1. Introduction
The early detection of graft damage is a major clinical concern in solid organ transplantation (TX). After liver transplantation, biopsy is considered the gold standard technique for evaluating organ rejection. However, this procedure is invasive and can result in clinical complications. Liver biopsy provides diagnostic confirmation of many graft complications, although it involves risk of morbidity (1%) and mortality (from 0.1 to 0.01%) to the patient [
1]. To adjust the treatment, continued assessment of graft integrity is essential for monitoring liver transplant patients. However, the performance of frequent serial liver biopsies should not be considered a possible approach, mainly because of an increase in patient risk exposure. Moreover, this is a costly and time-consuming procedure that is inconvenient for quick changes in medication treatment.
A new approach has gained importance in recent years with the detection of a noninvasive biomarker capable of rapidly detecting organ rejection: cell-free DNA (cfDNA) derived from graft cells. Thus, graft cfDNA in the circulation of liver transplant recipients has been proposed as a potential biomarker for organ rejection or cellular graft injury. The first evidence dates from 1998, when Lo et al. described the presence of chromosome Y-specific sequences in plasma obtained from a female host who received a kidney from a male donor [
2]. They proposed that the DNA released from necrotic or apoptotic cells during rejection may be a useful marker of graft damage. Since then, several groups have determined graft cfDNA using different techniques, such as whole-genome sequencing [
3,
4,
5,
6], digital PCR [
7,
8,
9], and real-time PCR [
10,
11,
12].
Real-time PCR (qPCR) of specific organ cfDNA has been proven to be a noninvasive and advantageous approach for monitoring either liver damage or rejection after TX in a specific subgroup of patients, such as a female host receiving an organ from a male donor [
10,
11]. To expand this methodology to the entire population, we proposed, in a previous work, a method based on a set of insertion–deletion (InDel) diallelic polymorphisms that are already described in the general population [
13]. With this approach, we were able to detect increases in donor-specific cfDNA (ds-cfDNA) in the recipient’s serum after either heart damage or heart transplantation rejection in a patient cohort.
The aim of this study was to detect ds-cfDNA present in the recipient’s serum after either liver damage or rejection using a qPCR-based method. For the proposed approach, the detection of a donor-recipient DNA mismatch for InDel sequences is key for monitoring organ health after TX. Thus, we first determined the donor–recipient DNA mismatch in liver transplant patients. We considered an informative mismatch in the amplification of the InDel sequence in the donor but not in the recipient. Thereafter, we monitored ds-cfDNA levels in the host serum during the first month of follow-up. We described a significant increase in ds-cfDNA related to liver damage on the day of rejection diagnosis and several days before. Moreover, the high negative predictive value obtained in this study suggests that ds-cfDNA quantification using qPCR may be helpful in deciding on the indication for biopsy in the setting of elevated liver biomarkers.
2. Materials and Methods
2.1. Study Subjects
All consecutive patients who underwent orthotropic liver TX at Virgen del Rocio University Hospital in Seville (Spain) during a 4-year period were offered inclusion in the study for the monitoring of ds-cfDNA during the first month. Patients were clinically evaluated at least 2 years after inclusion. After placement on the waiting list, the patients were informed about the study (unless they were in such a poor clinical status that precluded the individual from understanding the rationale behind the study). After agreement, the patients provided signed informed consent, and they were included in the study. Patients who underwent orthotropic liver TX fulfilled the following inclusion criteria: (a) be over 18 years old and (b) sign the informed consent.
During surgery, samples from both donors and recipients were collected. A blood sample from the central line was collected at two different times during surgery: a basal sample at the anhepatic phase and 15 min after graft reperfusion. When a recipient tissue sample was not available, a basal EDTA blood sample was used for obtaining DNA from peripheral blood mononuclear cells.
Patients from whom either a donor or a recipient tissue or blood sample could not be collected were excluded from the study. Cases with either absent or invalid serum samples at the moment of TX or with less than 1 week of follow-up were also excluded from the study. Finally, 97 liver-transplanted patients were included. The general characteristics of the patients are shown in
Table 1.
