Case Study on Increasing Breeding Value Estimation Reliability of Reproductive Traits in Serbian Highly Prolific Large White and Landrace Sows
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Phenotypes and Genotypes
2.2. Statistical Analyses
2.2.1. Applied Model
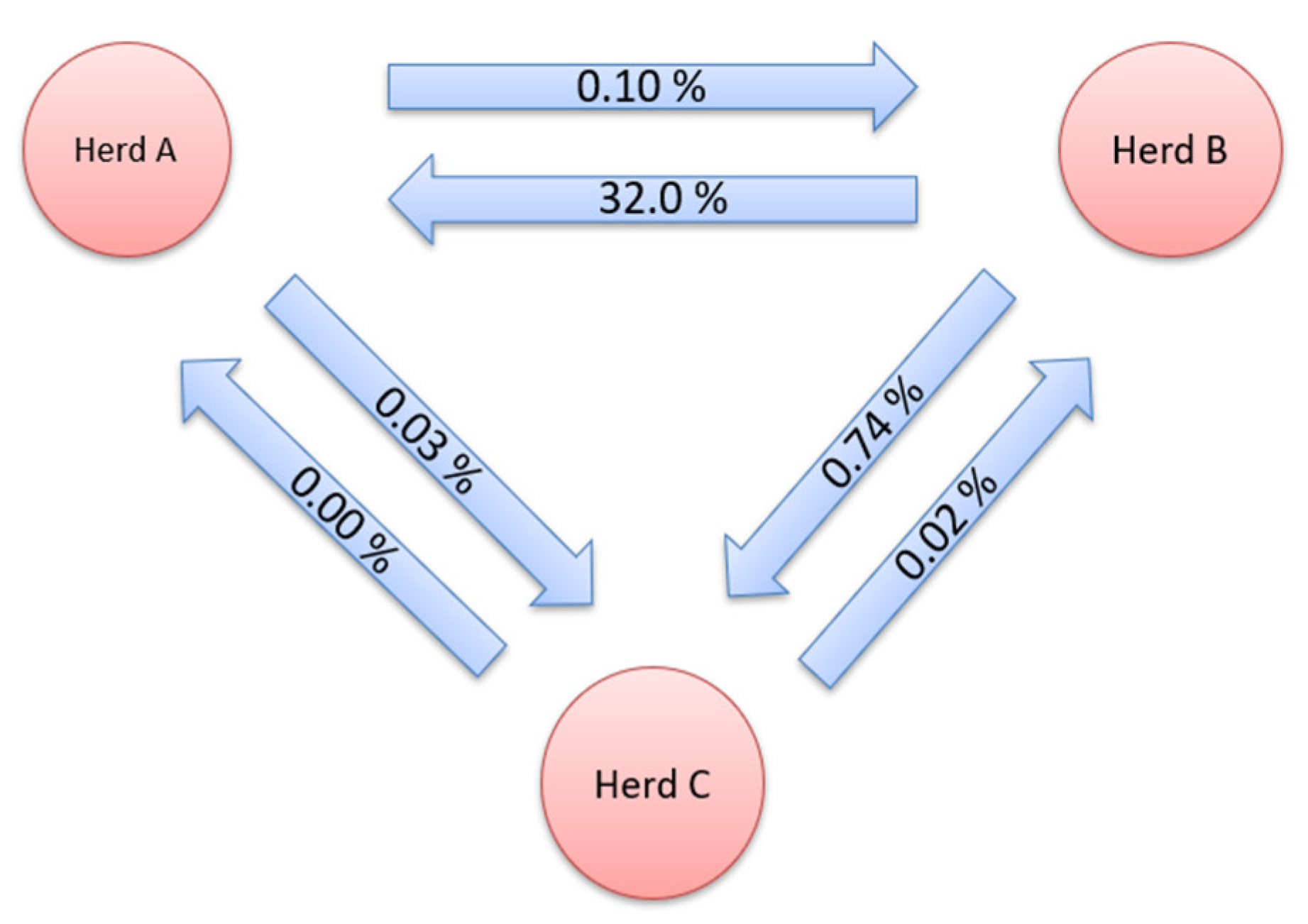
2.2.2. Gene Flow
2.2.3. Connectedness Rating
2.2.4. Reliability of Estimated Breeding Values
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mrode, R.A. Linear Models for the Prediction of Animal Breeding Values; CABI Publishing: Wallingford, UK, 2005; pp. 1–344. Available online: http://sherekashmir.informaticspublishing.com/278/1/9780851990002.pdf (accessed on 20 February 2022)ISBN 0-85199-000-2.
