Comparative Metabolome Analyses of Ivermectin-Resistant and -Susceptible Strains of Haemonchus contortus
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. H. contortus Strains and Samples
- (i)
- The Haecon-5 strain kindly presented by Professor Robin B. Gasser (University of Melbourne) and maintained in goats in Huazhong Agricultural, which is sensitive to all commercial anthelmintics.
- (ii)
- The Zhaosu strain, which was originally isolated in Zhaosu County, Xinjiang, China, as an IVM-resistant strain [22].
- (iii)
- Samples: Six goats (6-month-old goats free of parasites) were randomly divided into two groups (resistant strain and susceptible strain, three goats/group) and infected with the infective third-stage larvae (7000 L3s per goat) of the respective strain. The adult male and female worms of H. contortus were isolated from the abomasa of goats 45 days after infection. These adult worms were thoroughly washed in PBS and assigned into two groups with three biological replicates for each group. Each sample was made from 160 adult worms (80 males and 80 females/sample) of Haecon-5-S or Zhaosu-R. All samples were transferred to liquid nitrogen for storage until use.
2.2. Metabolite Extraction and HPLC-MS/MS Analysis
2.3. Data Analysis
3. Results
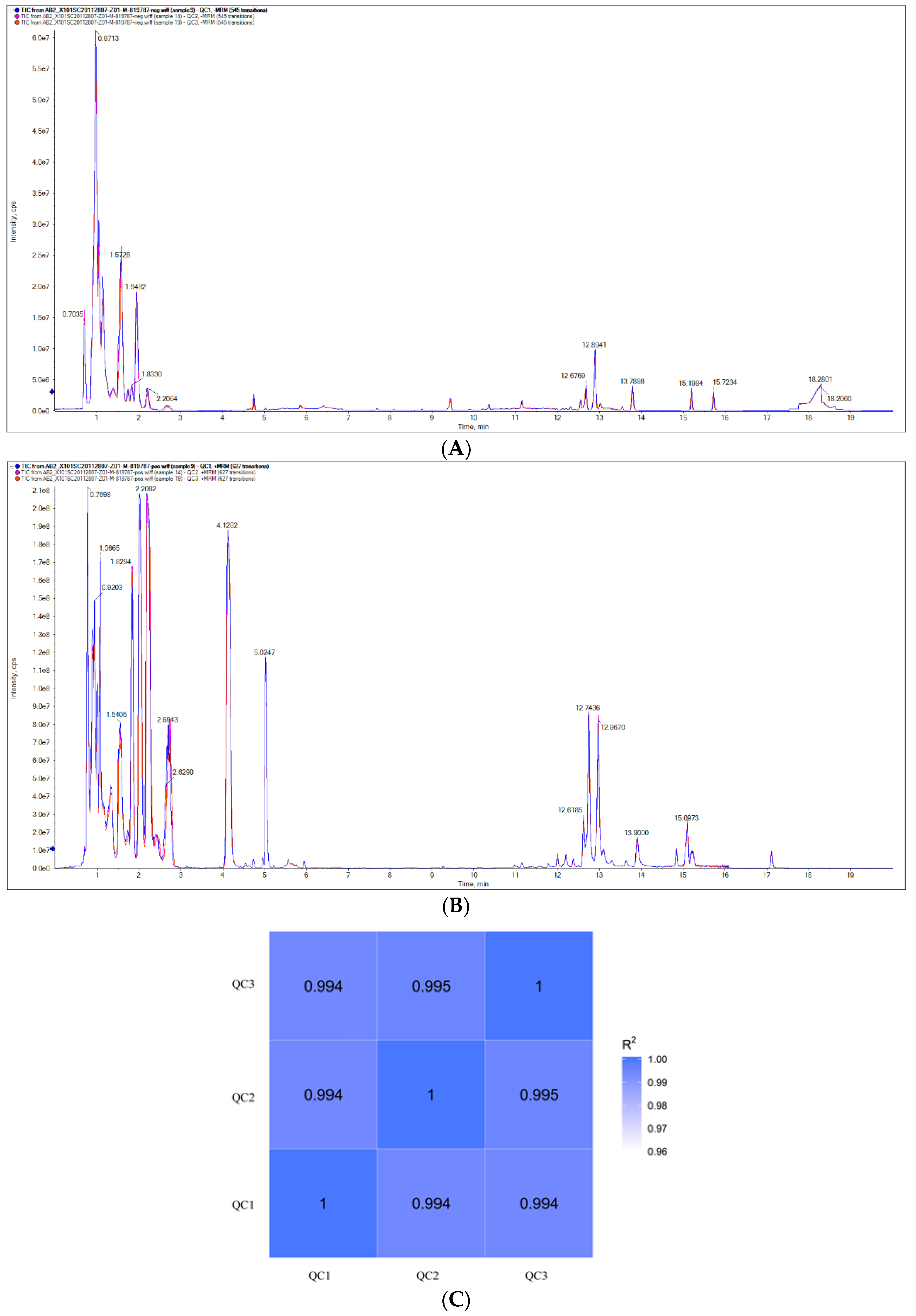
3.1. Data Analysis for Quality Control of Samples
3.2. Total Sample PCA and PLS-DA Analyses
3.3. Overall Metabolomics Analysis of H. contortus
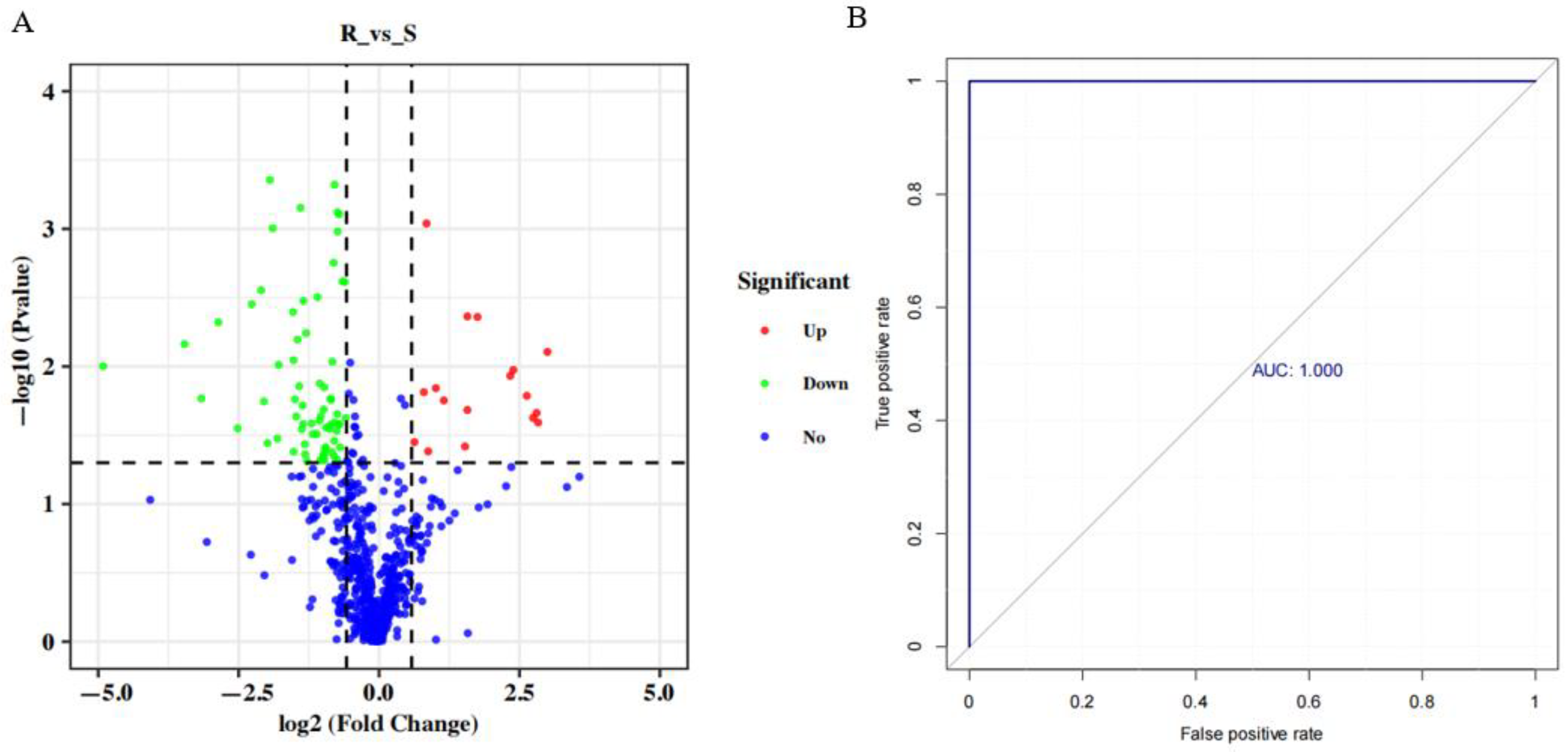
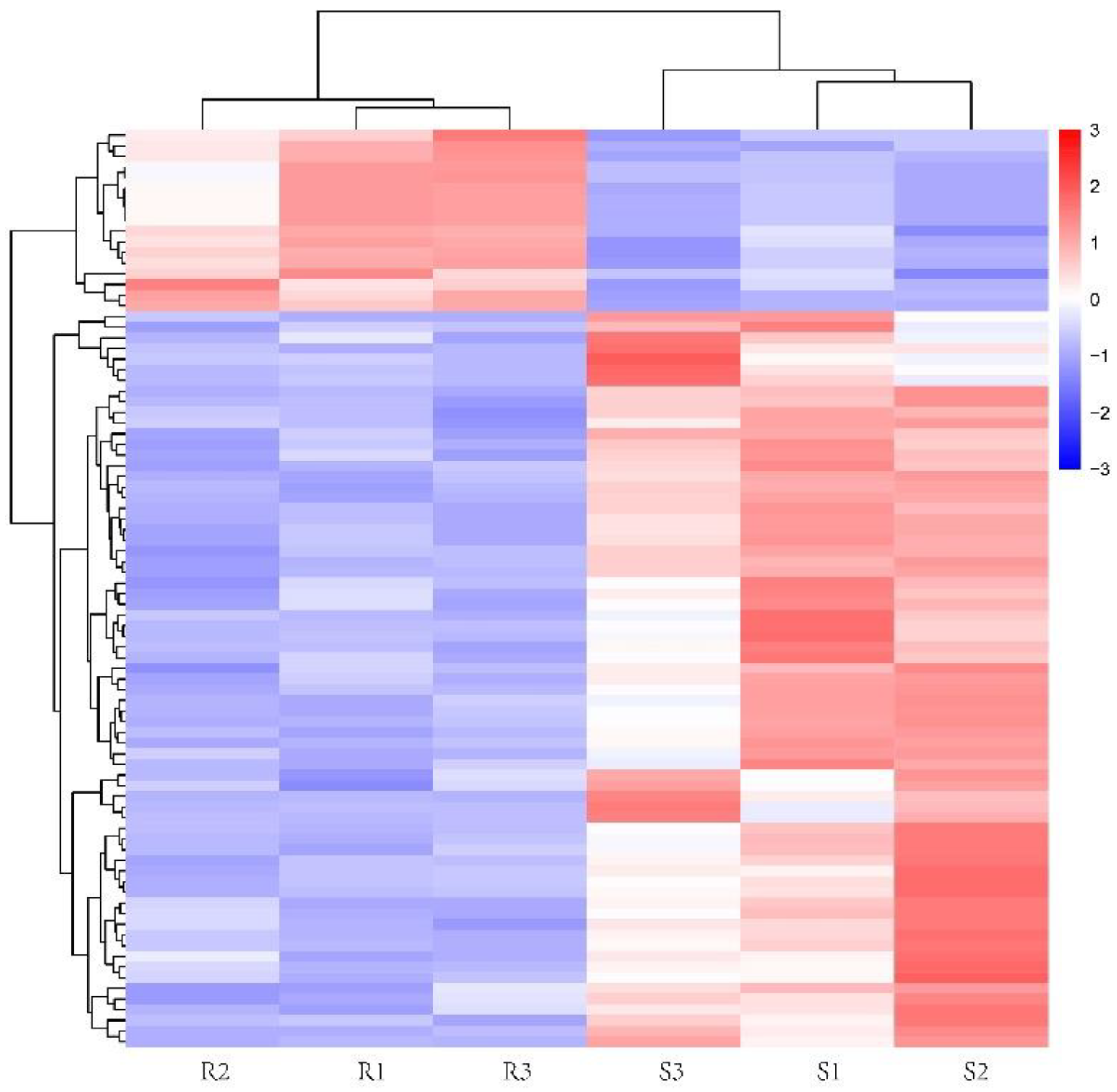
3.4. Differential Metabolite Analysis
