Fitness Cost of Antiretroviral Drug Resistance Mutations on the pol Gene during Analytical Antiretroviral Treatment Interruption among Individuals Experiencing Virological Failure
Abstract
1. Introduction
1.1. Origin of Study
1.2. Objectives
2. Results
2.1. Patients
2.2. Reversion Categories
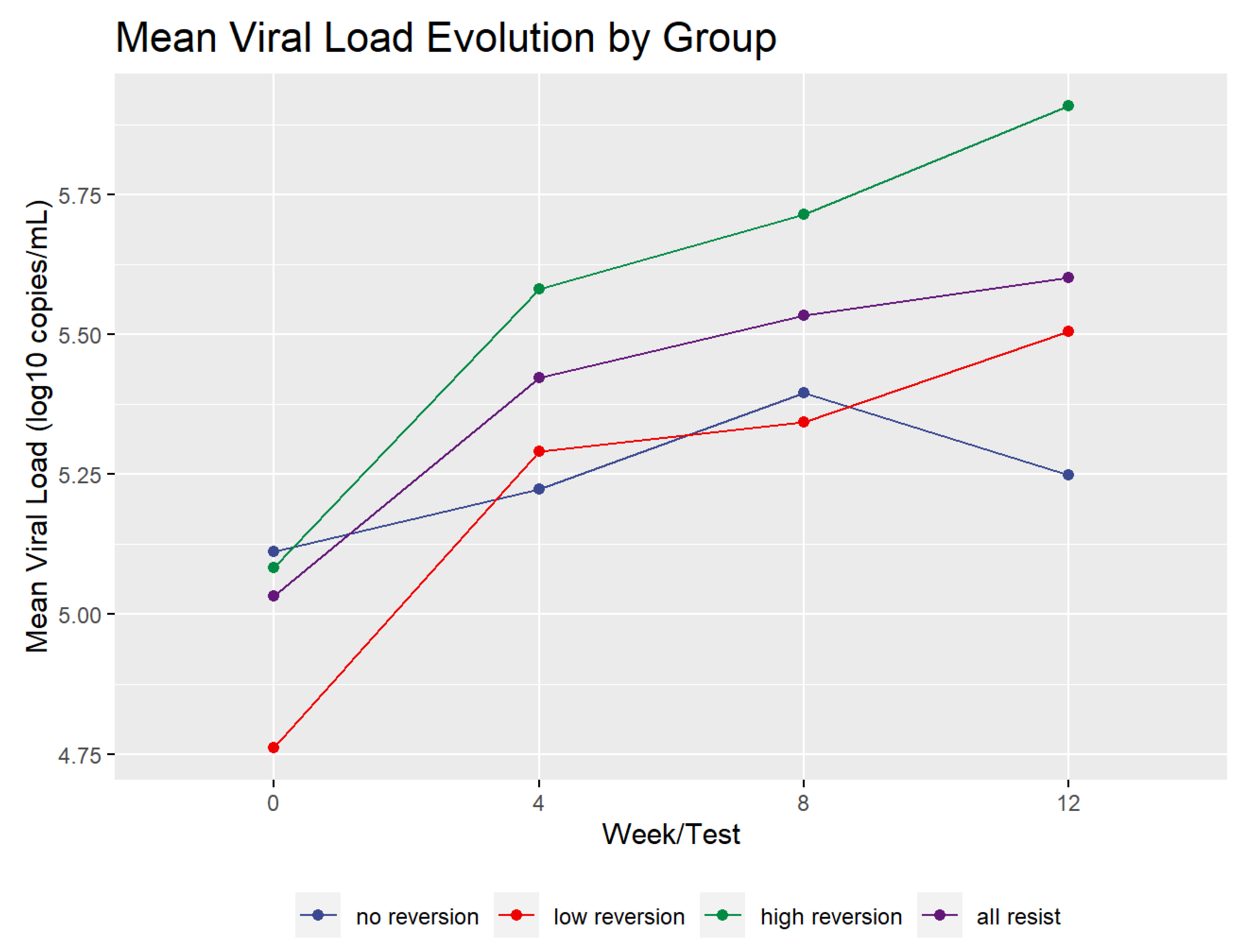
2.3. Viral Load Dynamics
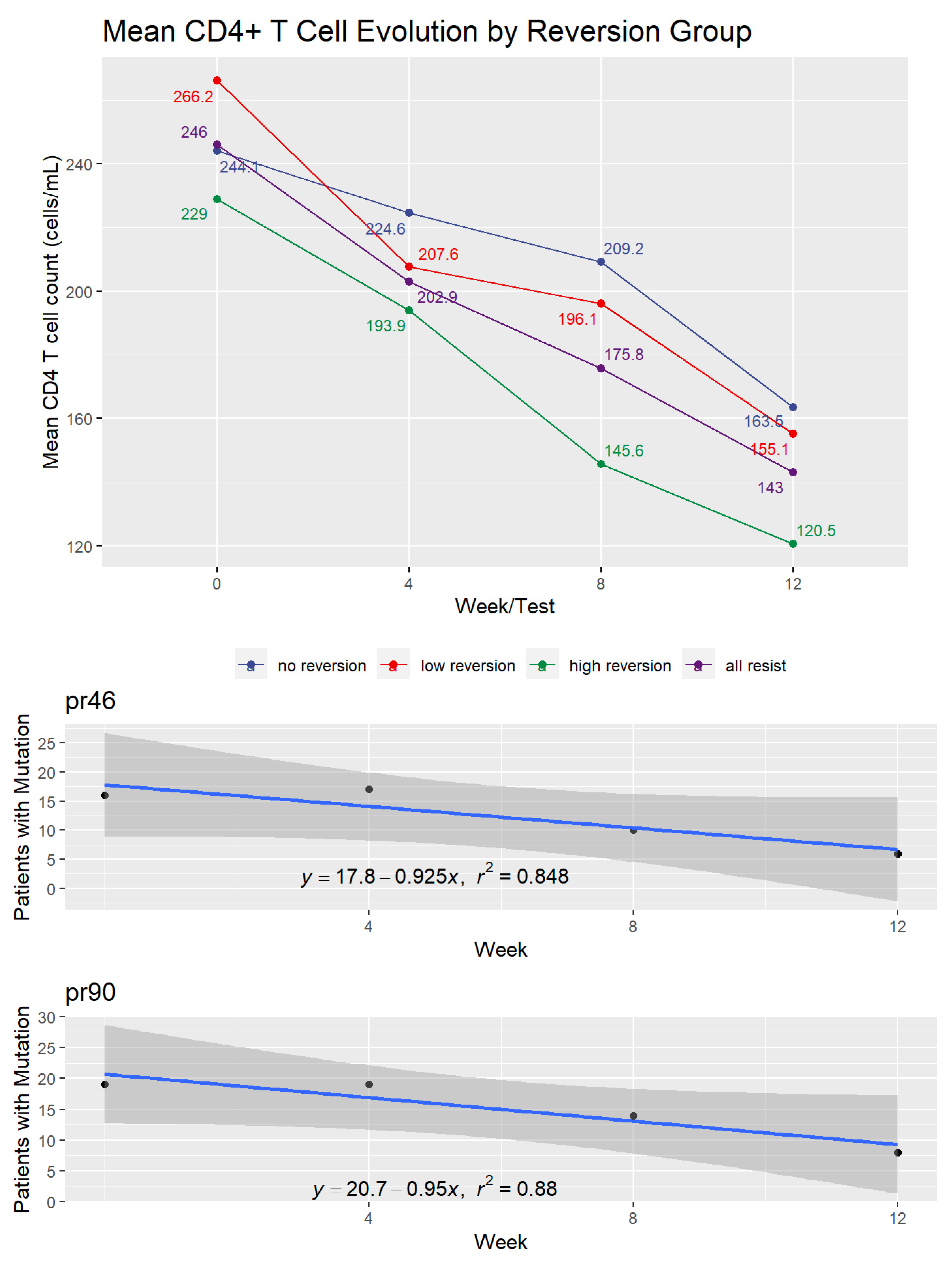
2.4. CD4+ and CD8+ T Cell Dynamics
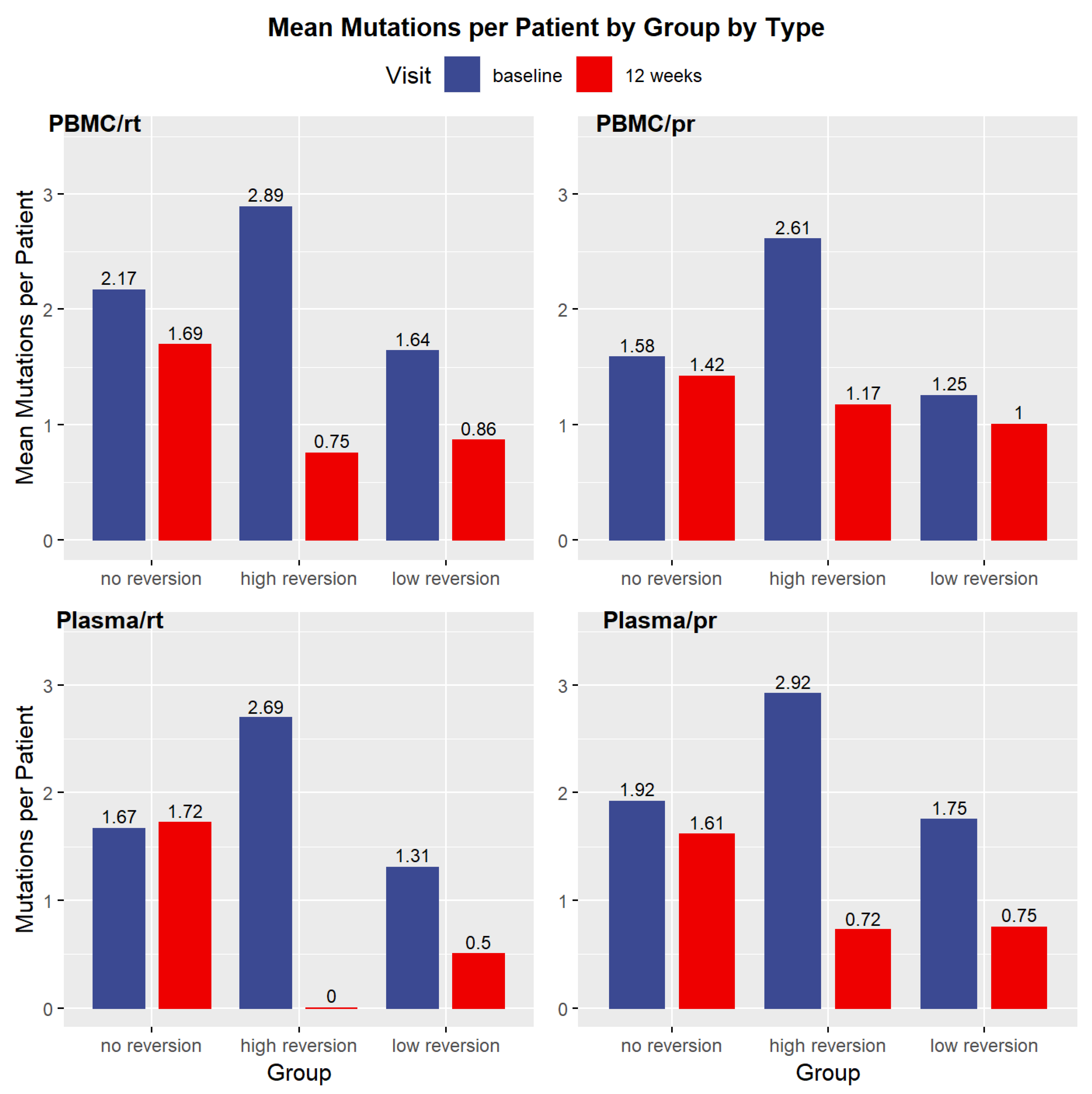
2.5. Reversion to Wild Type among Resistant Patients
2.6. Reversion to Wild Type in Plasma
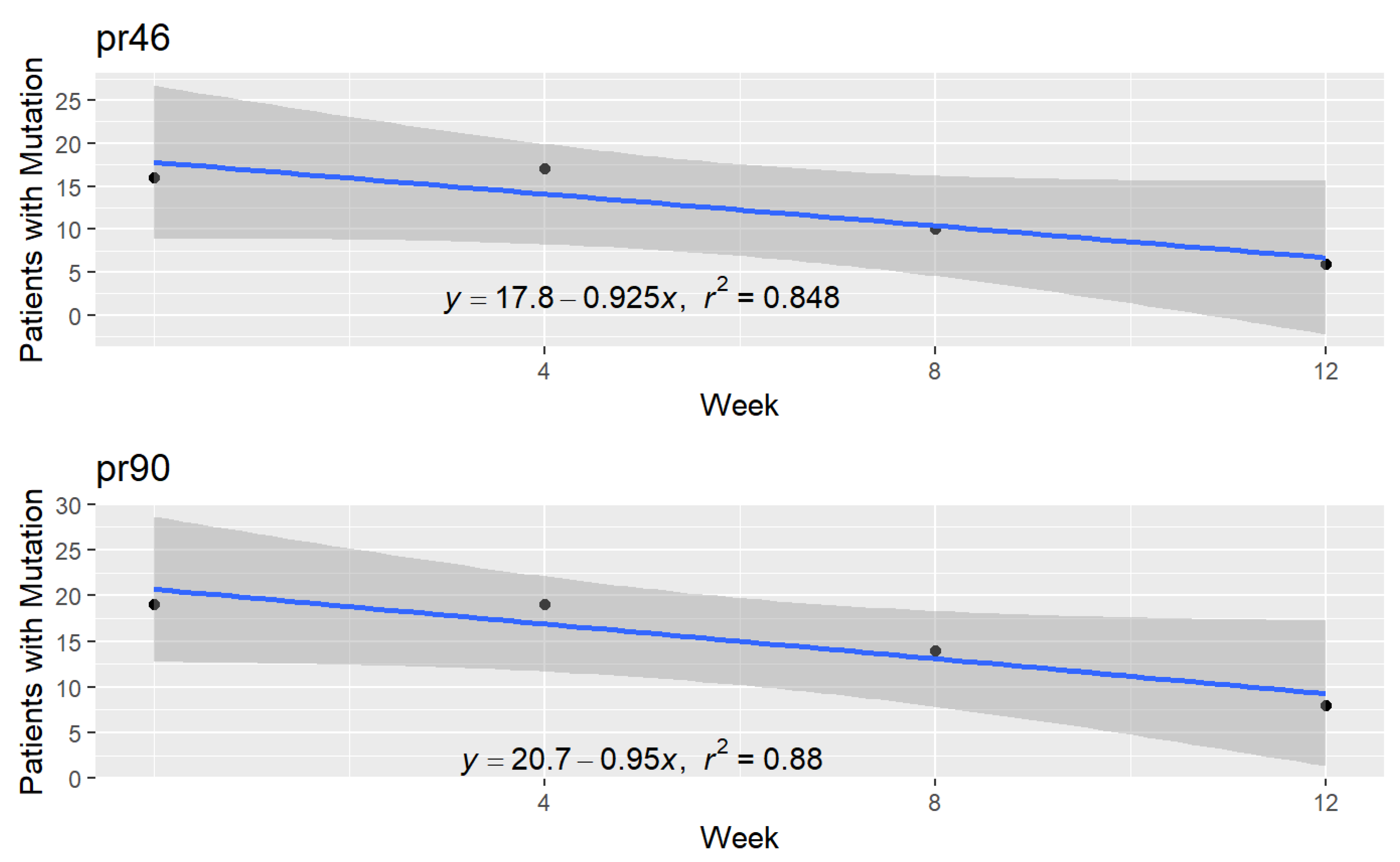
2.7. Protease Inhibitor (PI) Mutations at DNA Level
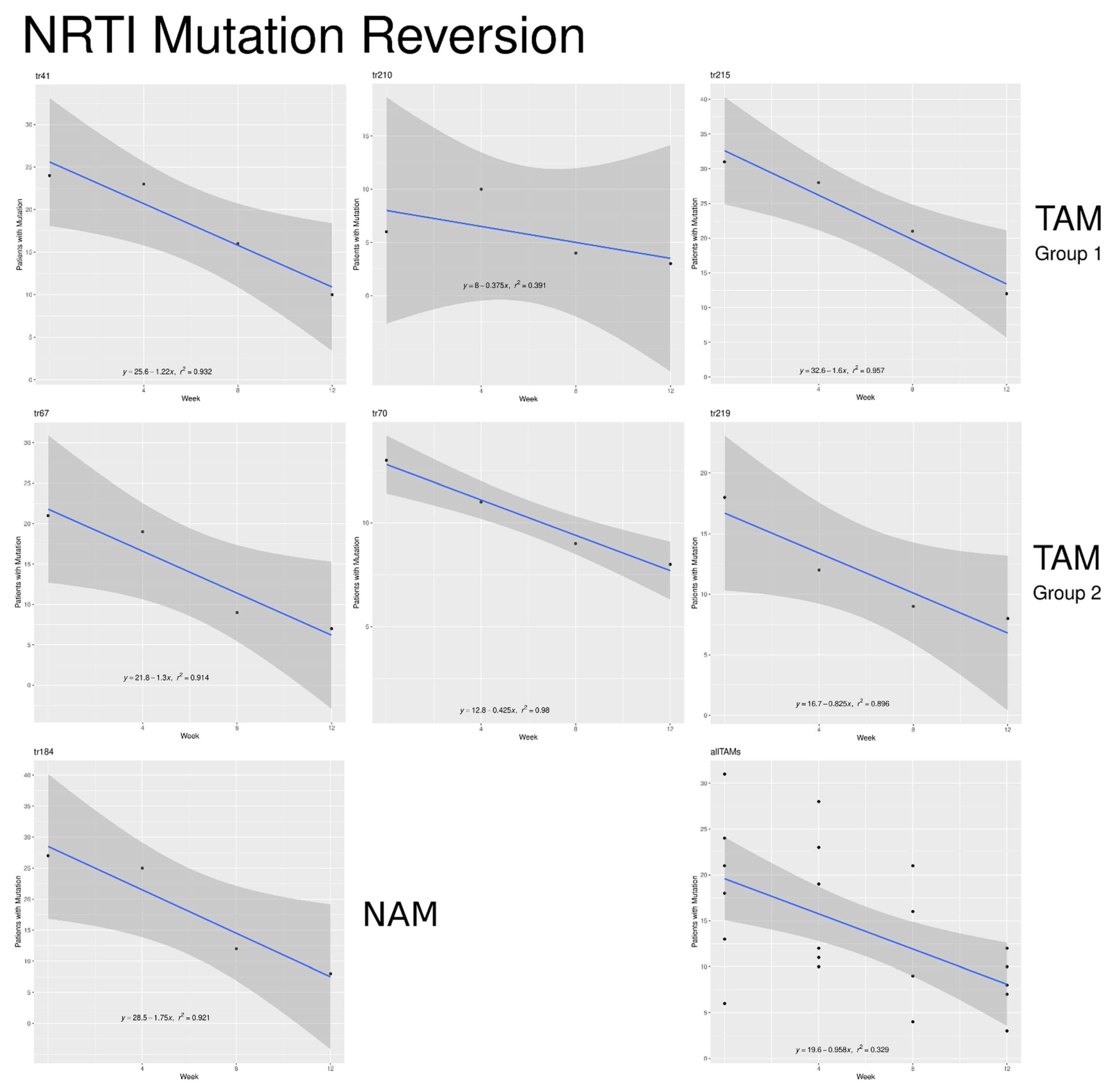
2.8. Nucleoside Reverse Transcriptase Inhibitor (NRTI) Mutations at DNA Level
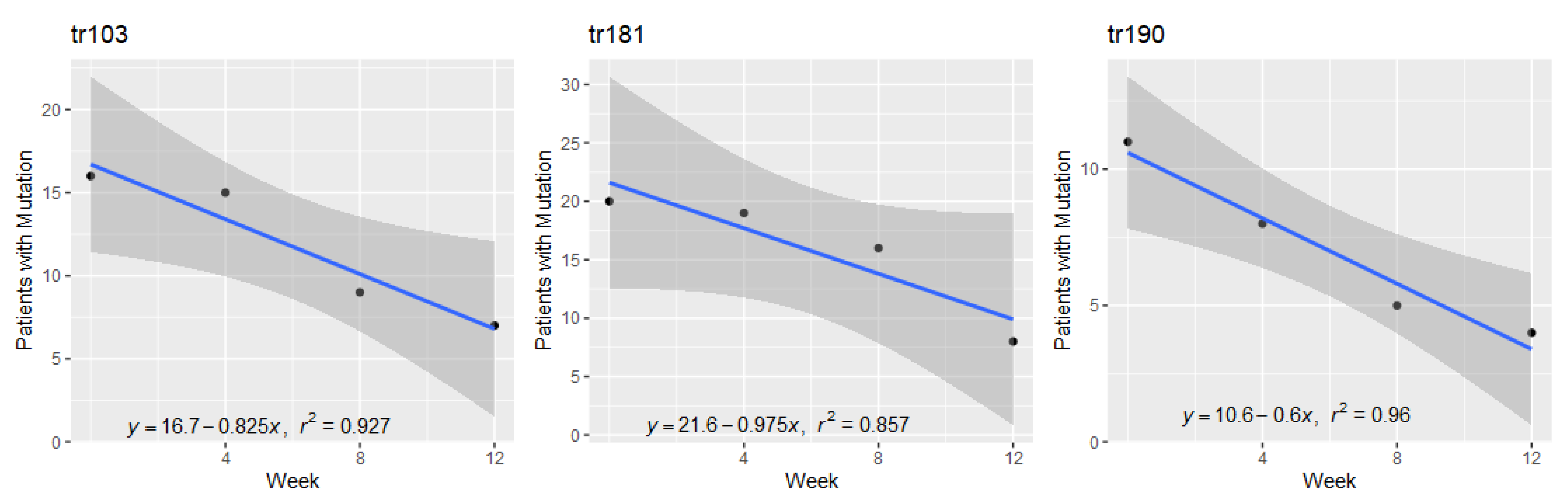
2.9. Non-Nucleoside Reverse Transcriptase Inhibitor (NNRTI) Mutations at DNA Level
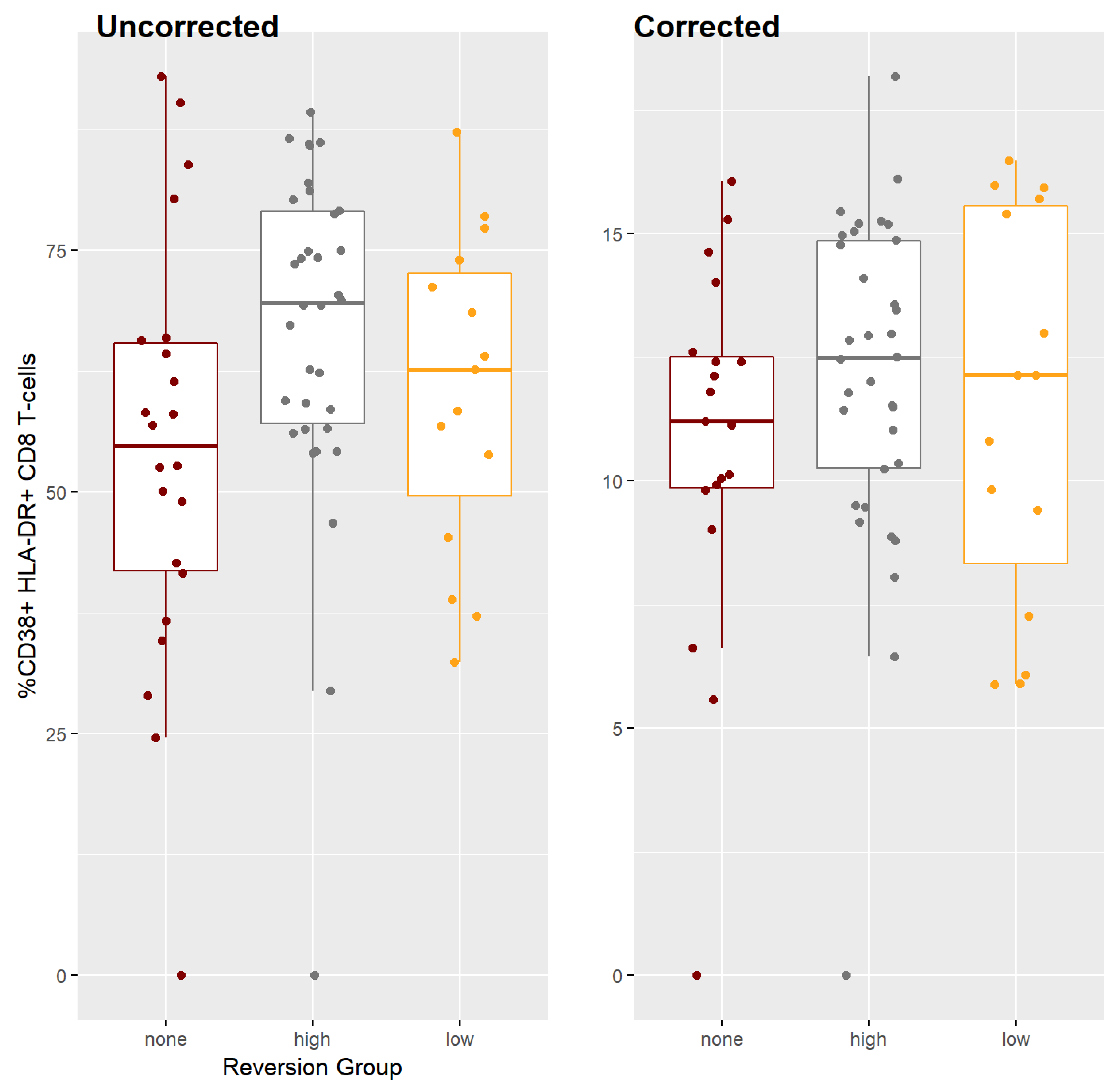
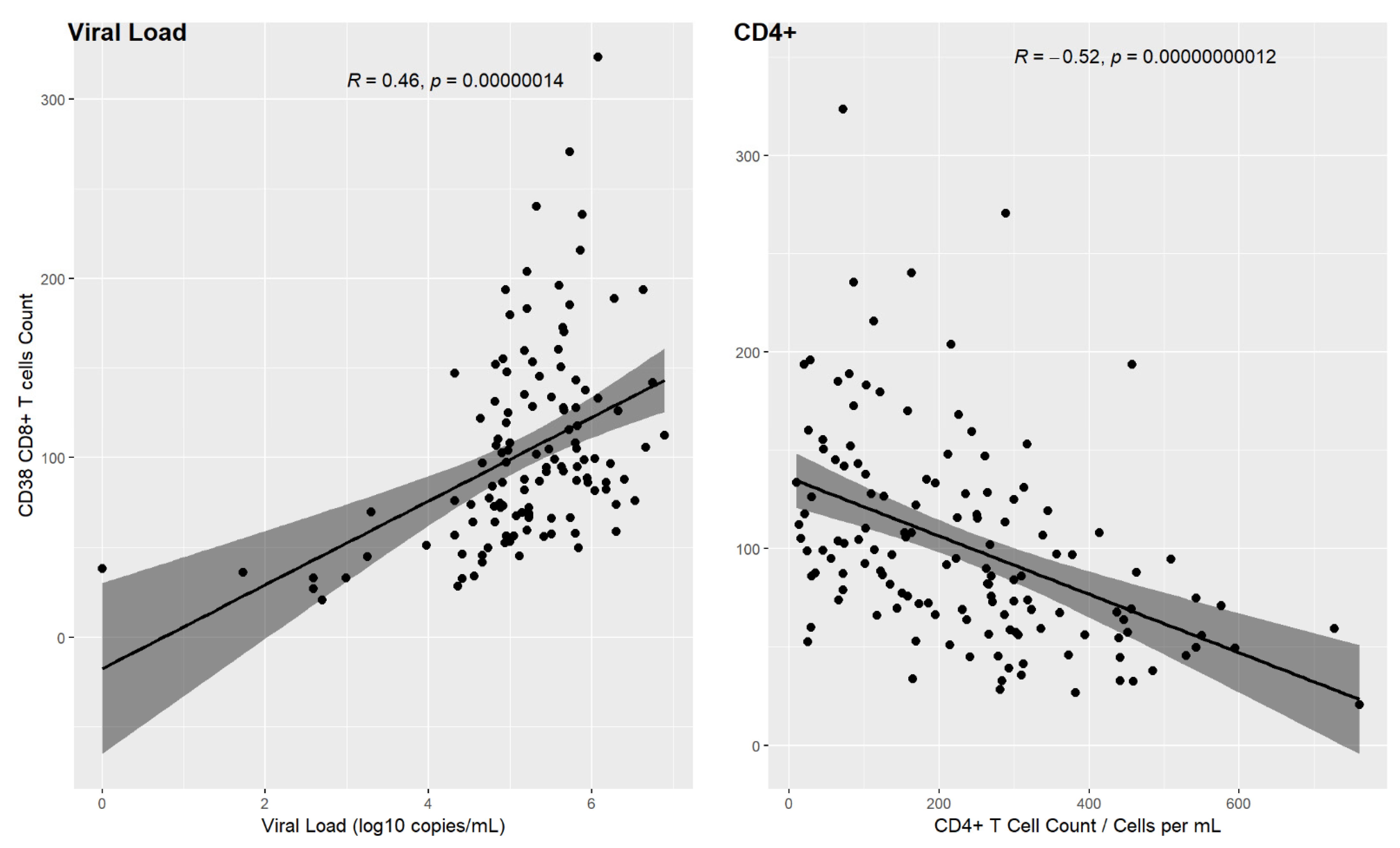
2.10. CD38 and HLA-DR T Cell Activation Markers
3. Discussion
3.1. Process of Reversion to Wild Type
3.2. Recombination
3.3. Persistence of Mutations versus Evolution to Wild Type
3.4. M184V as a Special Case of Mutation
3.5. Other Areas of the Viral Genome
4. Materials and Methods
4.1. Patients
4.2. Design
4.3. Plasma HIV Viral Load, CD4+ T Count, and Cell Activation Markers on CD4+ and CD8+ T Cells
4.4. Sequencing of the HIV-1 Pol Gene
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bajaria, S.H.; Webb, G.; Kirschner, D.E. Predicting differential responses to structured treatment interruptions during HAART. Bull. Math. Biol. 2004, 66, 1093–1118. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Margolick, J.B.; Conover, C.S.; Badri, S.; A Riddler, S.; Witt, M.D.; Jacobson, L.P. Interruption and discontinuation of highly active antiretroviral therapy in the multicenter AIDS cohort study. JAIDS J. Acquir. Immune Defic. Syndr. 2005, 38, 320–328. [Google Scholar]