- Henderson, C.R. Best Linear Unbiased Estimation and Prediction under a Selection Model. Biometrics 1975, 31, 423–447. [Google Scholar] [CrossRef] [PubMed]
- Noguera, J.L.; Varona, L.; Babot, D.; Estany, J. Multivariate analysis of litter size for multiple parities with production traits in pigs: II. Response to selection for litter size and correlated response to production traits. J. Anim. Sci. 2002, 80, 2548–2555. [Google Scholar] [CrossRef] [PubMed]
- Satoh, M. Comparison of genetic improvement for litter size at birth by direct and indirect selection in swine herd. Anim. Sci. J. 2006, 77, 566–573. [Google Scholar] [CrossRef]
- Dube, B.D.; Sendros, M.; Dzama, K. Estimation of genetic and phenotypic parameters for sow productivity traits in South African Large White pigs. S. Afr. J. Anim. Sci. 2012, 42, 389–397. [Google Scholar] [CrossRef]
- Luković, Z.; Radojković, D. Dispersion parameters for litter size in pigs. In Proceedings of the International Symposium Modern Trends in Livestock Production, Belgrade, Serbia, 2–4 October 2013; Available online: https://eprints.ugd.edu.mk/8382/1/PROCEEDINGS.pdf (accessed on 26 February 2022).
- Krupa, E.; Wolf, J. Simultaneous estimation of genetic parameters. Czech J. Anim. Sci. 2013, 58, 429–436. [Google Scholar] [CrossRef]
- Costa, E.V.; Ventura, H.T.; Figueredo, E.A.P.; Silva, F.F.; Gloria, L.S.; Godinho, R.M.; Resende, M.D.V.; Lopes, P.S. Multi-trait and repeatability models for genetic evaluation of litter traits in pigs. Rev. Bras. Saúde Prod. Anim. 2016, 17, 666–676. [Google Scholar] [CrossRef]
- Sell-Kubiak, E.; Knol, E.F.; Mulder, H.A. Selecting for changes in average “parity curve” pattern of litter size in Large White pigs. J. Anim. Breed. Genet. 2019, 136, 134–148. [Google Scholar] [CrossRef]
- Sell-Kubiak, E. Selection for litter size and litter birthweight in Large White pigs: Maximum, mean and variability of reproduction traits. Animal 2021, 15, 100352. [Google Scholar] [CrossRef]
- Lopez, B.I.M.; Seo, K. Genetic parameters for litter traits at different parities in purebred Landrace and Yorkshire pigs. Anim. Sci. J. 2019, 90, 1497–1502. [Google Scholar] [CrossRef]
- Konta, A.; Ogawa, S.; Kimata, M.; Ishii, K. Comparison of two models to estimate genetic parameters for number of born alive in pigs. Anim. Sci. J. 2020, 91, e13417. [Google Scholar] [CrossRef]
- Mathur, P.K.; Sullivan, B.P.; Chesnais, J.P. Measuring connectedness: Concept and application to a large in-461 dustry breeding program. In Proceedings of the 7th World Congress of Genetics Applied to Livestock Production, Montpellier, France, 19–23 August 2002; pp. 545–548. Available online: http://www.wcgalp.org/system/files/proceedings/2002/measuring-connectedness-concept-and-application-large-industry-breeding-program.pdf (accessed on 22 April 2022).
- Sun, C.Y.; Wang, C.K.; Wang, Y.C.; Zhang, Y.; Zhang, Q. Evaluation of connectedness between herds for three pig breeds in China. Animal 2009, 3, 482–485. [Google Scholar] [CrossRef]
- Soga, N. The Effect of Connectedness on the Bias and Accuracy for Prediction of Breeding Value in Swine Herds. Master’s Theses, Iowa State University, Ames, IO, USA, 2009; pp. 1–79. Available online: https://lib.dr.iastate.edu/etd/11116 (accessed on 10 April 2022).
- Soga, N.; Spangler, M.L.; Schwab, C.R.; Berger, P.J.; Baas, T.J. Comparison of Connectedness Measures and Changes in Connectedness of the U.S. Duroc Population. In Proceedings of the 9th World Congress on Genetics Applied to Livestock Production, Leipzig, Germany, 1–6 August 2010; Available online: https://digitalcommons.unl.edu/animalscifacpub/801 (accessed on 10 April 2022).
