Exploration of the Polymorphism Distribution of Bovine HMGA2 Gene in Worldwide Breeds and Its Associations with Ovarian Traits
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Bovine Ovary Collection
2.2. InDel Loci Selection and Primer Design
2.3. PCR Amplification and Genotyping
2.4. Statistical Analysis
2.5. Worldwide Bovine Breeds’ Sample Information
3. Results
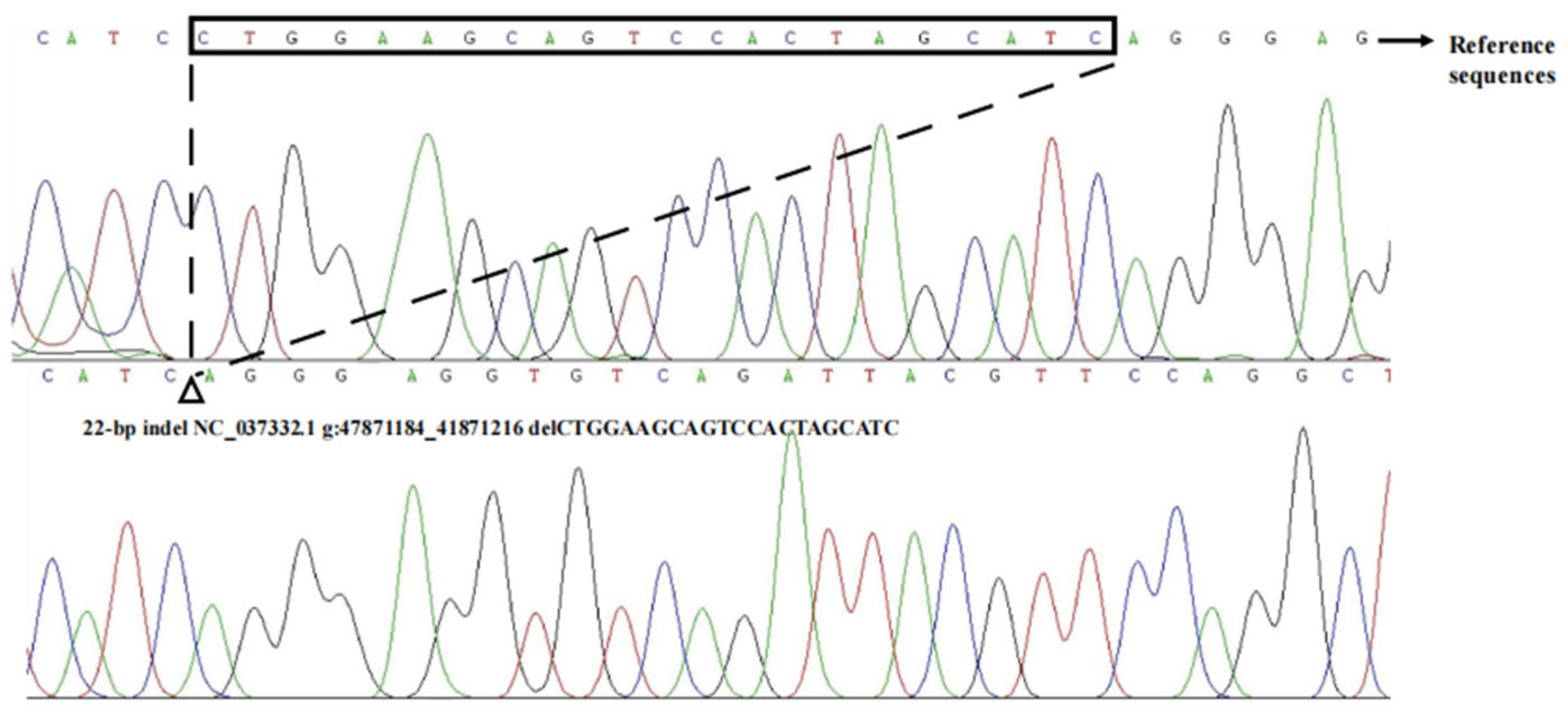
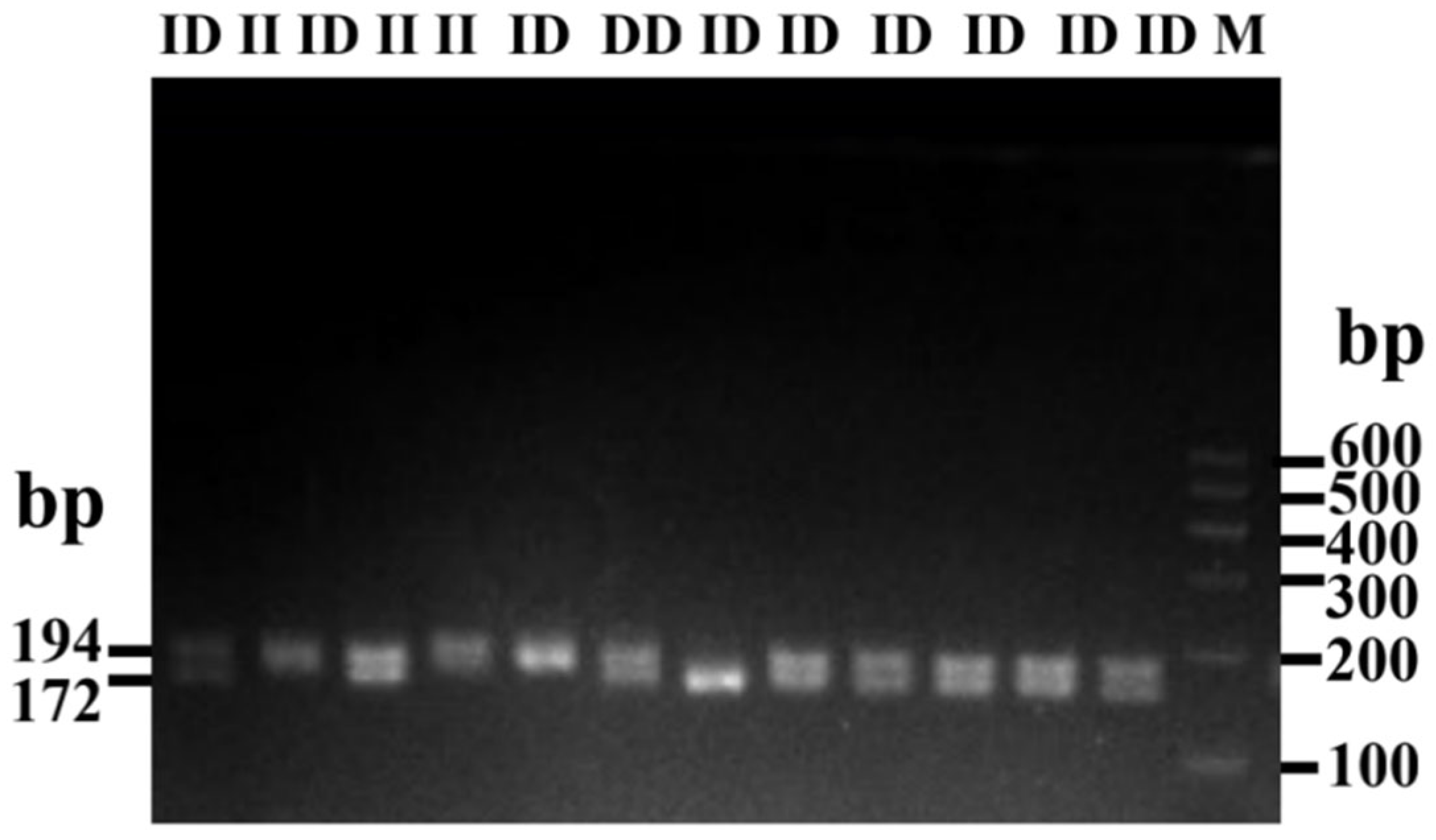
3.1. Identification of the Indels in Bovine HMGA2 Gene
3.2. Estimation of Polymorphism Parameters of Indel (P4-D22-bp) of HMGA2
3.3. Association Analysis of HMGA2 with Ovarian Traits
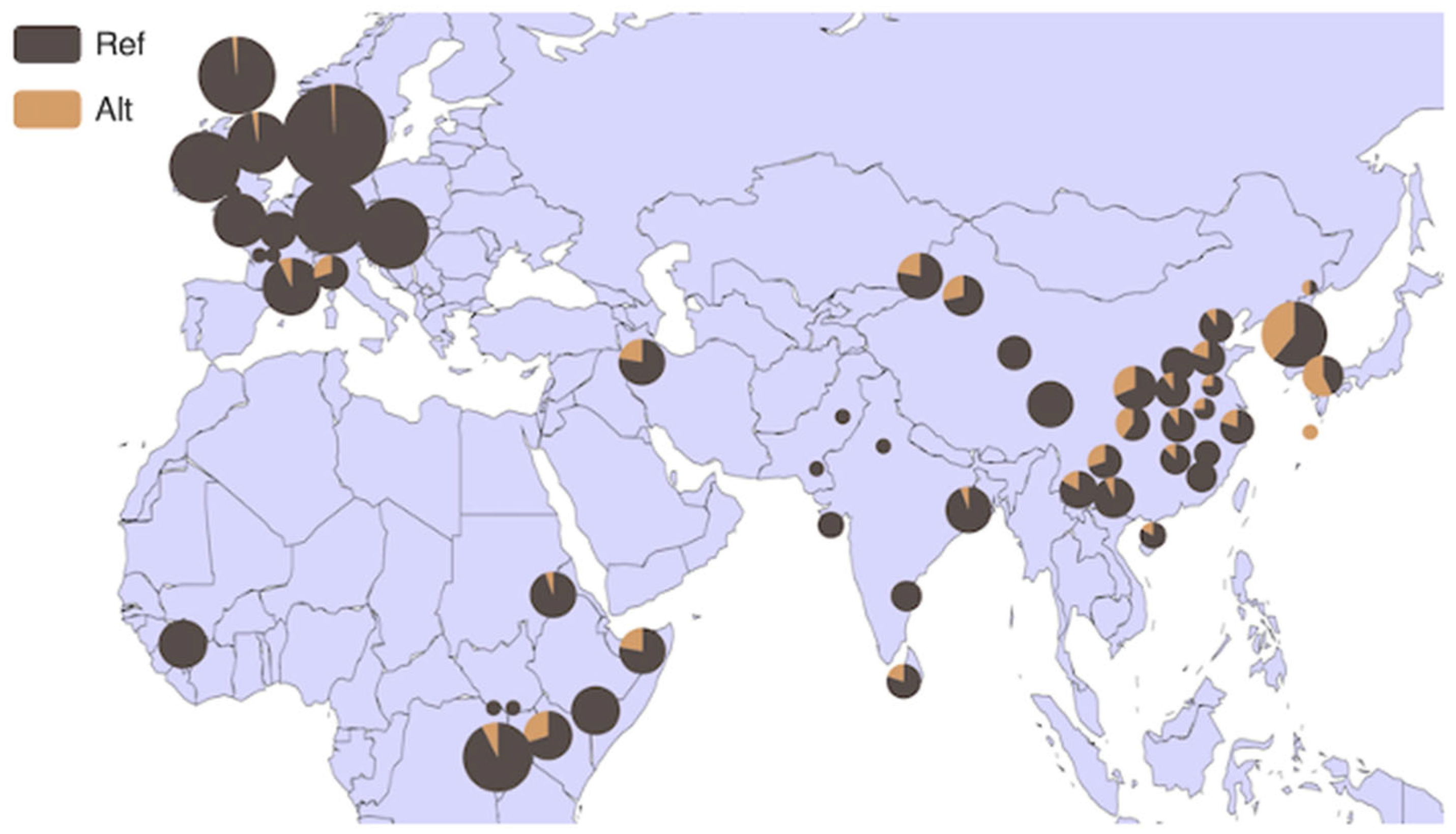
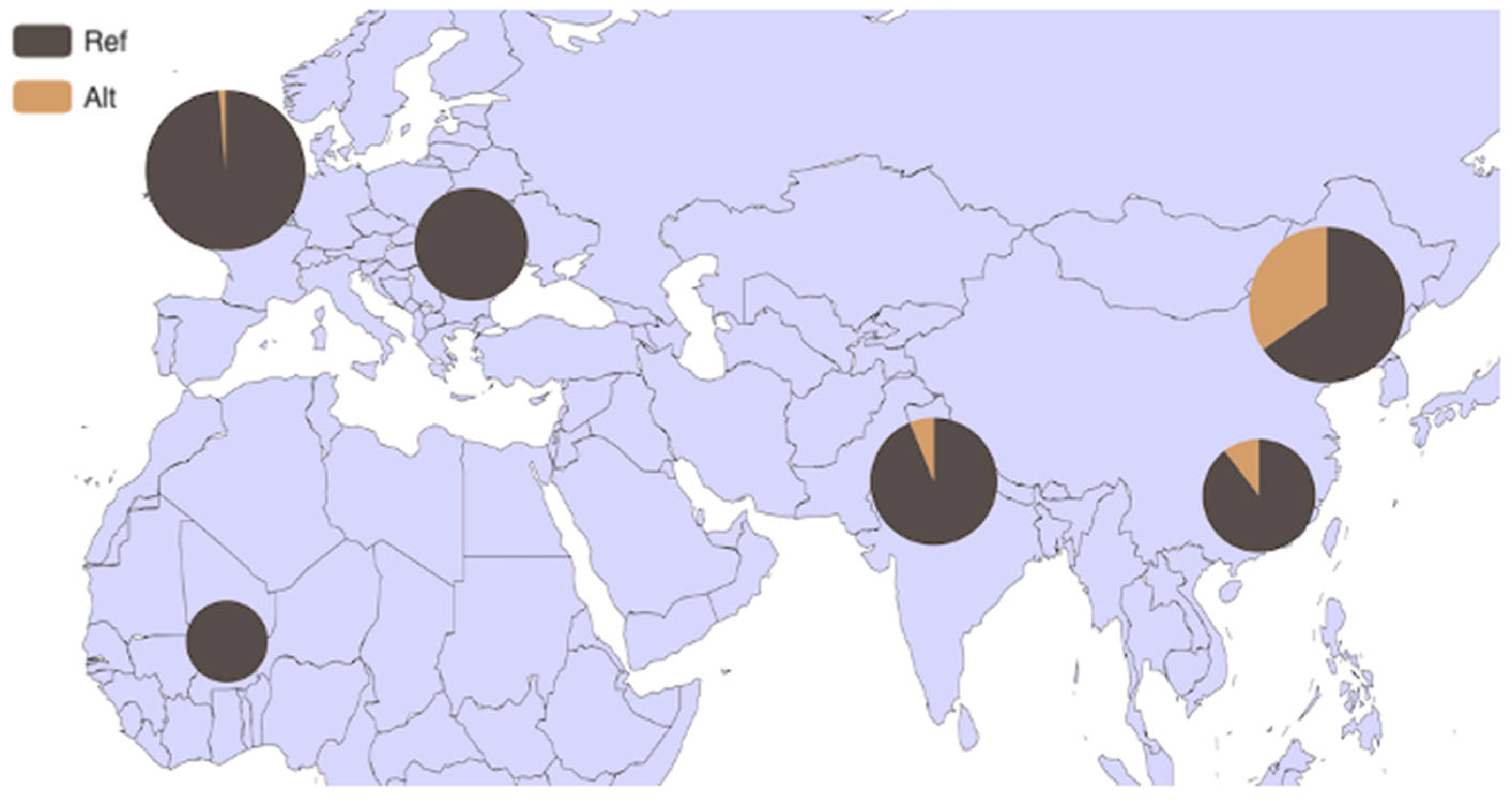
3.4. Analysis of HMGA2 Gene Distribution in 54 Bovine Breeds and 6 Core Bovine Breeds Worldwide
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Berglund, B. Genetic improvement of dairy cow reproductive performance. Reprod. Domest. Anim. 2008, 43 (Suppl. 2), 89–95. [Google Scholar] [CrossRef]
- Devoto, L.; Henríquez, S.; Kohen, P.; Strauss, J.F. The significance of estradiolmetabolites in human corpus luteum physiology. Steroids 2017, 123, 50–54. [Google Scholar] [CrossRef]
- Lande, R.; Thompson, R. Efficiency of marker-assisted selection in the improvement of quantitative traits. Genetics 1990, 124, 743–756. [Google Scholar] [CrossRef]
- Lin, S.; Zhang, H.; Hou, Y.; Liu, L.; Li, W.; Jiang, J.; Han, B.; Zhang, S.; Sun, D. SNV discovery and functional candidate gene identification for milk composition based on whole genome resequencing of Holstein bulls with extremely high and low breeding values. PLoS ONE 2019, 14, e0220629. [Google Scholar] [CrossRef]
