Transcriptomic Responses of Two Ecologically Divergent Populations of Japanese Mantis Shrimp (Oratosquilla oratoria) under Thermal Stress
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Approval and Participation Consent
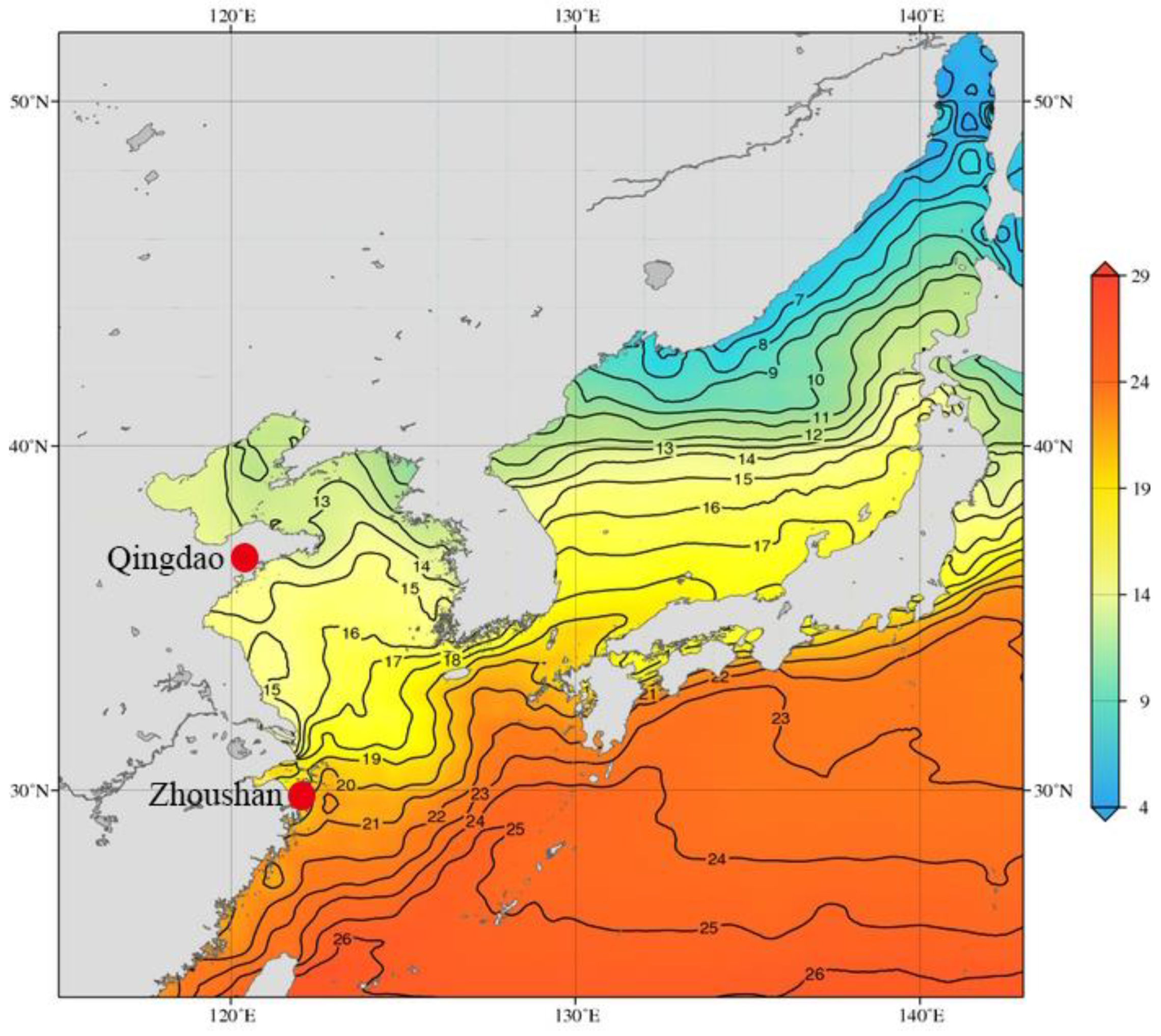
2.2. O. oratoria Maintenance and Heat Exposure
2.3. Total RNA Extraction and Illumina Sequencing
2.4. De Novo Assembly, Gene Expression Variation, and Gene Annotation
2.5. Testing for Population-Specific Genetic Differentiation
2.6. Identifying Expression Responses to Thermal Stress
2.7. Quantitative Reverse Transcription PCR (qRT-PCR) Validation
3. Results
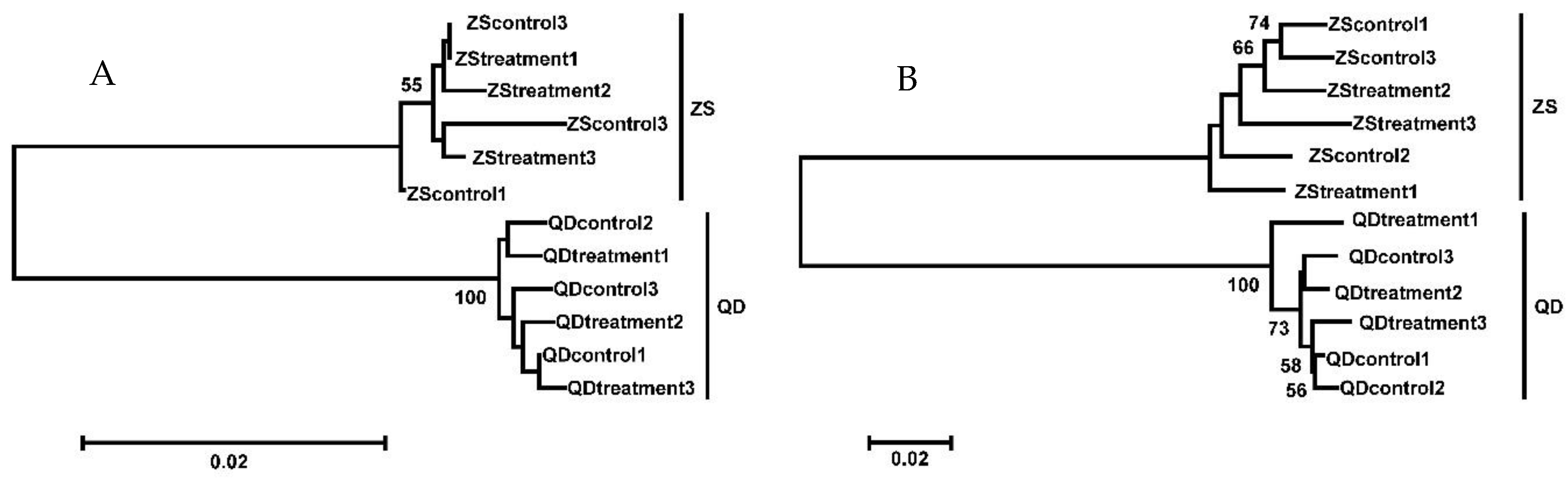
3.1. Phylogenetic Relationship, Illumina Sequencing, and Annotation of the O. oratoria Muscle Transcriptome
3.2. Quantifying Gene Expression Variation
3.3. Population-Specific Gene Expression
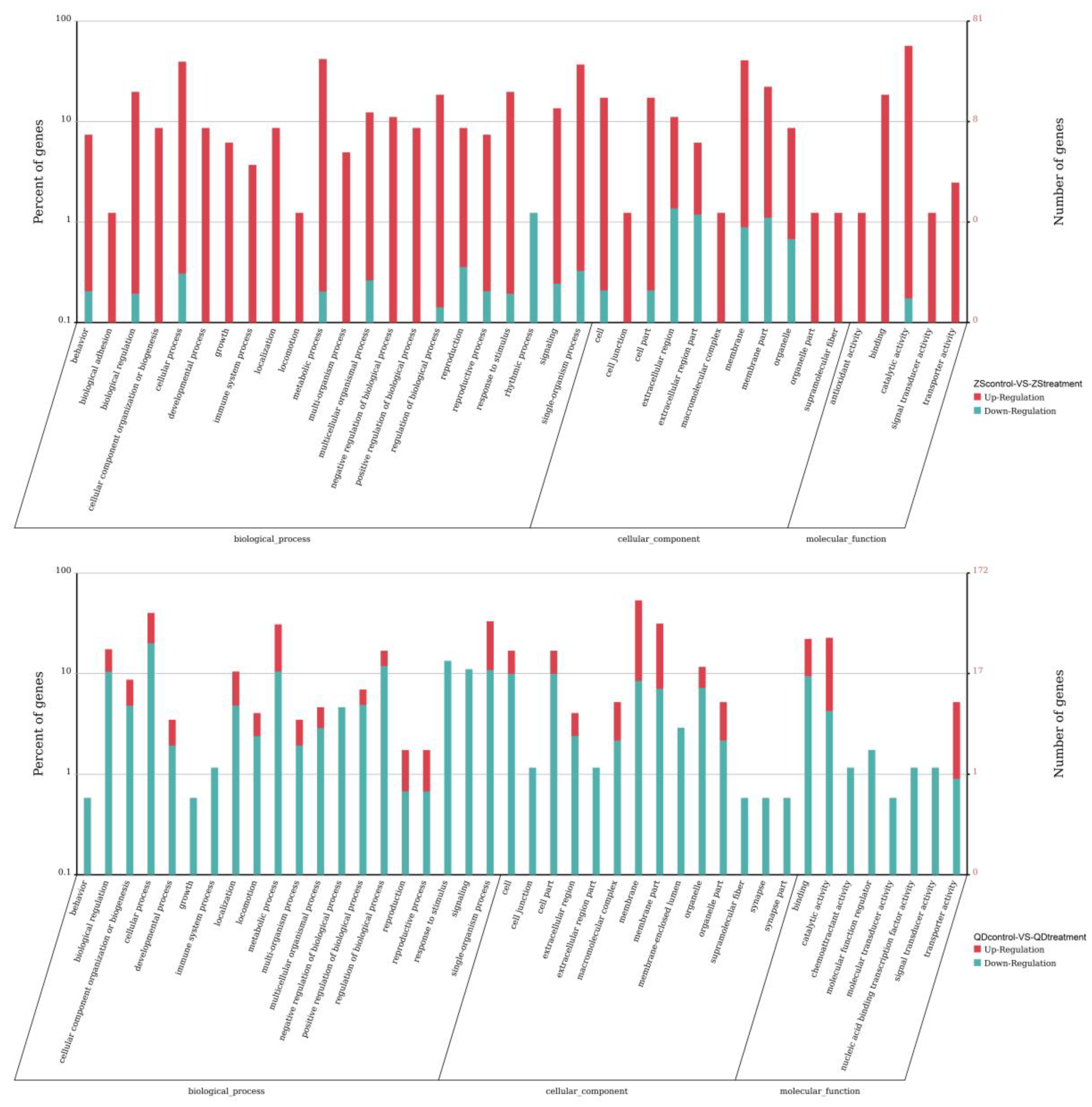
3.4. Differentially Expressed Responses of the Two Populations Exposed to the Same Thermal Scheme
3.5. Validation of the Transcriptome Data by qRT-PCR
4. Discussion
4.1. Gene Expression Variation between Geographic Populations
4.2. Differentially Expressed Responses of the Two Populations Exposed to Thermal Stress
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bopp, L.; Resplandy, L.; Orr, J.C.; Doney, S.C.; Dunne, J.P.; Gehlen, M.; Halloran, P.; Heinze, C.; Ilyina, T.; Séférian, R.; et al. Multiple stressors of ocean ecosystems in the 21st century: Projections with CMIP5 models. Biogeosciences 2013, 10, 6225–6245. [Google Scholar] [CrossRef]
- Dissanayake, A.; Ishimatsu, A. Synergistic effects of elevated CO2 and temperature on the metabolic scope and activity in a shallow-water coastal decapod (Metapenaeus joyneri; Crustacea: Penaeidae). ICES J. Mar. Sci. 2011, 68, 1147–1154. [Google Scholar] [CrossRef]
