The Effect of Coconut Oil Addition to Feed of Pigs on Rectal Microbial Diversity and Bacterial Abundance
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Diet and Animal Feeding
2.2. Nutritional Parameter Determination of Feed Rations
2.3. Sample Collection
2.4. Bacterial DNA Extraction
2.5. 16S rRNA Amplicon Sequencing
2.6. DNA Sequencing Data Analysis
2.7. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Willing, B.; Van Kessel, A. Intestinal microbiota differentially affect brush border enzyme activity and gene expression in the neonatal gnotobiotic pig. J. Anim. Physiol. Anim. Nutr. 2009, 93, 586–595. [Google Scholar] [CrossRef] [PubMed]
- Brestoff, J.R.; Artis, D. Commensal bacteria at the interface of host metabolism and the immune system. Nat. Immunol. 2013, 14, 676–684. [Google Scholar] [CrossRef] [PubMed]
- Jacobi, S.K.; Odle, J. Nutritional factors influencing intestinal health of the neonate. Adv. Nutr. 2012, 3, 687–696. [Google Scholar] [CrossRef] [PubMed]
- Che, L.; Chen, H.; Yu, B.; He, J.; Zheng, P.; Mao, X.; Yu, J.; Huang, Z.; Chen, D. Long-Term Intake of Pea Fiber Affects Colonic Barrier Function, Bacterial and Transcriptional Profile in Pig Model. Nutr. Cancer 2014, 66, 388–399. [Google Scholar] [CrossRef]
- Fouhse, J.; Zijlstra, R.; Willing, B. The role of gut microbiota in the health and disease of pigs. Anim. Front. 2016, 6, 30–36. [Google Scholar] [CrossRef]
- Imrich, I.; Mlyneková, E.; Mlynek, J.; Mad’arová, L.; Kanka, T.; Moravcová, L. Behavioural activity in pigs at habitual test in relation to production traits. Pol. J. Nat. Sci. 2012, 27, 257–267. [Google Scholar]
- Zeman, L.; Doležal, P.; Kopriva, A.; Mrkvicová, E.; Procházková, J.; Ryant, P.; Skládanka, J.; Straková, E.; Suchý, P.; Veselý, P.; et al. Nutrition and Feeding of Farm Animals; Profi Press: Prague, Czech Republic, 2016; p. 360. [Google Scholar]
- Van der Klis, J.; Jansman, A. Optimising nutrient digestion, absorption and gut barrier function in monogastrics: Reality or illusion. In Nutrition and Health of the Gastrointestinal Tract; Wageningen Academic Publishers: Wageningen, The Netherlands, 2002; pp. 15–36. [Google Scholar]
- Winter, S.E.; Winter, M.G.; Xavier, M.N.; Thiennimitr, P.; Poon, V.; Keestra, A.M.; Laughlin, R.C.; Gomez, G.; Wu, J.; Lawhon, S.D.; et al. Host-Derived Nitrate Boosts Growth of E. coli in the Inflamed Gut. Science 2013, 339, 708. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Liu, Q.; Zhang, W.M.; Zhang, Z.J.; Wang, W.L.; Zhuang, S. Gastrointestinal microbial diversity and short-chain fatty acid production in pigs fed different fibrous diets with or without cell wall-degrading enzyme supplementation. Livest. Sci. 2018, 207, 105–116. [Google Scholar] [CrossRef]