- Koup, R.A. Reconsidering Early HIV Treatment and Supervised Treatment Interruptions. PLoS Med. 2004, 1, e41. [Google Scholar] [CrossRef] [PubMed]
- Oxenius, A.; Price, D.A.; Günthard, H.F.; Dawson, S.J.; Fagard, C.; Perrin, L.; Fischer, M.; Weber, R.; Plana, M.; García, F.; et al. Stimulation of HIV-specific cellular immunity by structured treatment interruption fails to enhance viral control in chronic HIV infection. Proc. Natl. Acad. Sci. USA 2002, 99, 13747–13752. [Google Scholar] [CrossRef] [PubMed]
- Dybul, M.; Chun, T.-W.; Yoder, C.; Hidalgo, B.; Belson, M.; Hertogs, K.; Larder, B.; Dewar, R.L.; Fox, C.H.; Hallahan, C.W.; et al. Short-cycle structured intermittent treatment of chronic HIV infection with highly active antiretroviral therapy: Effects on virologic, immunologic, and toxicity parameters. Proc. Natl. Acad. Sci. USA 2001, 98, 15161–15166. [Google Scholar] [CrossRef]
- Devereux, H.L.; Youle, M.; Johnson, M.A.; Loveday, C. Rapid decline in detectability of HIV-1 drug resistance mutations after stopping therapy. AIDS 1999, 13, 123–127. [Google Scholar] [CrossRef]
- Verhofstede, C.; Van Wanzeele, F.; Van Der Gucht, B.; De Cabooter, N.; Plum, J. Interruption of reverse transcriptase inhibitors or a switch from reverse transcriptase to protease inhibitors resulted in a fast reappearance of virus strains with a reverse transcriptase inhibitor-sensitive genotype. AIDS 1999, 13, 2541–2546. [Google Scholar] [CrossRef] [PubMed]
- Izopet, J.; Massip, P.; Souyris, C.; Sandres, K.; Puissant-Lubrano, B.; Obadia, M.; Pasquier, C.; Bonnet, E.; Marchou, B.; Puel, J. Shift in HIV resistance genotype after treatment interruption and short-term antiviral effect following a new salvage regimen. AIDS 2000, 14, 2247–2255. [Google Scholar] [CrossRef]
- Deeks, S.G.; Grant, R.M.; Wrin, T.; Paxinos, E.E.; Liegler, T.; Hoh, R.; Martin, J.N.; Petropoulos, C.J. Persistence of drug-resistant HIV-1 after a structured treatment interruption and its impact on treatment response. AIDS 2003, 17, 361–370. [Google Scholar] [CrossRef]