- Sunday, J.M.; Bates, A.E.; Dulvy, N.K. Thermal tolerance and the global redistribution of animals. Nat. Clim. Chang. 2012, 2, 686–690. [Google Scholar] [CrossRef]
- Johansen, J.L.; Messmer, V.; Coker, D.J.; Hoey, A.S.; Pratchett, M.S. Increasing ocean temperatures reduce activity patterns of a large commercially important coral reef fish. Glob. Chang. Biol. 2014, 20, 1067–1074. [Google Scholar] [CrossRef] [PubMed]
- Pratchett, M.S. Dietary overlap among coral-feeding butterflyfishes (Chaetodontidae) at Lizard Island, northern Great Barrier Reef. Mar. Biol. 2005, 148, 373–382. [Google Scholar] [CrossRef]
- Tewksbury, J.J.; Huey, R.B.; Deutsch, C.A. Putting heat on tropical animals. Science 2008, 320, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Matos-Maraví, P.F.; Peña, C.; Willmott, K.R.; Freitas, A.V.L.; Wahlberg, N. Systematics and evolutionary history of butterflies in the “Taygetis clade” (Nymphalidae: Satyrinae: Euptychiina): Towards a better understanding of Neotropical biogeography. Mol. Phylogenet. Evol. 2013, 66, 54–68. [Google Scholar] [CrossRef] [PubMed]
- Yannic, G.; Pellissier, L.; Ortego, J.; Lecomte, N.; Couturier, S.; Cuyler, C.; Dussault, C.; Hundertmark, K.J.; Irvine, R.J.; Jenkins, D.A.; et al. Genetic diversity in caribou linked to past and future climate change. Nat. Clim. Chang. 2014, 4, 132–137. [Google Scholar] [CrossRef]
- Lemaire, P.; Bernard, E.; Martinez-Paz, J.; Chim, L. Combined effect of temperature and salinity on osmoregulation of juvenile and subadult Penaeus stylirostris. Aquaculture 2002, 209, 307–317. [Google Scholar] [CrossRef]
- Tiyagarajan, V.; Harder, T.; Qian, P.Y. Combined effects of temperature and salinity on larval development and attachment of the subtidal barnacle Balanus trigonus Darwin. J. Exp. Mar. Biol. Ecol. 2003, 287, 223–236. [Google Scholar] [CrossRef]
