hBN Nanoparticle-Assisted Rapid Thermal Cycling for the Detection of Acanthamoeba
Abstract
:1. Introduction
Nanoparticle Enhanced Thermal Conductivity
2. Materials and Methods
2.1. hBN Nanoparticle Characterization
2.2. Acanthamoeba castellanii Cell Cultures
2.3. Acanthamoeba DNA Extraction Using Chelex, Proteinase K, and Genet Bio Kit Methods
2.4. Preparation of hBN Nanofluid
2.5. PCR, Nano-PCR, and Gel Electrophoresis
2.6. Quantitative Analysis of Amplified Nano-PCR Products
3. Results
3.1. Nanoparticle Characterization
3.2. Thermal Conductivity
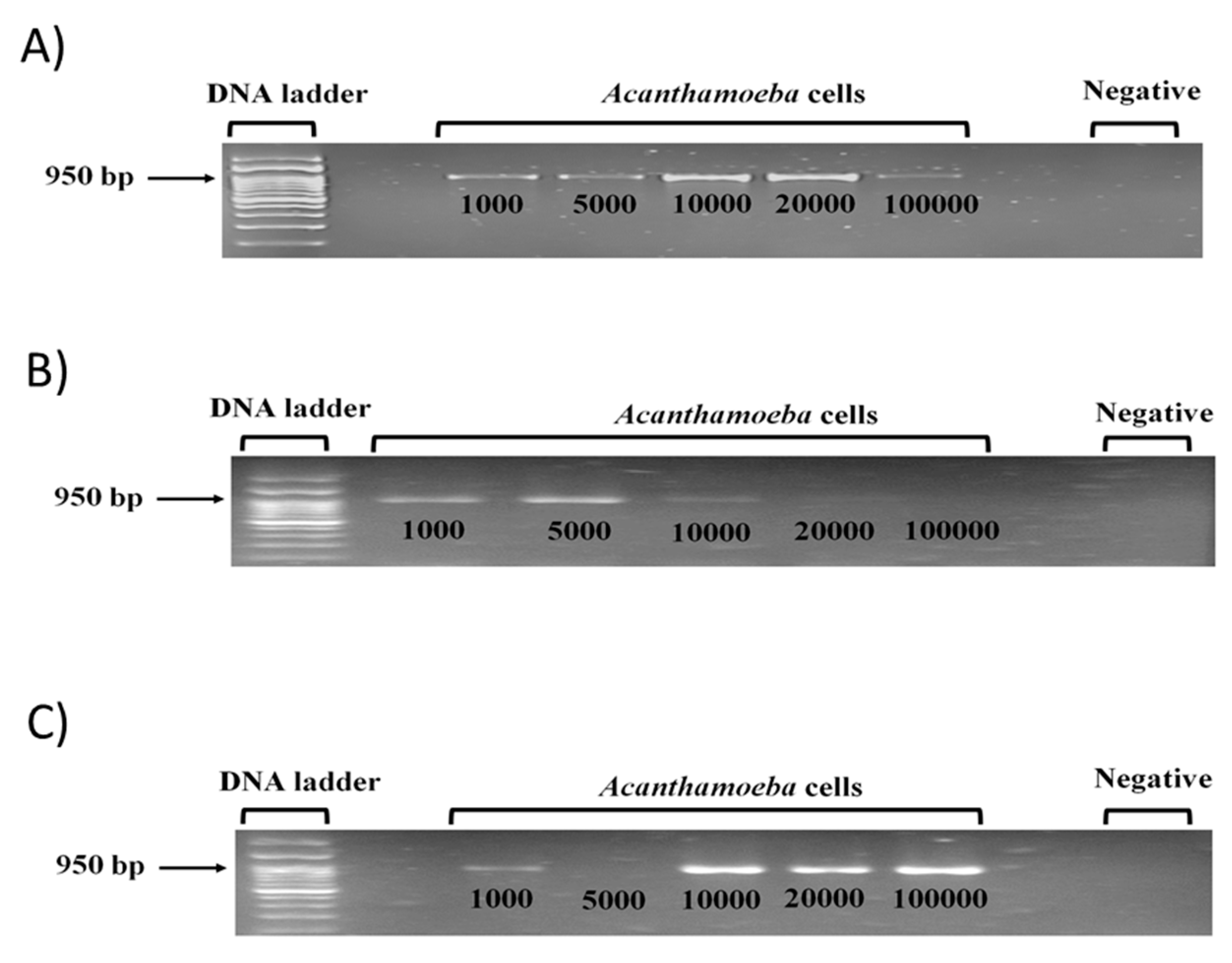
3.3. Chelex Assay with 10,000 Acanthamoeba Cells Was Optimized among Three DNA Extraction Methods Tested to Identify Low Cell Numbers
3.4. HBN Nanoparticle Enhances Acanthamoeba Detection in PCR and Increases PCR DNA Yield
3.5. hBN Nanoparticle Reduces PCR Temperature to Enhance Acanthamoeba Detection and Increases DNA Yield
3.6. hBN Nanoparticle Reduces PCR Step Time to Enhance Acanthamoeba Detection
3.7. hBN Nanoparticle Reduces PCR Cycle Time to Enhance Acanthamoeba Detection
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Centers for Disease Control and Prevention. Acanthamoeba Infection FAQs. Available online: https://www.cdc.gov/parasites/acanthamoeba/gen_info/acanthamoeba.html (accessed on 30 July 2020).
