Expression of Genes Involved in Anthracnose Resistance in Chili (Capsicum baccatum) ‘PBC80’-Derived Recombinant Inbred Lines
Abstract
:1. Introduction
2. Materials and Methods
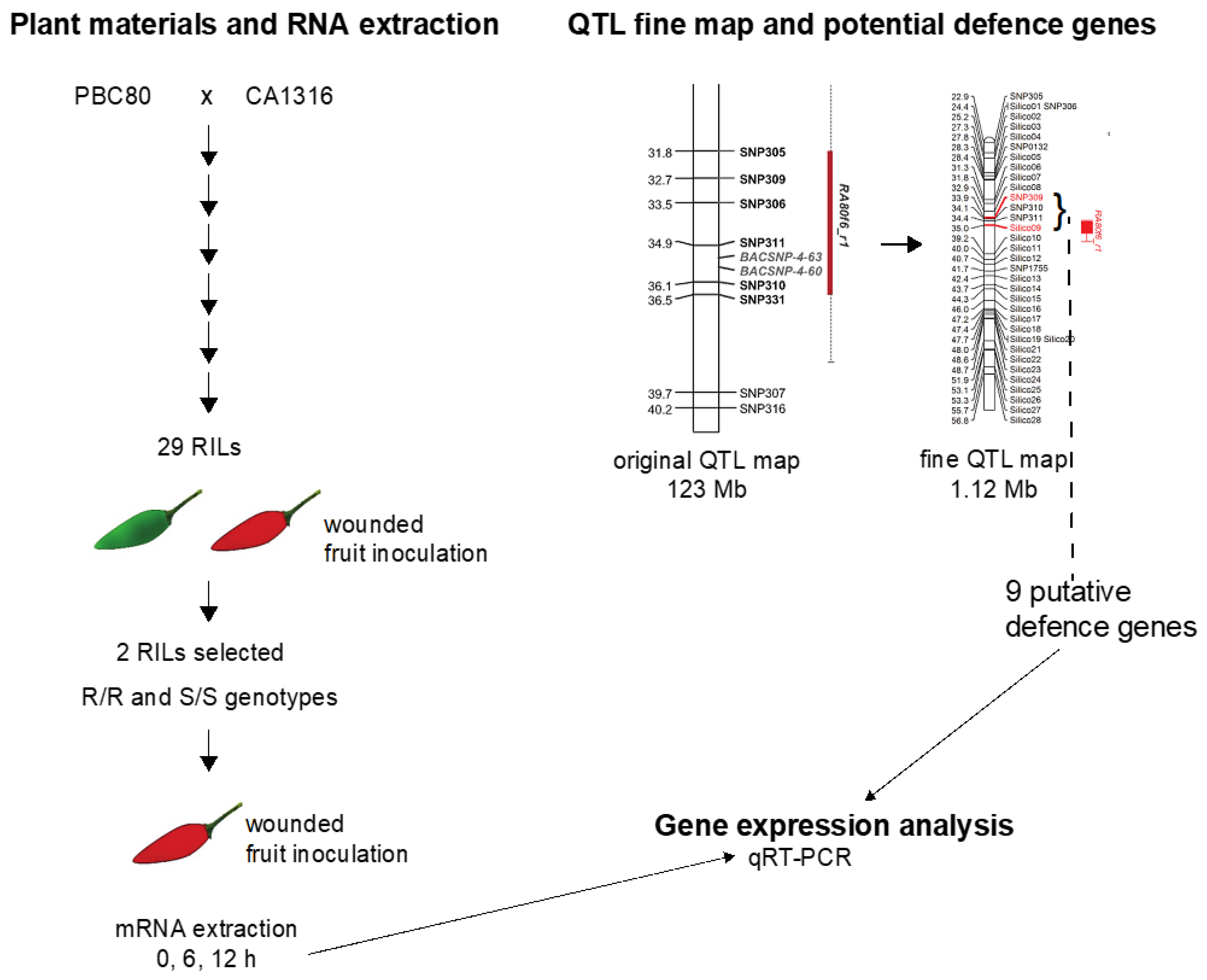
2.1. Fine Mapping in the QTL Region and Selection of Defense-Related Genes
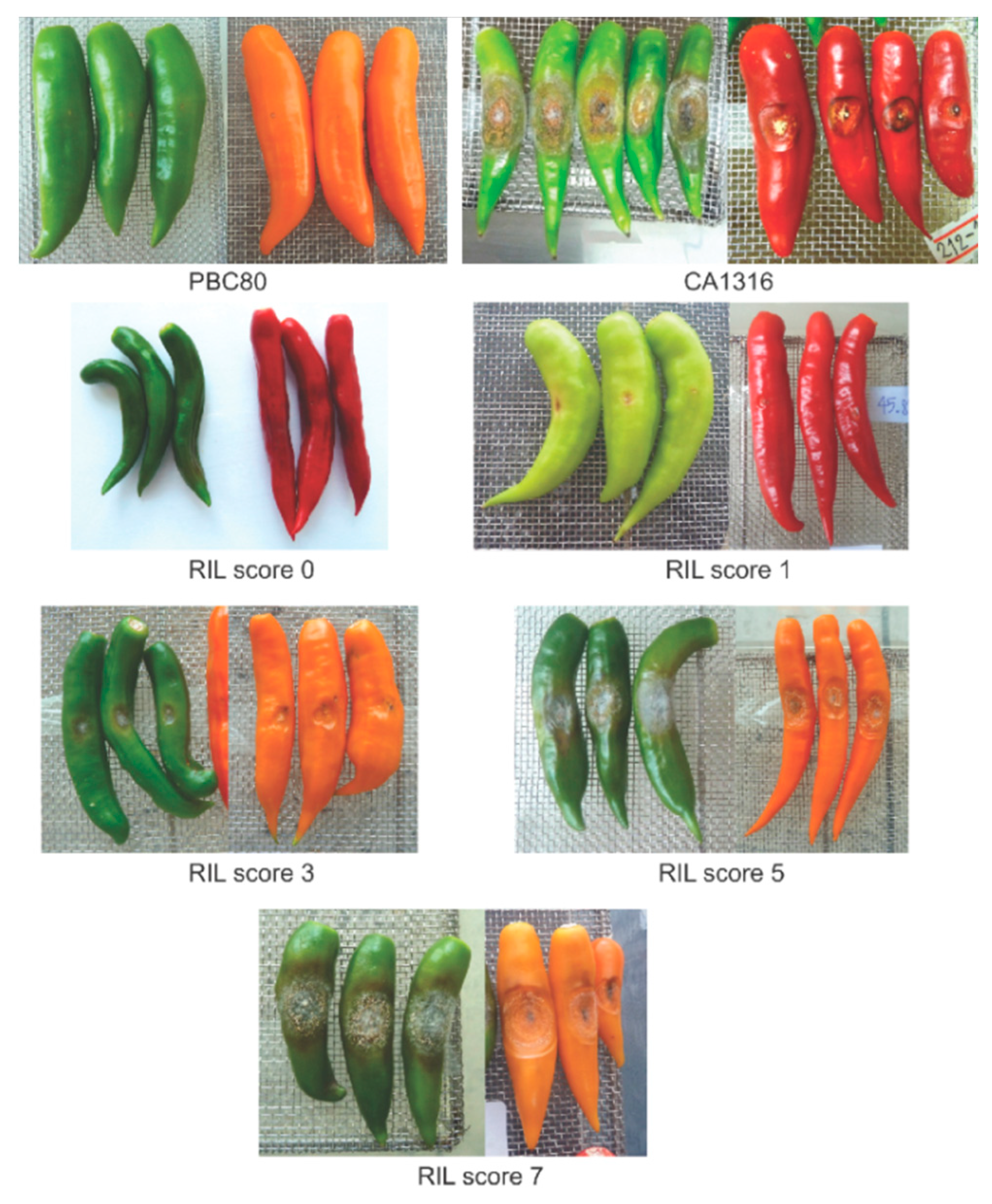
2.2. Selection of RILs with Resistance and Susceptibility to Anthracnose
2.3. Expression Study of the Putative Defense-Related Genes
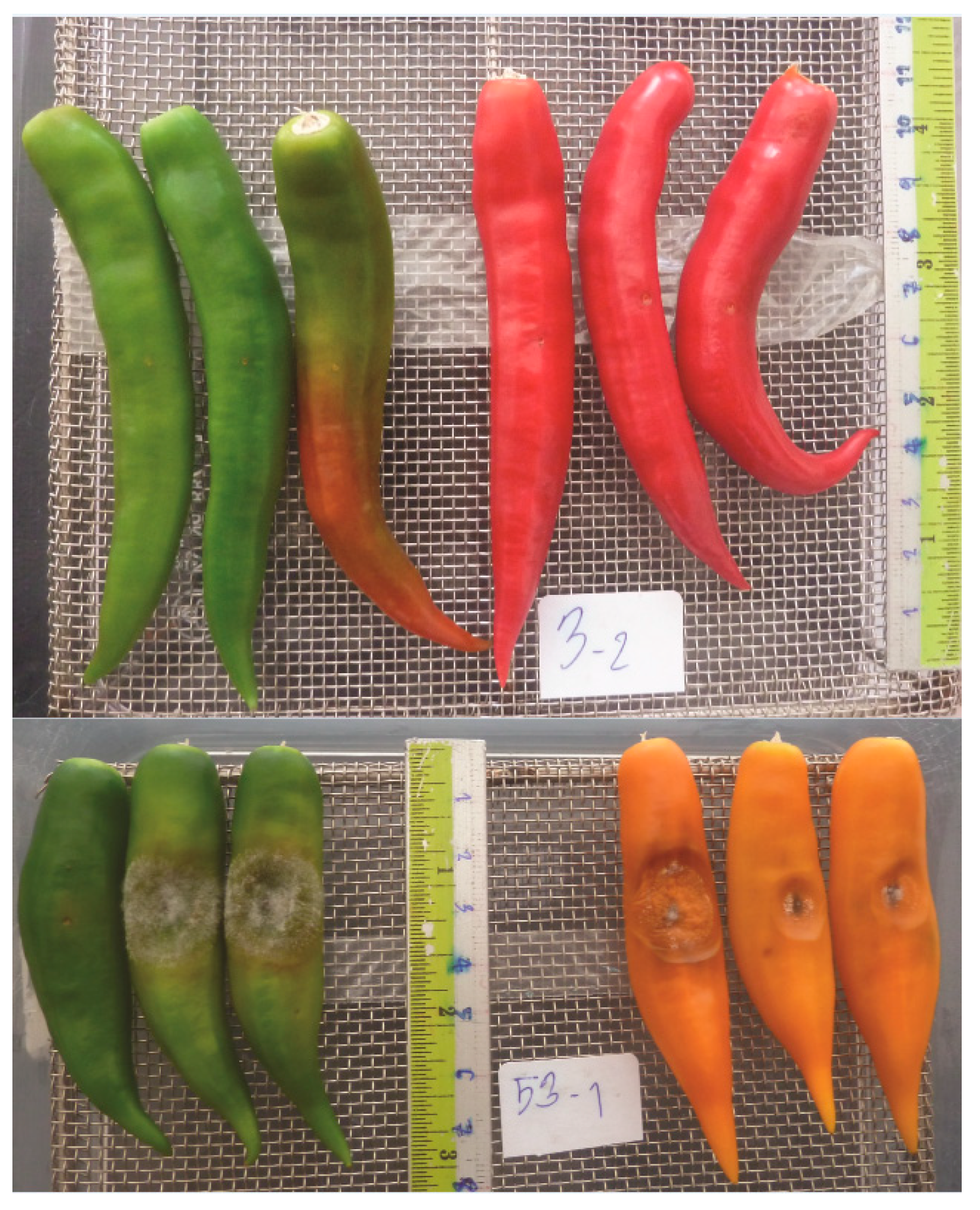
2.3.1. Fruit Inoculation
2.3.2. RNA Extraction and cDNA Synthesis
2.3.3. Designing the Primers Specific to the Targeted Defense-Related Genes
2.3.4. Gene Expression Analysis with qRT-PCR
3. Results
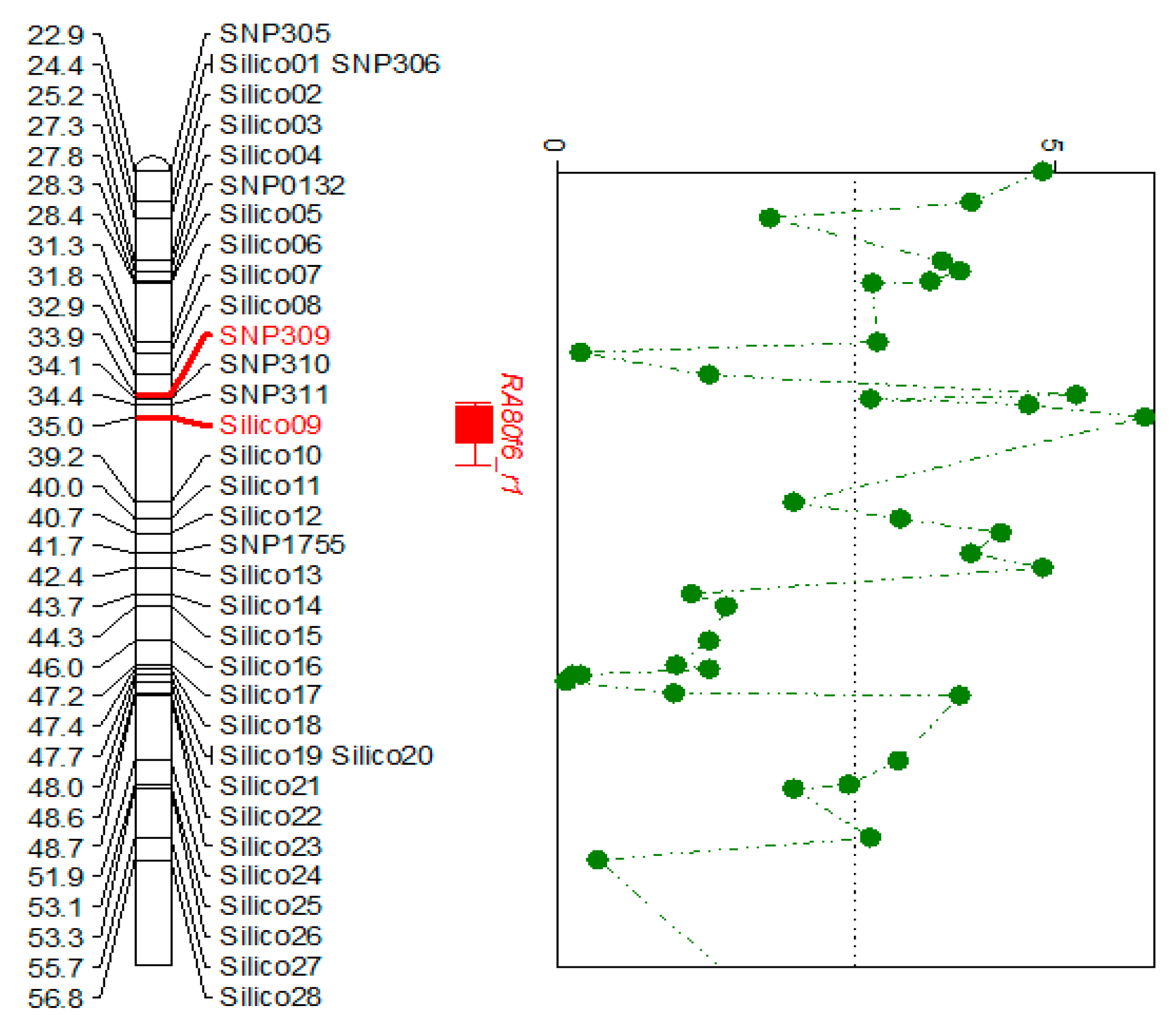
3.1. Fine Mapping and Defense-Related Gene Selection
3.2. Selection of the RILs with Resistance and Susceptibility to Anthracnose
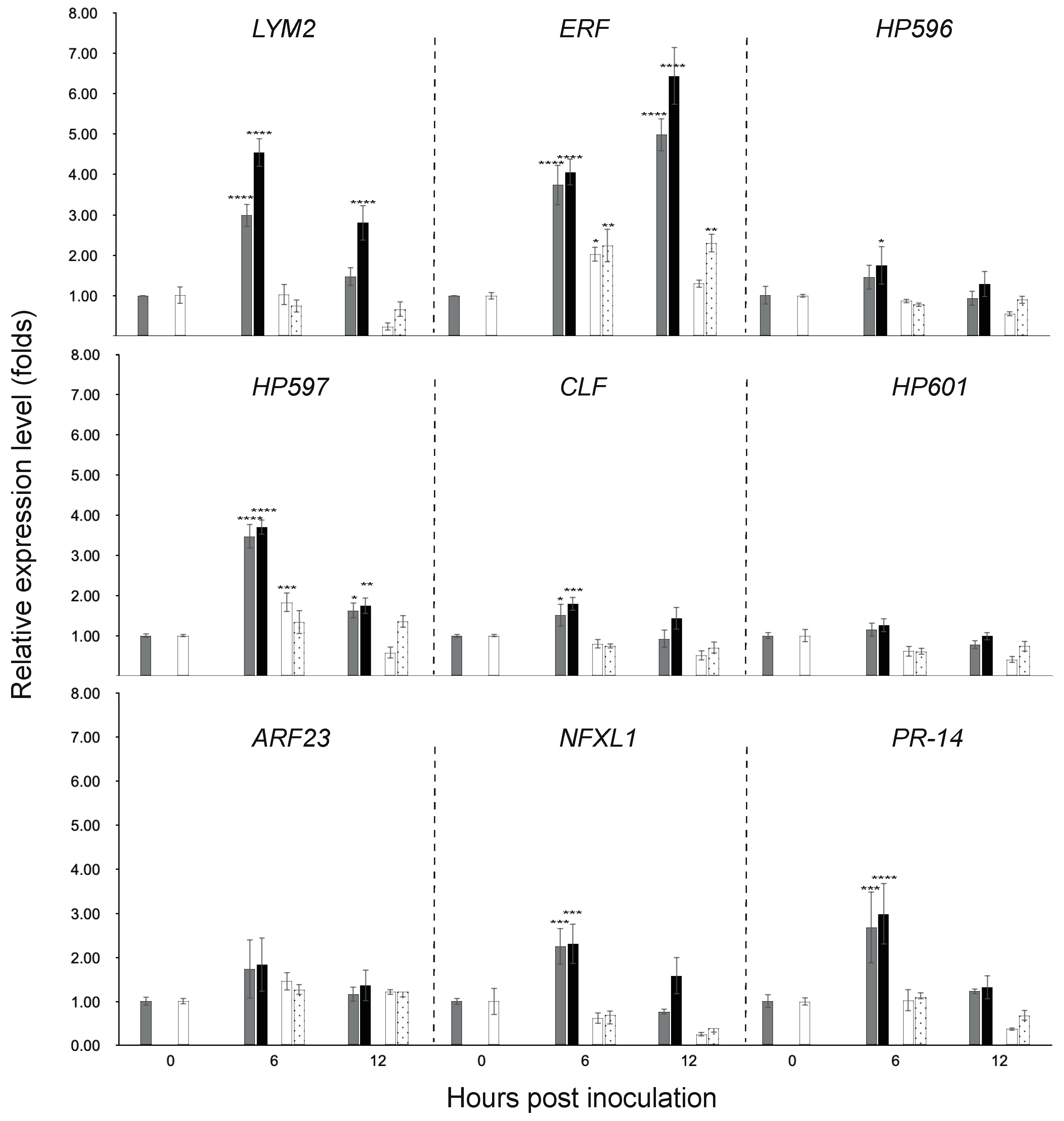
3.3. Gene Expression in the Chili Fruit Post Inoculation
4. Discussion
4.1. Defense Roles for the Identified Genes
4.2. Wound Response in Plant Defense Mechanism
4.3. Speculative Roles of LYM2 in HR
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mongkolporn, O. Capsicum: Breeding Strategies for Anthracnose Resistance; CRC Press: Boca Raton, FL, USA, 2018. [Google Scholar]
- Mongkolporn, O.; Taylor, P.W.J. Chili anthracnose: Colletotrichum taxonomy and pathogenicity. Plant Pathol. 2018, 67, 1255–1263. [Google Scholar] [CrossRef]
- Mongkolporn, O.; Montri, P.; Supakaew, T.; Taylor, P.W.J. Differential reactions on mature green and ripe chili fruit infected by three Colletotrichum species. Plant Dis. 2010, 94, 306–310. [Google Scholar] [CrossRef] [PubMed]
- St Clair, D.A. Quantitative disease resistance and quantitative resistance Loci in breeding. Annu. Rev. Phytopathol. 2010, 48, 247–268. [Google Scholar] [CrossRef] [PubMed]
- Pilet-Nayel, M.-L.; Moury, B.; Caffier, V.; Montarry, J.; Kerlan, M.-C.; Fournet, S.; Durel, C.-E.; Delourme, R. Quantitative resistance to plant pathogens in pyramiding strategies for durable crop protection. Front Plant Sci. 2017, 8, 1838. [Google Scholar] [CrossRef]
- Mahasuk, P.; Khumpeng, N.; Wasee, S.; Taylor, P.W.J.; Mongkolporn, O. Inheritance of resistance to anthracnose (Colletotrichum capsici) at seedling and fruiting stages in chili pepper (Capsicum spp.). Plant Breed. 2009, 128, 701–706. [Google Scholar] [CrossRef]
- Mahasuk, P.; Taylor, P.W.J.; Mongkolporn, O. Identification of two new genes conferring resistance to Colletotrichum acutatum in Capsicum baccatum L. Phytopathology 2009, 99, 1100–1104. [Google Scholar] [CrossRef]
- Mahasuk, P.; Chinthaisong, J.; Mongkolporn, O. Differential resistances to anthracnose in Capsicum baccatum as responding to two Colletotrichum pathotypes and inoculation methods. Breed. Sci. 2013, 63, 333–338. [Google Scholar] [CrossRef]
- Mahasuk, P.; Struss, D.; Mongkolporn, O. QTLs for resistance to anthracnose identified in two Capsicum sources. Mol. Breed. 2016, 36, 10. [Google Scholar] [CrossRef]
- Collard, B.C.; Jahufer, M.Z.Z.; Brouwer, J.B.; Pang, E.C.K. An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: The basic concepts. Euphytica 2005, 142, 169–196. [Google Scholar] [CrossRef]
- Kethom, W.; Mongkolporn, O. New QTLs for anthracnose resistance identified in Capsicum baccatum ‘PBC80’-derived recombinant inbred lines. Euphytica 2021, 217, 218. [Google Scholar] [CrossRef]