- Hanczakowska, E. The use of medium-chain fatty acids in piglet feeding—A review. Ann. Anim. Sci. 2017, 17, 967–977. [Google Scholar] [CrossRef]
- Puyalto, M.; Sol, C.; Mallo, J.J. Coconut oil: A new option for controlling pig pathogens. Pig Prog. 2016, 32, 35–36. [Google Scholar]
- AOAC. Official Methods of Analysis AOAC, 17th ed.; Association of Analytical Communities: Gaithersburg, MD, USA, 2000. [Google Scholar]
- Davídek, J.; Hrdlička, J.; Karvánek, M.; Pokorný, J.; Seifert, J.; Velíšek, J. Laboratory Handbook of Food Analysis; SNTL: Prague, Czech Republic, 1981. (In Czech) [Google Scholar]
- Juráček, M.; Vašeková, P.; Bíro, D.; Šimko, M.; Gálik, B.; Rolinec, M.; Hanušovský, O.; Kolláthová, R.; Kovár, P.; Andruška, N. The effect of different feeding system on fatty acids composition of cow’s milk. Acta Fytotech. Zootech. 2020, 23, 37–41. [Google Scholar] [CrossRef]
- Choudhury, R.; Middelkoop, A.; Bolhuis, J.E.; Kleerebezem, M. Legitimate and Reliable Determination of the Age-Related Intestinal Microbiome in Young Piglets; Rectal Swabs and Fecal Samples Provide Comparable Insights. Front. Microbiol. 2019, 10, 1886. [Google Scholar] [CrossRef] [PubMed]
- Wegl, G.; Rolinec, M.; Nagl, V.; Gierus, M.; Klose, V. Effect of oxytetracycline as well as an acid-based feed supplement on the prevalence and abundance of selected antibiotic resistance genes in weaned piglets. Anim. Husb. Dairy Vet. Sci. 2017, 1, 1–6. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108, 4516. [Google Scholar] [CrossRef]
- Větrovský, T.; Baldrian, P.; Morais, D. SEED 2: A user-friendly platform for amplicon high-throughput sequencing data analyses. Bioinformatics 2018, 34, 2292–2294. [Google Scholar] [CrossRef]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- Stoddard, S.F.; Smith, B.J.; Hein, R.; Roller, B.R.; Schmidt, T.M. rrn DB: Improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res. 2015, 43, D593–D598. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Oksanen, J.; Kindt, R.; Legendre, P.; O’Hara, B.; Stevens, M.H.H.; Oksanen, M.J.; Suggests, M. The vegan package. Community Ecol. Package 2007, 10, 631–637. [Google Scholar]
- Chen, J.; Bittinger, K.; Charlson, E.S.; Hoffmann, C.; Lewis, J.; Wu, G.D.; Collman, R.G.; Bushman, F.D.; Li, H. Associating microbiome composition with environmental covariates using generalized UniFrac distances. Bioinformatics 2012, 28, 2106–2113. [Google Scholar] [CrossRef]
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Guo, Y.; Sheng, Q.; Shyr, Y. Advanced heat map and clustering analysis using heatmap3. Biomed Res. Int. 2014, 2014, 986048. [Google Scholar] [CrossRef] [PubMed]