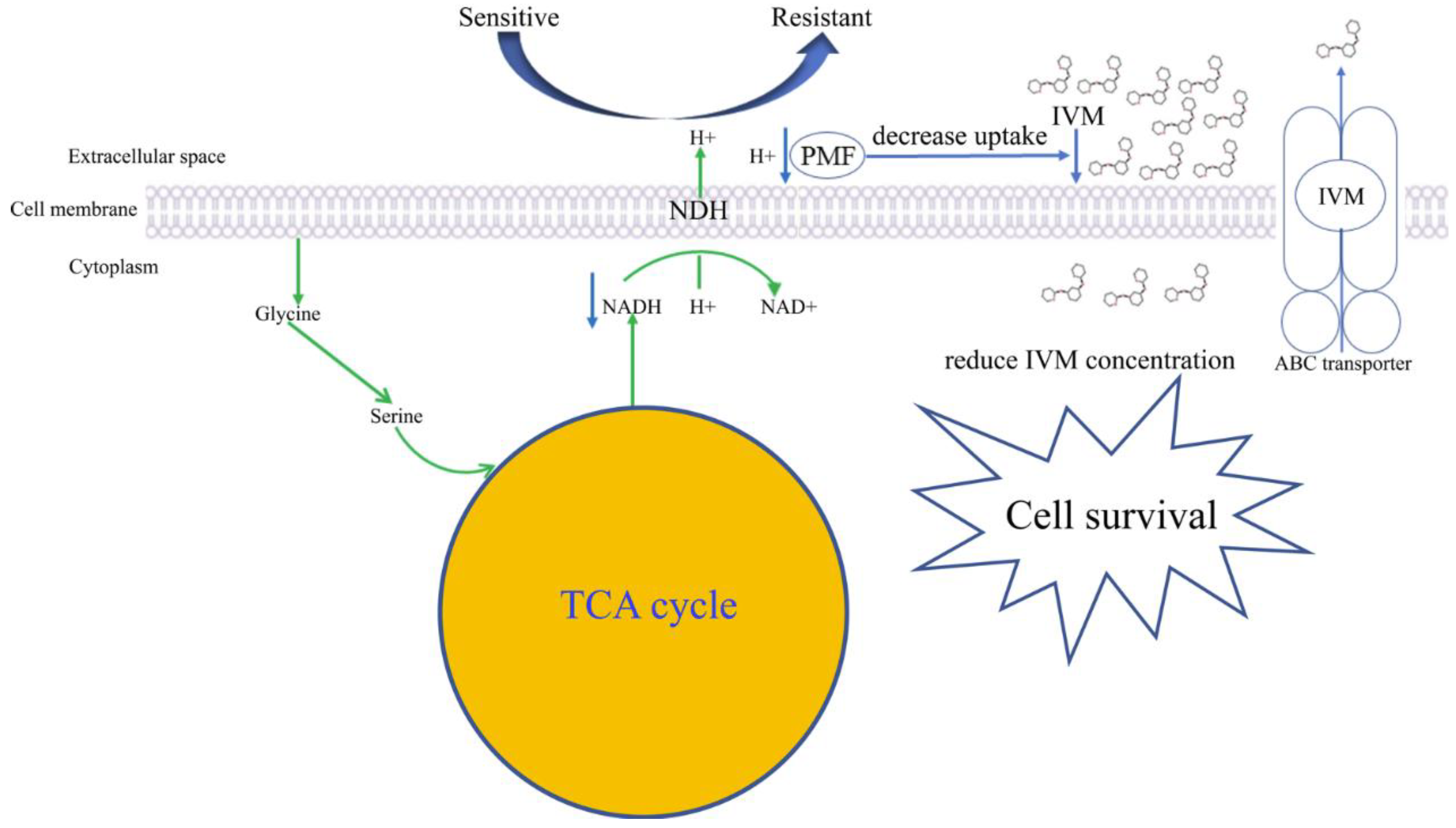
3.5. Functional Annotation Analysis of the Differential Metabolites
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Emery, D.L.; Hunt, P.W.; Le Jambre, L.F. Haemonchus contortus: The then and now and where to from here? Int. J. Parasitol. 2016, 46, 755–769. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Besier, R.B.; Kahn, L.P.; Sargison, N.D.; Van Wyk, J.A. Diagnosis, treatment and management of Haemonchus contortus in small ruminants. Adv. Parasitol. 2016, 93, 181–238. [Google Scholar]
- Mushonga, B.; Habumugisha, D.; Kandiwa, E.; Madzingira, O.; Samkange, A.; Segwagwe, B.E.; Jaja, I.F. Prevalence of Haemonchus contortus infections in sheep and goats in nyagatare district, Rwanda. J. Vet. Med. 2018, 2018, 3602081. [Google Scholar] [CrossRef] [Green Version]
- Naeem, M.; Iqbal, Z.; Roohi, N. Ovine haemonchosis: A review. Trop. Anim. Health. Prod. 2020, 53, 19. [Google Scholar] [CrossRef]
- Kaplan, R.M.; Vidyashankar, A.N. An inconvenient truth: Global worming and anthelmintic resistance. Vet. Parasitol. 2012, 186, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Gilleard, J.S.; Kotze, A.C.; Leathwick, D.; Nisbet, A.J.; Mcneilly, T.N.; Besier, B. A journey through 50 years of research relevant to the control of gastrointestinal nematodes in ruminant livestock and thoughts on future directions. Int. J. Parasitol. 2021, 51, 1133–1151. [Google Scholar] [CrossRef]
- Omura, S. Ivermectin: 25 years and still going strong. Int. J. Antimicrob. Agents 2008, 31, 91–99. [Google Scholar] [CrossRef]