2.2. Patient Surveillance and Treatment
Patients were evaluated clinically and biologically both during their stay in the intensive care unit (ICU) and the ward. Blood samples were drawn by peripheral venipuncture daily during the first three days, and every 2–3 days thereafter in a Vacuette® 9 mL Z Serum Separator Clot Activator (Greiner Bio-One, Kremsmünster, Austria). We monitored the cfDNA in the serum and the standard biochemical, hematological, and coagulation parameters as well as determined the immunosuppressive drug levels. For cfDNA determination, 10 mL of blood samples were drawn and centrifuged 5 min at 3500 rpm at room temperature within 6 h after extraction, and the serum was aliquoted and frozen at −80 °C for future determination. After the patients were discharged, they were followed up at scheduled visits, drawing a blood sample at every visit. DNA from frozen samples was obtained and stored for no longer than 6 months.
Immunosuppressive treatment for cohort patients consisted of Tacrolimus, mofetil mycophenolate, and steroids. A 7 ng/mL blood level of Calcineurin inhibitors was required during the first month, between 5–7 ng/mL during the first year and less than 5 ng/mL after the first year. With a protocol based on Basiliximab, tacrolimus delayed reduced dose, and mofetil mycophenolate and steroids were used in patients with chronic renal dysfunction.
2.3. Ethics Statement
Ethical approval for the study protocol was granted by the Medical Research Ethics Board of the Virgen del Rocío University Hospital of Seville. The clinical investigation was conducted according to the principles expressed in the Declaration of Helsinki.
2.4. General Workflow
We propose an approach to detect ds-cfDNA in the recipient based on the existence of long genomic sequences (InDels) which are either present or absent in the individual genotype. We have previously described a panel of 10 InDels that were useful in transplant monitoring to differentiate donor cfDNA from that of the recipient [
8,
13]. The panel consists of four null alleles (GSTM1, GSTT1, SRY and RhD) and six InDels (DCP1, Xq28, R271, rs4399, FVII, and THYR). The chromosome position, length of the insertion sequence and frequency of the different InDels is previously described [
8].
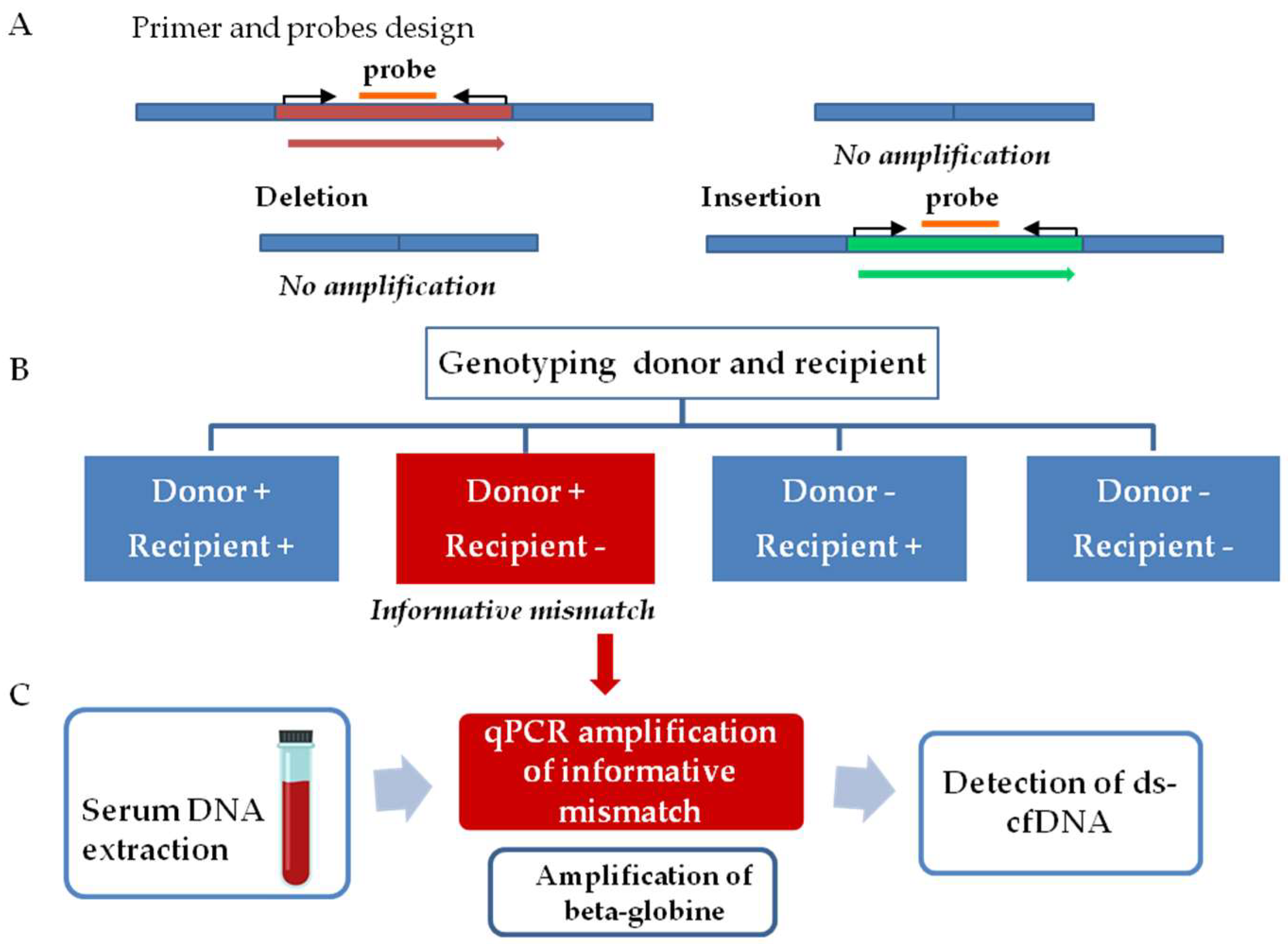
The workflow for graft injury evaluation involves two steps, as follows (
Figure 1). The first step comprised InDel identification. Thus, tissue samples from both donors and recipients were analyzed to determine a donor–recipient mismatch for the InDels analyzed. A mismatch was considered informative when a deleted sequence on the recipient DNA, but not in the donor DNA, was observed. The second step consists of the quantification of ds-cfDNA in the recipient’s serum. After the detection of an informative mismatch, cfDNA from sequential serum samples was analyzed by qPCR. The presence of elevated ds-cfDNA circulating in serum from the recipient is indicative of any type of transplanted liver complication. We also quantified beta-globin gene values as a control for general damage or patient clinical worsening.
