1. Introduction
Fish growth refers to the physiological process in which the nutrients in the food are digested and absorbed into their own substances and continuously accumulated in the individual, which is mainly affected by factors such as genes, the environment, and nutrition. In the same fish population, some individuals grow faster than others. Studies have shown that some individuals in the same Atlantic sturgeon (
Acipenser oxyrinchus) population have great differences in growth performance, especially in body length and weight [
1]. In the same growing environment, the growth rate of different individuals in the same fish population [
2] or different groups in the same fish population [
3] will have obvious differences, which is the result of the genetic performance of individual fish and their long-term adaptation to the growing environment. In the same fish population, the growth rate is mainly regulated by the expression of related genes, the endocrine system, and other factors [
4]. The growth rate of the same kind of fish is mainly regulated by external factors such as nutrition, breeding density, and temperature [
5]. In the artificial farming process, the faster the growth rate of fish, the shorter the time required to reach the market specifications, which can greatly reduce the cost of time, labor, and so on, and further improve the economic benefits of artificial fish farming. Therefore, growth traits are often regarded as one of the most important economic traits to measure fish in artificial culture. With the continuous development of the aquaculture industry, people pay more and more attention to growth traits of fish, and related studies on growth traits are increasing.
Transcriptome RNA sequencing (RNA-Seq) technology is mainly used to study gene function or gene structure at the transcriptional level, reveal the biological process of gene expression in a specific growth stage or under a specific growth environment, and analyze the differentially expressed genes (DEGs) of individuals. The development of transcriptome sequencing technology has laid a technical foundation for studying the differential expression of genes in specific tissues and cells in a certain growth environment or growth stage [
6]. At present, transcriptome analysis has been widely applied to the growth regulation of species. Sun et al. [
3] revealed the relevant mechanisms of fish growth regulation by comparing the transcriptome between the offspring and parents of hybrid grouper, providing a reference for a deeper understanding of the molecular mechanism and regulatory pathway of the growth advantage of hybrid offspring. Similar studies have also been applied to several fish such as black porgy (
Acanthopagrus schlegelii) [
2], black carp (
Mylopharyngodon piceus) [
7], and bighead carp (
Hypophthalmichthys nobilis) [
8]. In order to reveal the main regulatory genes and pathways associated with the growth traits of black porgy, Lin et al. [
2] conducted transcriptome sequencing analysis on the mixed samples of the brain, liver, and muscle tissue of 4-month-old black porgy with different growth rates, and concluded that the genes myosin heavy chain 4 (
MYH4), insulin-like growth factor 1 (
IGF-1), insulin-like growth factor 2 (
IGF-2), myosin light chain 3-like (
Myl3), myosin 6-like (
MYH6), and myoglobin (
MB) may have a negative impact on the growth of black porgy. At the same time, the important roles of the PPAR signaling pathway and GH-GHR-IGFs neuroendocrine pathway in the regulation of growth and development of black porgy were also revealed. Zhang et al. [
7] conducted transcriptome analysis on muscle and liver tissues of the same batch of black carp with different growth rates, and found that epidermal growth factor (
EGF) and vascular endothelial growth factor (
VEGF) were significantly up-regulated in muscle, suggesting that these genes might be related to the growth rate of black carp. Fu et al. [
8] used the RNA-Seq technology to further explore the regulatory role of the hypothalamus–pituitary–liver axis in the growth of bighead carp. They successfully identified three DEGs that were within the growth-related QTL related to the function of growth traits, and 164 lncRNAs specifically expressed in the liver and 749 lncRNAs specifically expressed in the hypothalamic–pituitary gland, which provided a large amount of basic data for revealing the genes associated with fish growth and their molecular mechanisms.