- Holman, D.B.; Brunelle, B.W.; Trachsel, J.; Allen, H.K. Meta-analysis To Define a Core Microbiota in the Swine Gut. mSystems 2017, 2. [Google Scholar] [CrossRef] [PubMed]
- Richards, J.D.; Gong, J.; de Lange, C.F.M. The gastrointestinal microbiota and its role in monogastric nutrition and health with an emphasis on pigs: Current understanding, possible modulations, and new technologies for ecological studies. Can. J. Anim. Sci. 2005, 85, 421–435. [Google Scholar] [CrossRef]
- Liao, S.F.; Nyachoti, M. Using probiotics to improve swine gut health and nutrient utilization. Anim. Nutr. 2017, 3, 331–343. [Google Scholar] [CrossRef]
- Bajagai, Y.S.; Klieve, A.V.; Dart, P.J.; Bryden, W.L. Probiotics in Animal Nutrition—Production, Impact and Regulation; FAO Animal Production and Health Paper; FAO: Rome, Italy, 2016. [Google Scholar]
- Gresse, R.; Chaucheyras Durand, F.; Dunière, L.; Blanquet-Diot, S.; Forano, E. Microbiota Composition and Functional Profiling Throughout the Gastrointestinal Tract of Commercial Weaning Piglets. Microorganisms 2019, 7, 343. [Google Scholar] [CrossRef]
- De Filippis, F.; Pellegrini, N.; Vannini, L.; Jeffery, I.B.; La Storia, A.; Laghi, L.; Serrazanetti, D.I.; Di Cagno, R.; Ferrocino, I.; Lazzi, C.; et al. High-level adherence to a Mediterranean diet beneficially impacts the gut microbiota and associated metabolome. Gut 2016, 65, 1812–1821. [Google Scholar] [CrossRef]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef]
- Biddle, A.; Stewart, L.; Blanchard, J.; Leschine, S. Untangling the Genetic Basis of Fibrolytic Specialization by Lachnospiraceae and Ruminococcaceae in Diverse Gut Communities. Diversity 2013, 5, 627–640. [Google Scholar] [CrossRef]
- Looft, T.; Allen, H.K.; Casey, T.A.; Alt, D.P.; Stanton, T.B. Carbadox has both temporary and lasting effects on the swine gut microbiota. Front. Microbiol. 2014, 5, 276. [Google Scholar] [CrossRef]
- Vital, M.; Howe, A.C.; Tiedje, J.M. Revealing the Bacterial Butyrate Synthesis Pathways by Analyzing (Meta)genomic Data. mBio 2014, 5, e00889-14. [Google Scholar] [CrossRef] [PubMed]
- Hamer, H.M.; Jonkers, D.; Venema, K.; Vanhoutvin, S.; Troost, F.J.; Brummer, R.J. Review article: The role of butyrate on colonic function. Aliment. Pharmacol. Ther. 2008, 27, 104–119. [Google Scholar] [CrossRef] [PubMed]
- Zábranský, L.; Šoch, M.; Čermák, B.; Lád, F.; Novák, P.; Kadlec, J.; Maršálek, M. Effect of Feed Supplements on the Occurrence of Coccidia Oocysts in the Digestive Tract of Pheasants. Acta Fytotech. Zootech. 2016, 19, 51–53. [Google Scholar] [CrossRef][Green Version]