- Larson, W.A.; Seeb, L.W.; Everett, M.V.; Waples, R.K.; Templin, W.D.; Seeb, J.E. Genotyping by sequencing resolves shallow population structure to inform conservation of Chinook salmon (Oncorhynchus tshawytscha). Evol. Appl. 2014, 7, 355–369. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.D.; Xue, D.X.; Wang, J.; Li, Y.L.; Liu, B.J.; Liu, J.X. Development and preliminary evaluation of a genomewide single nucleotide polymorphisms resource generated by RAD-seq for the small yellow croaker (Larimichthys polyactis). Mol. Ecol. Resour. 2015, 16, 755–768. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.H.; Cai, C.; Hu, X.X.; Liu, Y.; Guo, Y.N.; Hu, P.; Chen, Z.Z.; Peng, S.H.; Zhang, D.S.; Jiang, S.W.; et al. Evolutionary suppression of erythropoiesis via the modulation of TGF-β signalling in an Antarctic icefish. Mol. Ecol. 2015, 24, 4664–4678. [Google Scholar] [CrossRef] [PubMed]
- Schroder, K.; Irvine, K.M.; Taylor, M.S.; Bokil, N.J.; Cao, K.A.L.; Masterman, K.A.; Labzin, L.I.; Semple, C.A.; Kapetanovic, R.; Fairbairn, L.; et al. Conservation and divergence in Toll-like receptor 4-regulated gene expression in primary human versus mouse macrophages. Proc. Natl. Acad. Sci. USA 2012, 109, 944–953. [Google Scholar] [CrossRef] [PubMed]
- Tirosh, I.; Weinberger, A.; Carmi, M.; Barkai, N. A genetic signature of interspecies variations in gene expression. Nat. Genet. 2006, 38, 830–834. [Google Scholar] [CrossRef] [PubMed]
- Cheviron, Z.A.; Whitehead, A.; Brumfield, R.T. Transcriptomic variation and plasticity in rufous-collared sparrows (Zonotrichia capensis) along an altitudinal gradient. Mol. Ecol. 2010, 17, 4556–4569. [Google Scholar] [CrossRef]