Zig-zag eel (
M. armatus) belongs to the order Symbranchiformes, family Mastacembelidae, and genus
Mastacembelus. The meat of
M. armatus is tender and delicious with high protein content and unsaturated fatty acids, which makes it have very high research and development prospects [
9]. However, up to now, there are few reports on the growth of
M. armatus. Only Zhong et al. [
10,
11] cloned and expressed the growth hormone gene and myostatin (
MSTN) gene of
M. armatus. Thus, there is a serious lack of analytical studies on the growth and development of
M. armatus. This study mainly focuses on
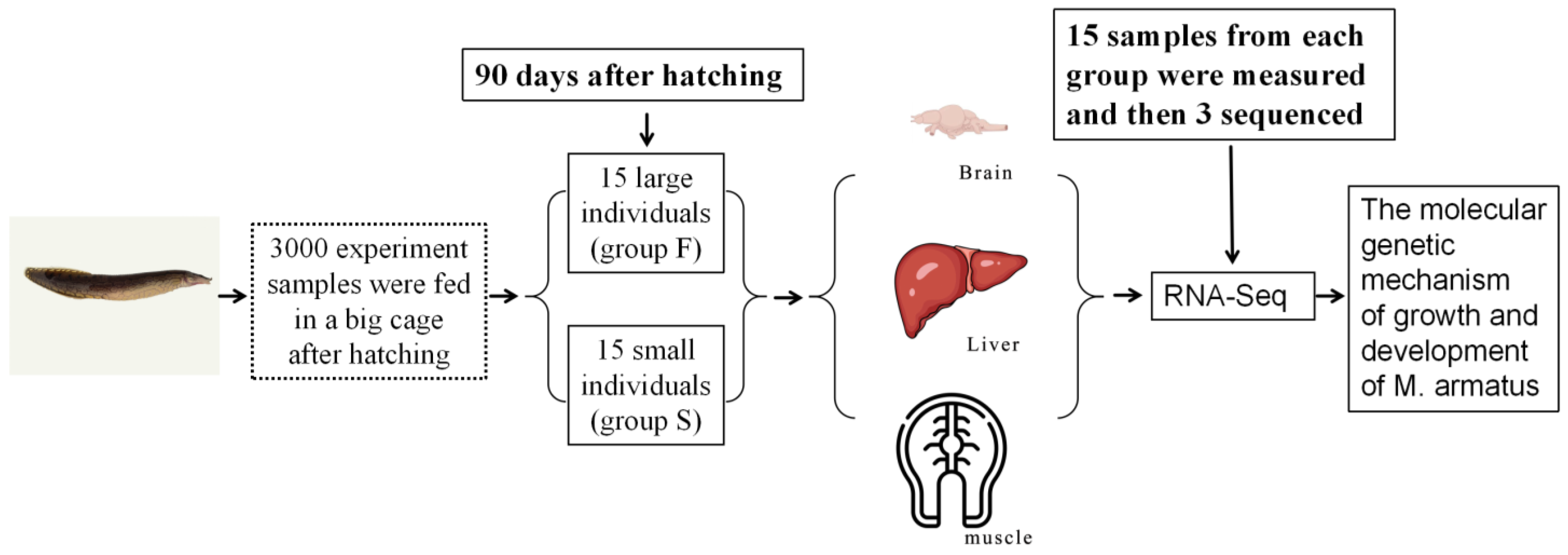
M. armatus with different growth rates, and first analyzes the molecular genetic mechanism of growth and development in
M. armatus with the help of RNA-Seq technology. The results will provide theoretical support for the genetic improvement of growth traits and molecular breeding of
M. armatus.
4. Discussion
In recent years, owing to the rapid development of transcriptome sequencing technology, researchers have been able to conduct in-depth studies on gene expression, gene function, signaling pathways, and other aspects [
21]. At present, transcriptome analysis technology has been widely used in related studies on developmental biology [
22,
23], environmental stress [
24], adaptive evolution [
25], immunology [
26], and growth rate [
27].