- Gershon, E.; Dekel, N. Newly Identified Regulators of Ovarian Folliculogenesis and Ovulation. Int. J. Mol. Sci. 2020, 21, 4565. [Google Scholar] [CrossRef]
- Bai, Y.; Zhang, F.; Zhang, H.; Xu, C.; Wu, L.; Xia, C. Follicular Fluid Metabolite Changes in Dairy Cows with Inactive Ovary Identified Using Untargeted Metabolomics. BioMed Res. Int. 2020, 2020, 9837543. [Google Scholar] [CrossRef]
- Aguiar, T.S.; Torrecilha, R.B.P.; Milanesi, M.; Utsunomiya, A.T.H.; Trigo, B.B.; Tijjani, A.; Musa, H.H.; Lopes, F.L.; Ajmone-Marsan, P.; Carvalheiro, R.; et al. Association of Copy Number Variation at Intron 3 of HMGA2 With Navel Length in Bos indicus. Front. Genet. 2018, 9, 627. [Google Scholar] [CrossRef] [PubMed]
- Lango Allen, H.; Estrada, K.; Lettre, G.; Berndt, S.I.; Weedon, M.N.; Rivadeneira, F.; Willer, C.J.; Jackson, A.U.; Vedantam, S.; Raychaudhuri, S.; et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 2010, 467, 832–838. [Google Scholar] [CrossRef]
- Wang, X.; Wang, J.; Zhao, J.; Wang, H.; Chen, J.; Wu, J. HMGA2 facilitates colorectal cancer progression via STAT3-mediated tumor-associated macrophage recruitment. Theranostics 2022, 12, 963–975. [Google Scholar] [CrossRef] [PubMed]
- Mansoori, B.; Mohammadi, A.; Ditzel, H.J.; Duijf, P.H.G.; Khaze, V.; Gjerstorff, M.F.; Baradaran, B. HMGA2 as a Critical Regulator in Cancer Development. Genes 2021, 12, 269. [Google Scholar] [CrossRef] [PubMed]