- Kotze, A.C.; Prichard, R.K. Anthelmintic Resistance in Haemonchus contortus: History, mechanisms and diagnosis. Adv. Parasitol. 2016, 93, 397–428. [Google Scholar] [PubMed]
- Rose, H.; Rinaldi, L.; Bosco, A.; Mavrot, F.; de Waal, T.; Skuce, P.; Charlier, J.; Torgerson, P.R.; Hertzberg, H.; Hendrickx, G.; et al. Widespread anthelmintic resistance in European farmed ruminants: A systematic review. Vet. Rec. 2015, 176, 546. [Google Scholar] [CrossRef] [Green Version]
- Jabbar, A.; Campbell, A.J.; Charles, J.A.; Gasser, R.B. First report of anthelmintic resistance in Haemonchus contortus in alpacas in Australia. Parasit. Vectors 2013, 6, 243. [Google Scholar] [CrossRef] [Green Version]
- Schnyder, M.; Torgerson, P.R.; Schönmann, M.; Kohler, L.; Hertzberg, H. Multiple anthelmintic resistance in Haemonchus contortus isolated from South African Boer goats in Switzerland. Vet. Parasitol. 2005, 128, 285–290. [Google Scholar] [CrossRef]
- Almeida, F.A.; Garcia, K.C.; Torgerson, P.R.; Amarante, A.F. Multiple resistance to anthelmintics by Haemonchus contortus and Trichostrongylus colubriformis in sheep in Brazil. Parasitol. Int. 2010, 59, 622–625. [Google Scholar] [CrossRef]
- Kotze, A.C.; Hunt, P.W.; Skuce, P.; von Samson-Himmelstjerna, G.; Martin, R.J.; Sager, H.; Krücken, J.; Hodgkinson, J.; Lespine, A.; Jex, A.R.; et al. Recent advances in candidate-gene and whole-genome approaches to the discovery of anthelmintic resistance markers and the description of drug/receptor interactions. Int. J. Parasitol. Drugs. Drug. Resist. 