The Role of ERG11 Point Mutations in the Resistance of Candida albicans to Fluconazole in the Presence of Lactate
Abstract
1. Introduction
2. Results
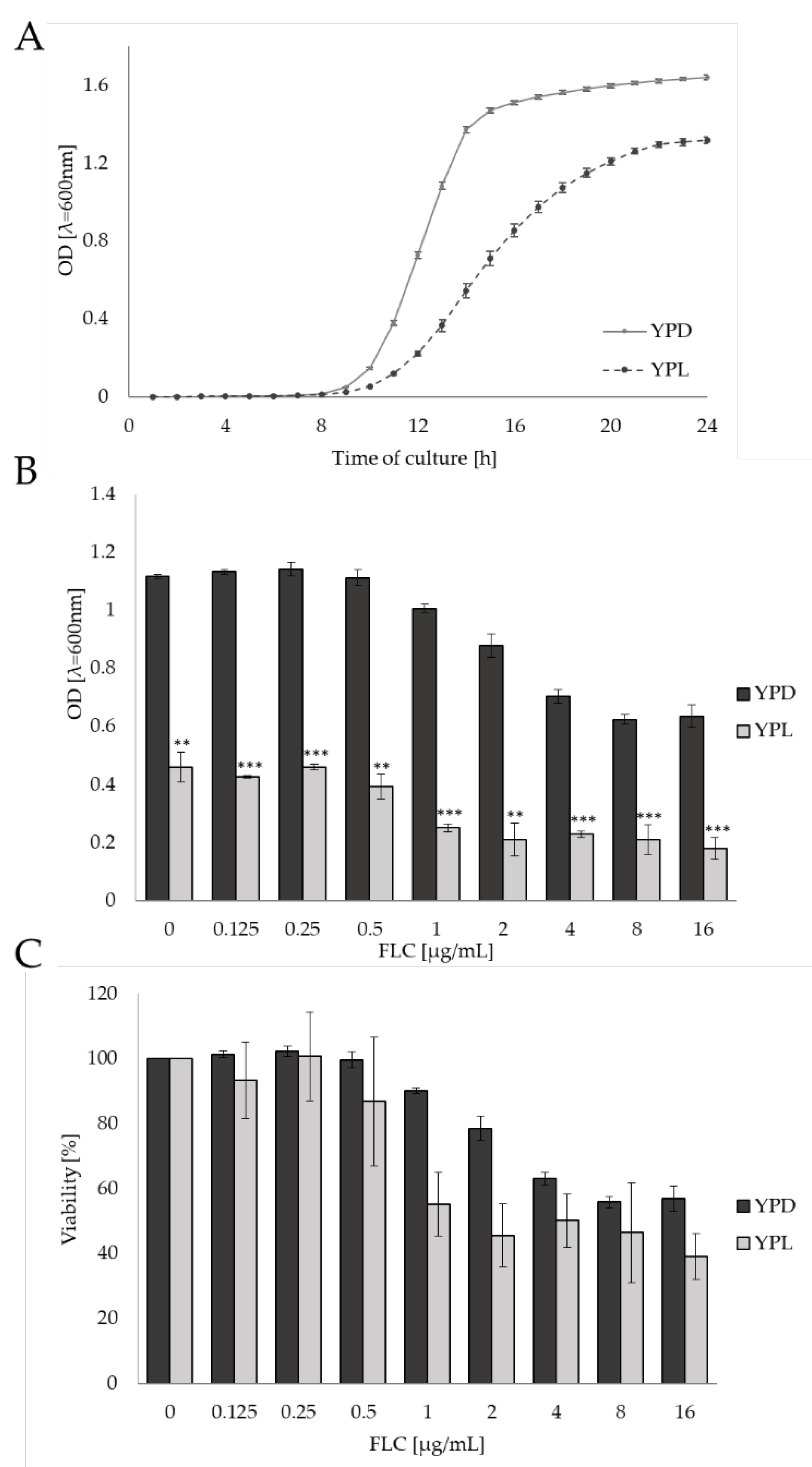
2.1. The Carbon Source Influences the Viability of C. albicans at Increasing Concentrations of Fluconazole
2.2. FLC and Lactate Synergistically Impact the Porosity of C. albicans CAF2-1
2.3. Agarose Gel Electrophoresis, Purification of the PCR Product and Sanger Sequencing
2.4. Identification of Point Mutations in ERG11 Gene
2.5. Identification of Amino Acid Substitutions as a Result of Aligning the Known Amino Acid Sequences of the Erg11 Protein of Selected Candida Species with the Amino Acid Sequence of the Erg11 Protein of C. albicans SC5314
3. Discussion
4. Materials and Methods
4.1. Materials
4.1.1. Strains and Growth Condition
4.1.2. Chemicals
4.2. Methods
4.2.1. Determination of C. albicans Growth Curve and Viability in Increasing Concentration of Fluconazole
4.2.2. Propidium Iodide (PI) Staining of C. albicans Cells
4.2.3. Cultures of C. albicans and Preparation of Material for DNA Isolation
4.2.4. DNA Isolation
4.2.5. Polymerase Chain Reaction (PCR)
4.2.6. Agarose Gel Electrophoresis, Purification of the PCR Product and Sanger Sequencing
4.2.7. The Identification of ERG11 Point Mutations among Amino Acid Sequences of C. albicans Strains Available in Databases
4.2.8. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mayer, F.L.; Wilson, D.; Hube, B. Candida albicans pathogenicity mechanisms. Virulence 2013, 4, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Lopes, J.P.; Lionakis, M.S. Pathogenesis and virulence of Candida albicans. Virulence 2022, 13, 89–121. [Google Scholar] [CrossRef] [PubMed]
- Pfaller, M.A.; Diekema, D.J. Epidemiology of invasive candidiasis: A persistent public health problem. Clin. Microbiol. Rev. 2007, 20, 133–163. [Google Scholar] [CrossRef] [PubMed]
- Quindós, G.; Miranda-Cadena, K.; San-Millán, R.; Borroto-Esoda, K.; Cantón, E.; Linares-Sicilia, M.J.; Hamprecht, A.; Montesinos, I.; Tortorano, A.M.; Prigitano, A.; et al. In vitro antifungal activity of ibrexafungerp (SCY-078) against contemporary blood isolates from medically relevant species of Candida: A european study. Front. Cell. Infect. Microbiol. 2022, 16, 906563. [Google Scholar] [CrossRef] [PubMed]
- Jallow, S.; Govender, N.P. Ibrexafungerp: A first-in-class oral triterpenoid glucan synthase inhibitor. J. Fungi 2021, 7, 163. [Google Scholar] [CrossRef]
