M.; Kowallik, V.; Oddie, M.A.Y.; Chantawannakul, P.; McAfee, A. Varroa destructor: A complex parasite, crippling honey bees worldwide. Trends Parasitol. 2020, 36, 592–606. [Google Scholar] [CrossRef]
- Rosenkranz, P.; Aumeier, P.; Ziegelmann, B. Biology and control of Varroa destructor. J. Invertebr. Pathol. 2010, 103, S96–S119. [Google Scholar] [CrossRef]
- Bartlett, L.J. Frontiers in effective control of problem parasites in beekeeping. Int. J. Parasitol. Parasites Wildl. 2022, 17, 263–272. [Google Scholar] [CrossRef]
- Jack, C.J.; Ellis, J.D. Integrated pest management control of Varroa destructor (Acari: Varroidae), the most damaging pest of Apis mellifera L. (Hymenoptera: Apidae) colonies. J. Insect Sci. 2021, 21, 6. [Google Scholar] [CrossRef]
- González-Cabrera, J.; Davies, T.G.; Field, L.M.; Kennedy, P.J.; Williamson, M.S. An amino acid substitution (L925V) associated with resistance to pyrethroids in Varroa destructor. PLoS ONE 2013, 8, e82941. [Google Scholar] [CrossRef] [PubMed]
- González-Cabrera, J.; Rodríguez-Vargas, S.; Davies, T.G.; Field, L.M.; Schmehl, D.; Ellis, J.D.; Krieger, K.; Williamson, M.S. Novel mutations in the voltage-gated sodium channel of pyrethroid-resistant Varroa destructor populations from the southeastern USA. PLoS ONE 2016, 11, e0155332. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Rodríguez, C.S.; Marin, O.; Calatayud, F.; Mahiques, M.J.; Mompo, A.; Segura, I.; Simo, E.; Gonzalez-Cabrera, J. Large-scale monitoring of resistance to coumaphos, amitraz, and pyrethroids in Varroa destructor. Insects 2021, 12, 27. [Google Scholar] [CrossRef]
- Hernández-Rodríguez, C.S.; Moreno-Martí, S.; Almecija, G.; Christmon, K.; Johnson, J.D.; Ventelon, M.; vanEngelsdorp, D.; Cook, S.C.; González-Cabrera, J. Resistance to amitraz in the parasitic honey bee mite Varroa destructor is associated with mutations in the β-adrenergic-like octopamine receptor. J. Pest Sci. 2021, 95, 1179–1195. [Google Scholar] [CrossRef]
- McGruddy, R.A.; Bulgarella, M.; Felden, A.; Baty, J.W.; Haywood, J.; Stahlmann-Brown, P.; Lester, P.J. Are increasing honey bee colony losses attributed to Varroa destructor in New Zealand driven by miticide resistance? J. Apic. Res. 2024, 63, 648–659. [Google Scholar] [CrossRef]
- Millán-Leiva, A.; Marín, Ó.; De la Rúa, P.; Muñoz, I.; Tsagkarakou, A.; Eversol, H.; Christmon, K.; vanEngelsdorp, D.; González-Cabrera, J. Mutations associated with pyrethroid resistance in the honey bee parasite Varroa destructor evolved as a series of parallel and sequential events. J. Pest Sci. 2021, 94, 1505–1517. [Google Scholar] [CrossRef]
- Rinkevich, F.D. Detection of amitraz resistance and reduced treatment efficacy in the Varroa mite, Varroa destructor, within commercial beekeeping operations. PLoS ONE 2020, 15, e0227264. [Google Scholar] [CrossRef]
- Gunes, N.; Aydın, L.; Belenli, D.; Hranitz, J.M.; Mengilig, S.; Selova, S. Stress responses of honey bees to organic acid and essential oil treatments against varroa mites. J. Apic. Res. 2017, 56, 175–181. [Google Scholar] [CrossRef]
- Ostermann, D.J.; Currie, R.W. Effect of formic acid formulations on honey bee (Hymenoptera: Apidae) colonies and influence of colony and ambient conditions on formic acid concentration in the hive. J. Econ. Entomol. 2004, 97, 1500–1508. [Google Scholar] [CrossRef]
- Rademacher, E.; Harz, M.; Schneider, S. Effects of oxalic acid on Apis mellifera (Hymenoptera: Apidae). Insects 2017, 8, 84. [Google Scholar] [CrossRef]