- Teixeira, D.; Munerato, P.; Komninakis, S.C.V.; Fusuma, E.E.; Janini, L.M.; Sucupira, M.C.A.; Diaz, R.S. The Detection ofin Vivoandin Vitro HIV Type 1 B/F Profiles in Brazil Using a Real-Time PCR Assay for Five HIV Type 1 Genomic Regions. AIDS Res. Hum. Retrovir. 2010, 26, 981–990. [Google Scholar] [CrossRef]
- Komninakis, S.V.; Santos, D.E.M.; Santos, C.; Oliveros, M.P.R.; Sanabani, S.; Diaz, R.S. HIV-1 Proviral DNA Loads (as Determined by Quantitative PCR) in Patients Subjected to Structured Treatment Interruption after Antiretroviral Therapy Failure. J. Clin. Microbiol. 2012, 50, 2132–2133. [Google Scholar] [CrossRef]
- Cuevas, J.M.; Geller, R.; Garijo, R.; López-Aldeguer, J.; Sanjuán, R. Extremely High Mutation Rate of HIV-1 In Vivo. PLoS Biol. 2015, 13, e1002251. [Google Scholar] [CrossRef] [PubMed]
- Dapp, M.; Heineman, R.H.; Mansky, L.M. Interrelationship between HIV-1 Fitness and Mutation Rate. J. Mol. Biol. 2012, 425, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Ananworanich, J.; Nuesch, R.; Le Braz, M.; Chetchotisakd, P.; Vibhagool, A.; Wicharuk, S.; Ruxrungtham, K.; Furrer, H.; Cooper, D.; Hirschel, B. Failures of 1 week on, 1 week off antiretroviral therapies in a randomized trial. AIDS 2003, 17, F33–F37. [Google Scholar] [CrossRef] [PubMed]
- Cohen, C.J.; Colson, A.E.; Sheble-Hall, A.G.; McLaughlin, K.A.; Morse, G.D. Pilot Study of a Novel Short-Cycle Antiretroviral Treatment Interruption Strategy: 48-Week Results of the Five-Days-On, Two-Days-Off (FOTO) Study. HIV Clin. Trials 2007, 8, 19–23. [Google Scholar] [CrossRef] [PubMed]
- Maggiolo, F.; Ripamonti, D.; Gregis, G.; Quinzan, G.; Callegaro, A.; Suter, F. Effect of Prolonged Discontinuation of Successful Antiretroviral Therapy on CD4 T Cells: A Controlled, Prospective Trial. AIDS Lond. Engl. 2004, 18, 439–446. [Google Scholar] [CrossRef]