- Lorenzo-Morales, J.; Khan, N.A.; Walochnik, J. An update on Acanthamoeba keratitis: Diagnosis, pathogenesis and treatment. Parasite 2015, 22, 10. [Google Scholar] [CrossRef] [Green Version]
- Boggild, A.K.; Martin, D.S.; Lee, T.Y.; Yu, B.; Low, D.E. Laboratory diagnosis of amoebic keratitis: Comparison of four diagnostic methods for different types of clinical specimens. J. Clin. Microbiol. 2009, 47, 1314–1318. [Google Scholar] [CrossRef] [Green Version]
- Hau, S.C.; Dart, J.K.; Vesaluoma, M.; Parmar, D.N.; Claerhout, I.; Bibi, K.; Larkin, D.F. Diagnostic accuracy of microbial keratitis with in vivo scanning laser confocal microscopy. Br. J. Ophthalmol. 2010, 94, 982–987. [Google Scholar] [CrossRef] [Green Version]
- Bhosale, N.K.; Parija, S.C.; Mandal, J.; Ahuja, S. Utility of Polymerase chain reaction in diagnosis of Acanthamoeba and Microsporidial keratitis. Int. J. Infect. Dis. 2016, 45, 354. [Google Scholar] [CrossRef] [Green Version]
- Visvesvara, G.S.; Moura, H.; Schuster, F.L. Pathogenic and opportunistic free-living amoebae: Acanthamoeba spp., Balamuthia mandrillaris, Naegleria fowleri, and Sappinia diploidea. Pathog. Dis. 2007, 50, 1–26. [Google Scholar] [CrossRef] [Green Version]
- Yuce, M.; Kurt, H.; Mokkapati, V.R.S.S.; Budak, H. Employment of nanomaterials in polymerase chain reaction: Insight into the impacts and putative operating mechanisms of nano-additives in PCR. Rsc Adv. 2014, 4, 36800–36814. [Google Scholar] [CrossRef]
- Zhao, G.; Zhai, H.; Yuan, Q.; Sun, S.; Liu, T.; Xie, L. Rapid and sensitive diagnosis of fungal keratitis with direct PCR without template DNA extraction. Clin. Microbiol. Infect. 2014, 20, O776–O782. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lorenz, T.C. Polymerase chain reaction: Basic protocol plus troubleshooting and optimization strategies. J. Vis. Exp. 2012, e3998. [Google Scholar] [CrossRef] [PubMed]
- Fu-Ming, S.; Xin, L.; Jia, L. Development of Nano-Polymerase Chain Reaction and Its Application. Chin. J. Anal. Chem. 2017, 45, 1745–1753. [Google Scholar] [CrossRef]
- Kim, H.R.; Baek, A.; Lee, I.J.; Kim, D.-E. Facilitation of Polymerase Chain Reaction with Poly (ethylene glycol)-Engrafted Graphene Oxide Analogous to a Single-Stranded-DNA Binding Protein. Acs Appl. Mater. Interfaces 2016, 8, 33521–33528. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Sun, L.; Hu, N.; Huang, G.; Weng, J. Graphene enhances the specificity of the polymerase chain reaction. Small 2012, 8, 2011–2015. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, N.; Wang, H.; Tang, C.; Lei, S.; Shen, W.; Wang, C.; Wang, G.; Wang, Z.; Wang, L. Toxicity evaluation of boron nitride nanospheres and water-soluble boron nitride in Caenorhabditis elegans. Int. J. Nanomed. 2017, 12, 5941–5957. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maxwell, J.C. A Treatise on Electricity and Magnetism; Clarendon: Oxford, UK, 1892; Volume 2, pp. 68–73. [Google Scholar]
- Keblinski, P.; Phillpot, S.; Choi, S.; Eastman, J. Mechanisms of heat flow in suspensions of nano-sized particles (nanofluids). Int. J. Heat Mass Transfer 2002, 45, 855–863. [Google Scholar] [CrossRef]
- Dhar, P.; Gupta, S.S.; Chakraborty, S.; Pattamatta, A.; Das, S.K. The role of percolation and sheet dynamics during heat conduction in poly-dispersed graphene nanofluids. Appl. Phys. Lett. 2013, 102, 163114. [Google Scholar] [CrossRef] [Green Version]
- Eastman, J. Novel Thermal Properties of Nanostructured Materials; Argonne National Lab.: Lemont, IL, USA, 1999. Available online: https://www.osti.gov/servlets/purl/10862 (accessed on 23 July 2020).