- Megahed, A.; Zeineldin, M.; Evans, K.; Maradiaga, N.; Blair, B.; Aldridge, B.; Lowe, J. Impacts of environmental complexity on respiratory and gut microbiome community structure and diversity in growing pigs. Sci. Rep. 2019, 9, 13773. [Google Scholar] [CrossRef] [PubMed]
- Guevarra, R.B.; Hong, S.H.; Cho, J.H.; Kim, B.-R.; Shin, J.; Lee, J.H.; Kang, B.N.; Kim, Y.H.; Wattanaphansak, S.; Isaacson, R.E.; et al. The dynamics of the piglet gut microbiome during the weaning transition in association with health and nutrition. J. Anim. Sci. Biotechnol. 2018, 9, 54. [Google Scholar] [CrossRef]
- Pajarillo, E.A.B.; Chae, J.P.; Balolong, M.P.; Kim, H.B.; Seo, K.-S.; Kang, D.-K. Characterization of the Fecal Microbial Communities of Duroc Pigs Using 16S rRNA Gene Pyrosequencing. Asian Australas J. Anim. Sci. 2015, 28, 584–591. [Google Scholar] [CrossRef]
- Ige, B.A. Probiotics use in intensive fish farming. Afr. J. Microbiol. 2013, 7, 2701–2711. [Google Scholar] [CrossRef]
- Dougal, K.; de la Fuente, G.; Harris, P.A.; Girdwood, S.E.; Pinloche, E.; Geor, R.J.; Nielsen, B.D.; Schott II, H.C.; Elzinga, S.; Newbold, C.J. Characterisation of the faecal bacterial community in adult and elderly horses fed a high fibre, high oil or high starch diet using 454 pyrosequencing. PLoS ONE 2014, 9, e87424. [Google Scholar] [CrossRef]
- Gueimonde, M.; Jalonen, L.; He, F.; Hiramatsu, M.; Salminen, S. Adhesion and competitive inhibition and displacement of human enteropathogens by selected lactobacilli. Food Res. Int. 2006, 39, 467–471. [Google Scholar] [CrossRef]
- Hopwood, D.; Hampson, D. Interactions between the intestinal microflora, diet and diarrhoea, and their influences on piglet health in the immediate post-weaning period. In Weaning the Pig: Concepts and Consequences; Wageningen Academic Publishers: Wageningen, The Netherlands, 2003; pp. 199–217. [Google Scholar] [CrossRef]
- Han, Y.-K.; Hwang, I.H.; Thacker, P.A. Use of a micro-encapsulated eucalyptus-medium chain fatty acid product as an alternative to zinc oxide and antibiotics for weaned pigs. J. Swine Health Prod. 2011, 19, 34–43. [Google Scholar]
- Hanczakowska, E.; Szewczyk, A.; Swiatkiewicz, M.; Okon, K. Short-and medium-chain fatty acids as a feed supplement for weaning and nursery pigs. Pol. J. Vet. Sci. 2013, 16, 647–654. [Google Scholar] [CrossRef] [PubMed]
- Zentek, J.; Ferrara, F.; Pieper, R.; Tedin, L.; Meyer, W.; Vahjen, W. Effects of dietary combinations of organic acids and medium chain fatty acids on the gastrointestinal microbial ecology and bacterial metabolites in the digestive tract of weaning piglets. J. Anim. Sci. 2013, 91, 3200–3210. [Google Scholar] [CrossRef] [PubMed]
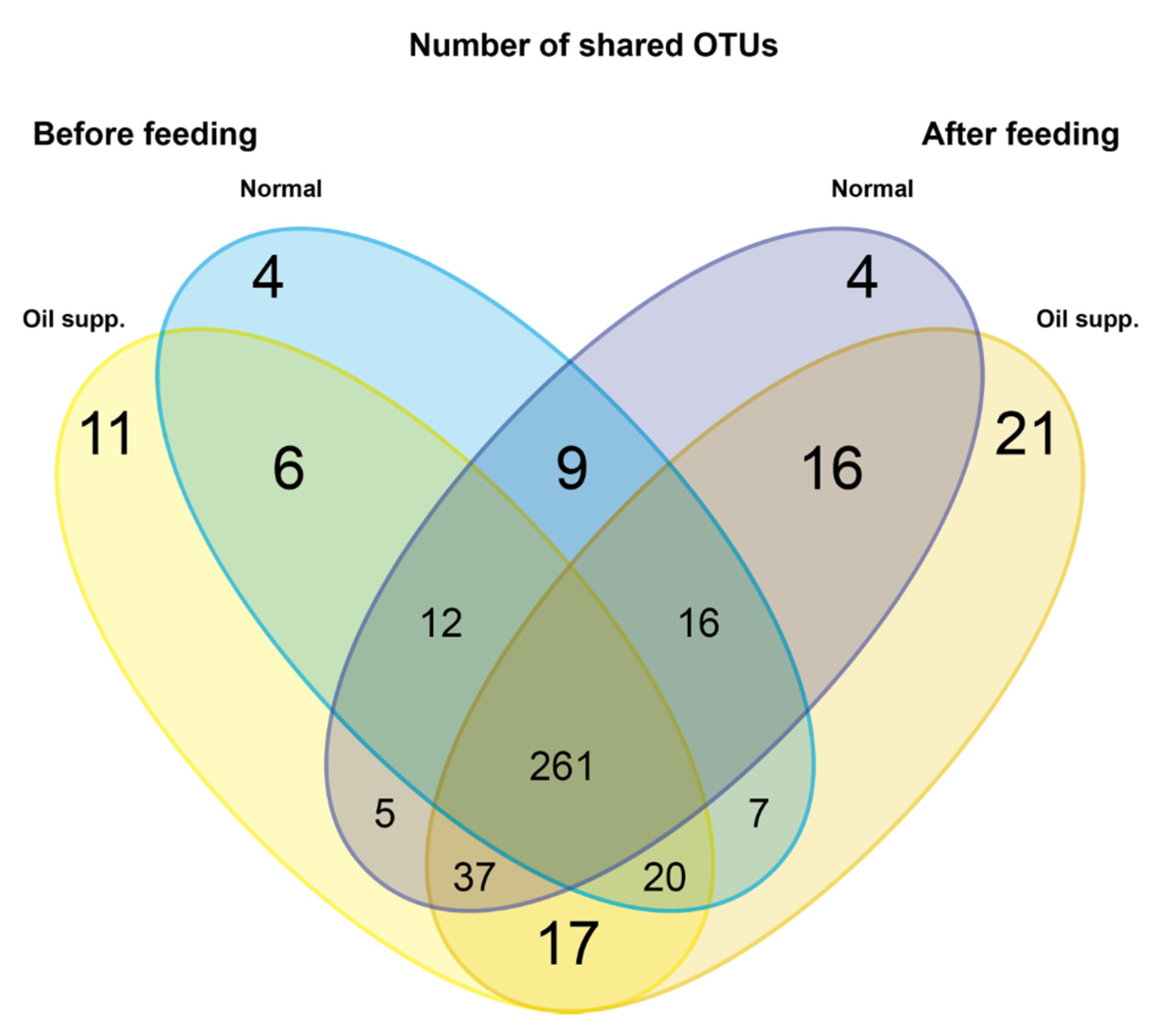