- de Oliveira Santos, G.C.; Vasconcelos, C.C.; Lopes, A.J.O.; de Sousa Cartágenes, M.D.S.; Filho, A.K.D.B.; do Nascimento, F.R.F.; Ramos, R.M.; Pires, E.R.R.B.; de Andrade, M.S.; Rocha, F.M.G.; et al. Candida infections and therapeutic strategies: Mechanisms of action for traditional and alternative agents. Front. Microbiol. 2018, 3, 1351. [Google Scholar] [CrossRef]
- Flowers, S.A.; Colón, B.; Whaley, S.G.; Schuler, M.A.; Rogers, P.D. Contribution of clinically derived mutations in ERG11 to azole resistance in Candida albicans. Antimicrob. Agents Chemother. 2015, 59, 450–460. [Google Scholar] [CrossRef]
- Fumiyoshi, A.; Toshiki, H. Mechanistic role of ergosterol in membrane rigidity and cycloheximide resistance in Saccharomyces cerevisiae. Biochim. Biophys. Acta (BBA)-Biomembr. 2009, 1788, 743–752. [Google Scholar]
- Dupont, S.; Lemetais, G.; Ferreira, T.; Cayot, P.; Gervais, P.; Beney, L. Ergosterol biosynthesis: A fungal pathway for life on land? Evolution 2012, 66, 2961–2968. [Google Scholar] [CrossRef]
- Bhattacharya, S.; Sae-Tia, S.; Fries, B.C. Candidiasis and mechanisms of antifungal resistance. Antibiotics 2020, 9, 312. [Google Scholar] [CrossRef]
- Lee, Y.; Puumala, E.; Robbins, N.; Cowen, L.E. Antifungal drug resistance: Molecular mechanisms in Candida albicans and beyond. Chem. Rev. 2021, 121, 3390–3411. [Google Scholar] [CrossRef] [PubMed]
- Xiang, M.J.; Liu, J.Y.; Ni, P.H.; Wang, S.; Shi, C.; Wei, B.; Ni, Y.X.; Ge, H.L. Erg11 mutations associated with azole resistance in clinical isolates of Candida albicans. FEMS Yeast Res. 2013, 13, 386–393. [Google Scholar] [CrossRef] [PubMed]
- Marichal, P.; Koymans, L.; Willemsens, S.; Bellens, D.; Verhasselt, P.; Luyten, W.; Borgers, M.; Ramaekers, F.C.S.; Odds, F.C.; Vanden Bossche, H. Contribution of mutations in the cytochrome P450 14alpha-demethylase (Erg11p, Cyp51p) to azole resistance in Candida albicans. Microbiology 1999, 145, 2701–2713. [Google Scholar] [CrossRef] [PubMed]
- Hargrove, T.Y.; Friggeri, L.; Wawrzak, Z.; Qi, A.; Hoekstra, W.J.; Schotzinger, R.J.; York, J.D.; Guengerich, F.P.; Lepesheva, G.I. Structural analyses of Candida albicans sterol 14α-demethylase complexed with azole drugs address the molecular basis of azole-mediated inhibition of fungal sterol biosynthesis. J. Biol. Chem. 2017, 292, 6728–6743. [Google Scholar] [CrossRef]
- Zeise, K.D.; Woods, R.J.; Huffnagle, G.B. Interplay between Candida albicans and lactic acid bacteria in the gastrointestinal tract: Impact on colonization resistance, microbial carriage, opportunistic infection, and host immunity. Clin. Microbiol. Rev. 2021, 34, 0032320. [Google Scholar] [CrossRef]
- Liang, W.; Guan, G.; Dai, Y.; Cao, C.; Tao, L.; Du, H.; Nobile, C.J.; Zhong, J.; Huang, G. Lactic acid bacteria differentially regulate filamentation in two heritable cell types of the human fungal pathogen Candida albicans. Mol. Microbiol. 2016, 102, 506–519. [Google Scholar]
