Evaluation of Virus-Free Manufacture of Recombinant Proteins Using CRISPR-Mediated Gene Disruption in Baculovirus-Infected Insect Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Culture Conditions
2.2. Plasmid Construction
2.3. Recombinant Baculovirus Generation, Amplification, and Quantification
2.4. Infections
2.5. Western Blot
2.6. Immunofluorescence
2.7. Flow Cytometry and Analysis
2.8. Quantification of Baculovirus Particles Using Flow Cytometry
2.9. Quantification of Gag-VLPs with Enzyme-Linked Immunosorbent Assay (ELISA)
3. Results
3.1. Targeting the gp64 ORF Is Site Specific
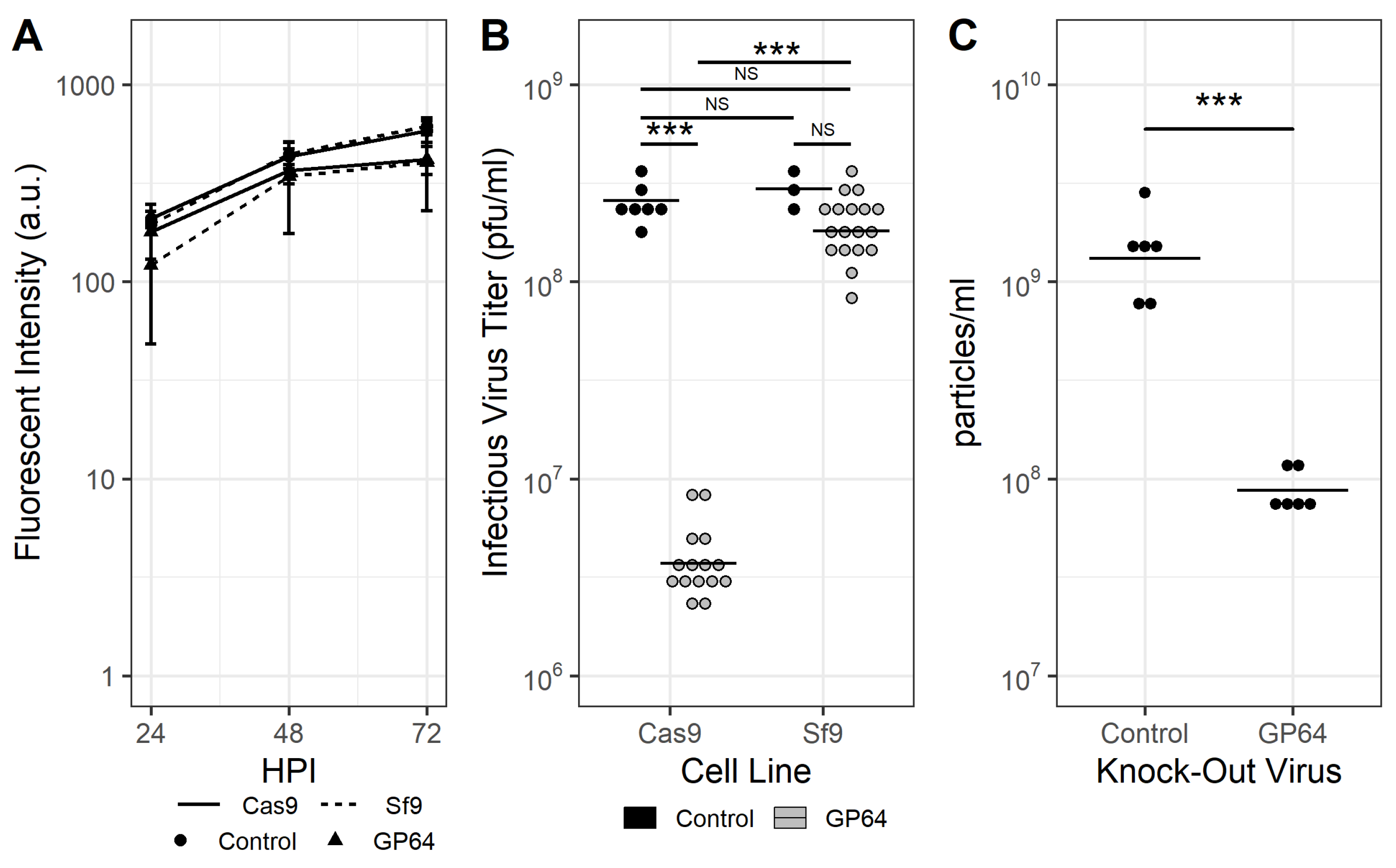
3.2. Cas9-Mediated Disruption of gp64 Impacts Progeny Virus Production but Not Late Gene Expression
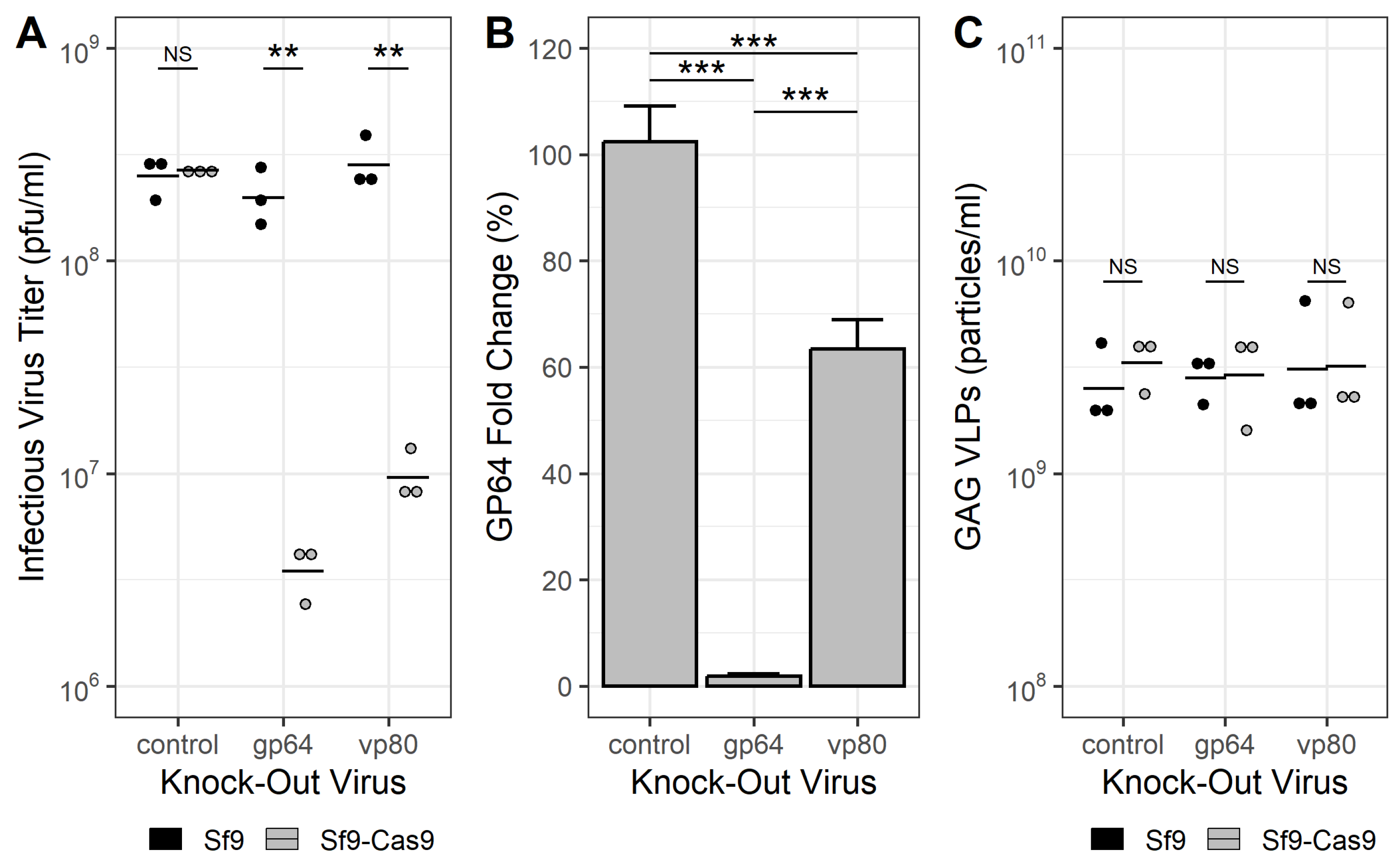
3.3. Production of HIV-1 Gag VLPs
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AcMNPV | Autographa californica multiple nucleopolyhedrovirus |
| APC | allophycocyanin |
| au | arbitrary units |
| BV | budded virus |
| BEVS | baculovirus expression vectors system |
| rBEV | recombinant baculovirus expression vector |
| eGFP | enhanced Green fluorescent protein |
| EPDA | end-point dilution assay |
| GFP | green fluorescent protein |
| hpi | hours post infection |
| hpt | hours post transfection |
| IQR | interquartile range |
| IVT | infectious virus titer |
| KOV | knockout virus |
| MOI | multiplicity of infection |
| ORF | open reading frame |
| PVDF | polyvinylidene difluoride |
| VLP | virus-like particle |
References
- Rohovie, M.J.; Nagasawa, M.; Swartz, J.R. Virus-like particles: Next-generation nanoparticles for targeted therapeutic delivery. Bioeng. Transl. Med. 2017, 2, 43–57. [Google Scholar] [CrossRef] [PubMed]
- Buonaguro, L.; Tagliamonte, M.; Tornesello, M.L.; Buonaguro, F.M. Developments in virus-like particle-based vaccines for infectious diseases and cancer. Expert Rev. Vaccines 2011, 10, 1569–1583. [Google Scholar] [CrossRef] [PubMed]
- Lua, L.H.; Connors, N.K.; Sainsbury, F.; Chuan, Y.P.; Wibowo, N.; Middelberg, A.P. Bioengineering virus-like particles as vaccines. Biotechnol. Bioeng. 2014, 111, 425–440. [Google Scholar] [CrossRef] [PubMed]