- Castagna, A.; Danise, A.; Menzo, S.; Galli, L.; Gianotti, N.; Carini, E.; Boeri, E.; Galli, A.; Cernuschi, M.; Hasson, H.; et al. Lamivudine monotherapy in HIV-1-infected patients harbouring a lamivudine-resistant virus: A randomized pilot study (E-184V study). AIDS 2006, 20, 795–803. [Google Scholar] [CrossRef] [PubMed]
- Günthard, H.; Frost, S.; Leigh-Brown, A.J.; Ignacio, C.C.; Kee, K.; Perelson, A.S.; Spina, C.A.; Havlir, D.V.; Hezareh, M.; Looney, D.J.; et al. Evolution of Envelope Sequences of Human Immunodeficiency Virus Type 1 in Cellular Reservoirs in the Setting of Potent Antiviral Therapy. J. Virol. 1999, 73, 9404–9412. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Ramratnam, B.; Tenner-Racz, K.; He, Y.; Vesanen, M.; Lewin, S.R.; Talal, A.; Racz, P.; Perelson, A.S.; Korber, B.; et al. Quantifying Residual HIV-1 Replication in Patients Receiving Combination Antiretroviral Therapy. N. Engl. J. Med. 1999, 340, 1605–1613. [Google Scholar] [CrossRef] [PubMed]
- Pariente, N.; Pernas, M.; de la Rosa, R.; Gómez-Mariano, G.; Fernandez, G.; Rubio, A.; López, M.; Benito, J.M.; Lopez-Galindez, C.; Leal, M.; et al. Long-term suppression of plasma viremia with highly active antiretroviral therapy despite virus evolution and very limited selection of drug-resistant genotypes. J. Med. Virol. 2004, 73, 350–361. [Google Scholar] [CrossRef]
- Wong, J.K.; Ignacio, C.C.; Torriani, F.; Havlir, D.; Fitch, N.J.; Richman, D.D. In Vivo Compartmentalization of Human Immunodeficiency Virus: Evidence from the Examination of Pol Sequences from Autopsy Tissues. J. Virol. 1997, 71, 2059–2071. [Google Scholar] [CrossRef] [PubMed]
- Kim, R.B.; Fromm, M.F.; Wandel, C.; Leake, B.; Wood, A.J.; Roden, D.M.; Wilkinson, G.R. The drug transporter P-glycoprotein limits oral absorption and brain entry of HIV-1 protease inhibitors. J. Clin. Investig. 1998, 101, 289–294. [Google Scholar] [CrossRef] [PubMed]