- Rasheed, A.K.; Khalid, M.; Rashmi, W.; Gupta, T.C.S.M.; Chan, A. Graphene based nanofluids and nanolubricants—Review of recent developments. Renew. Sustain. Energy Rev. 2016, 63, 346–362. [Google Scholar] [CrossRef]
- Hassani, S.; Saidur, R.; Mekhilef, S.; Hepbasli, A. A new correlation for predicting the thermal conductivity of nanofluids; using dimensional analysis. Int. J. Heat Mass Transf. 2015, 90, 121–130. [Google Scholar] [CrossRef]
- Ilhan, B.; Kurt, M.; Ertürk, H. Experimental investigation of heat transfer enhancement and viscosity change of hBN nanofluids. Exp. Therm. Fluid Sci. 2016, 77, 272–283. [Google Scholar] [CrossRef]
- Qayyum, M.S.; Hayat, H.; Matharu, R.K.; Tabish, T.A.; Edirisinghe, M. Boron nitride nanoscrolls: Structure, synthesis, and applications. Appl. Phys. Rev. 2019, 6, 021310. [Google Scholar] [CrossRef]
- Krishnam, M.; Bose, S.; Das, C. Boron nitride (BN) nanofluids as cooling agent in thermal management system (TMS). Appl. Therm. Eng. 2016, 106, 951–958. [Google Scholar] [CrossRef]
- Taha-Tijerina, J.; Peña-Paras, L.; Narayanan, T.N.; Garza, L.; Lapray, C.; Gonzalez, J.; Palacios, E.; Molina, D.; García, A.; Maldonado, D.; et al. Multifunctional nanofluids with 2D nanosheets for thermal and tribological management. Wear 2013, 302, 1241–1248. [Google Scholar] [CrossRef]
- Aravind, S.J.; Baskar, P.; Baby, T.T.; Sabareesh, R.K.; Das, S.; Ramaprabhu, S. Investigation of structural stability, dispersion, viscosity, and conductive heat transfer properties of functionalized carbon nanotube based nanofluids. J. Phys. Chem. C 2011, 115, 16737–16744. [Google Scholar] [CrossRef]
- Sissons, J.; Kim, K.S.; Stins, M.; Jayasekera, S.; Alsam, S.; Khan, N.A. Acanthamoeba castellanii Induces Host Cell Death via a Phosphatidylinositol 3-Kinase-Dependent Mechanism. Infect. Immun. 2005, 73, 2704–2708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siddiqui, R.; Saleem, S.; Khan, N.A. The effect of peptidic and non-peptidic proteasome inhibitors on the biological properties of Acanthamoeba castellanii belonging to the T4 genotype. Exp. Parasitol. 2016, 168, 16–24. [Google Scholar] [CrossRef] [PubMed]
- Khan, N.A.; Jarroll, E.L.; Paget, T.A. Acanthamoeba can be differentiated by the polymerase chain reaction and simple plating assays. Curr. Microbiol. 2001, 43, 204–208. [Google Scholar] [CrossRef] [PubMed]
- Gabriel, S.; Rasheed, A.K.; Siddiqui, R.; Appaturi, J.N.; Fen, L.B.; Khan, N.A. Development of nanoparticle-assisted PCR assay in the rapid detection of brain-eating amoebae. Parasitol. Res. 2018, 117, 1801–1811. [Google Scholar] [CrossRef] [PubMed]
- Khaliq, A.; Sonawane, P.J.; Sasi, B.K.; Sahu, B.S.; Pradeep, T.; Das, S.K.; Mahapatra, N.R. Enhancement in the efficiency of polymerase chain reaction by TiO2 nanoparticles: Crucial role of enhanced thermal conductivity. Nanotechnology 2010, 21, 255704. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abdul Khaliq, R.; Raed, K.; Hamzah Mohd, S.; Waleed Fekry, F. Enhancing the efficiency of polymerase chain reaction using graphene nanoflakes. Nanotechnology 2012, 23, 455106. [Google Scholar] [CrossRef]