- Sokolenko, S.; George, S.; Wagner, A.; Tuladhar, A.; Andrich, J.M.S.; Aucoin, M.G. Co-expression vs. co-infection using baculovirus expression vectors in insect cell culture: Benefits and drawbacks. Biotechnol. Adv. 2012, 30, 766–781. [Google Scholar] [CrossRef] [PubMed]
- Yee, C.M.; Zak, A.J.; Hill, B.D.; Wen, F. The Coming Age of Insect Cells for Manufacturing and Development of Protein Therapeutics. Ind. Eng. Chem. Res. 2018, 57, 10061–10070. [Google Scholar] [CrossRef]
- Khurana, S.; Wu, J.; Verma, N.; Verma, S.; Raghunandan, R.; Manischewitz, J.; King, L.R.; Kpamegan, E.; Pincus, S.; Smith, G.; et al. H5N1 Virus-Like Particle Vaccine Elicits Cross-Reactive Neutralizing Antibodies That Preferentially Bind to the Oligomeric Form of Influenza Virus Hemagglutinin in Humans. J. Virol. 2011, 85, 10945–10954. [Google Scholar] [CrossRef] [PubMed]
- Marek, M.; van Oers, M.M.; Devaraj, F.F.; Vlak, J.M.; Merten, O.W. Engineering of baculovirus vectors for the manufacture of virion-free biopharmaceuticals. Biotechnol. Bioeng. 2011, 108, 1056–1067. [Google Scholar] [CrossRef]
- Chaves, L.C.S.; Ribeiro, B.M.; Blissard, G.W. Production of GP64-free virus-like particles from baculovirus-infected insect cells. J. Gen. Virol. 2018, 99, 265–274. [Google Scholar] [CrossRef]
- Monsma, S.A.; Oomens, A.G.; Blissard, G.W. The GP64 envelope fusion protein is an essential baculovirus protein required for cell-to-cell transmission of infection. J. Virol. 1996, 70, 4607–4616. [Google Scholar] [CrossRef]
- Bruder, M.R.; Walji, S.D.; Aucoin, M.G. Comparison of CRISPR–Cas9 Tools for Transcriptional Repression and Gene Disruption in the BEVS. Viruses 2021, 13, 1925. [Google Scholar] [CrossRef]
- Stemmer, W.P.; Morris, S.K. Enzymatic inverse PCR: A restriction site independent, single-fragment method for high-efficiency, site-directed mutagenesis. BioTechniques 1992, 13, 214–220. [Google Scholar] [PubMed]