- Finzi, D.; Blankson, J.N.; Siliciano, J.D.; Margolick, J.B.; Chadwick, K.; Pierson, T.C.; A Smith, K.; Lisziewicz, J.; Lori, F.; Flexner, C.; et al. Latent infection of CD4+ T cells provides a mechanism for lifelong persistence of HIV-1, even in patients on effective combination therapy. Nat. Med. 1999, 5, 512–517. [Google Scholar] [CrossRef] [PubMed]
- Siliciano, R.F. Latency and reservoirs for HIV-1. AIDS 1999, 13, S49–S58. [Google Scholar] [PubMed]
- Munerato, P.; Sucupira, M.C.; Oliveros, M.P.R.; Janini, L.M.; de Souza, D.F.; Pereira, A.A.; Inocencio, L.A.; Diaz, R.S. HIV Type 1 Antiretroviral Resistance Mutations in Subtypes B, C, and F in the City of São Paulo, Brazil. AIDS Res. Hum. Retrovir. 2010, 26, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Deeks, S.G.; Wrin, T.; Liegler, T.; Hoh, R.; Hayden, M.; Barbour, J.D.; Hellmann, N.S.; Petropoulos, C.J.; McCune, J.M.; Hellerstein, M.K.; et al. Virologic and Immunologic Consequences of Discontinuing Combination Antiretroviral-Drug Therapy in HIV-Infected Patients with Detectable Viremia. N. Engl. J. Med. 2001, 344, 472–480. [Google Scholar] [CrossRef]
- Winters, M.A.; Baxter, J.D.; Mayers, D.L.; Wentworth, D.N.; Hoover, M.L.; Neaton, J.D.; Merigan, T.C. Frequency of Antiretroviral Drug Resistance Mutations in HIV-1 Strains from Patients Failing Triple Drug Regimens. The Terry Beirn Community Programs for Clinical Research on AIDS. Antivir. Ther. 2000, 5, 57–63. [Google Scholar]
- Domingo, E.; Escarmís, C.; Sevilla, N.; Moya, A.; Elena, S.; Quer, J.; Novella, I.S.; Holland, J.J. Basic concepts in RNA virus evolution. FASEB J. 1996, 10, 859–864. [Google Scholar] [CrossRef]
- Miller, V.; Sabin, C.; Hertogs, K.; Bloor, S.; Martinez-Picado, J.; D’Aquila, R.; Larder, B.; Lutz, T.; Gute, P.; Weidmann, E.; et al. Virological and immunological effects of treatment interruptions in HIV-1 infected patients with treatment failure. AIDS 2000, 14, 2857–2867. [Google Scholar] [CrossRef]
- Katlama, C.; Dominguez, S.; Gourlain, K.; Duvivier, C.; Delaugerre, C.; Legrand, M.; Tubiana, R.; Reynes, J.; Molina, J.-M.; Peytavin, G.; et al. Benefit of treatment interruption in HIV-infected patients with multiple therapeutic failures: A randomized controlled trial (ANRS 097). AIDS 2004, 18, 217–226. [Google Scholar] [CrossRef]