2.5. DNA Extraction from Tissue and Serum Samples
DNA from tissue samples was extracted using QIAcube (Qiagen GmbH, Hilden, Germany), according to the manufacturer’s protocol; 5 milligrams of donor and recipient biopsy samples were previously minced and incubated with 180 µL of ATL lysis buffer (animal tissue lysis buffer) and 20 µL of proteinase K for 2 hours at 56 °C. The DNA was eluted in a final volume of 100 μL and was frozen at −80 °C until determination.
DNA from 400 μL of serum samples was extracted using the automatized MagNaPure Compact Instrument (Roche Diagnostics, Basel, Switzerland) using the Magna Pure Compact Nucleic Acid Isolation Kit I, according to the protocol “Total NA Plasma 100 400 V3 1”. The DNA was eluted in a final volume of 50 μL and was frozen at −80 °C until either qPCR.
Quantification of the nucleic acids after DNA isolation was performed using Qubit 3.0 fluorometry (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions.
2.6. Determination of Donor–Recipient Mismatch and Monitoring Organ Specific cfDNA
Selected InDels were amplified from donor and recipient tissue DNA by qPCR assay using the Light-Cycler 480 Real-Time PCR instrument (Roche Diagnostics, Basel, Switzerland). We followed defined guidelines for qPCR analyisis [
14].
Briefly, 2 microliters of DNA were amplified in a final volume of 20 μL containing 200 nM of primers and 100 nM of the probe using the LC480 Probes Master Kit (Roche Diagnostics, Basel, Switzerland). The 2× concentrated master mix is optimized for a fixed MgCl
2 concentration, which works with nearly all primer combinations. No adjustment in the MgCl
2 concentration is needed to amplify different sequences. qPCR was performed at 95 °C for 5″ and at the specific InDel Tm for 20″ for 40 cycles. The standards for the calibration curve were based on the dilutions of human genomic DNA (Roche Diagnostics, Basel, Switzerland). Calibration curve slopes ranged from 3.34 to 3.85 and provided a PCR efficiency range close to 2 [1.81–1.99]. The lower standard provided a limit of detection of 1.5 genomic equivalents (GE)/mL serum. The error value (mean squared error of the single data points fit to the regression line), a measure of the accuracy of the quantification result, was always <0.2 [0.0004–0.01]. The primer and probe sequences, fragment length as well as annealing temperature have been previously described [
8].