- Krupa, E.; Žáková, E.; Krupová, Z.; Kasarda, R.; Svitáková, A. Evaluation of Genetic Connectedness of Landrace Herds in Czech Republic. In Proceedings of the 4th International Congress New Perspectives and Challenges of Sustainable Livestock Productions, Belgrade, Serbia, 7–9 October 2015; Available online: https://www.researchgate.net/publication/282755954_Evaluation_of_Genetic_Connectedness_of_Landrace_Herds_in_Czech_Republic (accessed on 9 March 2022).
- Skorput, D.; Gorjanc, G.; Lukovic, Z. Reliability of genetic evaluation using purebred and crossbred data from different pig farms. Anim. Sci. Pap. Rep. 2017, 35, 1, 47–56. [Google Scholar]
- Lopez, B.M.; Kang, H.S.; Viterbo, V.S.; Song, C.W.; Seo, K.S. Degree of connectedness among herds of three pig breeds in Korea. Indian J. Anim. Sci. 2018, 88, 724–726. Available online: https://www.researchgate.net/publication/326404664_Degree_of_connectedness_among_herds_of_three_pig_breeds_in_Korea (accessed on 29 March 2022).
- Zhang, H.; Liu, X.L.; Wang, C.; Li, J.; Yao-Sheng, C. Study on the measures of connectedness between herds. Agric. Sci. China 2004, 3, 143–148. [Google Scholar]
- Kennedy, B.W.; Trus, D. Considerations on genetic connectedness between management units under an animal model. J. Anim. Sci. 1993, 71, 2341–2352. [Google Scholar] [CrossRef]
- Mathur, P.K.; Sullivan, B.; Chesnais, J. A practical method for estimating connectedness in large. livestock populations with an application to Canadian swine herds. J. Anim. Sci. 1998, 76, 85. [Google Scholar]
- Sargolzaei, M.; Iwaisaki, H.; Colleau, J.J. CFC: A tool for monitoring genetic diversity. In Proceedings of the 8th World Congress on Genetics Applied to Livestock Production, Belo Horizonte, Brazil, 13–18 August 2006. [Google Scholar]
- The SAS System for Windows, Release 9.4; SAS Institute: Cary, NC, USA, 2013.
- Team, R Development Core. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2014. [Google Scholar]
- Bates, D.; Vazquez, A.I. Pedigree-Based Mixed-Effects Models. 2014. Available online: http://pedigreemm.r-forge.r-project.org/ (accessed on 1 February 2022).
- Coster, A. Pedigree Functions. 2012. Available online: https://cran.r-project.org/web/packages/pedigree/index.html (accessed on 1 February 2022).
- Bates, D.; Maechler, M. Modelling with Sparse and Dense Matrices. 2021. Available online: http://Matrix.R-forge.R-project.org/ (accessed on 1 February 2022).
- Mathur, P.K. Connectedness Programs. 2002. Available online: https://www.ccsi.ca/connectedness/ (accessed on 5 February 2022).
- Groeneveld, E.; Kovac, M.; Mielenz, N. VCE6 User’s Guide and Reference Manual; Institute of Farm Animal Genetics, FLI: Mariensee, Germany, 2010. [Google Scholar]
- Groeneveld, E.; Kovač, M.; Wang, T. PEST, A General Purpose BLUP Package for Multivariate Prediction and Estimation. In Proceedings of the 4th World Congress on Genetics Applied to Livestock Production, Edinburgh, Scotland, 23–27 July 1990. [Google Scholar]
- Radojković, D. Variability of Fertiliy Traits and Breeding Value Estimation Of Pigs. Ph.D. Theses, Faculty of Agriculture-University of Belgrade, Belgrade, Serbia, 2007; pp. 1–209. [Google Scholar]