- Boyko, A.R.; Quignon, P.; Li, L.; Schoenebeck, J.J.; Degenhardt, J.D.; Lohmueller, K.E.; Zhao, K.; Brisbin, A.; Parker, H.G.; von Holdt, B.M.; et al. A simple genetic architecture underlies morphological variation in dogs. PLoS Biol. 2010, 8, e1000451. [Google Scholar] [CrossRef]
- Makvandi-Nejad, S.; Hoffman, G.E.; Allen, J.J.; Chu, E.; Gu, E.; Chandler, A.M.; Loredo, A.I.; Bellone, R.R.; Mezey, J.G.; Brooks, S.A.; et al. Four loci explain 83% of size variation in the horse. PLoS ONE 2012, 7, e39929. [Google Scholar] [CrossRef]
- Chung, J.; Zhang, X.; Collins, B.; Sper, R.B.; Gleason, K.; Simpson, S.; Koh, S.; Sommer, J.; Flowers, W.L.; Petters, R.M.; et al. High mobility group A2 (HMGA2) deficiency in pigs leads to dwarfism, abnormal fetal resource allocation, and cryptorchidism. Proc. Natl. Acad. Sci. USA 2018, 115, 5420–5425. [Google Scholar] [CrossRef]
- Liu, M.; Hummitzsch, K.; Hartanti, M.D.; Rosario, R.; Bastian, N.A.; Hatzirodos, N.; Bonner, W.M.; Irving-Rodgers, H.F.; Laven, J.S.E.; Anderson, R.A.; et al. Analysis of expression of candidate genes for polycystic ovary syndrome in adult and fetal human and fetal bovine ovaries. Biol Reprod. 2020, 103, 840–853. [Google Scholar] [CrossRef] [PubMed]
- Ferrero, H. HMGA2 involvement in uterine leiomyomas development through angiogenesis activation. Fertil. Steril. 2020, 114, 974–975. [Google Scholar] [CrossRef]