For quantification of donor DNA in the recipient’s serum 5 microliters of DNA was amplified by qPCR assay using the Light-Cycler 480 Real-Time PCR instrument (Roche Diagnostics, Basel, Switzerland) following the same protocol as described above.
DNA from each patient was assayed in one round, and all samples were assayed in duplicate. No-template control was included to asses any possible contamination problem in the assay. The final concentration was calculated according to the following formula:
where
GE represents a genomic equivalent,
VDNA represents the total volume of cfDNA obtained after extraction from serum,
VPCR represents the sample volume used for PCR, and
Vext represents the volume of extracted serum. A conversion factor of 6.6 pg of DNA per diploid cell was used to express the cfDNA concentration as genome equivalents.
2.7. Statistical Analysis
All statistical analysis was performed with IBM SPSS version 26 (IBM, Armonk, NY, USA). Continuous data are presented by median and range. Comparisons between two continuous variables were performed by t-test for normal distribution. Mann–Whitney U-test was used to compare differences between two independent groups when the dependent variable was not normally distributed. Multiple comparison analysis was performed by ANOVA test and post hoc analysis by Bonferroni test.
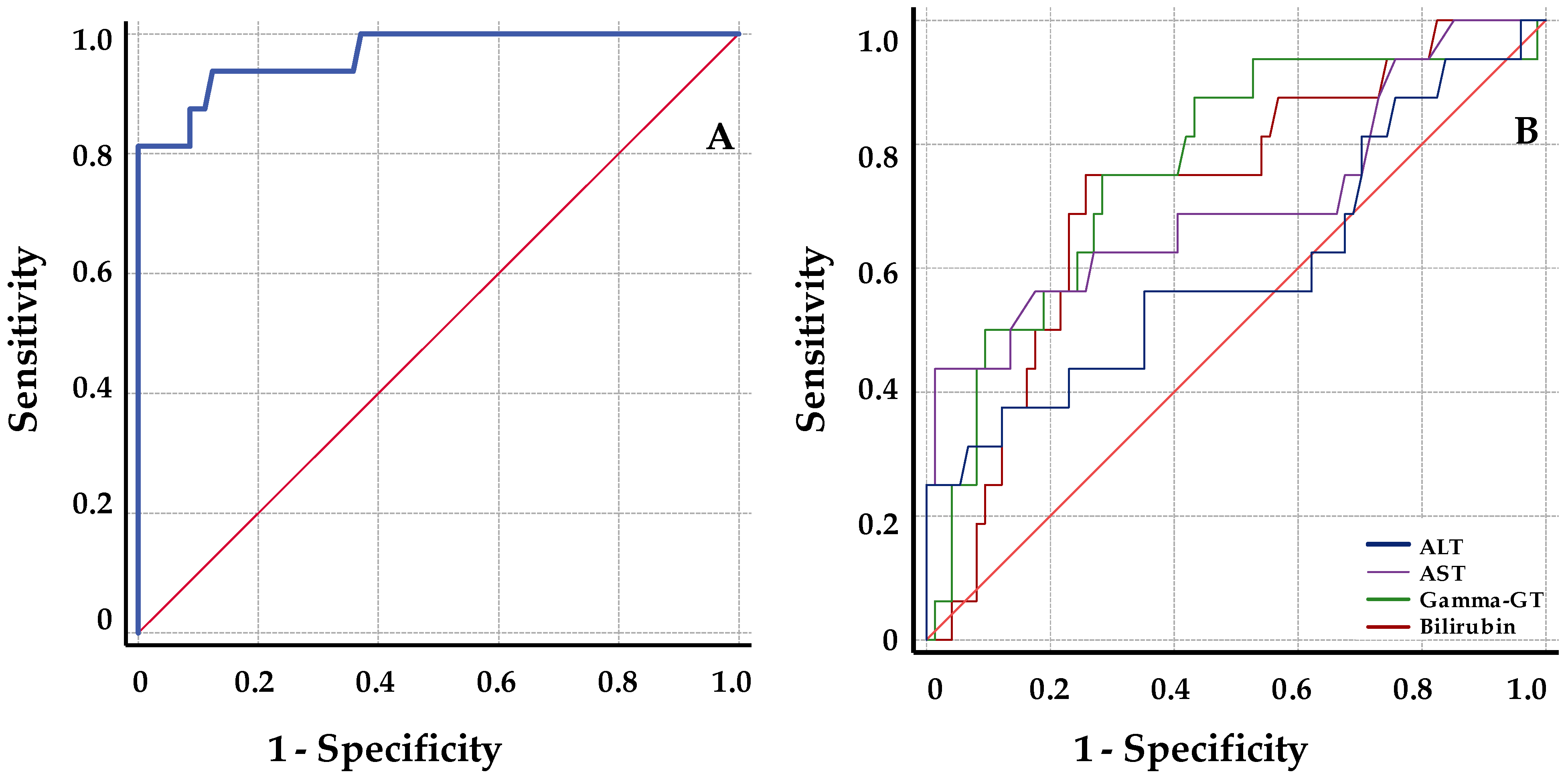
Receiver operator characteristic (ROC) analyses were performed to determine the best parameter to discriminate between biopsy-proven rejection (BPR) and stable samples. ROC curves were performed for classical hepatic markers and graft cfDNA increase in the same analysis. Samples were only included when a determination of all parameters studied was available. Threshold values for discrimination of BPR and stable samples were calculated based on the maximum Youden index. Based in the threshold value obtained, both the positive predictive value (PPV) and the negative predictive value (NPV) were calculated.
4. Discussion
Graft cfDNA released from damaged donor liver cells in the circulation of liver transplant recipients has been proposed as a potential noninvasive biomarker of organ rejection. In this study, transplanted liver rejection was evaluated using real-time PCR. We observed a significant increase in ds-cfDNA over basal levels at the BPR time point and several days before rejection.