- Venereo-Sanchez, A.; Gilbert, R.; Simoneau, M.; Caron, A.; Chahal, P.; Chen, W.; Ansorge, S.; Li, X.; Henry, O.; Kamen, A. Hemagglutinin and neuraminidase containing virus-like particles produced in HEK-293 suspension culture: An effective influenza vaccine candidate. Vaccine 2016, 34, 3371–3380. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.F.; Meghrous, J.; Kamen, A. Quantitation of baculovirus particles by flow cytometry. J. Virol. Methods 2002, 105, 321–330. [Google Scholar] [CrossRef]
- Oomens, A.G.P.; Blissard, G.W. Requirement for GP64 to drive efficient budding of Autographa californica multicapsid nucleopolyhedrovirus. Virology 1999, 254, 297–314. [Google Scholar] [CrossRef] [PubMed]
- Wagner, R.; Fließbach, H.; Wanner, G.; Motz, M.; Niedrig, M.; Deby, G.; Brunn, A.v.; Wolf, H. Studies on processing, particle formation, and immunogenicity of the HIV-1gag gene product: A possible component of a HIV vaccine. Arch. Virol. 1992, 127, 117–137. [Google Scholar] [CrossRef] [PubMed]
- Haynes, J.R. Influenza virus-like particle vaccines. Expert Rev. Vaccines 2014, 8, 435–445. [Google Scholar] [CrossRef]
- Gheysen, D.; Jacobs, E.; Foresta, F.d.; Thiriart, C.; Francotte, M.; Thines, D.; Wilde, M.D. Assembly and release of HIV-1 precursor Pr55gag virus-like particles from recombinant baculovirus-infected insect cells. Cell 1989, 59, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Garnier, L.; Ravallec, M.; Blanchard, P.; Chaabihi, H.; Bossy, J.P.; Devauchelle, G.; Jestin, A.; Cerutti, M. Incorporation of pseudorabies virus gD into human immunodeficiency virus type 1 Gag particles produced in baculovirus-infected cells. J. Virol. 1995, 69, 4060–4068. [Google Scholar] [CrossRef]
- Puente-Massaguer, E.; Gòdia, F.; Lecina, M. Development of a non-viral platform for rapid virus-like particle production in Sf9 cells. J. Biotechnol. 2020, 322, 43–53. [Google Scholar] [CrossRef]
- Puente-Massaguer, E.; Lecina, M.; Gòdia, F. Application of advanced quantification techniques in nanoparticle-based vaccine development with the Sf9 cell baculovirus expression system. Vaccine 2020, 38, 1849–1859. [Google Scholar] [CrossRef]