- Lourenço, A.; Pedro, N.A.; Salazar, S.B.; Mira, N.P. Effect of acetic acid and lactic acid at low pH in growth and azole resistance of Candida albicans and Candida glabrata. Front. Microbiol. 2019, 9, 3265. [Google Scholar] [CrossRef]
- Suchodolski, J.; Muraszko, J.; Bernat, P.; Krasowska, A. Lactate like fluconazole reduces ergosterol content in the plasma membrane and synergistically kills Candida albicans. Int. J. Mol. Sci. 2021, 22, 5219. [Google Scholar] [CrossRef]
- Dadar, M.; Tiwari, R.; Karthik, K.; Chakraborty, S.; Shahali, Y.; Dhama, K. Candida albicans—Biology, molecular characterization, pathogenicity, and advances in diagnosis and control—An update. Microb. Pathog. 2018, 117, 128–138. [Google Scholar] [CrossRef]
- Patil, S.; Rao, R.S.; Majumdar, B.; Anil, S. Clinical appearance of oral Candida infection and therapeutic strategies. Front. Microbiol. 2015, 6, 1391. [Google Scholar] [CrossRef]
- Howard, K.C.; Dennis, E.K.; Watt, D.S.; Garneau-Tsodikova, S. A comprehensive overview of the medicinal chemistry of antifungal drugs: Perspectives and promise. Chem. Soc. Rev. 2020, 49, 2426–2480. [Google Scholar] [CrossRef] [PubMed]
- Pricopie, A.I.; Focșan, M.; Ionuț, I.; Marc, G.; Vlase, L.; Găină, L.I.; Vodnar, D.C.; Elemer, S.; Barta, G.; Pîrnău, A.; et al. Novel 2,4-disubstituted-1,3-thiazole derivatives: Synthesis, anti-Candida activity evaluation and interaction with bovine serum albumine. Molecules 2020, 25, 1079. [Google Scholar]
- Lee, M.R.; Raman, N.; Ortiz-Bermúdez, P.; Lynn, D.M.; Palecek, S.P. 14-helical β-peptides elicit toxicity against C. albicans by forming pores in the cell membrane and subsequently disrupting intracellular organelles. Cell Chem. Biol. 2019, 26, 289–299. [Google Scholar] [CrossRef] [PubMed]
- Ene, I.V.; Adya, A.K.; Wehmeier, S.; Brand, A.C.; MacCallum, D.M.; Gow, N.A.; Brown, A.J. Host carbon sources modulate cell wall architecture, drug resistance and virulence in a fungal pathogen. Cell. Microbiol. 2012, 14, 1319–1335. [Google Scholar] [CrossRef] [PubMed]
- Alves, R.; Mota, S.; Silva, S.; Rodrigues, C.F.; Brown, A.J.P.; Henriques, M.; Casal, M.; Paiva, S. The carboxylic acid transporters Jen1 and Jen2 affect the architecture and fluconazole susceptibility of Candida albicans biofilm in the presence of lactate. Biofouling 2017, 33, 943–954. [Google Scholar] [CrossRef] [PubMed]
- Flint, H.J.; Scott, K.P.; Louis, P.; Duncan, S.H. The role of the gut microbiota in nutrition and health. Nat. Rev. Gastroenterol. Hepatol. 2012, 9, 577–589. [Google Scholar] [CrossRef]