- Pitsili, E.; Phukan, U.J.; Coll, N.S. Cell death in plant immunity. Cold Spring Harb. Perspect. Biol. 2020, 12, a036483. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Liu, Y.; Wang, J.; He, M.; Zhou, X.; Yang, C.; Yuan, C.; Wang, J.; Chern, M.; Yin, J.; et al. The durably resistant rice cultivar Digu activates defence gene expression before the full maturation of Magnaporthe oryzae appressorium. Mol. Plant Pathol. 2016, 17, 354–368. [Google Scholar] [CrossRef] [PubMed]
- Santos, M.d.L.; de Resende, M.L.V.; dos Santos Ciscon, B.A.; Freitas, N.C.; Pereira, M.H.d.B.; Reichel, T.; Mathioni, S.M. LysM receptors in Coffea arabica: Identification, characterization, and gene expression in response to Hemileia vastatrix. PLoS ONE 2022, 17, e0258838. [Google Scholar] [CrossRef] [PubMed]
- Auyong, A.S.; Ford, R.; Taylor, P.W. The role of cutinase and its impact on pathogenicity of Colletotrichum truncatum. J. Plant Pathol. Microbiol. 2015, 6, 259. [Google Scholar] [CrossRef]
- Cruz, V.M.V.; Kilian, A.; Dierig, D.A. Development of DArT marker platforms and genetic diversity assessment of the U.S. collection of the new oilseed crop Lesquerella and Related Species. PLoS ONE 2013, 8, e64062. [Google Scholar] [CrossRef]
- Wang, J.; Li, H.; Zhang, L.; Meng, L. Users’ Manual of QTL IciMapping; The Quantitative Genetics Group, Institute of Crop Science, Chinese Academy of Agricultural Sciences (CAAS): Beijing, China; Genetic Resources Program, International Maize and Wheat Improvement Center (CIMMYT): Texcoco, Mexico, 2016. [Google Scholar]
- Montri, P.; Taylor, P.W.J.; Mongkolporn, O. Pathotypes of Colletotrichum capsici, the causal agent of chili anthracnose, in Thailand. Plant Dis. 2009, 93, 17–20. [Google Scholar] [CrossRef]
- Chang, S.; Puryear, J.; Cairney, J. A simple and efficient method for isolating RNA from pine trees. Plant Mol. Biol. Rep. 1993, 11, 113–116. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: http://www.R-project.org (accessed on 22 January 2023).
- Kim, S.; Park, J.; Yeom, S.-I.; Kim, Y.-M.; Seo, E.; Kim, K.-T.; Kim, M.-S.; Lee, J.M.; Cheong, K.; Shin, H.-S.; et al. New reference genome sequences of hot pepper reveal the massive evolution of plant disease-resistance genes by retroduplication. Genome Biol. 2017, 18, 210. [Google Scholar] [CrossRef]
- Kim, K.H.; Yoon, J.B.; Park, H.G.; Park, E.W.; Kim, Y.H. Structural modifications and programmed cell death of chili pepper fruit related to resistance responses to Colletotrichum gloeosporioides infection. Phytopathology 2004, 94, 1295–1304. [Google Scholar] [CrossRef]
- Balint-Kurti, P. The plant hypersensitive response: Concepts, control and consequences. Mol. Plant Pathol. 2009, 20, 1163–1178. [Google Scholar] [CrossRef] [PubMed]
- Ngou, B.P.M.; Ding, P.; Jones, J.D.G. Thirty years of resistance: Zig-zag through the plant immune system. Plant Cell. 2022, 34, 1447–1478. [Google Scholar] [CrossRef] [PubMed]
- Ngou, B.P.M.; Ahn, H.K.; Ding, P.; Jones, J.D.G. Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 2021, 592, 110–115. [Google Scholar] [CrossRef] [PubMed]
- Khatib, M.; Lafitte, C.; Esquerré-Tugayé, M.-T.; Bottin, A.; Rickauer, M. The CBEL elicitor of Phytophthora parasitica var. nicotianae activates defence in Arabidopsis thaliana via three different signalling pathways. New Phytol. 2004, 162, 501–510. [Google Scholar] [CrossRef]
- Ma, Z.; Song, T.; Zhu, L.; Ye, W.; Wang, Y.; Shao, Y.; Dong, S.; Zhang, Z.; Dou, D.; Zheng, X.; et al. A Phytophthora sojae glycoside hydrolase 12 protein is a major virulence factor during soybean infection and is recognized as a PAMP. Plant Cell. 2015, 27, 2057–2072. [Google Scholar] [CrossRef]