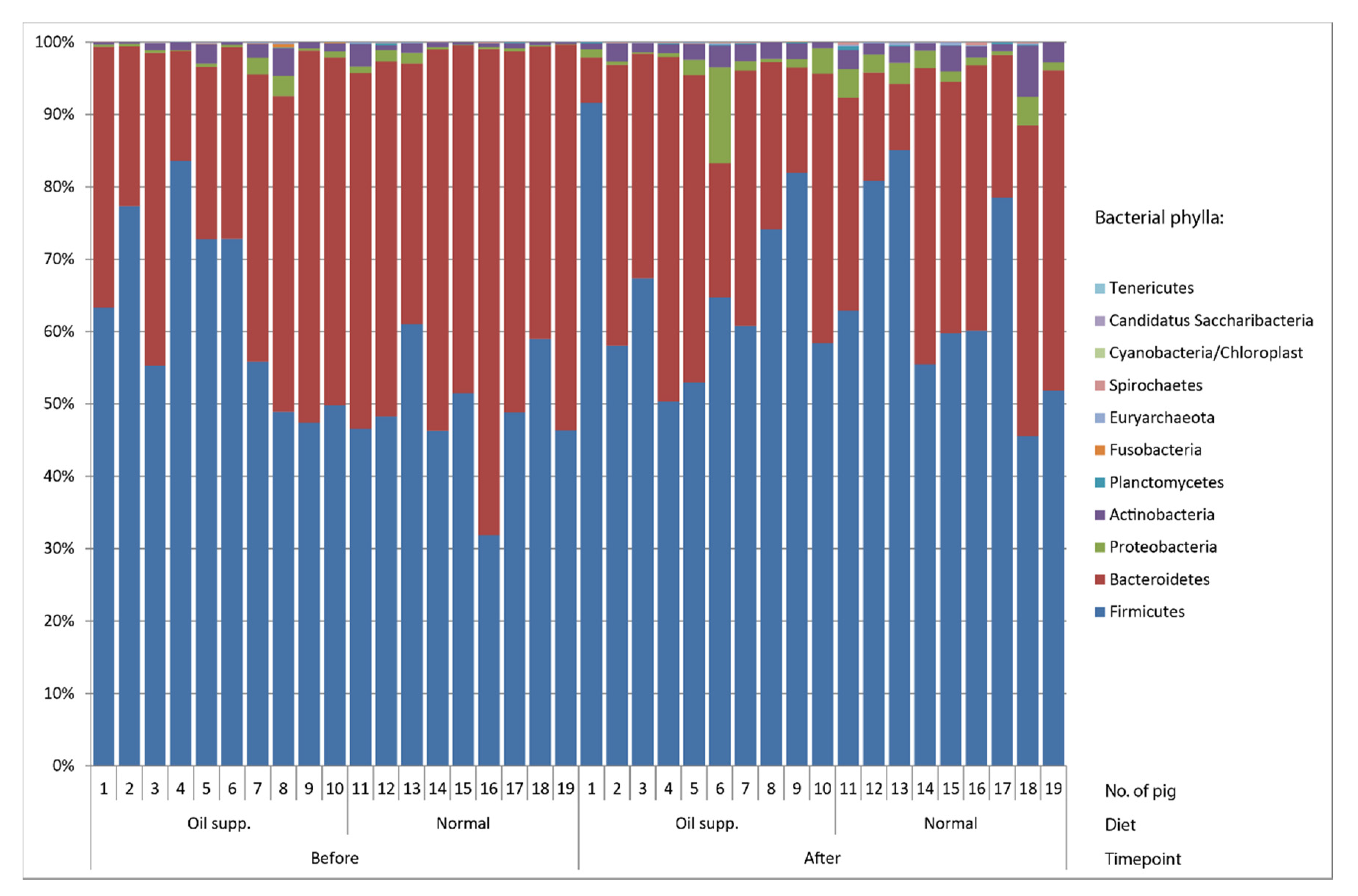
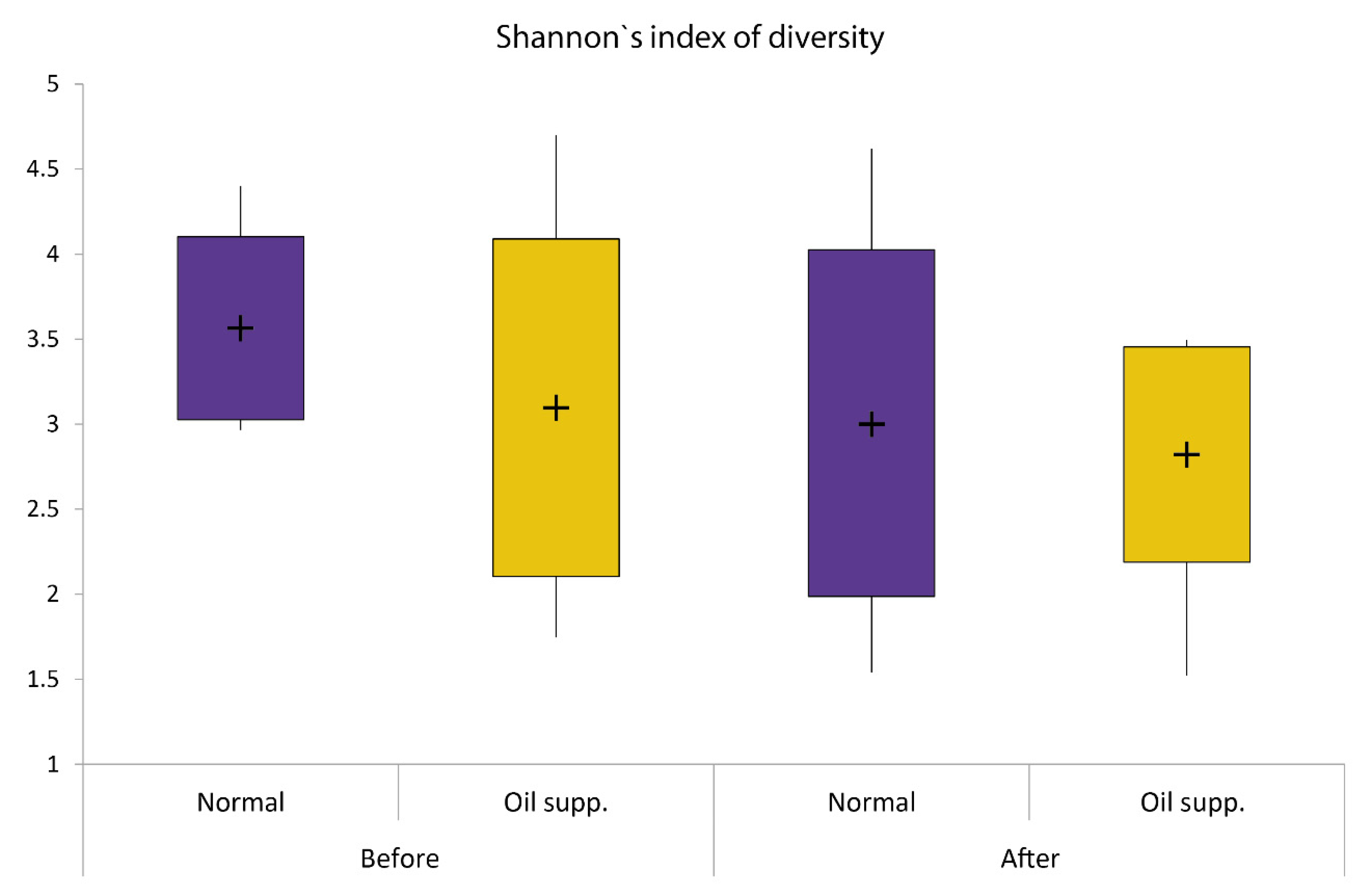
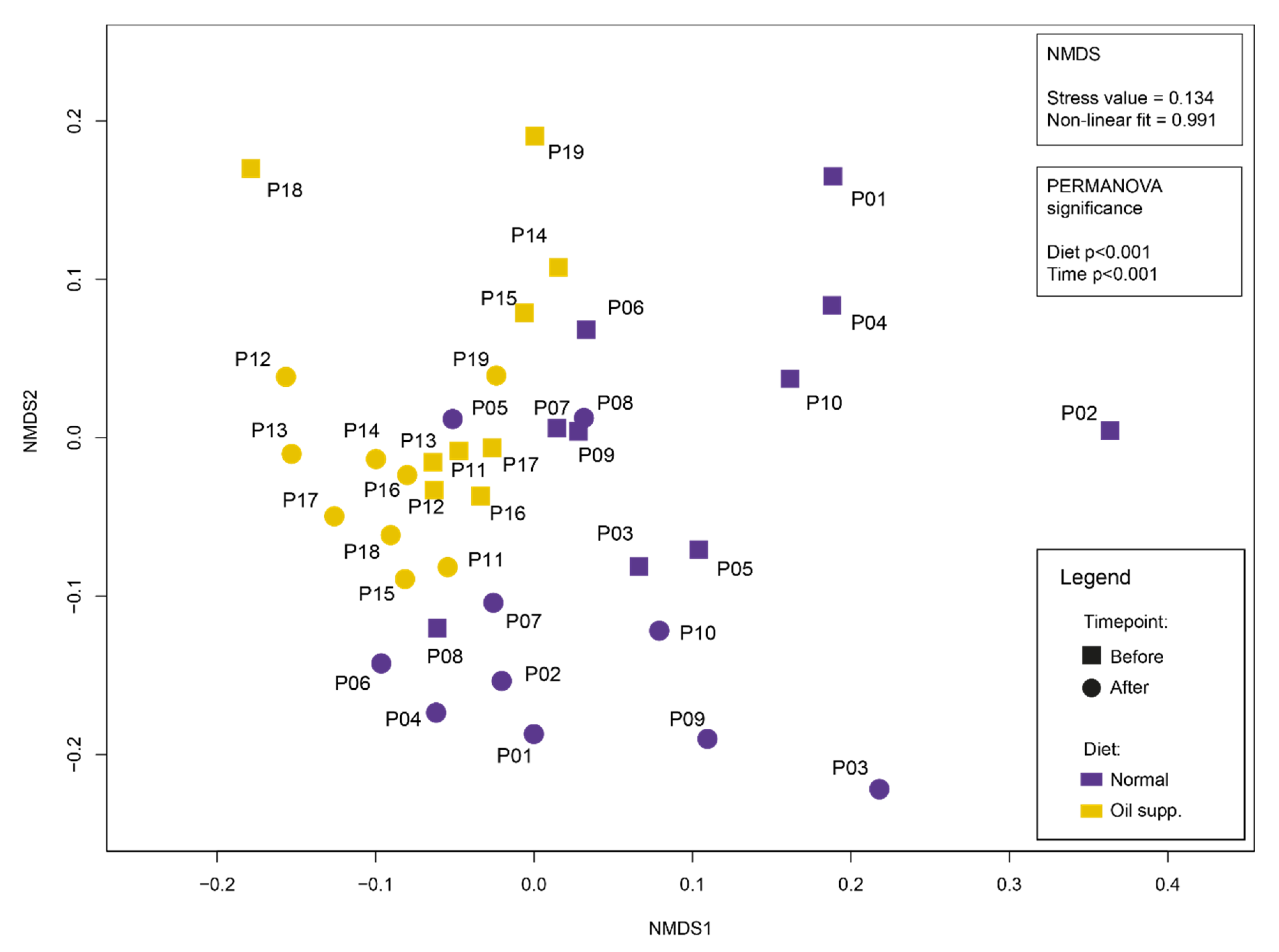
| Pigs Group | Normal | Oil Supplement |
|---|---|---|
| Composition of feed ration (%) | ||
| Commercial feed mixture | 100 | 99.70 |
| Commercial coconut oil | - | 0.30 |
| Nutritional parameters (%) | ||
| Dry matter | 89.30 | 89.10 |
| Crude protein | 15.90 | 16.27 |
| Lysine | 1.16 | 1.15 |
| Crude fat | 3.33 | 4.04 |
| Crude fiber | 4.23 | 4.76 |
| Nitrogen-free extract | 61.85 | 60.20 |
| Starch | 47.20 | 43.90 |
| Sugar | 2.98 | 2.40 |
| Ash | 3.99 | 3.83 |
| MEpigs (MJ.kg−1) | 15.70 | 15.80 |
| Proportion of fatty acids (g.100 g−1 Fatty acids) | ||
| C8:0 | - | 0.73 |
| C10:0 | - | 0.64 |
| C12:0 | 0.25 | 5.34 |
| C14:0 | 0.67 | 2.49 |
| C16:0 | 18.73 | 17.68 |
| C16:1 | 1.72 | 1.60 |
| C17:0 | 0.21 | 0.19 |
| C18:0 | 5.67 | 5.47 |
| C18:1cis n9 | 27.81 | 25.80 |
| C18:2cis n6 | 38.35 | 34.09 |
| C18:3 n3 | 3.34 | 2.94 |
| C20:0 | 0.20 | 0.19 |
| C20:1 n9 | 0.48 | 0.44 |
| C20:2 n6 | 0.18 | 0.16 |
| C20:4 n6 | 0.13 | 0.12 |
| C22:0 | 0.14 | 0.12 |
| Σ unidentified | 2.12 | 2.00 |
| PUFA | 42.00 | 37.31 |
| MUFA | 30.01 | 27.84 |
| SFA | 25.87 | 32.85 |
| Σn3/Σn6 | 0.09 | 0.09 |
| Σn6/Σn3 | 11.57 | 11.69 |
| Taxonomic Rank | Relative Abundance (%) | p-Value | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Phyllum | Class | Order | Family | Genus | OTU | Normal diet Before treatment | Normal diet After treatment | Oil supp. group Before treatment | Oil supp. group After treatment | In-time change in Normal diet group | In-time change in Oil supp. group | Difference Normal-Oil Before treatment | Difference Normal-Oil After treatment |
| Actinobacteria | 0.87 | 2.60 | 1.33 | 1.85 | 0.020 | 0.232 | 0.315 | 0.356 | |||||
| Actinobacteria | 0.87 | 2.60 | 1.33 | 1.85 | 0.020 | 0.232 | 0.315 | 0.356 | |||||
| Actinomycetales | 0.42 | 0.79 | 0.52 | 0.21 | 0.098 | 0.193 | 0.842 | 0.006 | |||||
| Corynebacteriaceae | 0.41 | 0.77 | 0.51 | 0.20 | 0.098 | 0.160 | 0.780 | 0.006 | |||||
| Corynebacterium | 0.41 | 0.77 | 0.51 | 0.20 | 0.098 | 0.160 | 0.780 | 0.006 | |||||
| Coriobacteriales | 0.45 | 1.79 | 0.79 | 1.42 | 0.004 | 0.002 | 0.315 | 0.780 | |||||
| Coriobacteriaceae | 0.45 | 1.79 | 0.79 | 1.42 | 0.004 | 0.002 | 0.315 | 0.780 | |||||
| Enterorhabdus | 0.03 | 0.36 | 0.12 | 0.53 | 0.012 | 0.004 | 0.010 | 0.211 | |||||
| OTU053 | 0.03 | 0.36 | 0.12 | 0.52 | 0.012 | 0.006 | 0.006 | 0.243 | |||||
| Olsenella | 0.31 | 1.15 | 0.52 | 0.45 | 0.203 | 0.695 | 0.842 | 0.211 | |||||
| OTU067 | 0.05 | 0.61 | 0.05 | 0.07 | 0.039 | 0.944 | 0.773 | 0.015 | |||||
| Bacteroidetes | 49.53 | 30.31 | 34.96 | 29.49 | 0.008 | 0.492 | 0.013 | 0.905 | |||||