- Kestens, L.; Vanham, G.; Vereecken, C.; Vandenbruaene, M.; Vercauteren, G.; Colebunders, R.L.; Gigase, P.L. Selective Increase of Activation Antigens HLA-DR and CD38 on CD4+ CD45RO+ T Lymphocytes during HIV-1 Infection. Clin. Exp. Immunol. 1994, 95, 436–441. [Google Scholar] [CrossRef]
- Hunt, P.W.; Martin, J.N.; Sinclair, E.; Bredt, B.; Hagos, E.; Lampiris, H.; Deeks, S.G. T Cell Activation Is Associated with Lower CD4+T Cell Gains in Human Immunodeficiency Virus–Infected Patients with Sustained Viral Suppression during Antiretroviral Therapy. J. Infect. Dis. 2003, 187, 1534–1543. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, R.M.; Mohri, H.; Ho, D.D.; Perelson, A.S. In vivo dynamics of T cell activation, proliferation, and death in HIV-1 infection: Why are CD4+ but not CD8+ T cells depleted? Proc. Natl. Acad. Sci. USA 2002, 99, 15572–15577. [Google Scholar] [CrossRef] [PubMed]
- Klatt, N.R.; Silvestri, G. CD4+ T Cells and HIV: A Paradoxical Pas de Deux. Sci. Transl. Med. 2012, 4, 123ps4. [Google Scholar] [CrossRef]
- Klatt, N.R.; Chomont, N.; Douek, D.C.; Deeks, S.G. Immune activation and HIV persistence: Implications for curative approaches to HIV infection. Immunol. Rev. 2013, 254, 326–342. [Google Scholar] [CrossRef]
- Ruggiero, A.; De Spiegelaere, W.; Cozzi-Lepri, A.; Kiselinova, M.; Pollakis, G.; Beloukas, A.; Vandekerckhove, L.; Strain, M.; Richman, D.; Phillips, A.; et al. During Stably Suppressive Antiretroviral Therapy Integrated HIV-1 DNA Load in Peripheral Blood is Associated with the Frequency of CD8 Cells Expressing HLA-DR/DP/DQ. EBioMedicine 2015, 2, 1153–1159. [Google Scholar] [CrossRef]
- HIV Drug Resistance Database [Internet] NRTI Resistance Notes [Updated 2016 May 31] 2021. Available online: https://hivdb.stanford.edu/dr-summary/resistance-notes/NRTI/ (accessed on 25 January 2019).