- Garbian, Y.; Maori, E.; Kalev, H.; Shafir, S.; Sela, I. Bidirectional transfer of RNAi between honey bee and Varroa destructor: Varroa gene silencing reduces Varroa population. PLoS Pathog. 2012, 8, e1003035. [Google Scholar] [CrossRef] [PubMed]
- Lim, Z.X.; Robinson, K.E.; Jain, R.G.; Chandra, G.S.; Asokan, R.; Asgari, S.; Mitter, N. Diet-delivered RNAi in Helicoverpa armigera—Progresses and challenges. J. Insect Physiol. 2016, 85, 86–93. [Google Scholar] [CrossRef] [PubMed]
- Lopez, S.B.G.; Guimarães-Ribeiro, V.; Rodriguez, J.V.G.; Dorand, F.A.P.S.; Salles, T.S.; sá-Guimarães, T.E.; Alvarenga, E.S.L.; Melo, A.C.A.; Almeida, R.V.; Moreira, M.F. RNAi-based bioinsecticide for Aedes mosquito control. Sci. Rep. 2019, 9, 4038. [Google Scholar] [CrossRef]
- Pallis, S.; Alyokhin, A.; Manley, B.; Rodrigues, T.; Barnes, E.; Narva, K. Effects of low doses of a novel dsRNA-based biopesticide (Calantha) on the Colorado potato beetle. J. Econ. Entomol. 2023, 116, 456–461. [Google Scholar] [CrossRef]
- Turner, C.T.; Davy, M.W.; MacDiarmid, R.M.; Plummer, K.M.; Birch, N.P.; Newcomb, R.D. RNA interference in the light brown apple moth, Epiphyas postvittana (Walker) induced by double-stranded RNA feeding. Insect Mol. Biol. 2006, 15, 383–391. [Google Scholar] [CrossRef]
- Siomi, H.; Siomi, M.C. On the road to reading the RNA-interference code. Nature 2009, 457, 396–404. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, N.; Hall, M.J.; Hellmich, R.L.; Coats, J.R.; Bradbury, S.P. Evaluating toxicity of Varroa mite (Varroa destructor)-active dsRNA to monarch butterfly (Danaus plexippus) larvae. PLoS ONE 2021, 16, e0251884. [Google Scholar] [CrossRef]
- Wenninger, E.J.; DeGrey, S.P.; Insinga, J.; Knopf, E.; Alyokhin, A.; Barnes, E.R.; Bradford, B.; Groves, R.L.; Manley, B.; Piaskowski, J. Responses of non-target arthropods to the dsRNA bioinsecticide Calantha™ and conventional insecticides targeting Colorado potato beetle, Leptinotarsa decemlineata (Say). Am. J. Potato Res. 2025. [Google Scholar] [CrossRef]
- Inberg, A.; Kapoor, M. Compositions and Methods for Controlling Arthropod Parasite and Pest. Infestations. Patent number US10907152B2, 2 February 2021. [Google Scholar]
- Masucci, J.D. Developing double-stranded RNA as a new Varroa control product. Am. Bee J. 2020, 160, 685–690. [Google Scholar]
- Wang, W.; Wan, P.; Lai, F.; Zhu, T.; Fu, Q. Double-stranded RNA targeting calmodulin reveals a potential target for pest management of Nilaparvata lugens. Pest Manag. Sci. 2018, 74, 1711–1719. [Google Scholar] [CrossRef]
- Chin, D.; Means, A.R. Calmodulin: A prototypical calcium sensor. Trends Cell Biol. 2000, 10, 322–328. [Google Scholar] [CrossRef] [PubMed]
- Klee, C.B.; Crouch, T.H.; Richman, P.G. Calmodulin. Annu. Rev. Biochem. 1980, 49, 489–515. [Google Scholar] [CrossRef]
- Kwadha, C.A.; Ong’amo, G.O.; Ndegwa, P.N.; Raina, S.K.; Fombong, A.T. The biology and control of the greater wax moth, Galleria mellonella. Insects 2017, 8, 61. [Google Scholar] [CrossRef]
- Dutta, T.K.; Mandal, A.; Kundu, A.; Phani, V.; Mathur, C.; Veeresh, A.; Sreevathsa, R. RNAi-mediated knockdown of gut receptor-like genes prohibitin and alpha-amylase altered the susceptibility of Galleria mellonella to Cry1AcF toxin. BMC Genom. 2022, 23, 601. [Google Scholar] [CrossRef] [PubMed]
- Grizanova, E.V.; Coates, C.J.; Butt, T.M.; Dubovskiy, I.M. RNAi-mediated suppression of insect metalloprotease inhibitor (IMPI) enhances Galleria mellonella susceptibility to fungal infection. Dev. Comp. Immunol. 2021, 122, 104126. [Google Scholar] [CrossRef]