- Radojković, D.; Savić, R.; Popovac, M.; Radović, Č.; Gogić, M. The share of variance components and correlations between sow production traits in different treatments of the litter size (the repeatability and multi-trait models). Contemp. Agric. 2018, 67, 207–214. [Google Scholar] [CrossRef]
- Kuehn, L.A.; Lewis, R.M.; Notter, D.R. Managing the risk of comparing estimated breeding values across flocks or herds through connectedness: A review and application. Genet. Sel. Evol. 2007, 39, 225–247. [Google Scholar] [CrossRef]
- Skorput, D.; Spehar, M.; Lukovic, Z. Connectedness between contemporary groups in Black Slavonian pig. Livest. Sci. 2018, 216, 6–8. [Google Scholar] [CrossRef]
- Skorput, D.; Gorjanc, G.; Lukovic, Z. Evaluation of Connectedness between the Management Units of Landrace Breed of Pigs in Croatia. In Proceedings of the 20th International Symposium“Animal Science Days”, Kranjska gora, Slovenia, 19–21 September 2012; pp. 181–185. [Google Scholar]
- Hui, Y.; Guosheng, S. Impact of phenotypic information of previous generations and depth of pedigree on estimates of genetic parameters and breeding values. Livest. Sci. 2016, 187, 61–67. [Google Scholar]
- Ibáñez-Escriche, N.; Reixach, J.; Lleonart, N.; Noguera, J.L. Genetic evaluation combining purebred and crossbred data in a pig breeding scheme. J. Anim. Sci. 2011, 89, 3881. [Google Scholar] [CrossRef] [PubMed]
- Melnikova, E.; Kabanov, A.; Nikitin, S.; Somova, M.; Kharitonov, S.; Otradnov, P.; Kostyunina, O.; Karpushkina, T.; Martynova, E.; Sermyagin, A. Application of Genomic Data for Reliability Improvement of Pig Breeding Value Estimates. Animals 2021, 11, 1557. [Google Scholar] [CrossRef] [PubMed]
- Bunter, K.; Connectedness in Across Herd Genetic Evaluations. AGBU Pig Genetics Workshop–November. 1995, pp. 67–73. Available online: http://agbu.une.edu.au/pig_genetics/pdf/1995/Paper%2011_Connectedness_Bunter_1995.pdf (accessed on 9 March 2022).
- Fouillou, M.N.; Clement, V.; Laloe, D. Measuring connectedness among herds in mixed linear models: From theory to practice in large-sized genetic evaluations. Genet. Sel. Evol. 2008, 40, 145–159. [Google Scholar] [CrossRef]

| Herd Name | Phenotypes | Litter Genotype | Pedigree File | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| N | -NBA | SD | L | LW | LXLW | LWXL | No. of Animals | % Base Animals | N2 | |
| A | 36,200 | 16.18 | 3.58 | 591 | 1111 | 21,948 | 12,550 | 15,478 | 12.96 | 10,192 |
| B | 7823 | 14.00 | 3.79 | 3510 | 4313 | / | / | 4886 | 20.91 | 3160 |
| C | 14,020 | 16.88 | 3.84 | 988 | 853 | 5801 | 6378 | 10,878 | 18.46 | 5609 |
| ABC | 58,043 | 16.05 | 3.77 | 5098 | 6277 | 27,749 | 18,928 | 23,453 | 13.02 | 18,708 |
| Herd | Estimates of Variance Components for Purebred Data | ||||
|---|---|---|---|---|---|
| Var(a) | Var(p) | Var(l) | Var(e) | Var(ph) | |
| A | 0.508 + 0.455 | 0.951 + 0.543 | 0.000 + 0.000 | 9.053 + 0.415 | 10.513 |
| B | 0.602 + 0.191 | 0.428 + 0.195 | 0.332 + 0.124 | 10.318 + 0.211 | 11.681 |
| C | 1.615 + 0.590 | 1.248 + 0.752 | 0.000 + 0.000 | 14.058 + 0.648 | 16.922 |
| ABC | 0.825 + 0.196 | 1.171 + 0.214 | 0.189 + 0.117 | 10.324 + 0.174 | 12.511 |
| h2 | p2 | l2 | e2 | ||
| A | 0.048 + 0.042 | 0.090 + 0.051 | 0.000 + 0.000 | 0.861 + 0.033 | |
| B | 0.051 + 0.016 | 0.036 + 0.016 | 0.028 + 0.010 | 0.883 + 0.014 | |
| C | 0.095 + 0.034 | 0.073 + 0.044 | 0.000 + 0.000 | 0.830 + 0.035 | |
| ABC | 0.065 + 0.015 | 0.093 + 0.017 | 0.015 + 0.009 | 0.825 + 0.012 | |