| Bacteroidia | 49.46 | 30.28 | 34.96 | 29.48 | 0.008 | 0.492 | 0.013 | 0.905 | |||||
| Bacteroidales | 49.46 | 30.28 | 34.96 | 29.48 | 0.008 | 0.492 | 0.013 | 0.905 | |||||
| Porphyromonadaceae | 0.42 | 1.13 | 0.49 | 1.10 | 0.020 | 0.037 | 0.905 | 0.905 | |||||
| Barnesiella | 0.35 | 0.93 | 0.38 | 0.90 | 0.098 | 0.037 | 0.842 | 0.905 | |||||
| OTU022 | 0.29 | 0.85 | 0.29 | 0.60 | 0.098 | 0.049 | 0.720 | 0.604 | |||||
| Prevotellaceae | 48.98 | 29.00 | 34.44 | 28.22 | 0.008 | 0.432 | 0.017 | 1.000 | |||||
| Alloprevotella | 1.14 | 0.18 | 0.35 | 1.04 | 0.012 | 0.037 | 0.182 | 0.001 | |||||
| OTU033 | 1.01 | 0.12 | 0.16 | 0.34 | 0.012 | 0.131 | 0.030 | 0.133 | |||||
| OTU066 | 0.05 | 0.05 | 0.09 | 0.59 | 0.834 | 0.084 | 0.189 | 0.008 | |||||
| Prevotella | 47.80 | 28.77 | 34.05 | 27.12 | 0.008 | 0.322 | 0.017 | 0.842 | |||||
| OTU002 | 12.16 | 3.29 | 8.62 | 5.11 | 0.004 | 0.275 | 0.243 | 0.315 | |||||
| OTU003 | 7.54 | 6.48 | 5.52 | 6.23 | 0.496 | 0.922 | 0.113 | 0.780 | |||||
| OTU004 | 4.25 | 3.74 | 3.38 | 2.47 | 0.910 | 0.432 | 0.243 | 0.278 | |||||
| OTU006 | 3.31 | 2.10 | 2.66 | 2.61 | 0.164 | 0.625 | 0.400 | 0.842 | |||||
| OTU008 | 3.06 | 0.44 | 2.40 | 0.03 | 0.129 | 0.002 | 0.968 | 0.013 | |||||
| OTU009 | 2.29 | 1.15 | 0.78 | 1.24 | 0.652 | 0.432 | 0.004 | 0.905 | |||||
| OTU010 | 1.08 | 2.81 | 0.68 | 0.61 | 0.129 | 0.922 | 0.133 | 0.053 | |||||
| OTU012 | 1.73 | 0.64 | 1.32 | 1.18 | 0.027 | 1.000 | 0.278 | 0.028 | |||||
| OTU015 | 0.98 | 0.80 | 1.00 | 1.10 | 0.496 | 0.846 | 0.720 | 0.604 | |||||
| OTU016 | 0.88 | 1.19 | 1.13 | 0.34 | 0.426 | 0.557 | 0.968 | 0.065 | |||||
| OTU017 | 2.67 | 0.01 | 0.58 | 0.05 | 0.107 | 0.193 | 0.837 | 0.263 | |||||
| OTU023 | 0.80 | 0.61 | 0.54 | 0.34 | 0.820 | 0.906 | 0.487 | 0.243 | |||||
| OTU024 | 0.73 | 0.67 | 0.48 | 0.33 | 0.820 | 0.322 | 0.079 | 0.079 | |||||
| OTU026 | 0.77 | 0.26 | 0.36 | 0.40 | 0.055 | 1.000 | 0.053 | 0.780 | |||||
| OTU027 | 1.11 | 0.12 | 0.40 | 0.15 | 0.008 | 0.049 | 0.053 | 0.967 | |||||
| OTU030 | 0.55 | 0.25 | 0.41 | 0.38 | 0.004 | 0.625 | 0.356 | 0.356 | |||||
| OTU031 | 0.31 | 0.72 | 0.06 | 0.23 | 0.496 | 0.084 | 0.165 | 0.356 | |||||
| OTU037 | 0.06 | 0.07 | 0.10 | 0.81 | 0.353 | 0.037 | 0.539 | 0.006 | |||||
| OTU039 | 0.52 | 0.09 | 0.22 | 0.12 | 0.008 | 0.160 | 0.156 | 0.400 | |||||
| OTU041 | 0.25 | 0.54 | 0.12 | 0.02 | 0.834 | 0.294 | 0.838 | 0.072 | |||||
| OTU045 | 0.00 | 0.00 | 0.35 | 0.58 | 0.201 | 0.813 | 0.007 | 0.256 | |||||
| Firmicutes | 48.86 | 64.46 | 62.71 | 66.03 | 0.027 | 0.625 | 0.013 | 0.780 | |||||
| Bacilli | 1.61 | 2.55 | 1.22 | 3.17 | 0.098 | 0.064 | 0.661 | 0.780 | |||||
| Lactobacillales | 1.61 | 2.55 | 1.22 | 3.17 | 0.098 | 0.064 | 0.661 | 0.780 | |||||