2014, 4, 164–184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blackhall, W.J.; Pouliot, J.F.; Prichard, R.K.; Beech, R.N. Haemonchus contortus: Selection at a glutamate-gated chloride channel gene in ivermectin- and moxidectin-selected strains. Exp. Parasitol. 1998, 90, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Williamson, S.M.; Storey, B.; Howell, S.; Harper, K.M.; Kaplan, R.M.; Wolstenholme, A.J. Candidate anthelmintic resistance-associated gene expression and sequence polymorphisms in a triple-resistant field isolate of Haemonchus contortus. Mol. Biochem. Parasitol. 2011, 180, 99–105. [Google Scholar] [CrossRef]
- Godoy, P.; Che, H.; Beech, R.N.; Prichard, R.K. Characterisation of P-glycoprotein-9.1 in Haemonchus contortus. Parasit. Vectors 2016, 9, 52. [Google Scholar] [CrossRef] [Green Version]
- Urdaneta-Marquez, L.; Bae, S.H.; Janukavicius, P.; Beech, R.; Dent, J.; Prichard, R. A dyf-7 haplotype causes sensory neuron defects and is associated with macrocyclic lactone resistance worldwide in the nematode parasite Haemonchus contortus. Int. J. Parasitol. 2014, 44, 1063–1107. [Google Scholar] [CrossRef] [PubMed]
- Dent, J.A.; Smith, M.M.; Vassilatis, D.K.; Avery, L. The genetics of ivermectin resistance in Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 2000, 97, 2674–2679. [Google Scholar] [CrossRef] [Green Version]
- Laing, R.; Kikuchi, T.; Martinelli, A.; Tsai, I.J.; Beech, R.N.; Redman, E.; Holroyd, N.; Bartley, D.J.; Beasley, H.; Britton, C.; et al. The genome and transcriptome of Haemonchus contortus, a key model parasite for drug and vaccine discovery. Genome Biol. 2013, 14, R88. [Google Scholar] [CrossRef] [Green Version]