- Hunt, P.W.; Brenchley, J.; Sinclair, E.; McCune, J.M.; Roland, M.; Page, K.; Hsue, P.; Emu, B.; Krone, M.; Lampiris, H.; et al. Relationship between T Cell Activation and CD4+T Cell Count in HIV-Seropositive Individuals with Undetectable Plasma HIV RNA Levels in the Absence of Therapy. J. Infect. Dis. 2008, 197, 126–133. [Google Scholar] [CrossRef] [PubMed]
- Li, J.Z.; Etemad, B.; Ahmed, H.; Aga, E.; Bosch, R.J.; Mellors, J.W.; Kuritzkes, D.R.; Lederman, M.M.; Para, M.; Gandhi, R.T. The size of the expressed HIV reservoir predicts timing of viral rebound after treatment interruption. AIDS 2015, 30, 343–353. [Google Scholar] [CrossRef] [PubMed]
- Albert, J.; Wahlberg, J.; Lundeberg, J.; Cox, S.; Sandström, E.; Wahren, B.; Uhlén, M. Persistence of azidothymidine-resistant human immunodeficiency virus type 1 RNA genotypes in posttreatment sera. J. Virol. 1992, 66, 5627–5630. [Google Scholar] [CrossRef] [PubMed]
- Deeks, S.G.; Hoh, R.; Neilands, T.B.; Liegler, T.; Aweeka, F.; Petropoulos, C.J.; Grant, R.M.; Martin, J.N. Interruption of Treatment with Individual Therapeutic Drug Classes in Adults with Multidrug-Resistant HIV-1 Infection. J. Infect. Dis. 2005, 192, 1537–1544. [Google Scholar] [CrossRef]
- Guimarães, A.P.; Sa-Filho, D.J.; Sucupira, M.C.; Janini, L.M.; Diaz, R.S. Profiling Resistance-Related Mutations in the Protease Region of the pol Gene: Single Genome Sequencing of HIV in Plasma and Peripheral Blood Mononuclear Cells. AIDS Res. Hum. Retrovir. 2008, 24, 969–971. [Google Scholar] [CrossRef]
- Silva, W.P.; Santos, D.E.M.; Leal, E.; Brunstein, A.; Sucupira, M.C.A.; Sabino, E.C.; Diaz, R.S. Reactivation of ancestral strains of HIV-1 in the gp120 V3 env region in patients failing antiretroviral therapy and subjected to structured treatment interruption. Virology 2006, 354, 35–47. [Google Scholar] [CrossRef][Green Version]
- Pelemans, H.; Esnouf, R.M.; Jonckheere, H.; De Clercq, E.; Balzarini, J. Mutational Analysis of Tyr-318 within the Non-nucleoside Reverse Transcriptase Inhibitor Binding Pocket of Human Immunodeficiency Virus Type I Reverse Transcriptase. J. Biol. Chem. 1998, 273, 34234–34239. [Google Scholar] [CrossRef]
- Wrin, T.; Gamarnik, A.; Huang, W. Drug Resistance Is Associated with Impaired PR and Reverse Transcriptase Function and Reduced Replication Capacity: Characterization of Recombinant Viruses Derived from 200 HIV-1 Infected Patients. Toronto, ON, Canada, 17 September 2000. [Google Scholar]
- Arruda, M.B.; Boullosa, L.T.; Cardoso, C.C.; Da Costa, C.M.; Brites, C.; De Lima, S.T.; Kaminski, H.T.; Aleixo, A.W.; Esposito, A.O.; Cavalcanti, A.M.; et al. Brazilian network for HIV Drug Resistance Surveillance (HIV-BresNet): A survey of treatment-naive individuals. J. Int. AIDS Soc. 2018, 21, e25032. [Google Scholar] [CrossRef]
- Jain, V.; Sucupira, M.C.; Bacchetti, P.; Hartogensis, W.; Diaz, R.S.; Kallas, E.G.; Janini, L.M.; Liegler, T.; Pilcher, C.D.; Grant, R.M.; et al. Differential Persistence of Transmitted HIV-1 Drug Resistance Mutation Classes. J. Infect. Dis. 2011, 203, 1174–1181. [Google Scholar] [CrossRef] [PubMed]
- Leda, A.R.; Hunter, J.; Oliveira, U.C.; Azevedo, I.J.; Sucupira, M.C.A.; Diaz, R.S. Insights about minority HIV-1 strains in transmitted drug resistance mutation dynamics and disease progression. J. Antimicrob. Chemother. 2018, 73, 1930–1934. [Google Scholar] [CrossRef] [PubMed]
- Baxter, J.; Dunn, D.; Marvig, R.; Bennedbaek, M.; Cozzi-Lepri, A. Pretreatment HIV Drug Resistance in the START Study Using Next Generation Sequencing. Top. Antivir. Med. 2018, 26 (Suppl. 222s). Available online: https://www.mendeley.com/catalogue/15a86a24-4afd-35e5-91e2-cf8fae51b860/ (accessed on 4 October 2021).
- Sustiva Packge Insert Revised 3/2016. Available online: https://www.Accessdata.Fda.Gov/Drugsatfda_Docs/Label/2016/020972s049-021360s038lbl.Pdf (accessed on 25 January 2019).
- Vuramune Package Insert Revised 11/2011. Available online: https://www.accessdata.fda.gov/drugsatfda_docs/label/2011/020636s039_020933s030lbl.pdf (accessed on 25 January 2019).
- Rodríguez-Nóvoa, S.; Barreiro, P.; Jiménez-Nácher, I.; Soriano, V. Overview of the pharmacogenetics of HIV therapy. Pharm. J. 2006, 6, 234–245. [Google Scholar] [CrossRef]
- de Goede, A.L.; van Deutekom, H.W.M.; Vrancken, B.; Schutten, M.; Allard, S.D.; van Baalen, C.A.; Osterhaus, A.D.M.E.; Thielemans, K.; Aerts, J.L.; Keşmir, C.; et al. HIV-1 evolution in patients undergoing immunotherapy with Tat, Rev, and Nef expressing dendritic cells followed by treatment interruption. AIDS 2013, 27, 2679–2689. [Google Scholar] [CrossRef] [PubMed]
- Diaz, R.S.; Zhang, L.; Busch, M.P.; Mosley, J.W.; Mayer, A. Divergence of HIV-1 quasispecies in an epidemiologic cluster. AIDS 1997, 11, 415–422. [Google Scholar] [CrossRef]
- Sucupira, M.C.A.; Caseiro, M.M.; Alves, K.; Tescarollo, G.; Janini, L.M.; Sabino, E.C.; Castelo, A.; Page-Shafer, K.; Diaz, R.S. High Levels of Primary Antiretroviral Resistance Genotypic Mutations And B/F Recombinants in Santos, Brazil. AIDS Patient Care STDs 2007, 21, 116–128. [Google Scholar] [CrossRef]
- Wensing, A.M.; Calvez, V.; Günthard, H.F.; Johnson, V.A.; Paredes, R.; Pillay, D.; Shafer, R.W.; Richman, U.D. 2017 Update of the Drug Resistance Mutations in HIV-1. Top. Antivir. Med. 2017, 24, 132–133. [Google Scholar]
- Caffo, B.S. Regression Models for Data Science in R; Leanpub: Victoria, BC, Canada, 2015; p. 137. [Google Scholar]
- Harrell, F.E. Regression Modeling Strategies; Springer Series in Statistics; Springer: New York, NY, USA, 2001; ISBN 978-1-4419-2918-1. [Google Scholar]
- Bland, J.M.; Altman, D.G. Statistics Notes: Survival probabilities (the Kaplan-Meier method). BMJ 1998, 317, 1572–1580. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, E.L.; Meier, P. Nonparametric Estimation from Incomplete Observations. J. Am. Stat. Assoc. 1958, 53, 457–481. [Google Scholar] [CrossRef]
- Rich, J.T.; Neely, J.G.; Paniello, R.C.; Voelker, C.C.J.; Nussenbaum, B.; Wang, E. A practical guide to understanding Kaplan-Meier curves. Otolaryngol. Neck Surg. 2010, 143, 331–336. [Google Scholar] [CrossRef]