| Lactobacillaceae | 1.60 | 2.52 | 1.16 | 3.10 | 0.098 | 0.049 | 0.720 | 0.604 | |||||
| Lactobacillus | 1.60 | 2.52 | 1.16 | 3.10 | 0.098 | 0.049 | 0.720 | 0.604 | |||||
| OTU013 | 0.71 | 0.75 | 0.63 | 1.58 | 0.734 | 0.027 | 0.968 | 0.035 | |||||
| OTU014 | 0.44 | 1.03 | 0.21 | 1.28 | 0.129 | 0.432 | 0.905 | 0.004 | |||||
| OTU025 | 0.43 | 0.70 | 0.26 | 0.15 | 0.129 | 0.105 | 0.720 | 0.010 | |||||
| Clostridia | 1.16 | 1.57 | 2.29 | 1.94 | 0.570 | 0.846 | 0.497 | 0.497 | |||||
| Clostridiales | 1.16 | 1.57 | 2.29 | 1.94 | 0.570 | 0.846 | 0.497 | 0.497 | |||||
| Lachnospiraceae | 0.52 | 0.71 | 0.88 | 0.51 | 0.496 | 0.131 | 0.400 | 0.604 | |||||
| Ruminococcaceae | 0.49 | 0.67 | 1.00 | 0.98 | 0.250 | 0.922 | 0.315 | 0.133 | |||||
| Negativicutes | 46.00 | 60.15 | 59.03 | 60.85 | 0.039 | 0.922 | 0.053 | 0.968 | |||||
| Selenomonadales | 46.00 | 60.15 | 59.03 | 60.85 | 0.039 | 0.922 | 0.053 | 0.968 | |||||
| Acidaminococcaceae | 0.38 | 0.59 | 1.21 | 1.33 | 0.164 | 0.846 | 0.278 | 0.356 | |||||
| Phascolarctobacterium | 0.21 | 0.28 | 0.86 | 1.20 | 0.496 | 0.695 | 0.356 | 0.113 | |||||
| OTU021 | 0.21 | 0.28 | 0.86 | 1.20 | 0.496 | 0.695 | 0.356 | 0.113 | |||||
| Veillonellaceae | 45.62 | 59.56 | 57.83 | 59.51 | 0.039 | 0.922 | 0.053 | 0.968 | |||||
| Dialister | 1.85 | 3.34 | 2.62 | 1.99 | 0.203 | 0.432 | 0.780 | 0.243 | |||||
| OTU005 | 1.79 | 3.21 | 2.48 | 1.83 | 0.203 | 0.432 | 0.842 | 0.211 | |||||
| Megasphaera | 37.27 | 52.80 | 50.08 | 55.24 | 0.027 | 0.375 | 0.182 | 0.780 | |||||
| OTU001 | 35.28 | 50.88 | 47.74 | 53.03 | 0.027 | 0.322 | 0.182 | 0.780 | |||||
| OTU020 | 0.54 | 0.26 | 0.65 | 0.59 | 0.098 | 0.492 | 0.842 | 0.156 | |||||
| Mitsuokella | 1.78 | 1.93 | 1.26 | 0.62 | 0.820 | 0.004 | 0.095 | 0.028 | |||||
| OTU018 | 0.88 | 0.59 | 0.49 | 0.13 | 0.129 | 0.002 | 0.079 | 0.095 | |||||
| OTU019 | 0.60 | 0.66 | 0.46 | 0.29 | 0.910 | 0.037 | 0.243 | 0.035 | |||||
| OTU029 | 0.18 | 0.62 | 0.13 | 0.17 | 0.250 | 0.846 | 0.278 | 0.013 | |||||
| Propionispira | 1.58 | 0.45 | 1.39 | 0.86 | 0.020 | 0.131 | 1.000 | 0.156 | |||||
| OTU007 | 1.57 | 0.45 | 1.38 | 0.85 | 0.020 | 0.131 | 1.000 | 0.156 | |||||
| Selenomonas | 2.96 | 0.95 | 2.26 | 0.53 | 0.004 | 0.010 | 0.315 | 0.243 | |||||
| OTU011 | 2.33 | 0.42 | 1.54 | 0.30 | 0.004 | 0.027 | 0.243 | 0.447 | |||||
| Proteobacteria | 0.60 | 2.21 | 0.82 | 2.42 | 0.004 | 0.275 | 0.447 | 0.211 | |||||
| Deltaproteobacteria | 0.14 | 0.63 | 0.43 | 0.90 | 0.020 | 0.193 | 0.133 | 0.549 | |||||
| Bdellovibrionales | 0.12 | 0.43 | 0.41 | 0.84 | 0.055 | 0.275 | 0.035 | 0.497 | |||||
| Bdellovibrionaceae | 0.12 | 0.43 | 0.41 | 0.84 | 0.055 | 0.275 | 0.035 | 0.497 | |||||