- Wang, Y.; Tyler, B.M.; Wang, Y. Defense and counterdefense during plant-pathogenic oomycete infection. Annu. Rev. Microbiol. 2019, 73, 667–696. [Google Scholar] [CrossRef]
- Kaku, H.; Nishizawa, Y.; Ishii-Minami, N.; Akimoto-Tomiyama, C.; Dohmae, N.; Takio, K.; Minami, E.; Shibuya, N. Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc. Natl. Acad. Sci. USA 2006, 103, 11086–11091. [Google Scholar] [CrossRef]
- Zhang, J.; Zhou, J.M. Plant immunity triggered by microbial molecular signatures. Mol. Plant 2010, 3, 783–793. [Google Scholar] [CrossRef]
- Hu, S.P.; Li, J.J.; Dhar, N.; Li, J.P.; Chen, J.Y.; Jian, W.; Dai, X.F.; Yang, X.Y. Lysin motif (LysM) proteins: Interlinking manipulation of plant immunity and fungi. Int. J. Mol. Sci. 2021, 22, 3114. [Google Scholar] [CrossRef]
- Liu, S.; Wang, J.; Han, Z.; Gong, X.; Zhang, H.; Chai, J. Molecular mechanism for fungal cell wall recognition by rice chitin receptor OsCEBiP. Structure 2016, 24, 1192–1200. [Google Scholar] [CrossRef]
- Narusaka, Y.; Shinya, T.; Narusaka, M.; Motoyama, N.; Shimada, H.; Murakami, K.; Shibuya, N. Presence of LYM2 dependent but CERK1 independent disease resistance in Arabidopsis. Plant Signal. Behav. 2013, 8, e25345. [Google Scholar] [CrossRef] [PubMed]
- Faulkner, C.; Petutschnig, E.; Benitez-Alfonso, Y.; Beck, M.; Robatzek, S.; Lipka, V.; Maule, A.J. LYM2-dependent chitin perception limits molecular flux via plasmodesmata. Proc. Natl. Acad. Sci. USA 2013, 110, 9166–9170. [Google Scholar] [CrossRef]
- Müssig, C.; Schröder, F.; Usadel, B.; Lisso, J. Structure and putative function of NFX1-like proteins in plants. Plant Biol. 2010, 12, 381–394. [Google Scholar] [CrossRef] [PubMed]
- Lisso, J.; Altmann, T.; Müssig, C. The AtNFXL1 gene encodes a NF-X1 type zinc finger protein required for growth under salt stress. FEBS Lett. 2006, 580, 4851–4856. [Google Scholar] [CrossRef]
- Levine, A.; Tenhaken, R.; Dixon, R.; Lamb, C. H2O2 from the oxidative burst orchestrates the plant hypersensitive disease resistance response. Cell 1994, 79, 583–593. Available online: http://bit.ly/2W6ZR2x (accessed on 28 April 2023). [CrossRef] [PubMed]
- Lamb, C.; Dixon, R.A. The oxidative burst in plant disease resistance. Annu. Rev. Plant Biol. 1997, 48, 251–275. [Google Scholar] [CrossRef]
- Mittler, R.; Vanderauwera, S.; Suzuki, N.; Miller, G.; Tognetti, V.B.; Vandepoele, K.; Gollery, M.; Shulaev, V.; Van Breusegem, F. ROS signaling: The new wave? Trends Plant Sci. 2011, 16, 300–309. [Google Scholar] [CrossRef]
- Gilroy, S.; Białasek, M.; Suzuki, N.; Górecka, M.; Devireddy, A.R.; Karpiński, S.; Mittler, R. ROS, calcium, and electric signals: Key mediators of rapid systemic signaling in plants. Plant Physiol. 2016, 171, 1606–1615. [Google Scholar] [CrossRef]
- Pitzschke, A.; Hirt, H. Mitogen-activated protein kinases and reactive oxygen species signaling in plants. Plant Physiol. 2006, 141, 351–356. [Google Scholar] [CrossRef]
- Choi, H.W.; Kim, Y.J.; Lee, S.C.; Hong, J.K.; Hwang, B.K. Hydrogen peroxide generation by the pepper extracellular peroxidase CaPO2 activates local and systemic cell death and defence response to bacterial pathogens. Plant Physiol. 2007, 145, 890–904. Available online: http://bit.ly/2WcpE9w (accessed on 28 April 2023). [CrossRef]