- Morris, M.R.J.; Richard, R.; Leder, E.H.; Barrett, R.D.H.; Aubin-Horth, N.; Rogers, S. Gene expression plasticity evolves in response to colonization of freshwater lakes in threespine stickleback. Mol. Ecol. 2014, 23, 3226–3240. [Google Scholar] [CrossRef]
- Guo, J.; Liu, R.; Huang, L.; Zheng, X.M.; Liu, P.L.; Du, Y.S.; Cai, Z.; Zhou, L.; Wei, X.H.; Zhang, F.M.; et al. Widespread and adaptive alterations in genome-wide gene expression associated with ecological divergence of two Oryza species. Mol. Biol. Evol. 2016, 33, 62–78. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.M.; Snyder, M. RNA-seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef]
- Humphreys, D.T.; Westman, B.J.; Martin, D.I.K.; Preiss, T. MicroRNAs control translation initiation by inhibiting eukaryotic initiation factor 4E/cap and poly(A) tail function. Proc. Natl. Acad. Sci. USA 2005, 102, 16961–16966. [Google Scholar] [CrossRef] [Green Version]
- Marshall, W.S.; Grosell, M. Ion transport, osmoregulation and acid-base balance. Physiol. Fishes 2006, 3, 177–230. [Google Scholar]
- Cheng, J.; Sha, Z.L. Cryptic diversity in the Japanese mantis shrimp Oratosquilla oratoria (Crustacea: Squillidae): Allopatric diversification, secondary contact and hybridization. Sci. Rep. 2017, 7, 1972. [Google Scholar] [CrossRef] [PubMed]
- Du, X.W.; Cai, S.S.; Yu, C.G.; Jiang, X.Q.; Lin, L.S.; Gao, T.X.; Han, Z.Q. Population genetic structure of mantis shrimps Oratosquilla oratoria: Testing the barrier effect of the Yangtze River Outflow. Biochem. Syst. Ecol. 2016, 66, 12–18. [Google Scholar] [CrossRef]
- Johnson, D.; Boyer, T. East Asian Seas Regional Climatology (Version 2). National Centers for Environmental Information, NOAA. 2015. Available online: https://www.nodc.noaa.gov/OC5/regional_climate/EASclimatology/ (accessed on 29 June 2018).