- Kong, H.-H.; Chung, D.-I. PCR and RFLP variation of conserved region of small subunit ribosomal DNA among Acanthamoeba isolates assigned to either A. castellanii or A. polyphaga. Korean J. Parasitol. 1996, 34, 127–134. [Google Scholar] [CrossRef]
- Yousuf, F.A.; Siddiqui, R.; Subhani, F.; Khan, N.A. Status of free-living amoebae (Acanthamoeba spp., Naegleria fowleri, Balamuthia mandrillaris) in drinking water supplies in Karachi, Pakistan. J. Water Health 2013, 11, 371–375. [Google Scholar] [CrossRef]
- Zhi, C.; Xu, Y.; Bando, Y.; Golberg, D. Highly Thermo-conductive Fluid with Boron Nitride Nanofillers. Acs Nano 2011, 5, 6571–6577. [Google Scholar] [CrossRef]
- Nazarov, A.S.; Demin, V.N.; Grayfer, E.D.; Bulavchenko, A.I.; Arymbaeva, A.T.; Shin, H.-J.; Choi, J.-Y.; Fedorov, V.E. Functionalization and Dispersion of Hexagonal Boron Nitride (h-BN) Nanosheets Treated with Inorganic Reagents. Chem. Asian J. 2012, 7, 554–560. [Google Scholar] [CrossRef] [PubMed]
- Lahiri, D.; Rouzaud, F.; Richard, T.; Keshri, A.K.; Bakshi, S.R.; Kos, L.; Agarwal, A. Boron nitride nanotube reinforced polylactide–polycaprolactone copolymer composite: Mechanical properties and cytocompatibility with osteoblasts and macrophages in vitro. Acta Biomater. 2010, 6, 3524–3533. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Wang, X. DNA Sequencing by Hexagonal Boron Nitride Nanopore: A Computational Study. Nanomaterial 2016, 6, 111. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Zou, X.; Zhou, G.; Liu, R.; Wu, J.; Li, J.; Duan, W. Adsorption of DNA/RNA nucleobases on hexagonal boron nitride sheet: An ab initio study. Phys. Chem. Chem. Phys. 2011, 13, 12225–12230. [Google Scholar] [CrossRef]







| Protist | Gene-Specific Primers | Primer Properties | Amplicon | Reference |
|---|---|---|---|---|
| Acanthamoeba | Forward primer 5′-TTTGAATTCGCTCCA ATAGCGTATATTAA-3′ Reverse primer 5′-TTTGAATTCAGAAA GAGCTATCAATCTGT-3′ | Tm: 55.1 °C GC content: 31% Tm: 55.1 °C GC content: 31% | 950 bp | (Kong and Chung, 1996) [31] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rasheed, A.K.; Siddiqui, R.; Ahmed, S.M.K.; Gabriel, S.; Jalal, M.Z.; John, A.; Khan, N.A. hBN Nanoparticle-Assisted Rapid Thermal Cycling for the Detection of Acanthamoeba. Pathogens 2020, 9, 824. https://doi.org/10.3390/pathogens9100824
Rasheed AK, Siddiqui R, Ahmed SMK, Gabriel S, Jalal MZ, John A, Khan NA. hBN Nanoparticle-Assisted Rapid Thermal Cycling for the Detection of Acanthamoeba. Pathogens. 2020; 9(10):824. https://doi.org/10.3390/pathogens9100824
Chicago/Turabian StyleRasheed, Abdul Khaliq, Ruqaiyyah Siddiqui, Salma Mohammed Kabir Ahmed, Shobana Gabriel, Mohammed Zayan Jalal, Akbar John, and Naveed Ahmed Khan. 2020. "hBN Nanoparticle-Assisted Rapid Thermal Cycling for the Detection of Acanthamoeba" Pathogens 9, no. 10: 824. https://doi.org/10.3390/pathogens9100824