- Doke, N.; Miura, Y.; Sanchez, L.M.; Park, H.J.; Noritake, T.; Yoshioka, H.; Kawakita, K. The oxidative burst protects plants against pathogen attack: Mechanism and role as an emergency signal for plant biodefence—A review. Gene 1996, 79, 45–51. Available online: http://bit.ly/2w6eNTP (accessed on 28 April 2023). [CrossRef]
- Lehmann, S.; Serrano, M.; L’Haridon, F.; Tjamos, S.E.; Metraux, J.-P. Reactive oxygen species and plant resistance to fungal pathogens. Phytochemistry 2015, 112, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Shetty, N.P.; Lyngs, H.J.J.; Jensen, D.; Collinge, D.B.; Shekar, H.C. Roles of reactive oxygen species in interactions between plants and pathogens. Eur. J. Plant Pathol. 2008, 121, 267–280. Available online: http://bit.ly/2xsyju3 (accessed on 29 April 2023). [CrossRef]
- Köhler, C.; Grossniklaus, U. Epigenetic inheritance of expression states in plant development: The role of polycomb group proteins. Curr. Opin. Cell Biol. 2002, 14, 773–779. [Google Scholar] [CrossRef] [PubMed]
- Schatlowski, N.; Creasey, K.; Goodrich, J.; Schubert, D. Keeping plants in shape: Polycomb-group genes and histone methylation. Semin. Cell Dev. Biol. 2008, 19, 547–553. [Google Scholar] [CrossRef]
- Schatlowski, N.; Stahl, Y.; Hohenstatt, M.L.; Goodrich, J.; Schubert, D. The CURLY LEAF interacting protein BLISTER controls expression of polycomb-group target genes and cellular differentiation of Arabidopsis thaliana. Plant Cell. 2010, 22, 2291–2305. [Google Scholar] [CrossRef]
- Yu, Y.; Bu, Z.; Shen, W.-H.; Dong, A. An update on histone lysine methylation in plants. Prog. Nat. Sci. 2009, 19, 407–413. [Google Scholar] [CrossRef]
- Kang, H.; Fan, T.; Wu, J.; Zhu, Y.; Shen, W.-H. Histone modification and chromatin remodeling in plant response to pathogens. Front. Plant Sci. 2022, 13, 986940. [Google Scholar] [CrossRef] [PubMed]
- Kleinmanns, J.A.; Schatlowski, N.; David, H.; Daniel, S. BLISTER regulates polycomb-target genes, represses stress-regulated genes and promotes stress responses in Arabidopsis thaliana. Front. Plant Sci. 2017, 8, 1530. [Google Scholar] [CrossRef]
- Liu, C.; Cheng, J.; Zhuang, Y.; Ye, L.; Li, Z.; Wang, Y.; Qi, M.; Xu, L.; Zhang, Y. Polycomb repressive complex 2 attenuates ABA-induced senescence in Arabidopsis. Plant J. 2019, 97, 368–377. [Google Scholar] [CrossRef]
- Wood, C.C.; Robertson, M.; Tanner, G.; Peacock, W.J.; Dennis, E.S.; Helliwell, C.A. The Arabidopsis thaliana vernalization response requires a polycomb-like protein complex that also includes VERNALIZATION INSENSITIVE 3. Proc. Natl. Acad. Sci. USA 2006, 103, 14631–14636. [Google Scholar] [CrossRef] [PubMed]
- Tomaštíková, E.D.; Hafrén, A.; Trejo-Arellano, M.S.; Rasmussen, S.R.; Sato, H.; Santos-González, J.; Köhler, C.; Hennig, L.; Hofius, D. Polycomb repressive complex 2 and KRYPTONITE regulate pathogen-induced programmed cell death in Arabidopsis. Plant Physiol. 2021, 185, 2003–2021. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.B.; Foley, R.C.; Oñate-Sánchez, L. Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 2002, 5, 430–436. [Google Scholar] [CrossRef] [PubMed]
- Berrocal-Lobo, M.; Molina, A.; Solano, R. Constitutive expression of ETHYLENE-RESPONSE-FACTOR1 in Arabidopsis confers resistance to several necrotrophic fungi. Plant J. 2002, 29, 23–32. [Google Scholar] [CrossRef]
- Berrocal-Lobo, M.; Molina, A. Ethylene Response Factor 1 mediates Arabidopsis resistance to the soilborne fungus Fusarium oxysporum. Mol. Plant-Microbe Interact. 2004, 17, 763–770. [Google Scholar] [CrossRef]