- Lohse, M.B.; Gulati, M.; Johnson, A.D.; Nobile, C.J. Development and regulation of single- and multi-species Candida albicans biofilms. Nat. Rev. Microbiol. 2018, 16, 19–31. [Google Scholar] [CrossRef]
- Winter, M.B.; Salcedo, E.C.; Lohse, M.B.; Hartooni, N.; Gulati, M.; Sanchez, H.; Takagi, J.; Hube, B.; Andes, D.R.; Johnson, A.D.; et al. Global identification of biofilm-specific proteolysis in Candida albicans. mBio 2016, 7, e01514-16. [Google Scholar] [CrossRef]
- Ballou, E.R.; Avelar, G.M.; Childers, D.S.; Mackie, J.; Bain, J.M.; Wagener, J.; Kastora, S.L.; Panea, M.D.; Hardison, S.E.; Walker, L.A.; et al. Lactate signalling regulates fungal β-glucan masking and immune evasion. Nat. Microbiol. 2016, 12, 16238. [Google Scholar] [CrossRef]
- Oliveira-Pacheco, J.; Alves, R.; Costa-Barbosa, A.; Cerqueira-Rodrigues, B.; Pereira-Silva, P.; Paiva, S.; Silva, S.; Henriques, M.; Pais, C.; Sampaio, P. The role of Candida albicans transcription factor RLM1 in response to carbon adaptation. Front. Microbiol. 2018, 29, 1127. [Google Scholar] [CrossRef]
- Dalle, F.; Wächtler, B.; L’Ollivier, C.; Holland, G.; Bannert, N.; Wilson, D.; Labruère, C.; Bonnin, A.; Hube, B. Cellular interactions of Candida albicans with human oral epithelial cells and enterocytes. Cell. Microbiol. 2010, 12, 248–271. [Google Scholar] [CrossRef] [PubMed]
- Ehrström, S.; Daroczy, K.; Rylander, E.; Samuelsson, C.; Johannesson, U.; Anzén, B.; Påhlson, C. Lactic acid bacteria colonization and clinical outcome after probiotic supplementation in conventionally treated bacterial vaginosis and vulvovaginal candidiasis. Microb. Infect. 2010, 12, 691–699. [Google Scholar] [CrossRef] [PubMed]
- White, T.C.; Holleman, S.; Dy, F.; Mirels, L.F.; Stevens, D.A. Resistance mechanisms in clinical isolates of Candida albicans. Antimicrob. Agents Chemother. 2002, 46, 1704–1713. [Google Scholar] [CrossRef]
- Asai, K.; Tsuchimori, N.; Okonogi, K.; Perfect, J.R.; Gotoh, O.; Yoshida, Y. Formation of azole-resistant Candida albicans by mutation of sterol 14-demethylase P450. Antimicrob. Agents Chemother. 1999, 43, 1163–1169. [Google Scholar] [CrossRef] [PubMed]
- Morio, F.; Loge, C.; Besse, B.; Hennequin, C.; Le Pape, P. Screening for amino acid substitutions in the Candida albicans Erg11 protein of azole-susceptible and azole-resistant clinical isolates: New substitutions and a review of the literature. Diagn. Microbiol. Infect. Dis. 2010, 66, 373–384. [Google Scholar] [CrossRef]