In recent years, several authors have proposed the quantification of ds-cfDNA in the circulation of transplant recipients as a potential biomarker for organ rejection or cellular graft injury [
4,
15,
16]. The direct assessment of liver integrity could lead to an earlier detection of acute rejection, which could provide prompt intervention and facilitate the adjustment of an effective therapy. Different techniques have been used for monitoring ds-cfDNA in the host serum, such as analyses of single-nucleotide polymorphisms (SNP), which discriminate between heterologous alleles in the recipient and the donor by means of digital PCR [
7]. Other methods include next-generation sequencing (NGS) analysis [
4,
5,
6] and evaluation of insertion/deletion polymorphisms [
8,
9,
12]. We developed a qPCR method for the quantification of serum ds-cfDNA in liver transplant patients based on the amplification of a panel of several InDels present in the donor but not in the recipient. Other research groups have analyzed InDel polymorphisms commonly found in the general population for the detection of ds-cfDNA based on digital PCR quantification [
9] or qPCR [
12]. As we propose herewith, Adamek et al. also designed primers and probes inside the insertion or deletion sequence for amplifying by qPCR only when the insertion is present [
12]. However, they included a first step for genotyping donor and recipient by a PCR followed by gel electrophoresis in order to distinguish the InDel polymorphisms by the amplicon size. We avoided the step of gel electrophoresis analysis, simplifying the procedure. On the other hand, a digital PCR approach may be very useful when the amount of ds-cfDNA after graft damage is too low and a more sensitive technique is needed [
8,
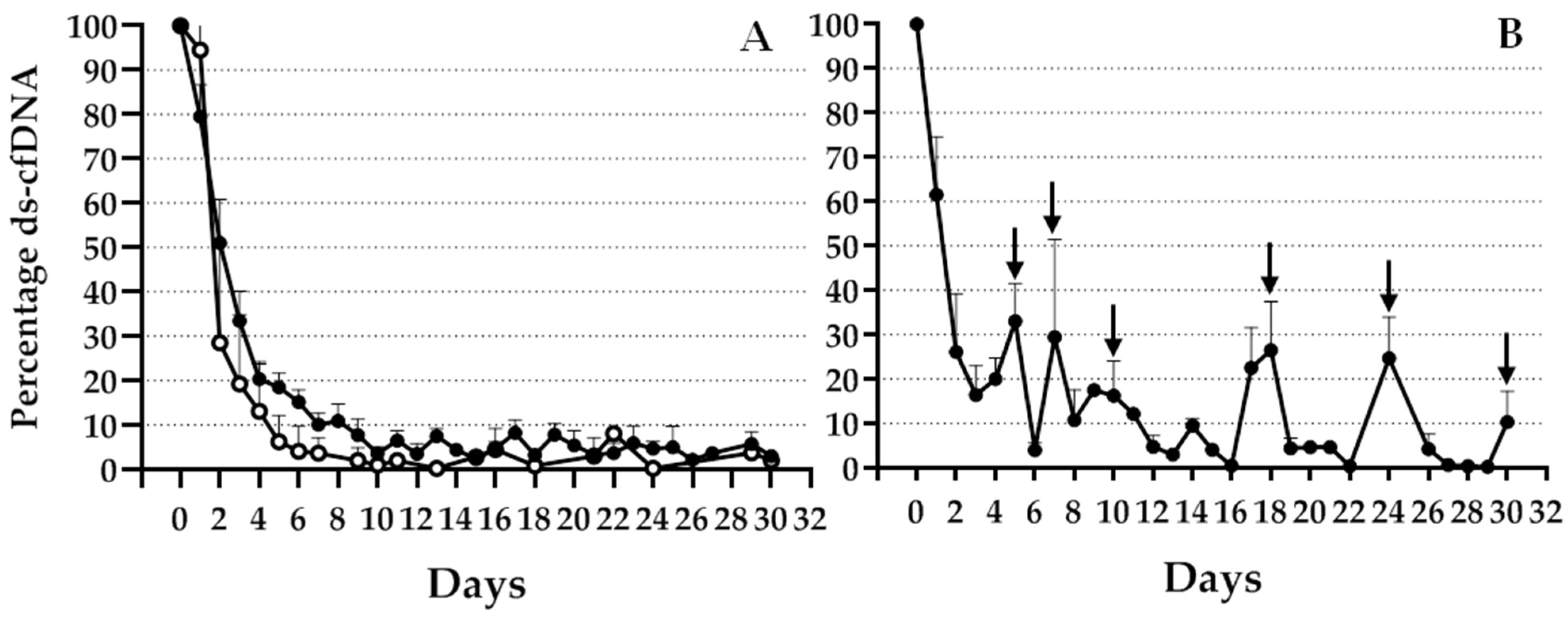