- Wei, J.J. HMGA2: A Biomarker in Gynecologic Neoplasia. J. Clin. Transl. Pathol. 2022, 2, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Zhao, H.; Zhao, S.G.; Wei, D.M.; Zhao, Y.R.; Huang, T.; Muhammad, T.; Yan, L.; Gao, F.; Li, L.; et al. The HMGA2-IMP2 Pathway Promotes Granulosa Cell Proliferation in Polycystic Ovary Syndrome. J. Clin. Endocrinol. Metab. 2019, 104, 1049–1059. [Google Scholar] [CrossRef]
- Neupane, M.; Geary, T.W.; Kiser, J.N.; Burns, G.W.; Hansen, P.J.; Spencer, T.E.; Neibergs, H.L. Loci and pathways associated with uterine capacity for pregnancy and fertility in beef cattle. PLoS ONE 2017, 12, e0188997. [Google Scholar] [CrossRef]
- Bachelot, A. Polycystic ovarian syndrome: Clinical and biological diagnosis. Ann. Biol. Clin. 2016, 74, 661–667. [Google Scholar] [CrossRef] [PubMed]
- Reynolds, L.P.; Redmer, D.A. Growth and development of the corpus luteum. J. Reprod. Fertil. Suppl. 1999, 54, 181–191. [Google Scholar] [CrossRef]
- Aljanabi, S.M.; Martinez, I. Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques. Nucleic Acids Res. 1997, 25, 4692–4693. [Google Scholar] [CrossRef]
- Akhatayeva, Z.; Mao, C.; Jiang, F.G.; Pan, C.Y.; Lin, C.J.; Hao, K.J.; Lan, T.X.; Chen, H.; Zhang, Q.F.; Lan, X.Y. Indel variants within the PRL and GHR genes associated with sheep litter size. Reprod. Domest. Anim. 2020, 55, 1470–1478. [Google Scholar] [CrossRef]
- Hui, Y.; Zhang, Y.; Wang, K.; Pan, C.; Chen, H.; Qu, L.; Song, X.; Lan, X. Goat DNMT3B: An indel mutation detection, association analysis with litter size and mRNA expression in gonads. Theriogenology 2020, 147, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhang, Z.; He, Z.; Tang, W.; Li, T.; Zeng, Z.; He, L.; Shi, Y.Y. A partition-ligation-combination-subdivision EM algorithm for haplotype inference with multiallelic markers: Update of the SHEsis. Cell Res. 2009, 19, 519–523. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.; Fu, W.; Zhao, J.; Shen, J.; Chen, Q.; Zheng, Z.; Chen, H.; Sonstegard, T.S.; Lei, C.; Jiang, Y. BGVD: An Integrated Database for Bovine Sequencing Variations and Selective Signatures. Genom. Proteom. Bioinform. 2020, 18, 186–193. [Google Scholar] [CrossRef]
- Cao, C.; Zhou, Q.; Kang, Y.; Zhanerke, A.; Liu, P.; Bai, Y.; Li, R.; Jiang, Y.; Zhang, Q.; Lan, X.; et al. A repertoire of single nucleotide polymorphisms (SNPs) of major fecundity BMPR1B gene among 75 sheep breeds worldwide. Theriogenology 2024, 219, 59–64. [Google Scholar] [CrossRef] [PubMed]
- Sharma, R.K.; Singh, P.; Setia, A.; Sharma, A.K. Insecticides and ovarian functions. Environ. Mol. Mutagen. 2020, 61, 369–392. [Google Scholar] [CrossRef] [PubMed]
- Nagai, K.; Yanagawa, Y.; Katagiri, S.; Nagano, M. The relationship between antral follicle count in a bovine ovary and developmental competence of in vitro-grown oocytes derived from early antral follicles. Biomed. Res. 2016, 37, 63–71. [Google Scholar] [CrossRef]
- Franchi, F.F.; Hernandes, M.P.; Coalho Ferreira, A.L.; Vieira de Lima, V.A.; de Oliveira Mendes, L.; Musa de Aquino, A.; Scarano, W.R.; César de Souza Castilho, A. Fractal analysis and histomolecular phenotyping provides insights into extracellular matrix remodeling in the developing bovine fetal ovary. Biochem. Biophys. Res. Commun. 2020, 523, 823–828. [Google Scholar] [CrossRef]