- Cheng, M.C.; Liao, P.M.; Kuo, W.W.; Lin, T.P. The Arabidopsis ETHYLENE RESPONSE FACTOR1 regulates abiotic stress-responsive gene expression by binding to different cis-acting elements in response to different stress signals. Plant Physiol. 2013, 162, 1566–1582. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Munné-Bosch, S. Ethylene response factors: A key regulatory hub in hormone and stress signaling. Plant Physiol. 2015, 169, 32–41. [Google Scholar] [CrossRef]
- Xing, L.; Di, Z.; Yang, W.; Liu, J.; Li, M.; Wang, X.; Cui, C.; Wang, X.; Wang, X.; Zhang, R.; et al. Overexpression of ERF1-V from Haynaldia villosa can enhance the resistance of wheat to powdery mildew and increase the tolerance to salt and drought stresses. Front. Plant Sci. 2017, 8, 1948. [Google Scholar] [CrossRef]
- Yang, C.; Li, W.; Cao, J.; Meng, F.; Yu, Y.; Huang, J.; Jiang, L.; Liu, M.; Zhang, Z.; Chen, X.; et al. Activation of ethylene signaling pathways enhances disease resistance by regulating ROS and phytoalexin production in rice. Plant J. 2017, 89, 338–353. [Google Scholar] [CrossRef]
- Liu, F.; Zhang, X.; Lu, C.; Zeng, X.; Li, Y.; Fu, D.; Wu, G. Non-specific lipid transfer proteins in plants: Presenting new advances and an integrated functional analysis. J. Exp. Bot. 2015, 66, 5663–5681. [Google Scholar] [CrossRef]
- Van Loon, L.C.; Van Strien, E.A. The families of pathogenesis-related proteins, their activities, and comparative analysis of PR-1 type proteins. Physiol. Mol. Plant Pathol. 1999, 55, 85–97. Available online: https://dspace.library.uu.nl/bitstream/handle/1874/1590/PMPP-van+Loon-1999.pdf?sequence=1 (accessed on 25 April 2023). [CrossRef]
- Wang, S.Y.; Wu, J.H.; Ng, T.B.; Ye, X.Y.; Rao, P.F. A non-specific lipid transfer protein with antifungal and antibacterial activities from the mung bean. Peptides 2004, 25, 1235–1242. [Google Scholar] [CrossRef]
- Patkar, R.N.; Chattoo, B.B. Transgenic indica rice expressing ns-LTP-like protein shows enhanced resistance to both fungal and bacterial pathogens. Mol. Breed. 2006, 17, 159–171. [Google Scholar] [CrossRef]
- Kirubakaran, S.I.; Begum, S.M.; Ulaganathan, K.; Sakthivel, N. Characterization of a new antifungal lipid transfer protein from wheat. Plant Physiol. Biochem. 2018, 46, 918–927. [Google Scholar] [CrossRef] [PubMed]
- Caaveiro, J.M.; Molina, A.; González-Mañas, J.M.; Rodríguez-Palenzuela, P.; Garcia-Olmedo, F.; Goñi, F.M. Differential effects of five types of antipathogenic plant peptides on model membranes. FEBS Lett. 1997, 410, 338–342. [Google Scholar] [CrossRef] [PubMed]
- Regente, M.C.; De La Canal, L. Purification, characterization and antifungal properties of a lipid-transfer protein from sunflower (Helianthus annuus) seeds. Physiol. Plant. 2000, 110, 158–163. [Google Scholar] [CrossRef]
- Savatin, D.V.; Gramegna, G.; Modesti, V.; Cervone, F. Wounding in the plant tissue: The defense of a dangerous passage. Front. Plant Sci. 2014, 5, 470. [Google Scholar] [CrossRef]
- Hou, S.; Liu, Z.; Shen, H.; Wu, D. Damage-associated molecular pattern-triggered immunity in plants. Front. Plant Sci. 2019, 10, 646. [Google Scholar] [CrossRef]
- Vega-Muñoz, I.; Duran-Flores, D.; Fernández-Fernández, A.D.; Heyman, J.; Ritter, A.; Stael, S. Breaking bad news: Dynamic molecular mechanisms of wound response in plants. Front. Plant Sci. 2020, 11, 610445. [Google Scholar] [CrossRef]