- Podrabsky, J.E.; Somero, G.N. Changes in gene expression associated with acclimation to constant temperatures and fluctuating daily temperatures in an annual killifish Austrofundulus limnaeus. J. Exp. Biol. 2004, 207, 2237–2254. [Google Scholar] [CrossRef] [PubMed]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I form diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar] [PubMed]
- Lui, K.K.Y.; Leung, P.T.Y.; Ng, W.C.; Leung, K.M.Y. Genetic variation of Oratosquilla oratoria (Crustacea: Stomatopoda) across Hong Kong waters elucidated by mitochondrial DNA control region sequences. J. Mar. Biol. Assoc. UK 2010, 90, 623–631. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Feng, D.; Li, Q.; Yu, H.; Zhao, X.L.; Kong, L.F. Comparative transcriptome analysis of the Pacific oyster Crassostrea gigas characterized by shell colors: Identification of genetic bases potentially involved in pigmentation. PLoS ONE 2015, 10, e0145257. [Google Scholar] [CrossRef]
- Smith, N.G.C.; Eyre-Walker, A. Adaptive protein evolution in Drosophila. Nature 2002, 415, 1022–1024. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Dai, W.; Kang, J.; Yang, L.; He, S. Comprehensive transcriptome analysis of six catfish species from an altitude gradient reveals adaptive evolution in Tibetan fishes. G3 Genes Genomes Genet. 2016, 6, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, L.; Zhou, K.; Zhang, Y.; Song, Z.; He, S. Evidence for adaptation to the Tibetan Plateau inferred from Tibetan loach transcriptomes. Genome Biol. Evol. 2015, 7, 2970–2982. [Google Scholar] [CrossRef] [PubMed]
- Backström, N.; Zhang, Q.; Edwards, S.V. Evidence from a house finch (Haemorhous mexicanus) spleen transcriptome for adaptive evolution and biased gene conversion in passerine birds. Mol. Biol. Evol. 2013, 30, 1046–1050. [Google Scholar] [CrossRef] [PubMed]
- Logan, C.A.; Buckley, B.A. Transcriptomic responses to environmental temperature in eurythermal and stenothermal fishes. J. Exp. Biol. 2015, 218, 1915–1924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodnick, K.J.; Gamperl, A.K.; Lizars, K.R.; Bennett, M.T.; Rausch, R.N.; Keeley, E.R. Thermal tolerance and metabolic physiology among redband trout populations in south-eastern Oregon. J. Fish Biol. 2004, 64, 310–335. [Google Scholar] [CrossRef]
- Eliason, E.J.; Clark, T.D.; Hague, M.J.; Hanson, L.M.; Gallagher, Z.S.; Jeffries, K.M.; Gale, M.K.; Patterson, D.A.; Hinch, S.G.; Farrell, A.P. Differences in thermal tolerance among sockeye salmon populations. Science 2011, 109, 109–112. [Google Scholar] [CrossRef]
- Passow, C.N.; Brown, A.P.; Arias-Rodriquez, L.; Yee, M.; Sockell, A.; Schartl, M.; Warren, W.C.; Bustamante, C.; Kelley, J.L.; Tobler, M. Complexities of gene expression patterns in natural populations of an extremophile fish (Poecilia mexicana, Poeciliidae). Mol. Ecol. 2017, 16, 4211–4225. [Google Scholar] [CrossRef]
- Pörtner, H.O. Climate variations and the physiological basis of temperature dependent biogeography: Systemic to molecular hierarchy of thermal tolerance in animals. Comp. Biochem. Phys. A 2002, 4, 739–761. [Google Scholar] [CrossRef]
- Pörtner, H.O.; Peck, L.; Somero, G. Thermal limits and adaptation in marine Antarctic ectotherms: An integrative view. Philos. Trans. R. Soc. A 2007, 1488, 2233–2258. [Google Scholar] [CrossRef]
- Sala-Rabanal, M.; Sánchez, J.; Ibarz, A.; Fernández-Borràs, J.; Blasco, J.; Gallardo, M.A. Effects of low temperatures and fasting on hematology and plasma composition of Gilthead sea bream (Sparus aurata). Fish Physiol. Biochem. 2004, 2, 105–115. [Google Scholar] [CrossRef]