- Lan, K.; Shen, C.; Li, J.; Zhang, S.; Lan, X.; Pan, C.; Wang, Y. A novel indel within the bovine SEPT7 gene is associated with ovary length. Anim. Biotechnol. 2023, 34, 8–14. [Google Scholar] [CrossRef]
- Li, J.; Zhang, S.; Shen, C.; Niu, Z.; Yang, H.; Zhang, K.; Liu, Z.; Wang, Y.; Lan, X. Indel mutations within the bovine HSD17B3 gene are significantly associated with ovary morphological traits and mature follicle number. J. Steroid Biochem. Mol. Biol. 2021, 209, 105833. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Jiang, H.; Xiao, L.; Yang, X. MicroRNA-33b-5p is overexpressed and inhibits GLUT4 by targeting HMGA2 in polycystic ovarian syndrome: An in vivo and in vitro study. Oncol. Rep. 2018, 39, 3073–3085. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, A.; Liu, Z.; Gellert, L.; Zou, X.; Yang, G.; Lee, P.; Yang, X.; Wei, J.J. HMGA2: A biomarker significantly overexpressed in high-grade ovarian serous carcinoma. Mod. Pathol. 2010, 23, 673–681. [Google Scholar] [CrossRef]
- Wu, J.; Liu, Z.; Shao, C.; Gong, Y.; Hernando, E.; Lee, P.; Narita, M.; Muller, W.; Liu, J.; Wei, J.J. HMGA2 overexpression-induced ovarian surface epithelial transformation is mediated through regulation of EMT genes. Cancer Res. 2011, 71, 349–359. [Google Scholar] [CrossRef]
- Hock, R.; Witte, F.; Brocher, J.; Schütz, M.; Scheer, U. Expression of HMGA2 variants during oogenesis and early embryogenesis of Xenopus laevis. Eur. J. Cell Biol. 2006, 85, 519–528. [Google Scholar] [CrossRef]