- Benedetti, V.P.; Savi, D.C.; Aluizio, R.; Adamoski, D.; Kava, V.; Galli-Terasawa, L.V.; Glienke, C. ERG11 gene polymorphisms and susceptibility to fluconazole in Candida isolates from diabetic and kidney transplant patients. Rev. Soc. Bras. Med. Trop. 2019, 52, 20180473. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.J.; Kim, Y.J.; Yong, D.; Byun, J.H.; Kim, T.S.; Chang, Y.S.; Choi, M.J.; Byeon, S.A.; Won, E.J.; Kim, S.H.; et al. Fluconazole-Resistant Candida parapsilosis bloodstream isolates with Y132F mutation in ERG11 gene, South Korea. Emerg. Infect. Dis. 2018, 24, 1768–1770. [Google Scholar] [CrossRef] [PubMed]
- Rizzato, C.; Poma, N.; Zoppo, M.; Posteraro, B.; Mello, E.; Bottai, D.; Lupetti, A.; Sanguinetti, M.; Tavanti, A. CoERG11 A395T mutation confers azole resistance in Candida orthopsilosis clinical isolates. J. Antimicrob. Chemother. 2018, 73, 1815–1822. [Google Scholar] [CrossRef] [PubMed]
- Castanheira, M.; Deshpande, L.M.; Messer, S.A.; Rhomberg, P.R.; Pfaller, M.A. Analysis of global antifungal surveillance results reveals predominance of Erg11 Y132F alteration among azole-resistant Candida parapsilosis and Candida tropicalis and country-specific isolate dissemination. Int. J. Antimicrob. Agents 2020, 55, 105799. [Google Scholar] [CrossRef] [PubMed]
- Vijayaraman, R.S.; Kalva, S.; Ramaraj, V.K.; Kindo, A.J. Interaction of fluconazole with Candida auris and Candida albicans lanosterol 14 α-demethylase (Erg11). RJLBPCS 2019, 5, 787–798. [Google Scholar]
- Vandeputte, P.; Larcher, G.; Bergès, T.; Renier, G.; Chabasse, D.; Bouchara, J.P. Mechanisms of azole resistance in a clinical isolate of Candida tropicalis. Antimicrob. Agents Chemother. 2005, 49, 4608–4615. [Google Scholar] [CrossRef] [PubMed]
- Manastır, L.; Ergon, M.C.; Yucesoy, M. Investigation of mutations in Erg11 gene of fluconazole resistant Candida albicans isolates from Turkish hospitals. Mycoses 2011, 54, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Pradhan, A.; Avelar, G.M.; Bain, J.M.; Childers, D.; Pelletier, C.; Larcombe, D.E.; Shekhova, E.; Netea, M.G.; Brown, G.D.; Erwig, L. Non-canonical signalling mediates changes in fungal cell wall PAMPs that drive im-mune evasion. FEMS Yeast Res. 2019, 10, 5315. [Google Scholar]
- Xu, Y.; Chen, L.; Li, C. Susceptibility of clinical isolates of Candida species to fluconazole and detection of Candida albicans ERG11 mutations. J. Antimicrob. Chemother. 2008, 61, 798–804. [Google Scholar] [CrossRef] [PubMed]
- Strzelczyk, J.K.; Slemp-Migiel, A.; Rother, M.; Gołąbek, K.; Wiczkowski, A. Nucleotide substitutions in the Candida albicans ERG11 gene of azole-susceptible and azole-resistant clinical isolates. Acta Biochim. Pol. 2013, 60, 547–552. [Google Scholar] [CrossRef]