- Lück, S.; Kreszies, T.; Strickert, M.; Schweizer, P.; Kuhlmann, M.; Douchkov, D. siRNA-Finder (si-Fi) software for RNAi-target design and off-target prediction. Front. Plant Sci. 2019, 10, 1023. [Google Scholar] [CrossRef] [PubMed]
- Azlan, A.; Dzaki, N.; Azzam, G. Argonaute: The executor of small RNA function. J. Genet. Genom. 2016, 43, 481–494. [Google Scholar] [CrossRef]
- Schwarz, D.S.; Hutvágner, G.; Du, T.; Xu, Z.; Aronin, N.; Zamore, P.D. Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003, 115, 199–208. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Mondal, M.; Peter, J.; Scarbrough, O.; Flynt, A. Environmental RNAi pathways in the two-spotted spider mite. BMC Genom. 2021, 22, 42. [Google Scholar] [CrossRef] [PubMed]
- Santos, D.; Mingels, L.; Vogel, E.; Wang, L.; Christiaens, O.; Cappelle, K.; Wynant, N.; Gansemans, Y.; Van Nieuwerburgh, F.; Smagghe, G.; et al. Generation of virus- and dsRNA-derived siRNAs with species-dependent length in insects. Viruses 2019, 11, 738. [Google Scholar] [CrossRef]
- Rodrigues, T.B.; Mishra, S.K.; Sridharan, K.; Barnes, E.R.; Alyokhin, A.; Tuttle, R.; Kokulapalan, W.; Garby, D.; Skizim, N.J.; Tang, Y.W.; et al. First sprayable double-stranded RNA-based biopesticide product targets proteasome subunit beta type-5 in Colorado potato beetle (Leptinotarsa decemlineata). Front. Plant Sci. 2021, 12, 728652. [Google Scholar] [CrossRef] [PubMed]
- Chejanovsky, N.; Ophir, R.; Schwager, M.S.; Slabezki, Y.; Grossman, S.; Cox-Foster, D. Characterization of viral siRNA populations in honey bee colony collapse disorder. Virology 2014, 454–455, 176–183. [Google Scholar] [CrossRef]
- Mondal, M.; Brown, J.K.; Flynt, A. Exploiting somatic piRNAs in Bemisia tabaci enables novel gene silencing through RNA feeding. Life Sci. Alliance 2020, 3, e202000731. [Google Scholar] [CrossRef] [PubMed]
- Sabin, L.R.; Zheng, Q.; Thekkat, P.; Yang, J.; Hannon, G.J.; Gregory, B.D.; Tudor, M.; Cherry, S. Dicer-2 processes diverse viral RNA species. PLoS ONE 2013, 8, e55458. [Google Scholar] [CrossRef]
- Pettis, J.S.; Chen, Y.P.; Ellis, J.; Evans, J.D.; Rennich, K.D.; Vanengelsdorp, D.; Shimanuki, H. Diseases and Pests of Honey Bees. In The Hive and the Honey Bee; Graham, J.M., Ed.; Dadant & Sons, Inc.: Hamilton, IL, USA, 2015; pp. 833–879. [Google Scholar]
- Simão, F.A.; Waterhouse, R.M.; Ioannidis, P.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 2015, 31, 3210–3212. [Google Scholar] [CrossRef] [PubMed]
- Zdobnov, E.M.; Tegenfeldt, F.; Kuznetsov, D.; Waterhouse, R.M.; Simao, F.A.; Ioannidis, P.; Seppey, M.; Loetscher, A.; Kriventseva, E.V. OrthoDB v9.1: Cataloging evolutionary and functional annotations for animal, fungal, plant, archaeal, bacterial and viral orthologs. Nucleic Acids Res. 2017, 45, D744–D749. [Google Scholar] [CrossRef]
- Jorjão, A.L.; Oliveira, L.D.; Scorzoni, L.; Figueiredo-Godoi, L.M.A.; Cristina, A.P.M.; Jorge, A.O.C.; Junqueira, J.C. From moths to caterpillars: Ideal conditions for Galleria mellonella rearing for in vivo microbiological studies. Virulence 2018, 9, 383–389. [Google Scholar] [CrossRef]
- Ellis, J.D.; Graham, J.R.; Mortensen, A. Standard methods for wax moth research. J. Apic. Res. 2015, 52, 1–17. [Google Scholar] [CrossRef]
- Firacative, C.; Khan, A.; Duan, S.; Ferreira-Paim, K.; Leemon, D.; Meyer, W. Rearing and maintenance of Galleria mellonella and its application to study fungal virulence. J. Fungi 2020, 6, 130. [Google Scholar] [CrossRef]
- Pinheiro, J.; Bates, D. nlme: Linear and Nonlinear Mixed Effects Models; R Package Version 3.1-162; R Core Team: Vienna, Austria, 2023; Available online: https://CRAN.R-project.org/package=nlme (accessed on 1 December 2024).