- Van Laere, A.S.; Nguyen, M.; Braunschweig, M.; Nezer, C.; Collette, C.; Moreau, L.; Archibald, A.L.; Haley, C.S.; Buys, N.; Tally, M.; et al. A regulatory mutation in IGF2 causes a major QTL effect on muscle growth in the pig. Nature 2003, 425, 832–836. [Google Scholar] [CrossRef]
- Zheng, J.; Deng, T.; Jiang, E.; Li, J.; Wijayanti, D.; Wang, Y.; Ding, X.; Lan, X. Genetic variations of bovine PCOS-related DENND1A gene identified in GWAS significantly affect female reproductive traits. Gene 2021, 802, 145867. [Google Scholar] [CrossRef] [PubMed]


| Loci | Rs Numbers | Primer Sequences (5′-3′) | Product Size (bp) | Tm (°C) | Region |
|---|---|---|---|---|---|
| P1-I8-bp | rs516271779 | F: GGATTTGCGAGCAAGTCT | 111/95 | TD-PCR | 5′UTR |
| R: TACCCTCCTGGCAGATTG | |||||
| P2-D12-bp | rs797520182 | F: CATTCTCCAGCCCTACCTCG | 172 | TD-PCR | 5′UTR |
| R: GTTGTCCCTGGGCTGAAGTG | |||||
| P3-I23-bp | rs797609300 | F: CAACGGTTCTGTTGAGCAGC | 120 | TD-PCR | Intron |
| R: CACCGGTTGGCTCCTAGTTT | |||||
| P4-D22-bp | rs133750033 | F:AGAGCCTTTCTAGCAGTAGGTG | 172/194 | TD-PCR | Intron |
| R: CAAGGTAAAGGGGGCTTTGG |
| Sample Size | Genotypic Frequencies | Allelic Frequencies | HWE p Value | Population Parameter Estimates | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| II | ID | DD | I | D | Ho | He | Ne | PIC | ||
| 634 | 0.180 | 0.340 | 0.480 | 0.350 | 0.650 | p < 0.05 | 0.543 | 0.457 | 1.840 | 0.352 |
| Sample Sizes | Traits (Units) | Observed Genotypes (Mean ± SE) | p-Values | ||
|---|---|---|---|---|---|
| II (n) | ID (n) | DD (n) | |||
| 339 | Ovarian length (mm) | 42.65 A ± 0.60 | 40.68 B ± 1.11 | 37.67 AB ± 1.39 | 0.001 |