9]. However, for liver transplantation qPCR has been proven to be a suitable technique capable of detecting ds-cfDNA in the recipient cfDNA. The proposed InDel panel allowed the monitoring of most of the patients from our population after TX, although the frequency and informativeness of the different InDels were variable. To avoid the limitation of a lack of an informative donor–recipient mismatch for some patients, we considered adding further loci to the genotyping panel. Moreover, although in our area it is not common to receive a liver from a related donor, in this case, the probability of a lack of an informative locus increases. Thus, to fully extend the monitoring to the entire population, we expanded the panel to detect a higher number of polymorphic InDels. For all included InDels, we observed an early increase during the first 24 h after TX, with a different decay after this time point, depending on the clinical evolution of the patient.
A small number of patients experienced transient acute rejection during the first month after TX. As previously described, although there was an elevation of InDel percentage over basal levels at the time point of BPR, values returned to baseline after successful treatment. Since ds-cfDNA is quickly cleared from the serum, it is considered an acute specific biomarker, in contrast to the markers that gradually decrease over a long period, as seen with liver function markers such as aminotransferases enzymes. We observed significant differences in ds-cfDNA increase over basal levels, not only at the BPR time point but also several days before that. Other studies using NGS or SNP analysis [
4,
16] also observed an increase in ds-cfDNA prior to rejection diagnosis. Differences were also found when functional liver markers were analyzed (ALT, AST, gamma-GT, and bilirubin), although discrimination between the basal, Pre-R, and R groups was clearly more significant for graft cfDNA. Most patients with rejection during the first month of follow-up showed an elevation in serum liver biomarker levels over baseline levels. However, this elevation was also observed in many patients with good evolution, and in most cases, the liver marker levels did not decrease to basal levels until several months after TX. Thus, in the ANOVA comparative analysis of samples under or close to BPR and basal samples, we included the vales obtained 2 years after TX to achieve stable basal levels. In fact, only gamma-GT and bilirubin values showed significant differences between basal and BPR sample values.