| Herd. | Estimates of Variance Components for Purebred and Crossbreed Data | ||||
|---|---|---|---|---|---|
| Var(a) | Var(p) | Var(l) | Var(e) | Var(ph) | |
| A | 0.912 + 0.123 | 0.601 + 0.090 | 0.090 + 0.050 | 7.787 + 0.070 | 9.393 |
| B | / | / | / | / | / |
| C | 0.955 + 0.198 | 1.107 + 0.199 | 0.120 + 0.126 | 10.219 + 0.160 | 12.403 |
| ABC | 0.970 + 0.089 | 0.727 + 0.075 | 0.181 + 0.039 | 8.754 + 0.058 | 10.634 |
| h2 | p2 | l2 | e2 | ||
| A | 0.097 + 0.012 | 0.064 + 0.009 | 0.009 + 0.005 | 0.829 + 0.007 | |
| B | / | / | / | / | |
| C | 0.077 + 0.015 | 0.089 + 0.016 | 0.009 + 0.010 | 0.823 + 0.011 | |
| ABC | 0.091 + 0.008 | 0.068 + 0.007 | 0.017 + 0.003 | 0.823 + 0.005 | |
| Dataset | Herd | A | B | C |
|---|---|---|---|---|
| A | 1.000000 | 0.887295 | 0.875809 | |
| Large White | B | 0.875334 | 1.000000 | 0.983121 |
| C | 0.864112 | 0.983272 | 1.000000 | |
| Landrace | A | 1.000000 | 0.274184 | 0.251712 |
| B | 0.269284 | 1.000000 | 0.894574 | |
| C | 0.245873 | 0.895578 | 1.000000 |
| Dataset | Herd | A | B | C |
|---|---|---|---|---|
| A | 1.000000 | 0.869794 | 0.860125 | |
| Purebred | B | 0.838694 | 1.000000 | 0.987166 |
| C | 0.828842 | 0.986661 | 1.000000 | |
| A | 1.000000 | 0.971128 | 0.994608 | |
| Purebred and crossbred | B | 0.976018 | 1.000000 | 0.980043 |
| C | 0.994738 | 0.975265 | 1.000000 |
| Herd | Reliability | ||||
|---|---|---|---|---|---|
| N | SD | Min | Max | ||
| A | 1194 | 0.5198505 | 0.1327724 | 0.3415963 | 0.8014904 |
| B | 4887 | 0.7101317 | 0.0744571 | 0.3778204 | 0.9441333 |
| C | 3091 | 0.8672705 | 0.0213980 | 0.7678735 | 0.9430393 |
| ABC | 7481 | 0.7426409 | 0.0886325 | 0.5907634 | 0.9690157 |
| Herd | Reliability | ||||
|---|---|---|---|---|---|
| N | SD | Min | Max | ||
| A | 14,376 | 0.8224012 | 0.0411112 | 0.7261741 | 0.9791662 |
| B | / | / | / | / | / |
| C | 10,880 | 0.8122522 | 0.0419156 | 0.6077485 | 0.9695336 |
| ABC | 23,453 | 0.8460510 | 0.0414397 | 0.7423716 | 0.9885285 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stojiljković, N.; Radojković, D.; Luković, Z.; Gogić, M.; Radović, Č.; Popovac, M.; Škorput, D. Case Study on Increasing Breeding Value Estimation Reliability of Reproductive Traits in Serbian Highly Prolific Large White and Landrace Sows. Animals 2022, 12, 2688. https://doi.org/10.3390/ani12192688
Stojiljković N, Radojković D, Luković Z, Gogić M, Radović Č, Popovac M, Škorput D. Case Study on Increasing Breeding Value Estimation Reliability of Reproductive Traits in Serbian Highly Prolific Large White and Landrace Sows. Animals. 2022; 12(19):2688. https://doi.org/10.3390/ani12192688
Chicago/Turabian StyleStojiljković, Nenad, Dragan Radojković, Zoran Luković, Marija Gogić, Čedomir Radović, Mladen Popovac, and Dubravko Škorput. 2022. "Case Study on Increasing Breeding Value Estimation Reliability of Reproductive Traits in Serbian Highly Prolific Large White and Landrace Sows" Animals 12, no. 19: 2688. https://doi.org/10.3390/ani12192688
APA StyleStojiljković, N., Radojković, D., Luković, Z., Gogić, M., Radović, Č., Popovac, M., & Škorput, D. (2022). Case Study on Increasing Breeding Value Estimation Reliability of Reproductive Traits in Serbian Highly Prolific Large White and Landrace Sows. Animals, 12(19), 2688. https://doi.org/10.3390/ani12192688