- Wu, Y.; Gao, Y.; Zhan, Y.; Kui, H.; Liu, H.; Yan, L.; Kemmerling, B.; Zhou, J.-M.; He, K.; Li, J. Loss of the common immune coreceptor BAK1 leads to NLR-dependent cell death. Proc. Natl. Acad. Sci. USA 2020, 117, 27044–27053. [Google Scholar] [CrossRef]
- Zhang, Z.; Wu, Y.; Gao, M.; Zhang, J.; Kong, Q.; Liu, Y.; Ba, H.; Zhou, J.; Zhang, Y. Disruption of PAMP-Induced MAP Kinase cascade by a Pseudomonas syringae effector activates plant immunity mediated by the NB-LRR protein SUMM2. Cell Host Microbe 2012, 11, 253–263. [Google Scholar] [CrossRef] [PubMed]
- Pruitt, R.N.; Locci, F.; Wanke, F.; Zhang, L.; Saile, S.C.; Joe, A.; Karelina, D.; Hua, C.; Fröhlich, K.; Wan, W.-L.; et al. The EDS1-PAD4-ADR1 node mediates Arabidopsis pattern-triggered immunity. Nature 2021, 598, 495–499. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Jiang, Z.; Bi, G.; Nomura, K.; Liu, M.; Wang, Y.; Cai, B.; Zhou, J.M.; He, S.Y.; Xin, X.F. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 2021, 592, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Son, S.; Kim, S.; Lee, K.S.; Oh, J.; Choi, I.; Do, J.W.; Yoon, J.B.; Han, J.; Choi, D.; Park, S.R. Identification of the Capsicum baccatum NLR protein CbAR9 conferring disease resistance to anthracnose. Int. J. Mol. Sci. 2021, 22, 12612. [Google Scholar] [CrossRef]


| Gene ID | Gene Description | Forward/Reverse Primer 5′–3′ | Product Size (bp) |
|---|---|---|---|
| CQW23_09568 | LysM domain-containing GPI-anchored protein 2 (LYM2) | CCCGATCTCTCTTCTCATACAAATGC GGCAGACTTAAGATCCCATCCACAC | 133 |
| CQW23_09584 | Ethylene-responsive transcription factor (ERF) | GGGAAGTTGAGATTGTGAGAAGCA AGGGAGTGAGAATGAGAAGCTGG | 172 |
| CQW23_09596 | Hypothetical protein (HP596) | TCTTTGTCTGAGGTTCCATCGG ACCTTACTACTCTATGCCTTCAAAG | 76 |
| CQW23_09597 | Hypothetical protein (HP597) | CCCAATGAAGAGGATGGCTCTGGT GCAACATCGATTGAACCCCAGAAAC | 200 |
| CQW23_09600 | Histone-lysine N-methyltransferase (CLF) | TTCCTCTGAAGATGCAACTGTG AAGATCCTTCGTCAGATTCTCC | 148 |
| CQW23_09601 | Hypothetical protein (HP601) | TCTTGCTGTGGATCTGTTGCTG TCCTTGCTTTTTGTCTCTGCGG | 96 |
| CQW23_09609 | Auxin response factor 23 (ARF23) | AAAGGTCCGAGCAATCAAAGGG TGCCATCCCTCTCTCTAGAAGC | 93 |
| CQW23_09618 | NF-X1-type zinc finger protein (NFXL1) | TGCTTTTGTGGGAAGAGGCAAG GCAGGGACAACTTCTAGCTGGA | 210 |
| CQW23_09644 | Non-specific lipid-transfer protein 2 (PR-14) | ACAAAGGCAAGGTTTCTGCTCTC GCGATTACATCATCACAACCACCC | 72 |
| CQW23_20069 | Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) | AGACCTTGGAGTTGCAGATGTG TGAGACCGTGACAATGCTAACC | 78 |
| CQW23_12274 | Elongation factor 1-alpha (EF-1α) | TCTCCGACGACAAACCTCAAGC CGCCATTCCTGAATTGTGTGATAGGG | 80 |
| Marker | Physical Position (bp) | LOD 1 | PVE (%) 2 | Add 3 |
|---|---|---|---|---|
| SNP305 | 22,920,913 | 4.88 | 20.87 | −0.21 |
| Silico01 | 24,353,170 | 4.16 | 18.07 | −0.20 |
| SNP306 | 24,353,190 | 4.16 | 18.07 | −0.20 |
| Silico02 | 25,181,503 | 2.14 | 9.78 | −0.15 |
| Silico03 | 27,298,089 | 3.87 | 16.92 | −0.19 |
| Silico04 | 27,762,754 | 4.05 | 17.67 | −0.20 |
| SNP0132 | 28,348,529 | 3.75 | 16.48 | −0.19 |
| Silico05 | 28,350,263 | 3.17 | 14.10 | −0.18 |
| Silico06 | 31,275,816 | 3.22 | 14.33 | −0.18 |
| Silico07 | 31,813,244 | 0.24 | 1.14 | −0.05 |
| Silico08 | 32,891,183 | 1.53 | 7.08 | −0.13 |
| SNP309 | 33,876,713 | 5.22 | 22.14 | −0.22 |
| SNP310 | 34,133,643 | 3.15 | 14.02 | −0.17 |
| SNP311 | 34,378,585 | 4.74 | 20.34 | −0.21 |
| Silico09 | 34,998,101 | 5.91 | 24.67 | −0.23 |
| Silico10 | 39,171,666 | 2.38 | 10.80 | −0.15 |