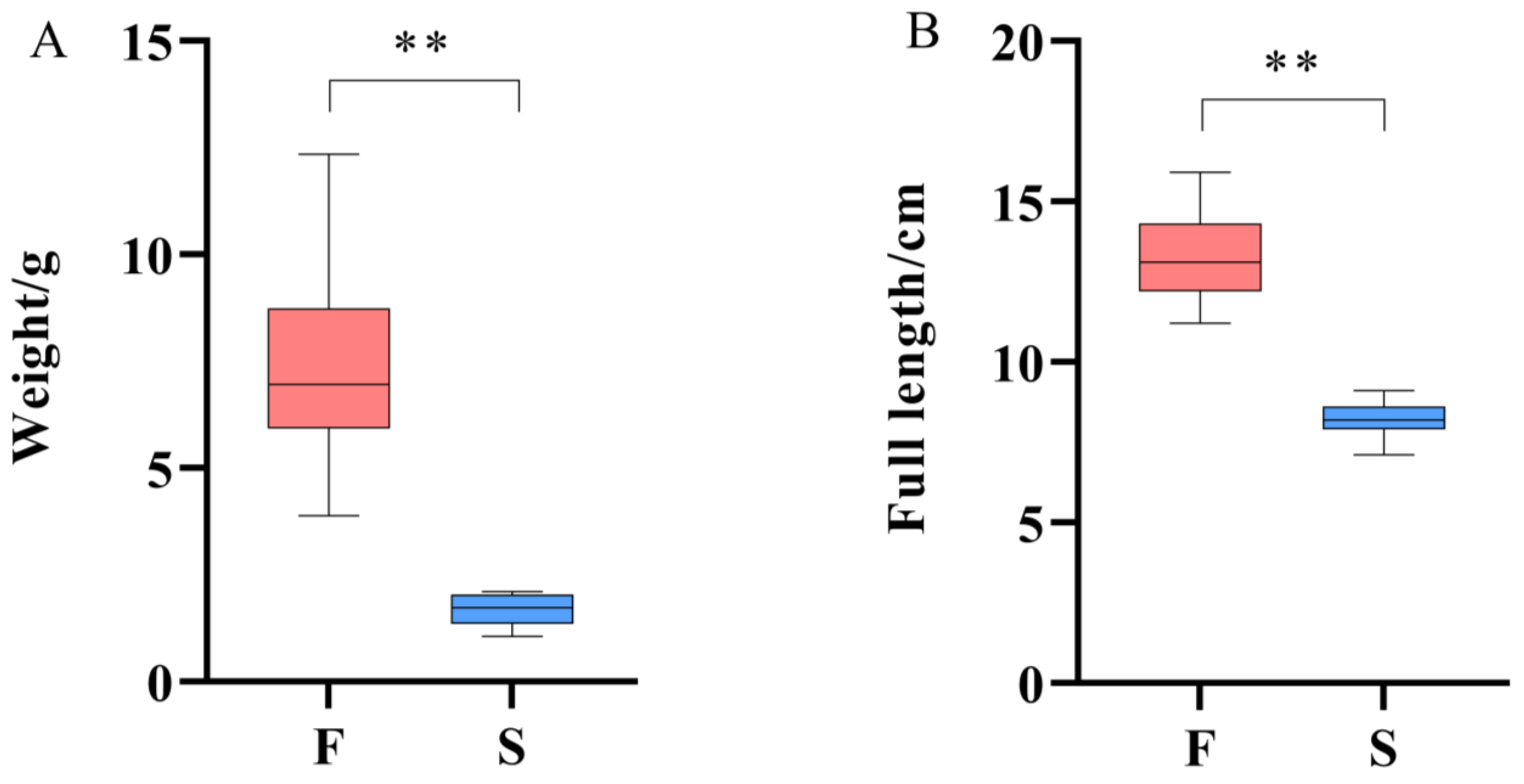
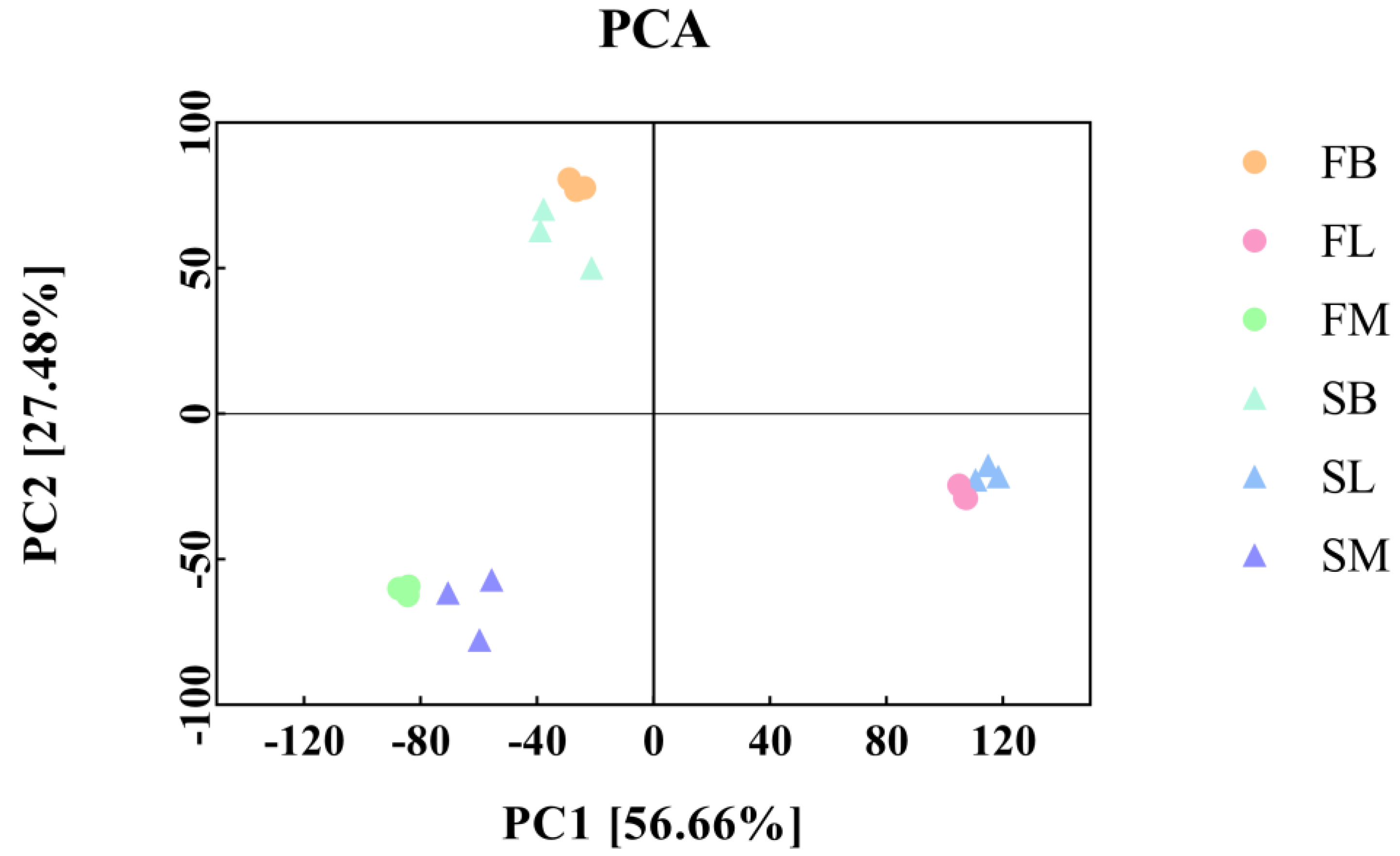
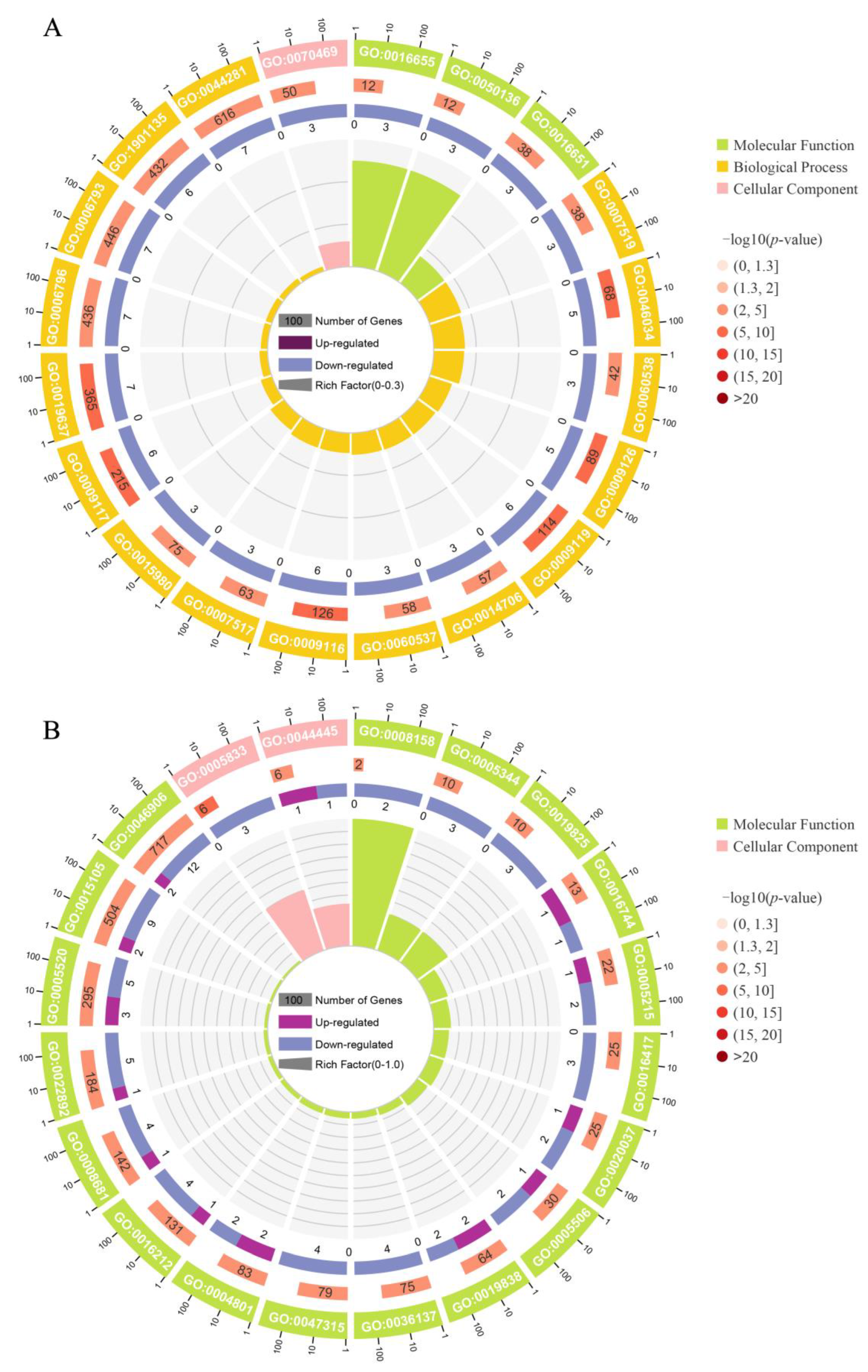
This study found that under the same breeding conditions, there were significant differences in the growth rate of the same batch of M. armatus seedlings. Transcriptome sequencing was performed on brain, liver, and muscle tissue samples of 3-month-old M. armatus with different growth rates by high-throughput sequencing technology. A total of 2887 DEGs were screened, including IGFBP1B, ACLY, ACTA1, IGFBP1A, MYL1, MB, and other key genes involved in regulating body growth.
The effects of growth hormone on cell growth are mainly mediated by the growth hormone/somatomedin (
GH/IGF) growth-axis [
28]. It is a major activator of the growth hormone/somatomedin (
GH/IGF) growth-axis, stimulating
IGF-I secretion in liver or other target tissues [
29,
30]. The original auxin hypothesis put forward that
GH mainly promoted the growth and differentiation of cells by stimulating the liver to secrete
IGF-I, and then
IGF-I is circulated to the target organ through the blood circulation [
31]. Interestingly, the role of growth hormone in promoting individual growth seems to be due to the direct effect of growth hormone on target tissues, rather than the circulating
IGF-I mediated by liver production [
32]. Studies have shown that growth hormone can not only stimulate the secretion of
IGF-I in liver tissues to increase the level of
IGF-I in plasma, but it can also affect the
IGF-I produced by target tissues to act in an autocrine or paracrine way [
33]. Meanwhile, based on relevant experimental data of gene knockout mice, circulating
IGF-I is mainly derived from