- Derkacz, D.; Bernat, P.; Krasowska, A. K143R Amino Acid Substitution in 14-α-Demethylase (Erg11p) Changes Plasma Membrane and Cell Wall Structure of Candida albicans. Int. J. Mol. Sci. 2022, 23, 1631. [Google Scholar] [CrossRef]
- Carolus, H.; Pierson, S.; Lagrou, K.; Van Dijck, P. Amphotericin B and other polyenes-discovery, clinical use, mode of action and drug resistance. J. Fungi 2020, 6, 321. [Google Scholar] [CrossRef]
- Fonzi, W.A.; Irwin, M.Y. Isogenic strain construction and gene mapping in Candida albicans. Genetics 1993, 134, 717–728. [Google Scholar] [CrossRef]
- Suchodolski, J.; Feder-Kubis, J.; Krasowska, A. Antifungal activity of ionic liquids based on (−)-menthol: A mechanism study. Microbiol. Res. 2017, 197, 56–64. [Google Scholar] [CrossRef]
- Dal Mas, C.; Rossato, L.; Shimizu, T.; Oliveira, E.B.; da Silva Junior, P.I.; Meis, J.F.; Colombo, A.L.; Hayashi, M.A.F. Effects of the natural peptide crotamine from a South American rattlesnake on Candida auris, an emergent multidrug antifungal resistant human pathogen. Biomolecules 2019, 9, 205. [Google Scholar] [CrossRef]
- Pfaller, M.A.; Messer, S.A.; Gee, S.; Joly, S.; Pujol, C.; Sullivan, D.J.; Coleman, D.C.; Soll, D.R. In vitro susceptibilities of Candida dubliniensis isolates tested against the new triazole and echinocandin antifungal agents. J. Clin. Microbiol. 1999, 37, 870–872. [Google Scholar] [CrossRef] [PubMed]
- Marcos-Arias, C.; Eraso, E.; Madariaga, L.; Quindós, G. In vitro activities of natural products against oral Candida isolates from denture wearers. BMC Complement. Altern. Med. 2011, 11, 119. [Google Scholar] [CrossRef] [PubMed]
- Borst, A.; Raimer, M.T.; Warnock, D.W.; Morrison, C.J.; Arthington-Skaggs, B.A. Rapid acquisition of stable azole resistance by Candida glabrata isolates obtained before the clinical introduction of fluconazole. Antimicrob. Agents Chemother. 2005, 49, 783–787. [Google Scholar] [CrossRef] [PubMed]
- Shimokawa, O.; Nakayama, H. Estimation of minimum sterol 14α demethylation-inhibitory concentration of azoles in Candida yeasts using acetate mediated growth inhibition: Potential utility in susceptibility testing. J. Clin. Microbiol. 2000, 38, 2893–2896. [Google Scholar] [CrossRef]
- Costa, L.J.; Birman, E.G.; Alves, S.H.; Cury, A.E. Antifungal susceptibility of Candida albicans isolated from oral mucosa of patients with cancer. Rev. Odontol. Univ. São Paulo 1999, 13, 219–223. [Google Scholar] [CrossRef]
- Asadzadeh, M.; Ahmad, S.; Al-Sweih, N.; Khan, Z. Epidemiology and molecular basis of resistance to fluconazole among clinical Candida parapsilosis isolates in Kuwait. Microb. Drug Resist. 2017, 23, 966–972. [Google Scholar] [CrossRef] [PubMed]