| 339 | Ovarian width (mm) | 26.59 a ± 0.50 | 25.38 b ± 0.65 | 25.03 ab ± 0.87 | 0.172 |
| 339 | Ovarian height (mm) | 24.70 B ± 0.58 | 25.31 AB ± 0.72 | 27.82 A ± 1.13 | 0.026 |
| 338 | Ovarian weight (g) | 11.91 ± 0.38 | 11.52 ± 0.55 | 10.80 ± 0.68 | 0.356 |
| 341 | Number of corpus lutea | 1.83 a ± 0.08 | 1.75 ab ± 0.11 | 1.57 b ± 0.13 | 0.288 |
| 340 | Diameter of corpus luteum (mm) | 10.45 ± 0.54 | 10.30 ± 0.77 | 10.59 ± 0.88 | 0.971 |
| 285 | Number of mature follicles | 0.43 B ± 0.05 | 0.44 A ± 0.08 | 0.50 A ± 0.11 | 0.827 |
| 98 | Diameter of mature follicle (mm) | 12.99 B ± 0.58 | 13.11 A ± 0.98 | 11.68 A ± 1.15 | 0.539 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, S.; Zhu, L.; Yang, Y.; Bi, Y.; Li, J.; Wang, Y.; Pan, C.; Wang, S.; Lan, X. Exploration of the Polymorphism Distribution of Bovine HMGA2 Gene in Worldwide Breeds and Its Associations with Ovarian Traits. Animals 2024, 14, 796. https://doi.org/10.3390/ani14050796
Shen S, Zhu L, Yang Y, Bi Y, Li J, Wang Y, Pan C, Wang S, Lan X. Exploration of the Polymorphism Distribution of Bovine HMGA2 Gene in Worldwide Breeds and Its Associations with Ovarian Traits. Animals. 2024; 14(5):796. https://doi.org/10.3390/ani14050796
Chicago/Turabian StyleShen, Siyuan, Leijing Zhu, Yuanzhe Yang, Yi Bi, Jie Li, Yongsheng Wang, Chuanying Pan, Shuilian Wang, and Xianyong Lan. 2024. "Exploration of the Polymorphism Distribution of Bovine HMGA2 Gene in Worldwide Breeds and Its Associations with Ovarian Traits" Animals 14, no. 5: 796. https://doi.org/10.3390/ani14050796