- Leaver, M.J.; Boukouvala, E.; Antonopoulou, E.; Diez, A.; Favre-Krey, L.; Ezaz, M.T.; Bautista, J.M.; Tocher, D.R.; Krey, G. Three peroxisome proliferator-activated receptor isotypes from each of two species of marine fish. Endocrinology 2005, 7, 3150–3162. [Google Scholar] [CrossRef] [PubMed]
- Savir, Y.; Tlusty, T. The ribosome as an optimal decoder: A lesson in molecular recognition. Cell 2013, 2, 471–479. [Google Scholar] [CrossRef] [PubMed]
- Brandman, O.; Stewart-Ornstein, J.; Wong, D.; Larson, A.; Williams, C.C.; Li, G.; Zhou, S.; King, D.; Shen, P.S.; Weibezahn, J.; et al. A ribosome-bound quality control complex triggers degradation of nascent peptides and signals translation stress. Cell 2012, 5, 1042–1054. [Google Scholar] [CrossRef] [PubMed]
- Shen, P.S.; Park, J.; Qin, Y.; Li, X.; Parsawar, K.; Larson, M.H.; Cox, J.; Cheng, Y.F.; Lambowitz, A.M.; Weissman, J.S.; et al. Rqc2p and 60S ribosomal subunits mediate mRNA-independent elongation of nascent chains.elongation of nascent chains. Science 2015, 6217, 75–78. [Google Scholar] [CrossRef] [PubMed]
- Ron, D.; Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat. Rev. Mol. Cell Biol. 2007, 7, 519–529. [Google Scholar] [CrossRef] [PubMed]
- Imrie, D.; Sadler, K.C. Stress management: How the unfolded protein response impacts fatty liver disease. J. Hepatol. 2012, 57, 1147–1151. [Google Scholar] [CrossRef] [Green Version]
- Hoekstra, H.E.; Hirschmann, R.J.; Bundey, R.A.; Insel, P.A.; Crossland, J.P. A single amino acid mutation contributes to adaptive beach mouse color pattern. Science 2006, 5783, 101–104. [Google Scholar] [CrossRef]
- Mininni, A.N.; Milan, M.; Ferraresso, S.; Petochi, T.; Di, M.P.; Marino, G.; Livi, S.; Romualdi, C.; Barqelloni, L.; Patarnello, T. Liver transcriptome analysis in gilthead sea bream upon exposure to low temperature. BMC Genom. 2014, 15, 765. [Google Scholar] [CrossRef]
- Cassinelli, J.D.; Moffitt, C.M. Comparison of growth and stress in resident redband trout held in laboratory simulations of montane and desert summer temperature cycles. Trans. Am. Fish. Soc. 2010, 2, 339–352. [Google Scholar] [CrossRef]
- Narum, S.R.; Campbell, N.R.; Meyer, K.A.; Miller, M.R.; Hardy, R.W. Thermal adaptation and acclimation of ectotherms from differing aquatic climates. Mol. Ecol. 2013, 11, 3090–9097. [Google Scholar] [CrossRef] [PubMed]
- Somero, G.N.; Barnes, B.; Gordon, M.; Sato, K.; Hoppeler, H. The physiology of climate change: How potentials for acclimatization and genetic adaptation will determine ‘winners’ and ‘losers’. J. Exp. Biol. 2010, 6, 912–920. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Back, S.H.; Hur, J.; Lin, Y.H.; Gildersleeve, R.; Shan, J.X.; Yuan, C.L.; Krokowski, D.; Wang, S.Y.; Hatzoglou, M.; et al. ER-stressinduced transcriptional regulation increases protein synthesis leading to cell death. Nat. Cell Biol. 2013, 5, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Pearson, G.; Robinson, F.; Beers, G.T.; Xu, B.E.; Karandikar, M.; Berman, K.; Cobb, M.H. Mitogen-activated protein (MAP) kinase pathways: Regulation and physiological functions. Endocr. Rev. 2001, 2, 153–183. [Google Scholar]
- Cargnello, M.; Roux, P.P. Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol. Mol. Biol. R. 2011, 1, 50–83. [Google Scholar] [CrossRef] [PubMed]
- Wilson, C.H.; Ali, E.S.; Scrimgeour, A.N.; Martin, A.M.; Hua, J.; Tallis, G.A.; Rychkov, G.Y.; Barritt, G.J. Steatosis inhibits liver cell store-operated Ca2+ entry and reduces ER Ca2+ through a protein kinase C-dependent mechanism. Biochem. J. 2015, 2, 379–390. [Google Scholar] [CrossRef] [PubMed]