- Barchiesi, F.; Calabrese, D.; Sanglard, D.; Falconi Di Francesco, L.; Caselli, F.; Giannini, D.; Giacometti, A.; Gavaudan, S.; Scalise, G. Experimental induction of fluconazole resistance in Candida tropicalis ATCC 750. Antimicrob. Agents Chemother. 2000, 44, 1578–1584. [Google Scholar] [CrossRef]
- Wang, B.; Huang, L.H.; Zhao, J.X.; Wei, M.; Fang, H.; Wang, D.Y.; Wang, H.F.; Yin, J.G.; Xiang, M. ERG11 mutations associated with azole resistance in Candida albicans isolates from vulvovaginal candidosis patients. Asian Pac. J. Trop. Biomed. 2015, 5, 909–914. [Google Scholar] [CrossRef]
- Ge, S.H.; Wan, Z.; Li, J.; Xu, J.; Li, R.Y.; Bai, F.Y. Correlation between azole susceptibilities, genotypes, and ERG11 mutations in Candida albicans isolates associated with vulvovaginal candidiasis in China. Antimicrob. Agents Chemother. 2010, 54, 3126–3131. [Google Scholar] [CrossRef]


| Concentration of Fluconazole [µg/mL] | |||||||
|---|---|---|---|---|---|---|---|
| 0 | 0.25 | 1 | 4 | 16 | |||
| Carbon source | Glucose | 8 h | C30T, T819M, T826G, A828T | C30T, A31G, T37A, T819M, T826G, A828T | A105T, C746A, A751T, T826G, A828T, T1558C | A35T, T37A, A797G, T826G, A828T, T1558C | C30A, A31G, T37A, A383C, T826G, A828T, T1470C, T1558C |
| 14 h | C30T, A35T, T37A, A790G, G791A, G794A | A35T, T37A, A657G, T768A, T826G, A828T, T1470C | A35T, A105T, C746A, T826G, A828T | T33G, A35T, A383C, A750T, T826G, A828T, T1470C, T1558C | T27G, C30T, A34T, A357C, A383C, T826G, A828T, T1470C, T1558C | ||
| 24 h | C30W, A31G, T32S, A90G, T92G, T826G, A828T | C30W, A31G, T32S, T822M, G823Y, A824G, T826G, A828T | T33G, A35T, T37A, G673C, T826G, A828T, A1440G, A1516G | A31G, T32S, G791A, G796A, A797G, T826G, A828T, A1516G | C30T, T32S, A35T, T826G, A828T, T1470A, A1516G | ||
| Lactate | 8 h | A31G, T32S, T826G, A828T | T32G, T33S, A34T, T37A, C799R, T826G, A828T | C30W, A31G, T32S, T826G, A828T | G29W, A35T, T37A, T826G, A828T, T1470A | C30A, A31S, T36A, T826G, A828T, T1470C | |
| 14 h | T33G, A34T, G791A, A792G, G794A, T826G, A828T | T33G, A34T, T826G, A828T, T1470C | A34T, G673T, T826G, A828T | C30T, A34T, A383C, T826G, A828T, T1470C | T26G, C30T, A383C, A776T, A790G, T826G, A828T | ||
| 24 h | A31G, T32S, A790G, T826G, A828T, A1440G, T1470C | C658T, A776T, G794A, T822G, T826G, A828T | A31G, T33G, A793G, T826G, A828T | A31G, T32S, A383C, A780C, A785T, T826G, A828T | G28A, A35T, T36A, A383C, A780C, G796A, A798G, C799A, T826G, A828T, A1551T | ||
| Concentration of Fluconazole [µg/mL] | |||||||
|---|---|---|---|---|---|---|---|
| 0 | 0.25 | 1 | 4 | 16 | |||
| Carbon source | Glucose | 8 h | N273K, L276V | I11V, Y13N, N273K, L276V | L35F, A249D, K251STOP, L276V, W520R | N12I, Y13N, E266G, L276V, W520R | I11V, Y13N, K128T, D275G, L276V, W520R |