- Crawley, M.J. Death and Failure Data. In Statistics: An Introduction Using R; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2011; pp. 263–268. ISBN 978-1-119-94175-0. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2016. [Google Scholar]
- Huber, W.; Carey, V.J.; Gentleman, R.; Anders, S.; Carlson, M.; Carvalho, B.S.; Bravo, H.C.; Davis, S.; Gatto, L.; Girke, T.; et al. Orchestrating High-Throughput Genomic Analysis with Bioconductor. Nat. Methods 2015, 12, 115–121. [Google Scholar] [CrossRef]
- Charif, D.; Lobry, J.R. SeqinR 1.0-2: A contributed package to the R project for statistical computing devoted to biological sequences retrieval and analysis. In Structural Approaches to Sequence Evolution: Molecules, Networks, Populations; Bastolla, U., Porto, M., Roman, H.E., Vendruscolo, M., Eds.; Biological and Medical Physics, Biomedical Engineering; Springer Verlag: New York, NY, USA, 2007; pp. 207–232. [Google Scholar]
| Week | Control | All Resistant Subjects | No Reversion | Low Reversion | High Reversion |
|---|---|---|---|---|---|
| 0 | 5.230 | 5.021 (0.556) | 5.111 (0.730) | 5.082 (0.354) | 4.761 (0.532) |
| 4 | 6.072 | 5.386 (0.511) | 5.222 (0.609) | 5.290 (0.614) | 5.580 (0.275) |
| 8 | 5.841 | 5.516 (0.537) | 5.395 (0.610) | 5.343 (0.622) | 5.714 (0.368) |
| 12 | 5.962 | 5.580 (0.597) | 5.247 (0.621) | 5.505 (0.513) | 5.908 (0.456) |
| Overall Increase | 0.732 | 0.568 | 0.136 | 0.744 | 0.826 |
| p-value | N/A | <1.0 × 10−6 * | 0.3021 | 0.0024 * | <1.0 × 10−6 * |
| n | 2 | 36 | 13 | 15 | 8 |
| Codon | AA–Pre-ATI | AA–Post ATI | Patients with Mutation | Present in IAS List |
|---|---|---|---|---|
| pr10 | I | L | 8 | Yes |
| pr20 | R | K | 5 | Yes |
| pr20 | I | K | 2 | Yes |
| pr24 | I | L | 2 | Yes |
| pr33 | F | L | 2 | Yes |
| pr35 | D | E | 2 | No |
| pr36 | I | M | 3 | Yes |
| pr46 | I | M | 6 | Yes |
| pr48 | V | G | 3 | Yes |
| pr54 | V | I | 7 | Yes |
| pr57 | K | R | 2 | No |
| pr58 | E | Q | 2 | No |
| pr62 | V | I | 6 | Yes |
| pr63 | P | L | 5 | No |
| pr71 | V | A | 6 | Yes |
| pr73 | S | G | 2 | Yes |
| pr73 | T | G | 2 | Yes |
| pr82 | A | V | 7 | Yes |
| pr84 | V | I | 7 | Yes |
| pr84 | I | V | 2 | No |
| pr90 | M | L | 6 | Yes |
| pr93 | L | I | 2 | Yes |
| tr18 | R | K | 2 | No |
| tr35 | V | I | 2 | No |
| tr41 | L | M | 7 | Yes |
| tr67 | N | D | 10 | Yes |
| tr70 | R | K | 3 | Yes |
| tr103 | N | K | 2 | Yes |
| tr106 | V | I | 3 | Yes |
| tr118 | I | V | 3 | No |
| tr173 | K | E | 2 | No |
| tr181 | C | Y | 5 | Yes |
| tr184 | V | M | 10 | Yes |
| tr190 | A | G | 2 | Yes |
| tr210 | W | L | 3 | Yes |
| tr211 | K | R | 3 | No |
| tr211 | Q | K | 2 | No |
| tr215 | Y | T | 9 | Yes |
| tr215 | F | T | 2 | Yes |
| tr218 | E | D | 2 | No |
| Codon | ARV Type | Weeks to 0 Patients | R2 | Mutation Survival Probability |
|---|---|---|---|---|
| pr10 | PI | 44.000 | 0.143 | 0.184 |
| pr20 | PI | 37.538 | 0.573 | 0.282 |
| pr24 | PI | 17.667 | 0.800 | 0.333 |
| pr30 | PI | 16.000 | 0.600 | 0.000 |
| pr36 | PI | 98.500 | 0.246 | 0.483 |
| pr46 | PI | 19.077 | 0.848 | 0.250 |
| pr48 | PI | 16.000 | 0.720 | 0.188 |
| pr53 | PI | 24.000 | 0.263 | 0.200 |
| pr54 | PI | 34.261 | 0.354 | 0.423 |
| pr63 | PI | 97.667 | 0.288 | 0.647 |
| pr71 | PI | 33.692 | 0.676 | 0.391 |
| pr73 | PI | 15.000 | 0.690 | 0.000 |
| pr77 | PI | NA | 0.400 | 0.318 |
| pr82 | PI | 22.400 | 0.641 | 0.300 |
| pr84 | PI | 19.889 | 0.800 | 0.294 |
| pr88 | PI | NA | 0.833 | 0.176 |
| pr90 | PI | 21.500 | 0.880 | 0.300 |
| tr41 | NRTI–TAM 1 | 20.898 | 0.932 | 0.317 |
| tr67 | NRTI–TAM 2 | 16.357 | 0.914 | 0.217 |
| tr70 | NRTI–TAM 2 | 30.118 | 0.980 | 0.320 |
| tr184 | NRTI | 16.000 | 0.916 | 0.214 |
| tr210 | NRTI–TAM 1 | 23.143 | 0.391 | 0.167 |
| tr215 | NRTI–TAM 1 | 19.824 | 0.957 | 0.355 |
| tr219 | NRTI–TAM 2 | 20.242 | 0.896 | 0.250 |
| tr103 | NNRTI | 20.242 | 0.927 | 0.353 |
| tr181 | NNRTI | 21.714 | 0.857 | 0.259 |
| tr190 | NNRTI | 17.481 | 0.960 | 0.250 |
| Class | Mean | Standard Deviation | Number of Mutations in Study |
|---|---|---|---|
| PI | 34.5 | 27.27 | 17 |
| NRTI | 20.9 | 4.76 | 7 |
| NNRTI | 19.8 | 2.15 | 3 |
| Reversion Group | N | Mean Cell Activation | Std Dev Cell Activation |
|---|---|---|---|
| Expression of Cell Activation Proteins | |||
| No reversion | 6 | 54.17 | 22.17 |
| High reversion | 10 | 66.56 | 17.86 |
| Low reversion | 4 | 60.44 | 16.47 |
| Cell Activation Proteins Controlled for Log Viral Load | |||
| No reversion | 6 | 10.78 | 3.74 |
| High reversion | 10 | 12.07 | 3.41 |
| Low reversion | 4 | 11.466 | 3.95 |
| Measure | No Reversion | High Reversion | p-Value of Difference |
|---|---|---|---|
| cd8/cd38/hla-dr | 21.637 | 13.252 | 0.3272 |
| Corrected for Viral Load | 4.916 | 0.658 | 0.1797 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hunter, J.R.; dos Santos, D.E.M.; Munerato, P.; Janini, L.M.; Castelo, A.; Sucupira, M.C.; Truong, H.-H.M.; Diaz, R.S. Fitness Cost of Antiretroviral Drug Resistance Mutations on the pol Gene during Analytical Antiretroviral Treatment Interruption among Individuals Experiencing Virological Failure. Pathogens 2021, 10, 1425. https://doi.org/10.3390/pathogens10111425
Hunter JR, dos Santos DEM, Munerato P, Janini LM, Castelo A, Sucupira MC, Truong H-HM, Diaz RS. Fitness Cost of Antiretroviral Drug Resistance Mutations on the pol Gene during Analytical Antiretroviral Treatment Interruption among Individuals Experiencing Virological Failure. Pathogens. 2021; 10(11):1425. https://doi.org/10.3390/pathogens10111425
Chicago/Turabian StyleHunter, James R., Domingos E. Matos dos Santos, Patricia Munerato, Luiz Mario Janini, Adauto Castelo, Maria Cecilia Sucupira, Hong-Ha M. Truong, and Ricardo Sobhie Diaz. 2021. "Fitness Cost of Antiretroviral Drug Resistance Mutations on the pol Gene during Analytical Antiretroviral Treatment Interruption among Individuals Experiencing Virological Failure" Pathogens 10, no. 11: 1425. https://doi.org/10.3390/pathogens10111425
APA StyleHunter, J. R., dos Santos, D. E. M., Munerato, P., Janini, L. M., Castelo, A., Sucupira, M. C., Truong, H.-H. M., & Diaz, R. S. (2021). Fitness Cost of Antiretroviral Drug Resistance Mutations on the pol Gene during Analytical Antiretroviral Treatment Interruption among Individuals Experiencing Virological Failure. Pathogens, 10(11), 1425. https://doi.org/10.3390/pathogens10111425