- Ali, E.S.; Hua, J.; Wilson, C.H.; Tallis, G.A.; Zhou, F.H.; Rychkov, G.Y.; Barritt, G.J. The glucagon-like peptide-1 analogue exendin-4 reverses impaired intracellular Ca2+ signalling in steatotic hepatocytes. BBA-Mol. Cell Res. 2016, 9, 2135–2146. [Google Scholar]
- Ibarz, A.; Blasco, J.; Gallardo, M.A.; Fernández-Borràs, J. Energy reserves and metabolic status affect the acclimation of gilthead sea bream (Sparus aurata) to cold. Comp. Biochem. Phys. A 2010, 3, 319–326. [Google Scholar] [CrossRef]
- Narum, S.R.; Campbell, N.R. Transcriptomic response to heat stress among ecologically divergent populations of redband trout. BMC Genom. 2015, 16, 103. [Google Scholar] [CrossRef]
- Koolhaas, J.M. Coping style and immunity in animals: Making sense of individual variation. Brain Behav. Immun. 2008, 5, 662–667. [Google Scholar] [CrossRef]
- Schulte, P.M. What is environmental stress? Insights from fish living in variable environments. J. Exp. Biol. 2014, 217, 23–34. [Google Scholar] [CrossRef] [PubMed]





| ID | Swiss-Annotation | Primer(5′ to 3′) | Product Length |
|---|---|---|---|
| CL17736.Contig.3_All | HSP6C_DROME | For_ATAATGCCGACGGGAAACC | 140 bp |
| Rev_ATATCAACGAGGTATGACTGA | |||
| CL3018.Contig.1_All | H3V1_DICDI | For_ATGGTCGTTGCTGAGTGG | 185 bp |
| Rev_CAACGCCACAGCAAAT | |||
| Unigene53382_All | ORCT_DROME | For_ CCACAAGGTTGGGTCT | 146 bp |
| Rev_ TTGGCTCTAGGTGAAT | |||
| Unigene16740_All | Y7791_DICDI | For_ ACCCTCCTACTGGTCTTCC | 196 bp |
| Rev_ TCGGTGATGTATTGAGGC | |||
| Unigene80423_All | CFDP2_TRAJA | For_ ATCCGAAATAGTCGTCAC | 111 bp |
| Rev_ ATCGTCCTGAAGTGGC |
| Populations | Replicate | Condition | Raw Reads | Raw Bases | Clean Reads | Clean Bases |
|---|---|---|---|---|---|---|
| Qingdao | 1 | 20 °C/24 h | 91,494,516 | 13,724,177,400 | 70,228,416 | 10,534,262,400 |
| Qingdao | 2 | 20 °C/24 h | 84,027,732 | 12,604,159,800 | 69,060,346 | 10,359,051,900 |
| Qingdao | 3 | 20 °C/24 h | 98,810,632 | 14,821,594,800 | 71,406,246 | 10,710,936,900 |
| Qingdao | 1 | 28 °C/24h | 80,275,070 | 12,041,260,500 | 69,696,038 | 10,454,405,700 |
| Qingdao | 2 | 28 °C/24h | 87,961,408 | 13,194,211,200 | 73,770,504 | 11,065,575,600 |
| Qingdao | 3 | 28 °C/24h | 101,108,472 | 15,166,270,800 | 68,445,158 | 10,266,773,700 |
| Zhoushan | 1 | 20 °C/24 h | 88,024,914 | 13,203,737,100 | 73,164,030 | 10,974,604,500 |
| Zhoushan | 2 | 20 °C/24 h | 80,841,856 | 12,126,278,400 | 68,454,208 | 10,268,131,200 |
| Zhoushan | 3 | 20 °C/24 h | 90,334,502 | 13,550,175,300 | 74,637,442 | 11,195,616,300 |
| Zhoushan | 1 | 28 °C/24h | 88,929,736 | 13,339,460,400 | 74,887,104 | 11,233,065,600 |
| Zhoushan | 2 | 28 °C/24h | 82,744,124 | 12,411,618,600 | 62,347,700 | 9,352,155,000 |
| Zhoushan | 3 | 28 °C/24h | 76,745,954 | 11,511,893,100 | 66,281,820 | 9,942,273,000 |
| GO ID | Description | Cluster Frequency | Genome Frequency | p-Value |
|---|---|---|---|---|
| QDcontrol-VS-ZScontrol (20 °C) | ||||
| GO:0016491 | oxidoreductase activity | 269 out of 1679 genes | 1055 out of 8386 genes | 0.00146 |
| GO:0003995 | acyl-CoA dehydrogenase activity | 13 out of 1679 genes | 22 out of 8386 genes | 0.04377 |
| QDtreatment-VS-ZStreatment (28 °C) | ||||
| GO:0005737 | cytoplasm | 798 out of 1635 genes | 3039 out of 8684 genes | 1.74 × 10−36 |
| GO:0044444 | cytoplasmic part | 638 out of 1635 genes | 2357 out of 8684 genes | 1.6 × 10−30 |
| GO:0044391 | ribosomal subunit | 195 out of 1635 genes | 520 out of 8684 genes | 5.27 × 10−22 |
| GO:0005840 | ribosome | 253 out of 1635 genes | 748 out of 8684 genes | 1.17 × 10−23 |
| GO:0022626 | cytosolic ribosome | 157 out of 1635 genes | 404 out of 8684 genes | 5.44 × 10−19 |