- Hollowell, H.; Rieske, L.K. Southern pine beetle-specific RNA interference exhibits no effect on model nontarget insects. J. Pest Sci. 2022, 95, 1429–1441. [Google Scholar] [CrossRef]
- Taning, C.N.T.; Gui, S.; De Schutter, K.; Jahani, M.; Castellanos, N.L.; Christiaens, O.; Smagghe, G. A sequence complementarity-based approach for evaluating off-target transcript knockdown in Bombus terrestris, following ingestion of pest-specific dsRNA. J. Pest Sci. 2021, 94, 487–503. [Google Scholar] [CrossRef]
- Dubovskiy, I.M.; Grizanova, E.V.; Whitten, M.M.; Mukherjee, K.; Greig, C.; Alikina, T.; Kabilov, M.; Vilcinskas, A.; Glupov, V.V.; Butt, T.M. Immuno-physiological adaptations confer wax moth Galleria mellonella resistance to Bacillus thuringiensis. Virulence 2016, 7, 860–870. [Google Scholar] [CrossRef] [PubMed]
- Bulgarella, M.; Baty, J.W.; McGruddy, R.; Lester, P.J. Gene silencing for invasive paper wasp management: Synthesized dsRNA can modify gene expression but did not affect mortality. PLoS ONE 2023, 18, e0279983. [Google Scholar] [CrossRef]
- Morse, D.L.; Carroll, D.; Weberg, L.; Borgstrom, M.C.; Ranger-Moore, J.; Gillies, R.J. Determining suitable internal standards for mRNA quantification of increasing cancer progression in human breast cells by real-time reverse transcriptase polymerase chain reaction. Anal. Biochem. 2005, 342, 69–77. [Google Scholar] [CrossRef]
- Quinn, O.; Gruber, M.A.M.; Brown, R.L.; Baty, J.W.; Bulgarella, M.; Lester, P.J. A metatranscriptomic analysis of diseased social wasps (Vespula vulgaris) for pathogens, with an experimental infection of larvae and nests. PLoS ONE 2018, 13, e0209589. [Google Scholar] [CrossRef] [PubMed]
- Weiner, S.A.; Geffre, A.G.; Toth, A.L. Functional genomics in the wild: A case study with paper wasps shows challenges and prospects for RNA interference in ecological systems. Genome 2017, 61, 266–272. [Google Scholar] [CrossRef]
- Sammataro, D.; Gerson, U.; Needham, G. Parasitic mites of honey bees: Life history, implications, and impact. Annu. Rev. Entomol. 2000, 45, 519–548. [Google Scholar] [CrossRef]
- Chantawannakul, P.; Ramsey, S.; vanEngelsdorp, D.; Khongphinitbunjong, K.; Phokasem, P. Tropilaelaps mite: An emerging threat to European honey bee. Curr. Opin. Insect Sci. 2018, 26, 69–75. [Google Scholar] [CrossRef]
- Roberts, J.M.K.; Schouten, C.N.; Sengere, R.W.; Jave, J.; Lloyd, D. Effectiveness of control strategies for Varroa jacobsoni and Tropilaelaps mercedesae in Papua New Guinea. Exp. Appl. Acarol. 2020, 80, 399–407. [Google Scholar] [CrossRef] [PubMed]
- Whyard, S.; Singh, A.D.; Wong, S. Ingested double-stranded RNAs can act as species-specific insecticides. Insect Biochem. Mol. Biol. 2009, 39, 824–832. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Peng, Y.; Zhang, H.; Wang, K.; Zhao, C.; Zhu, G.; Reddy Palli, S.; Han, Z. Off-target effects of RNAi correlate with the mismatch rate between dsRNA and non-target mRNA. RNA Biol. 2021, 18, 1747–1759. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, S.J.; Reeves, P.T.; Hoang, B.T.; Mitter, N. A perspective on RNAi-based biopesticides. Front. Plant Sci. 2020, 11, 51. [Google Scholar] [CrossRef]