- Yoo, H.C.; Han, J.M. Amino acid metabolism in cancer drug resistance. Cells 2022, 11, 140. [Google Scholar] [CrossRef]
- Stokes, J.M.; Lopatkin, A.J.; Lobritz, M.A.; Collins, J.J. Bacterial metabolism and antibiotic efficacy. Cell. Metab. 2019, 30, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Tuersong, W.; Zhou, C.; Wu, S.; Qin, P.; Wang, C.; Di, W.; Liu, L.; Liu, H.; Hu, M. Comparative analysis on transcriptomics of ivermectin resistant and susceptible strains of Haemonchus contortus. Parasit. Vectors 2022, 15, 159. [Google Scholar] [CrossRef] [PubMed]
- Want, E.J.; Masson, P.; Michopoulos, F.; Wilson, I.D.; Theodoridis, G.; Plumb, R.S.; Shockcor, J.; Loftus, N.; Holmes, E.; Nicholson, J.K. Global metabolic profiling of animal and human tissues via UPLC-MS. Nat. Protoc. 2012, 8, 17–32. [Google Scholar] [CrossRef]
- Luo, P.; Dai, W.; Yin, P.; Zeng, Z.; Kong, H.; Zhou, L.; Wang, X.; Chen, S.; Lu, X.; Xu, G. Multiple reaction monitoring-ion pair finder: A systematic approach to transform nontargeted mode to pseudo-targeted mode for metabolomics study based on liquid chromatography-mass spectrometry. Anal. Chem. 2015, 87, 5050–5055. [Google Scholar] [CrossRef]
- Yang, J.H.; Bening, S.C.; Collins, J.J. Antibiotic efficacy-context matters. Curr. Opin. Microbiol. 2017, 39, 73–80. [Google Scholar] [CrossRef] [PubMed]