IGF-I produced by the liver, and the influence of circulating
IGF-I on growth may be little or even nonexistent, while locally produced
IGF-I in target tissues may be the reason why
IGF-I promotes growth [
34]. In an mRNA localization experiment of the
IGF-I and
IGF-I receptor in gilthead sea bream (
Sparus aurata), Funkenstein et al. [
35] found that in the early growth and development of fish larvae,
IGF-I might stimulate cell proliferation in target tissues by paracrine or autocrine means, while the liver did not play an important role in
IGF-I production. When we subsequently verified the expression of growth-related genes in
M. armatus by RT-qPCR, it was found that the expression of
GH in group F was significantly higher than that in group S in brain tissue, and there was no difference in the expression of
IGF-I between group F and group S in liver tissue, while the expression of
IGF-I in the muscle of group F was significantly higher than that in group S. It seemed that the growth-promoting effect of
GH might be directly exerted on the growth tissues in the early developmental stage of
M. armatus, rather than mediated by liver secretion of
IGF-I. Meanwhile, the expression of
GH and
GHR in the muscle tissue of group F was significantly higher than that in group S, indicating that, like gilthead sea bream [
32,
35], in the early developmental stage of
M. armatus,
GH acted directly by binding to the
GH receptor (
GHR) in the target tissue (e.g., muscle tissue), and induced the target tissue to secrete
IGF-I in an autocrine/paracrine manner to participate in the regulation of tissue growth and differentiation. Notably, Silva et al. [