| Silico11 | 40,015,431 | 3.45 | 15.26 | −0.18 |
| Silico12 | 40,678,856 | 4.46 | 19.27 | −0.20 |
| SNP1755 | 41,686,959 | 4.16 | 18.07 | −0.20 |
| Silico13 | 42,390,824 | 4.88 | 20.87 | −0.21 |
| Silico14 | 43,749,964 | 1.35 | 6.28 | −0.12 |
| Silico15 | 44,285,168 | 1.70 | 7.83 | −0.13 |
| Silico16 | 46,002,881 | 1.53 | 7.11 | −0.13 |
| Silico17 | 47,161,782 | 1.20 | 5.62 | −0.11 |
| Silico18 | 47,414,359 | 1.53 | 7.11 | −0.13 |
| Silico19 | 47,677,887 | 0.16 | 0.79 | −0.04 |
| Silico20 | 47,681,951 | 0.24 | 1.14 | −0.05 |
| Silico21 | 48,000,095 | 0.09 | 0.43 | −0.03 |
| Silico22 | 48,586,639 | 1.18 | 5.51 | −0.11 |
| Silico23 | 48,746,637 | 4.05 | 17.67 | −0.20 |
| Silico24 | 51,871,096 | 3.43 | 15.17 | −0.18 |
| Silico25 | 53,148,783 | 2.93 | 13.11 | −0.17 |
| Silico26 | 55,311,209 | 2.38 | 10.80 | −0.15 |
| Silico27 | 55,657,165 | 3.15 | 14.02 | −0.17 |
| Silico28 | 56,845,351 | 0.41 | 1.95 | −0.07 |
| SNP7049 | 65,265,181 | 2.38 | 10.80 | −0.15 |
| Silico29 | 65,700,039 | 2.16 | 9.86 | −0.15 |
| Silico30 | 65,987,191 | 0.00 | 0.04 | −0.01 |
| Silico31 | 68,981,148 | 2.64 | 11.91 | −0.16 |
| Silico32 | 71,888,214 | 2.87 | 12.87 | −0.17 |
| Silico33 | 72,249,488 | 3.48 | 15.36 | −0.19 |
| Silico34 | 75,024,622 | 3.44 | 15.20 | −0.18 |
| Silico35 | 78,008,814 | 3.44 | 15.20 | −0.18 |
| Silico36 | 78,028,368 | 3.73 | 16.38 | −0.19 |
| Silico37 | 78,035,410 | 0.43 | 2.05 | −0.07 |
| Silico38 | 78,164,097 | 2.16 | 9.86 | −0.15 |
| BACSNP_4_63 | 84,305,804 | 3.14 | 14.00 | −0.18 |
| Silico39 | 85,431,446 | 3.14 | 14.00 | −0.18 |
| Silico40 | 86,667,602 | 3.14 | 14.00 | −0.18 |
| Silico41 | 91,189,255 | 2.38 | 10.80 | −0.15 |
| Silico42 | 92,568,227 | 1.93 | 8.86 | −0.14 |
| Silico43 | 92,939,149 | 1.19 | 5.55 | −0.11 |
| Silico44 | 100,038,292 | 2.38 | 10.80 | −0.16 |
| Silico45 | 100,064,499 | 3.73 | 16.38 | −0.19 |
| Silico46 | 101,835,880 | 1.96 | 8.97 | −0.14 |
| Silico47 | 102,448,025 | 2.89 | 12.94 | −0.17 |
| Silico48 | 103,063,217 | 1.36 | 6.34 | −0.12 |
| Silico49 | 103,132,977 | 2.41 | 10.94 | −0.15 |
| Silico50 | 103,371,531 | 2.87 | 12.87 | −0.17 |
| Silico51 | 103,567,675 | 3.15 | 14.02 | −0.17 |
| Silico52 | 103,579,898 | 1.96 | 8.97 | −0.14 |
| Silico53 | 104,238,202 | 1.39 | 6.45 | −0.12 |
| Silico54 | 106,569,759 | 2.16 | 9.86 | −0.15 |
| Silico55 | 111,192,822 | 3.73 | 16.38 | −0.19 |
| BACSNP_4_60 | 113,019,674 | 2.16 | 9.86 | −0.15 |
| Silico56 | 114,140,855 | 2.41 | 10.94 | −0.15 |
| Silico57 | 115,808,681 | 0.80 | 3.76 | −0.09 |
| Silico58 | 125,855,206 | 3.18 | 14.16 | −0.17 |
| Silico59 | 129,409,957 | 3.15 | 14.02 | −0.17 |
| Silico60 | 132,674,360 | 1.59 | 7.35 | −0.13 |
| Silico61 | 133,136,427 | 2.38 | 10.80 | −0.15 |
| Silico62 | 134,969,221 | 1.77 | 8.13 | −0.13 |
| Silico63 | 136,379,454 | 3.44 | 15.20 | −0.18 |
| Silico64 | 137,027,139 | 2.87 | 12.87 | −0.17 |
| Silico65 | 139,918,888 | 2.00 | 9.15 | −0.14 |
| Silico66 | 142,635,277 | 2.89 | 12.94 | −0.17 |
| Silico67 | 144,664,453 | 3.74 | 16.43 | −0.19 |
| Silico68 | 144,808,619 | 2.64 | 11.89 | −0.17 |
| Silico69 | 144,901,329 | 2.93 | 13.11 | −0.17 |
| SNP331 | 146,776,687 | 4.74 | 20.34 | −0.21 |
| Gene | Physical Position (bp) 1 | Gene Function 2 |
|---|---|---|
| LYM2 | 27,166,666–27,169,869 | Involved in defense response as chitin-binding protein. |
| ERF | 31,226,500–31,227,246 | Transcriptional activator that may involve in disease resistance pathways. |
| HP596 | 33,798,986–33,799,247 | Unknown. |
| HP597 | 34,375,190–34,380,266 | Unknown. |