- Allison, K.R.; Brynildsen, M.P.; Collins, J.J. Metabolite-enabled eradication of bacterial persisters by aminoglycosides. Nature 2011, 473, 216–220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meylan, S.; Porter, C.B.M.; Yang, J.H.; Belenky, P.; Gutierrez, A.; Lobritz, M.A.; Park, J.; Kim, S.H.; Moskowitz, S.M.; Collins, J.J. Carbon sources tune antibiotic susceptibility in pseudomonas aeruginosa via tricarboxylic acid cycle control. Cell. Chem. Biol. 2017, 24, 195–206. [Google Scholar] [CrossRef]
- Yang, J.H.; Bhargava, P.; McCloskey, D.; Mao, N.; Palsson, B.O.; Collins, J.J. Antibiotic-induced changes to the host metabolic environment inhibit drug efficacy and alter immune function. Cell. Host. Microbe 2017, 22, 757–765. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peng, B.; Su, Y.B.; Li, H.; Han, Y.; Guo, C.; Tian, Y.M.; Peng, X.X. Exogenous alanine and/or glucose plus kanamycin kills antibiotic-resistant bacteria. Cell. Metab. 2015, 21, 249–262. [Google Scholar] [CrossRef] [Green Version]
- Yong, Y.; Zhou, Y.; Liu, K.; Liu, G.; Wu, L.; Fang, B. Exogenous citrulline and glutamine contribute to reverse the resistance of Salmonella to apramycin. Front. Microbiol. 2021, 12, 759170. [Google Scholar] [CrossRef]
- Ye, J.Z.; Lin, X.M.; Cheng, Z.X.; Su, Y.B.; Li, W.X.; Ali, F.M.; Zheng, J.; Peng, B. Identification and efficacy of glycine, serine and threonine metabolism in potentiating kanamycin-mediated killing of Edwardsiella piscicida. J. Proteom. 2018, 183, 34–44. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, R.; Xiao, X.; Wang, Z. Bacterial metabolism-inspired molecules to modulate antibiotic efficacy. J. Antimicrob. Chemother. 2019, 74, 3409–3417. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.Z.; Su, Y.B.; Lin, X.M.; Lai, S.S.; Li, W.X.; Ali, F.; Zheng, J.; Peng, B. Alanine enhances Aminoglycosides-induced ROS production as revealed by proteomic analysis. Front. Microbiol. 2018, 9, 29. [Google Scholar] [CrossRef] [PubMed]