36] created double transgenic zebrafish (
Danio rerio) and the utilization of the highly expressed
GHR in zebrafish muscle tissue reduced the negative effects of overexpressed
GH on reproduction. Creating double transgenic
M. armatus overexpressing both the growth hormone (
GH) and its receptor (
GHR) may not only effectively reduce the negative effects of growth hormone, but also further achieve maximal growth and minimal side effects and significantly improve the growth rate of
M. armatus in commercial aquaculture settings.
Under the stimulation of growth hormone (
GH), the liver secretes insulin-like growth factor (
IGF)-
I [
37].
IGFBP-1 can bind to
IGF and regulate the role of
IGF in the growth and development of somatic cells [
38]. Due to its high affinity for
IGF-I,
IGFBP-1 can prevent
IGF-I from interacting with other receptors and inhibit the effect of
IGF-I after binding with
IGF-I [
39]. Since the
IGFBP-1 of fish has no nuclear localization sequence,
IGFBP-1 plays a role mainly by regulating the action of
IGF-1 [
40]. Studies have shown that circulating
IGFBP-1 can inhibit the interaction between the
IGF-1 and
IGF receptor on pituitary cells [
40], and
IGFBP-1A can inhibit the action of
IGF-1 [
41]. Because
IGFBP-1B has a higher dissociation rate,
IGFBP-1B has a much lower affinity for
IGF-I and
-II than
IGFBP-1A, and both
IGFBP-1A and
IGFBP-1B inhibit the
IGF-1-induced cell proliferation process during embryonic development in zebrafish (
Danio rerio). Overexpression of
IGFBP-1B can significantly reduce the growth and development rate of embryos [
38]. It is concluded that
IGFBP-1A and
IGFBP-1B may play an important role in the growth regulation of