- Morse, R.A.; Gary, N.E. Insect invaders of the honeybee colony. Bee World 1961, 42, 179–181. [Google Scholar] [CrossRef]
- Strange, J.P.; Baur, A.; Koch, J.B. A scientific note on Bombus (Psithyrus) insularis invasions of bumblebee nests and honeybee hives in the western USA. Apidologie 2014, 45, 554–556. [Google Scholar] [CrossRef]
- McGruddy, R.A.; Smeele, Z.E.; Manley, B.; Masucci, J.D.; Haywood, J.; Lester, P.J. RNA interference as a next-generation control method for suppressing Varroa destructor reproduction in honey bee (Apis mellifera) hives. Pest Manag. Sci. 2024, 80, 4770–4778. [Google Scholar] [CrossRef]
- Merk, J.; Anastasi, M.; McGruddy, R.A.; Manley, B.; Felden, A.; Lester, P.J. Longevity and Foraging Performance of Honey Bees Treated with an RNAi-based Varroa destructor Biopesticide; Victoria University of Wellington: Wellington, New Zealand, 2025. [Google Scholar]

| Species | Common Name | GenBank or RefSeqGenome/ Transcriptome ID | 21-Mer Hits | |
|---|---|---|---|---|
| Genome (Total/ Efficient) | Transcriptome (Unique) | |||
| Arachnida | ||||
| Mesostigmata | ||||
| Stratiolaelaps scimitus | Stratiolaelaps mite | GCA_019614645.1 | 9/6 | - |
| Tropilaelaps mercedesae | Tropilaelaps mite | GCA_002081605.1 | 27/19 | 18 |
| Varroa destructor | Honey bee mite | GCF_002443255.1 | 281/143 | 278 |
| Varroa jacobsoni | Varroa mite | GCF_002532875.1/GCF_002532875.2 | 260/122 | 257 |
| Trombidiformes | ||||
| Acarapis woodi | Honey bee tracheal mite | GCA_023170135.1 | 0 | - |
| Tetranychus urticae | Two-spotted spider mite | GCF_000239435.1 | 1/1 | 1 |
| Insecta | ||||
| Blattodea | ||||
| Blattella germanica | German cockroach | GCA_000762945.2 | 0 | - |
| Coleoptera | ||||
| Aethina tumida | Small hive beetle | GCF_024364675.1 | 0 | 0 |
| Dermaptera | ||||
| Forficula auricularia | European earwig | GCA_024734495.1 | 1/1 | - |
| Diptera | ||||
| Drosophila melanogaster | Fruit fly | GCF_000001215.4 | 10/5 | 10 |
| Hermetia illucens | Black soldier fly | GCF_905115235.1 | 0 | 0 |
| Hymenoptera | ||||
| Apis cerana | Asiatic honey bee | GCF_001442555.1 | 0 | 0 |
| Apis dorsata | Giant honey bee | GCF_000469605.1 | 0 | 0 |
| Apis florea | Little honey bee | GCF_000184785.3 | 0 | 0 |
| Apis mellifera | Honey bee | GCF_003254395.2 | 0 | 0 |
| Bombus ignitus | Fiery-tailed bumble bee | GCA_014825875.1 | 0 | - |
| Bombus impatiens | Common eastern bumble bee | GCF_000188095.3 | 0 | 0 |
| Bombus lapidarius | Red tailed bumble bee | GCA_936014575.1 | 0 | - |
| Bombus pascuorum | Common carder bee | GCF_905332965.1 | 0 | 0 |
| Bombus sylvicola | Forest bumble bee | GCA_019677175.1 | 0 | - |
| Bombus terrestris | Buff-tailed bumble bee | GCF_910591885.1 | 0 | 0 |
| Linepithema humile | Argentine ant | GCF_000217595.1 | 0 | 0 |
| Monomorium pharaonis | Pharaoh ant | GCF_013373865.1 | 0 | 0 |
| Polistes canadensis | Red paper wasp | GCF_001313835.1 | 0 | 0 |
| Polistes dominula | European paper wasp | GCF_001465965.1 | 0 | 0 |
| Polistes fuscatus | Common paper wasp | GCF_010416935.1 | 0 | 0 |
| Solenopsis invicta | Red fire ant | GCF_016802725.1 | 0 | 0 |