- Pranzini, E.; Pardella, E.; Paoli, P.; Fendt, S.M.; Taddei, M.L. Metabolic reprogramming in anticancer drug resistance: A focus on amino acids. Trends. Cancer 2021, 7, 682–699. [Google Scholar] [CrossRef] [PubMed]
- Lieu, E.L.; Nguyen, T.; Rhyne, S.; Kim, J. Amino acids in cancer. Exp. Mol. Med. 2020, 52, 15–30. [Google Scholar] [CrossRef]
- Yang, J.H.; Wright, S.N.; Hamblin, M.; McCloskey, D.; Alcantar, M.A.; Schrübbers, L.; Lopatkin, A.J.; Satish, S.; Nili, A.; Palsson, B.O.; et al. A white-box machine learning approach for revealing antibiotic mechanisms of action. Cell 2019, 177, 1649–1661. [Google Scholar] [CrossRef] [PubMed]


| Type | Quantity |
|---|---|
| Amino Acids and Derivatives | 154 |
| Organic Acids and Derivatives | 105 |
| Nucleotides and Derivates | 74 |
| Fatty Acyls | 50 |
| Carbohydrates and Derivatives | 42 |
| Hormones | 36 |
| Phospholipids | 26 |
| Carnitines | 22 |
| Organic heterocyclic compounds | 20 |
| Sugar Acids and Derivatives | 14 |
| Vitamins | 13 |
| Eicosanoids | 12 |
| Others | 11 |
| Bile Acids | 11 |
| Benzoic Acid and Derivatives | 11 |
| Indole and Derivatives | 10 |
| Benzene and substituted derivatives | 10 |
| Cholines | 9 |
| Pyridine and Derivatives | 9 |
| Sugar Alcohols | 7 |
| TCA Cycle | 7 |
| Phenols and Derivatives | 7 |
| Polyamines | 7 |
| Phenolamides | 5 |
| Prenol lipids | 4 |
| Cinnamic acids and derivatives | 3 |
| Pyrimidines and pyrimidine derivatives | 3 |