M. armatus seedlings, and the expression levels of
IGFBP-1A and
IGFBP-1B are significantly increased in slow-growing individuals. In this experiment,
IGFBP-1A and
IGFBP-1B were significantly down-regulated in fast-growing
M. armatus seedlings, which was consistent with the above research results.
Myoglobin (
MB) is an oxygen-binding protein, which can effectively promote oxygen diffusion in the muscle and is an efficient oxygen carrier [
42]. It has been thought to be a protein only found in muscle cells [
43], but later studies have shown that
MB is not only expressed in the muscles of the Tibetan Plateau endemic schizothoracine fish (
Schizopygopsis pylzovi), but also in the liver, muscle, kidney, brain, eyes, and skin [
44].
MB, which is rich in bioavailable iron [
45], can also clear active oxygen in vivo [
46], store oxygen and promote intracellular oxygen diffusion [
43], and regulate the concentration of nitric oxide in the sarcoplasm [
42]. Transcriptome analysis of mixed tissues of black porgy (
Acanthopagrus schlegelii) [
2] showed that the expression of
MB was significantly down-regulated in the slow-growing group. At the same time, some studies have shown that while the cultured Pacific bluefin tuna (
Thunnus orientalis) grows, its
MB content also increases continuously (
p ≤ 0.05) [
47]. Through transcriptome studies on grass carp (
Ctenopharyngodon idella) with different growth rates, it was found that the expression of
MB in fast-growing individuals was significantly higher than that in slow-growing individuals [
48]. In this study,
MB was significantly upregulated in the muscle tissue of fast-growing
M. armatus individuals, similar to the results of the above study.