| Vampirovibrio | 0.12 | 0.43 | 0.41 | 0.84 | 0.055 | 0.275 | 0.035 | 0.497 | |||||
| Epsilonproteobacteria | 0.05 | 0.66 | 0.10 | 0.44 | 0.012 | 0.275 | 0.437 | 0.182 | |||||
| Campylobacterales | 0.05 | 0.66 | 0.10 | 0.44 | 0.012 | 0.275 | 0.437 | 0.182 | |||||
| Campylobacteraceae | 0.05 | 0.66 | 0.10 | 0.41 | 0.012 | 0.275 | 0.437 | 0.182 | |||||
| Campylobacter | 0.05 | 0.66 | 0.10 | 0.41 | 0.012 | 0.275 | 0.437 | 0.182 | |||||
| OTU050 | 0.05 | 0.52 | 0.10 | 0.41 | 0.164 | 0.275 | 0.437 | 1.000 | |||||
| Gammaproteobacteria | 0.40 | 0.91 | 0.28 | 1.07 | 0.055 | 0.922 | 0.549 | 0.079 | |||||
| Enterobacteriales | 0.09 | 0.21 | 0.02 | 0.74 | 0.426 | 1.000 | 0.966 | 0.066 | |||||
| Enterobacteriaceae | 0.09 | 0.21 | 0.02 | 0.74 | 0.426 | 1.000 | 0.966 | 0.066 | |||||
| Escherichia/Shigella | 0.00 | 0.12 | 0.02 | 0.74 | 0.036 | 0.726 | 0.131 | 0.537 | |||||
| OTU028 | 0.00 | 0.12 | 0.02 | 0.74 | 0.036 | 0.726 | 0.131 | 0.537 | |||||
| Gammaproteobacteria | 0.40 | 0.91 | 0.28 | 1.07 | 0.055 | 0.922 | 0.549 | 0.079 | |||||
| Pseudomonadales | 0.20 | 0.50 | 0.12 | 0.02 | 0.426 | 0.037 | 0.182 | 0.003 | |||||
| Moraxellaceae | 0.20 | 0.50 | 0.12 | 0.02 | 0.426 | 0.037 | 0.182 | 0.003 | |||||
| Psychrobacter | 0.20 | 0.50 | 0.12 | 0.02 | 0.426 | 0.027 | 0.182 | 0.002 | |||||
| OTU043 | 0.20 | 0.50 | 0.12 | 0.02 | 0.426 | 0.027 | 0.182 | 0.002 | |||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rolinec, M.; Medo, J.; Gábor, M.; Miluchová, M.; Bíro, D.; Šimko, M.; Juráček, M.; Hanušovský, O.; Schubertová, Z.; Gálik, B. The Effect of Coconut Oil Addition to Feed of Pigs on Rectal Microbial Diversity and Bacterial Abundance. Animals 2020, 10, 1764. https://doi.org/10.3390/ani10101764
Rolinec M, Medo J, Gábor M, Miluchová M, Bíro D, Šimko M, Juráček M, Hanušovský O, Schubertová Z, Gálik B. The Effect of Coconut Oil Addition to Feed of Pigs on Rectal Microbial Diversity and Bacterial Abundance. Animals. 2020; 10(10):1764. https://doi.org/10.3390/ani10101764
Chicago/Turabian StyleRolinec, Michal, Juraj Medo, Michal Gábor, Martina Miluchová, Daniel Bíro, Milan Šimko, Miroslav Juráček, Ondrej Hanušovský, Zuzana Schubertová, and Branislav Gálik. 2020. "The Effect of Coconut Oil Addition to Feed of Pigs on Rectal Microbial Diversity and Bacterial Abundance" Animals 10, no. 10: 1764. https://doi.org/10.3390/ani10101764
APA StyleRolinec, M., Medo, J., Gábor, M., Miluchová, M., Bíro, D., Šimko, M., Juráček, M., Hanušovský, O., Schubertová, Z., & Gálik, B. (2020). The Effect of Coconut Oil Addition to Feed of Pigs on Rectal Microbial Diversity and Bacterial Abundance. Animals, 10(10), 1764. https://doi.org/10.3390/ani10101764