- Fuenmayor, J.; Gòdia, F.; Cervera, L. Production of virus-like particles for vaccines. New Biotechnol. 2017, 39, 174–180. [Google Scholar] [CrossRef] [PubMed]


| Plasmid Construct | Sequence (5′-3′) | Use (Template) |
|---|---|---|
| Retarget sgRNAs | gttttagagctagaaatagcaagttaaaataagg | retarget sgRNA † (fwd primer) |
| cggtggtcgagcacga | retarget sgRNA † (rev primer) | |
| p6.9GAG-sgRNA | cgaccccagcagccagtaaggcgcgccatgaatc | p6.9-sgRNA backbone |
| catgtttaaattgtgtaatttatgtagctgtaatttttacc | ||
| acagctacataaattacacaatttaaacatgggcgccagagcc | HIV-1 gag ORF | |
| cgattcatggcgcgccttactggctgctggggtcg |
| Gene | Protospacer Sequence (5′-3′) | PAM | Strand |
|---|---|---|---|
| GP64-1 | GGAAACGCTGCAAAAGGACG | TGG | Template |
| GP64-2 | GTTGTAGTCCGTCTCCACGA | TGG | Nontemplate |
| VP80-1 | GCCCGCCGCAATCGCCGCCG | CGG | Template |
| VP80-2 | TCGCTGGATGTTACCCGCGG | CGG | Nontemplate |
| rBEV | Sf9-Cas9 | Sf9 | ||||
|---|---|---|---|---|---|---|
| FL. Intensity (au) | IVT (pfu/mL) | Particles/mL | FL. Intensity (au) | IVT (pfu/mL) | Particles/mL | |
| Control | - | |||||
| GP64 | - | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bruder, M.R.; Aucoin, M.G. Evaluation of Virus-Free Manufacture of Recombinant Proteins Using CRISPR-Mediated Gene Disruption in Baculovirus-Infected Insect Cells. Vaccines 2023, 11, 225. https://doi.org/10.3390/vaccines11020225
Bruder MR, Aucoin MG. Evaluation of Virus-Free Manufacture of Recombinant Proteins Using CRISPR-Mediated Gene Disruption in Baculovirus-Infected Insect Cells. Vaccines. 2023; 11(2):225. https://doi.org/10.3390/vaccines11020225
Chicago/Turabian StyleBruder, Mark R., and Marc G. Aucoin. 2023. "Evaluation of Virus-Free Manufacture of Recombinant Proteins Using CRISPR-Mediated Gene Disruption in Baculovirus-Infected Insect Cells" Vaccines 11, no. 2: 225. https://doi.org/10.3390/vaccines11020225
APA StyleBruder, M. R., & Aucoin, M. G. (2023). Evaluation of Virus-Free Manufacture of Recombinant Proteins Using CRISPR-Mediated Gene Disruption in Baculovirus-Infected Insect Cells. Vaccines, 11(2), 225. https://doi.org/10.3390/vaccines11020225