| Vespa velutina | Yellow-legged hornet | GCF_912470025.1 | 0 | 0 |
| Vespula germanica | German wasp | GCA_905340365.1 | 0 | - |
| Vespula vulgaris | Common wasp | GCA_905475345.1 | 0 | 0 |
| Lepidoptera | ||||
| Achroia grisella | Lesser wax moth | GCF_030625045.1 | 0 | 0 |
| Danaus plexippus | Monarch butterfly | GCF_009731565.1 | 1/1 | 1 |
| Galleria mellonella | Greater wax moth | GCF_026898425.1 | 0 | 0 |
| Pieris rapae | Cabbage white butterfly | GCF_905147795.1 | 0 | 0 |
| Vanessa cardiu | Painted lady | GCF_905220365.1 | 0 | 0 |
| Orthoptera | ||||
| Acheta domestica | House cricket | GCA_014858955.1 | 0 | - |
| Locusta migratoria | Migratory locust | GCA_026315105.1 | 0 | - |
| Teleogryllus occipitalis | Asian cricket | GCA_011170035.1 | 0 | 0 |
| Phasmatodea | ||||
| Clitarchus hookeri | Smooth stick insect | GCA_002778355.1 | 0 | - |
| Treatment | Replicate | Starting Number of F1 Larvae/Number of Larvae That Pupated | Number of F1 Adults | Starting Number of F2 Larvae/Number of Larvae That Pupated | Number of F2 Adults |
|---|---|---|---|---|---|
| Control, no dsRNA | 1 | 25/17 | 17 | 30/30 | 29 |
| 2 | 25/17 | 16 | 30/29 | 27 | |
| 3 | 25/25 | 25 | 30/30 | 30 | |
| 4 | 25/25 | 21 | 30/30 | 30 | |
| 2 g/L non-targeting | 1 | 25/19 | 17 | 30/26 | 24 |
| dsRNA | 2 | 25/21 | 19 | 30/23 | 22 |
| 3 | 25/25 | 24 | 30/30 | 30 | |
| 4 | 25/21 | 17 | 30/30 | 28 | |
| 2 g/L of vadescana | 1 | 25/19 | 17 | 30/30 | 30 |
| 2 | 25/17 | 17 | 30/30 | 30 | |
| 3 | 25/24 | 23 | 30/30 | 30 | |
| 4 | 25/24 | 24 | 30/30 | 30 | |
| 4 g/L of vadescana | 1 | 25/19 | 17 | 30/22 | 22 |
| 2 | 25/21 | 20 | 30/29 | 28 | |
| 3 | 25/22 | 21 | 30/30 | 30 | |
| 4 | 25/20 | 19 | 30/30 | 30 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bulgarella, M.; Reason, A.; Baty, J.W.; McGruddy, R.A.; Gordon, E.R.L.; Devisetty, U.K.; Lester, P.J. In Silico Analysis of Potential Off-Target Effects of a Next-Generation dsRNA Acaricide for Varroa Mites (Varroa destructor) and Lack of Effect on a Bee-Associated Arthropod. Insects 2025, 16, 317. https://doi.org/10.3390/insects16030317
Bulgarella M, Reason A, Baty JW, McGruddy RA, Gordon ERL, Devisetty UK, Lester PJ. In Silico Analysis of Potential Off-Target Effects of a Next-Generation dsRNA Acaricide for Varroa Mites (Varroa destructor) and Lack of Effect on a Bee-Associated Arthropod. Insects. 2025; 16(3):317. https://doi.org/10.3390/insects16030317
Chicago/Turabian StyleBulgarella, Mariana, Aiden Reason, James W. Baty, Rose A. McGruddy, Eric R. L. Gordon, Upendra K. Devisetty, and Philip J. Lester. 2025. "In Silico Analysis of Potential Off-Target Effects of a Next-Generation dsRNA Acaricide for Varroa Mites (Varroa destructor) and Lack of Effect on a Bee-Associated Arthropod" Insects 16, no. 3: 317. https://doi.org/10.3390/insects16030317
APA StyleBulgarella, M., Reason, A., Baty, J. W., McGruddy, R. A., Gordon, E. R. L., Devisetty, U. K., & Lester, P. J. (2025). In Silico Analysis of Potential Off-Target Effects of a Next-Generation dsRNA Acaricide for Varroa Mites (Varroa destructor) and Lack of Effect on a Bee-Associated Arthropod. Insects, 16(3), 317. https://doi.org/10.3390/insects16030317