| CLF | 34,927,068–34,936,068 | Involved in chromosome silencing, histone methylation, regulation of gene expression by genetic imprinting, cell differentiation, etc. |
| HP601 | 34,997,949–35,013,770 | Unknown. |
| ARF23 | 36,381,979–36,386,588 | Transcriptional activator that may involve in disease resistance pathways. |
| NFXL1 | 36,512,044–36,515,448 | Promotes H2O2 production, defense response to bacterium, response to microbial phytotoxin, response to salt stress, salicylic acid biosynthetic process, etc. |
| PR-14 | 40,270,316–40,270,594 | Transfer lipids across membranes. May play a role in plant defense or in the biosynthesis of cuticle layers. |
| RIL Code | Anthracnose Severity Score (0–9) | |
|---|---|---|
| Mature Green | Ripe | |
| G1-017130 | 0 | 0 |
| G1-017135 | 0 | 5 |
| G1-017147 | 5 | 0 |
| G1-017233 | 5 | 5 |
| G1-017257 | 5 | 5 |
| G1-017261 | 5 | 3 |
| G1-017264 | 5 | 5 |
| G1-017267 | 5 | 7 |
| G1-017325 | 1 | 5 |
| G1-017339 | 5 | 0 |
| G1-017396 | 5 | 5 |
| G1-017399 | 0 | 0 |
| G1-017415 | 3 | 0 |
| G1-017416 | 3 | 0 |
| G1-017425 | 0 | 0 |
| G1-017431 | 5 | 1 |
| G1-017433 | 0 | 5 |
| G1-017473 | 0 | 1 |
| G1-017531 | 1 | 1 |
| G1-017539 | 5 | 0 |
| G1-017540 | 5 | 3 |
| G1-017541 | 5 | 5 |
| G1-017544 | 5 | 0 |
| G1-017579 | 3 | 0 |
| G1-017588 | 5 | 5 |
| G1-017593 | 5 | 5 |
| G1-017623 | 5 | 0 |
| G1-017679 | 5 | 5 |
| G1-017845 | 0 | 0 |
| Genotype + Inoculation | LYM2 | ERF | ||
|---|---|---|---|---|
| 6 h | 12 h | 6 h | 12 h | |
| RR + water | 2.99 ± 0.28 bx | 1.47 ± 0.22 cy | 3.74 ± 0.51 cy | 4.98 ± 0.40 bx |
| RR + MJ5 | 4.53 ± 0.35 ax | 2.80 ± 0.44 by | 4.05 ± 0.33 bcy | 6.43 ± 0.71 ax |
| SS + water | 1.03 ± 0.26 cdx | 0.24 ± 0.10 ey | 2.03 ± 0.18 dex | 1.30 ± 0.10 efx |
| SS + MJ5 | 0.75 ± 0.16 dex | 0.67 ± 0.19 dex | 2.25 ± 0.42 dex | 2.30 ± 0.24 dx |
| Genotype + inoculation | HP597 | CLF | ||
| 6 h | 12 h | 6 h | 12 h | |
| RR + water | 3.47 ± 0.30 ax | 1.62 ± 0.20 by | 1.52 ± 0.29 ax | 0.92 ± 0.23 cdy |
| RR + MJ5 | 3.70 ± 0.19 ax | 1.75 ± 0.20 by | 1.80 ± 0.17 ax | 1.43 ± 0.28 bcx |
| SS + water | 1.83 ± 0.24 bx | 0.58 ± 0.15 dy | 0.8 ± 0.11 cdx | 0.52 ± 0.12 dx |
| SS + MJ5 | 1.34 ± 0.29 bcx | 1.36 ± 0.16 bcx | 0.75 ± 0.06 cdx | 0.71 ± 0.15 cdx |
| Genotype + inoculation | NFXL1 | PR-14 | ||
| 6 h | 12 h | 6 h | 12 h | |
| RR + water | 2.25 ± 0.42 ax | 0.77 ± 0.07 cy | 2.68 ± 0.82 ax | 1.23 ± 0.06 by |
| RR + MJ5 | 2.31 ± 0.46 ax | 1.58 ± 0.42 abx | 2.98 ± 0.69 ax | 1.33 ± 0.28 by |
| SS + water | 0.62 ± 0.13 cx | 0.26 ± 0.04 cx | 1.02 ± 0.25 bx | 0.37 ± 0.04 bx |
| SS + MJ5 | 0.70 ± 0.22 cx | 0.40 ± 0.10 cx | 1.09 ± 0.04 bx | 0.68 ± 0.13 bx |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kethom, W.; Taylor, P.W.J.; Mongkolporn, O. Expression of Genes Involved in Anthracnose Resistance in Chili (Capsicum baccatum) ‘PBC80’-Derived Recombinant Inbred Lines. Pathogens 2023, 12, 1306. https://doi.org/10.3390/pathogens12111306
Kethom W, Taylor PWJ, Mongkolporn O. Expression of Genes Involved in Anthracnose Resistance in Chili (Capsicum baccatum) ‘PBC80’-Derived Recombinant Inbred Lines. Pathogens. 2023; 12(11):1306. https://doi.org/10.3390/pathogens12111306
Chicago/Turabian StyleKethom, Wassana, Paul W. J. Taylor, and Orarat Mongkolporn. 2023. "Expression of Genes Involved in Anthracnose Resistance in Chili (Capsicum baccatum) ‘PBC80’-Derived Recombinant Inbred Lines" Pathogens 12, no. 11: 1306. https://doi.org/10.3390/pathogens12111306