| Organic nitrogen compounds | 2 |
| Ketones | 2 |
| Purines and purine derivatives | 2 |
| Glycerolipids | 2 |
| Co-Enzyme Factors | 2 |
| Alcohols and polyols | 2 |
| Benzene and substituted derivatives | 2 |
| Amines | 2 |
| Esters | 1 |
| Organic oxygen compounds | 1 |
| Aldehydes | 1 |
| Lignans, neolignans, and related compounds | 1 |
| Ethers | 1 |
| Steroids and steroid derivatives | 1 |
| Phenylpropanoids and polyketides | 1 |
| total | 705 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tuersong, W.; Liu, X.; Wang, Y.; Wu, S.; Qin, P.; Zhu, S.; Liu, F.; Wang, C.; Hu, M. Comparative Metabolome Analyses of Ivermectin-Resistant and -Susceptible Strains of Haemonchus contortus. Animals 2023, 13, 456. https://doi.org/10.3390/ani13030456
Tuersong W, Liu X, Wang Y, Wu S, Qin P, Zhu S, Liu F, Wang C, Hu M. Comparative Metabolome Analyses of Ivermectin-Resistant and -Susceptible Strains of Haemonchus contortus. Animals. 2023; 13(3):456. https://doi.org/10.3390/ani13030456
Chicago/Turabian StyleTuersong, Waresi, Xin Liu, Yifan Wang, Simin Wu, Peixi Qin, Shengnang Zhu, Feng Liu, Chunqun Wang, and Min Hu. 2023. "Comparative Metabolome Analyses of Ivermectin-Resistant and -Susceptible Strains of Haemonchus contortus" Animals 13, no. 3: 456. https://doi.org/10.3390/ani13030456
APA StyleTuersong, W., Liu, X., Wang, Y., Wu, S., Qin, P., Zhu, S., Liu, F., Wang, C., & Hu, M. (2023). Comparative Metabolome Analyses of Ivermectin-Resistant and -Susceptible Strains of Haemonchus contortus. Animals, 13(3), 456. https://doi.org/10.3390/ani13030456