| 14 h | N12I, Y13N, R264E, R265K | N12I, Y13N, L276V | N12I, L35F, A249D, L276V | I11M, N12I, K128T, Q250H, L276V, W520R | D9E, N12Y, K119N, K128T, L276V, W520R | ||
| 24 h | I11A/I11G, F31V, L276V | I11A/I11G, D275R/D275C, L276V | I11M, N12I, Y13N, D225H, L276V, S506G | I11A/I11G, R265K, E266R, L276V, S506G | I11T/I11S, N12I, L276V, N490K, S506G | ||
| Lactate | 8 h | I11A/I11G, L276V | I11S/I11R, N12Y, Y13N, R267S/R267G, L276V | I11A/I11G, L276V | G10D/G10V, N12I, Y13N, L276V, N490K | I11L/I11V, N12L, L276V | |
| 14 h | I11M, N12Y, R265G, R265K, L276V | I11M, N12Y, L276V | N12Y, D225Y, L276V | N12Y, K128T, L276V | D9Q, N12Y, K128T, K259I, R265G, L276V | ||
| 24 h | I11A/I11G, R265G, L276V | K259I, R265G, L276V | I11V, R265G, L276V | I11A/I11G, K128T, E260D, K262I, L276V | G10S, N12I, K128T, E260D, E266K, R267S, L276V, E517D | ||
| Fragment | Name | Sequence | Product Size |
|---|---|---|---|
| A | ERG11-FA | 5′- ATGGCTATTGTTGAAACTGTC-3′ | 785 bp |
| ERG11-RA | 5′- CGTTCTCTTCTCAGTTTAATTTC-3′ | ||
| B | ERG11-FB | 5′- GAAGAGAACGTGGTGATATTGATC-3′ | 826 bp |
| ERG11-RB | 5′- CACTGAATCGAAAGAAAGTTGCC-3′ |
| Step | Program | Cycles | |||
|---|---|---|---|---|---|
| ZL-ERG11A | ZL-ERG11B | ||||
| Time | Temperature | Time | Temperature | ||
| Initial denaturation | 5 min | 98 °C | 5 min | 98 °C | - |
| Denaturation | 30 s | 98 °C | 28 s | 98 °C | 35 |
| Annealing | 30 s | 51 °C | 29 s | 55 °C | |
| Extension | 30 s | 72 °C | 30 s | 72 °C | |
| Final extension | 5 min | 72 °C | 5 min | 72 °C | - |
| ∞ | 4 °C | ∞ | 4 °C | - | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Urbanek, A.K.; Łapińska, Z.; Derkacz, D.; Krasowska, A. The Role of ERG11 Point Mutations in the Resistance of Candida albicans to Fluconazole in the Presence of Lactate. Pathogens 2022, 11, 1289. https://doi.org/10.3390/pathogens11111289
Urbanek AK, Łapińska Z, Derkacz D, Krasowska A. The Role of ERG11 Point Mutations in the Resistance of Candida albicans to Fluconazole in the Presence of Lactate. Pathogens. 2022; 11(11):1289. https://doi.org/10.3390/pathogens11111289
Chicago/Turabian StyleUrbanek, Aneta K., Zofia Łapińska, Daria Derkacz, and Anna Krasowska. 2022. "The Role of ERG11 Point Mutations in the Resistance of Candida albicans to Fluconazole in the Presence of Lactate" Pathogens 11, no. 11: 1289. https://doi.org/10.3390/pathogens11111289
APA StyleUrbanek, A. K., Łapińska, Z., Derkacz, D., & Krasowska, A. (2022). The Role of ERG11 Point Mutations in the Resistance of Candida albicans to Fluconazole in the Presence of Lactate. Pathogens, 11(11), 1289. https://doi.org/10.3390/pathogens11111289