Our results showed that ds-cfDNA levels discriminated BPR with an AUC of 96.5 obtained from the ROC analysis. Compared with functional liver markers, the results of ROC analysis indicated a higher diagnostic sensitivity for ds-cfDNA percentage. Maximizing both sensitivity and specificity, a threshold cutoff of 8.6% provided an estimated NPV of 99% and a PPV of 60%. The predictive values of rejection for defined thresholds have been well established in several reports regarding kidney and heart transplants. Thus, the pooled NPV described from studies in kidney and heart transplant patients ranged from 75% to 98%, and the combined PPV reported was rather low, ranging from 12% to 77% [
6,
17].
Few reports have analyzed the predictive values of liver TX [
18,
19]. The NPV values were always approximately 100%, similar to the value obtained in our study (99%; 95% CI = 93–100). We believe that the NPV values obtained, together with those previously published, are clinically promising. Thus, the high NPV suggests that this test may be helpful in avoiding unnecessary biopsies only triggered by elevated liver biomarker values.
The PPV reported for liver BPR was in the average range of 55%–62.5%. The data presented showed a PPV within this range (60%; 95% CI = 41–77). These relatively low PPV values may be explained by two facts: the relatively low prevalence of rejection for liver TX, and because the ds-cfDNA quantification is not a specific marker of rejection, it may be associated with general liver damage. In this way, the PPV obtained suggest that ds-cfDNA may be an adjuvant value that, together with other clinical evidences of rejection, provides valuable information about the liver health in order to make the decision to perform the biopsy.
Individually designed treatment is an important issue that must be addressed. In this context, individualized immunosuppressant dose evaluations using ds-cfDNA detection should be considered. Thus, monitoring ds-cfDNA may be a useful tool for patient stratification, which may allow for selection of patients with higher levels of ds-cfDNA and probably greater rejection risk, which may require more intensive therapy. In addition, during immunosuppressant drug minimization attempts [
20], ds-cfDNA levels may guide the dose reduction protocol. Moreover, ds-cfDNA quantification could help in assessing the efficacy of drug dose changes or testing a possible underdose of immunosuppressant drug due to drug toxicity or poor adherence to the treatment. Thus, another advantage of ds-cfDNA quantification is its potential to identify patients with low immunosuppression levels. It could be shown that in a subgroup of patients with tacrolimus concentrations below the therapeutic range, ds-cfDNA may be significantly increased.
The different approaches used to quantify ds-cfDNA in the recipient circulation have some limitations that might reduce their clinical effectiveness. The goal of testing is to improve graft survival, but it must have a reasonable turnaround time and must be available at a reasonable cost. The test described here involves a rapid turnaround process that may allow for the adoption of decisions and modifications in the clinical management of patients after TX. One limitation of this approach may be the low amount of ds-cfDNA that is usually diluted in the recipient cfDNA. Thus, different organs may release different amounts of cfDNA [
21]. Although we were able to quantify ds-cfDNA in liver transplant patients by qPCR [
11], a more sensitive technique such as digital PCR is needed in heart transplant patients, as reported in a previous study [
8]. In addition, the variable fragmentation of cfDNA should be considered in order to design the optimal PCR conditions for amplification [
22]. Finally, increases in ds-cfDNA may be the consequence of other liver damage in addition to rejection, including immunosuppressive drug toxicity, post-operative complications, or recurrence of primary disease. Therefore, ds-cfDNA in recipient’s serum by itself may not be a definitive diagnostic test for identification of rejection and should be considered as an adjunct rather than a definitive tool to replace biopsy.
The results from this prospective study validated and extended prior reports suggesting that ds-cfDNA measured in liver transplant patients may be a useful marker of graft integrity, helping to identify patients with acute rejection better than other conventional liver biomarkers. The negative predictive value obtained in this study confirms that ds-cfDNA quantification using qPCR may be a helpful adjuvant test to prevent rejection, avoiding unnecessary biopsies due to elevated liver biomarker values. Further research should be conducted to increase the number of patients to validate the proposed approach. Moreover, an extended panel would allow us to detect a higher number of polymorphic InDels.