| GO:0003735 | structural constituent of ribosome | 233 out of 1585 genes | 695 out of 8386 genes | 9.98 × 10−19 |
| GO:0043228 | non-membrane-bounded organelle | 416 out of 1635 genes | 1487 out of 8684 genes | 1.36 × 10−20 |
| GO:0043232 | intracellular non-membrane-bounded organelle | 416 out of 1635 genes | 1487 out of 8684 genes | 1.36 × 10−20 |
| GO:0005198 | structural molecule activity | 268 out of 1585 genes | 847 out of 8386 genes | 6.25 × 10−18 |
| GO:0044445 | cytosolic part | 161 out of 1635 genes | 431 out of 8684 genes | 1.78 × 10−19 |
| GO ID | Description | Cluster Frequency | Genome Frequency | p-Value |
|---|---|---|---|---|
| ZScontrol-VS-ZStreatment | ||||
| GO:0004698 | calcium-dependent protein kinase C activity | 5 out of 49 genes | 8 out of 8386 genes | 4.05 × 10−8 |
| GO:0019992 | diacylglycerol binding | 5 out of 49 genes | 8 out of 8386 genes | 4.05 × 10−8 |
| GO:0009931 | calcium-dependent protein serine/threonine kinase activity | 5 out of 49 genes | 10 out of 8386 genes | 1.81 × 10−9 |
| GO:0010857 | calcium-dependent protein kinase activity | 5 out of 49 genes | 10 out of 8386 genes | 1.81 × 10−9 |
| GO:0004697 | protein kinase C activity | 5 out of 49 genes | 11 out of 8386 genes | 3.29 × 10−7 |
| GO:0034389 | lipid particle organisation | 5 out of 54 genes | 11 out of 8179 genes | 1.94 × 10−8 |
| GO:0007252 | I-kappaB phosphorylation | 5 out of 54 genes | 12 out of 8179 genes | 3.29 × 10−6 |
| GO:0003824 | catalytic activity | 46 out of 49 genes | 4939 out of 8386 genes | 4.59 × 10−6 |
| GO:0009950 | dorsal/ventral axis specification | 5 out of 54 genes | 13 out of 8179 genes | 5.33 × 10−6 |
| GO:0002793 | positive regulation of peptide secretion | 5 out of 54 genes | 14 out of 8179 genes | 8.24 × 10−6 |
| QDcontrol-VS-QDtreatment | ||||
| GO:0031435 | mitogen-activated protein kinase kinase binding | 4 out of 75 genes | 5 out of 8386 genes | 6.52 × 10−6 |
| GO:0030968 | endoplasmic reticulum unfolded protein response | 5 out of 88 genes | 11 out of 8179 genes | 5.74 × 10−5 |
| GO:0034620 | cellular response to unfolded protein | 5 out of 88 genes | 11 out of 8179 genes | 5.74 × 10−5 |
| GO:0035967 | cellular response to topologically incorrect protein | 5 out of 88 genes | 11 out of 8179 genes | 5.74 × 10−5 |
| GO:0004721 | phosphoprotein phosphatase activity | 9 out of 75 genes | 109 out of 8386 genes | 1.04 × 10−6 |
| GO:0016020 | membrane | 92 out of 115 genes | 5303 out of 8684 genes | 0.00046 |
| GO:0006986 | response to unfolded protein | 5 out of 88 genes | 20 out of 8179 genes | 0.00065 |
| GO:0035966 | response to topologically incorrect protein | 5 out of 88 genes | 20 out of 8179 genes | 0.00065 |
| GO:0016791 | phosphatase activity | 9 out of 75 genes | 158 out of 8386 genes | 0.00084 |
| GO:0042578 | phosphoric ester hydrolase activity | 9 out of 75 genes | 196 out of 8386 genes | 0.00470 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lou, F.; Han, Z.; Gao, T. Transcriptomic Responses of Two Ecologically Divergent Populations of Japanese Mantis Shrimp (Oratosquilla oratoria) under Thermal Stress. Animals 2019, 9, 399. https://doi.org/10.3390/ani9070399
Lou F, Han Z, Gao T. Transcriptomic Responses of Two Ecologically Divergent Populations of Japanese Mantis Shrimp (Oratosquilla oratoria) under Thermal Stress. Animals. 2019; 9(7):399. https://doi.org/10.3390/ani9070399
Chicago/Turabian StyleLou, Fangrui, Zhiqiang Han, and Tianxiang Gao. 2019. "Transcriptomic Responses of Two Ecologically Divergent Populations of Japanese Mantis Shrimp (Oratosquilla oratoria) under Thermal Stress" Animals 9, no. 7: 399. https://doi.org/10.3390/ani9070399






