An Outbreak of Clostridium (Clostridioides) difficile Infections within an Acute and Long-Term Care Wards Due to Moxifloxacin-Resistant PCR Ribotype 176 Genotyped as PCR Ribotype 027 by a Commercial Assay
Abstract
:1. Introduction
2. Material and Methods
2.1. Diagnostics of CDI
2.2. Molecular Typing of Isolates
2.3. Antimicrobial Susceptibility Testing
3. Results
3.1. Presumptive Ribotype 027 CDI Outbreak Was Identified from Stool Samples
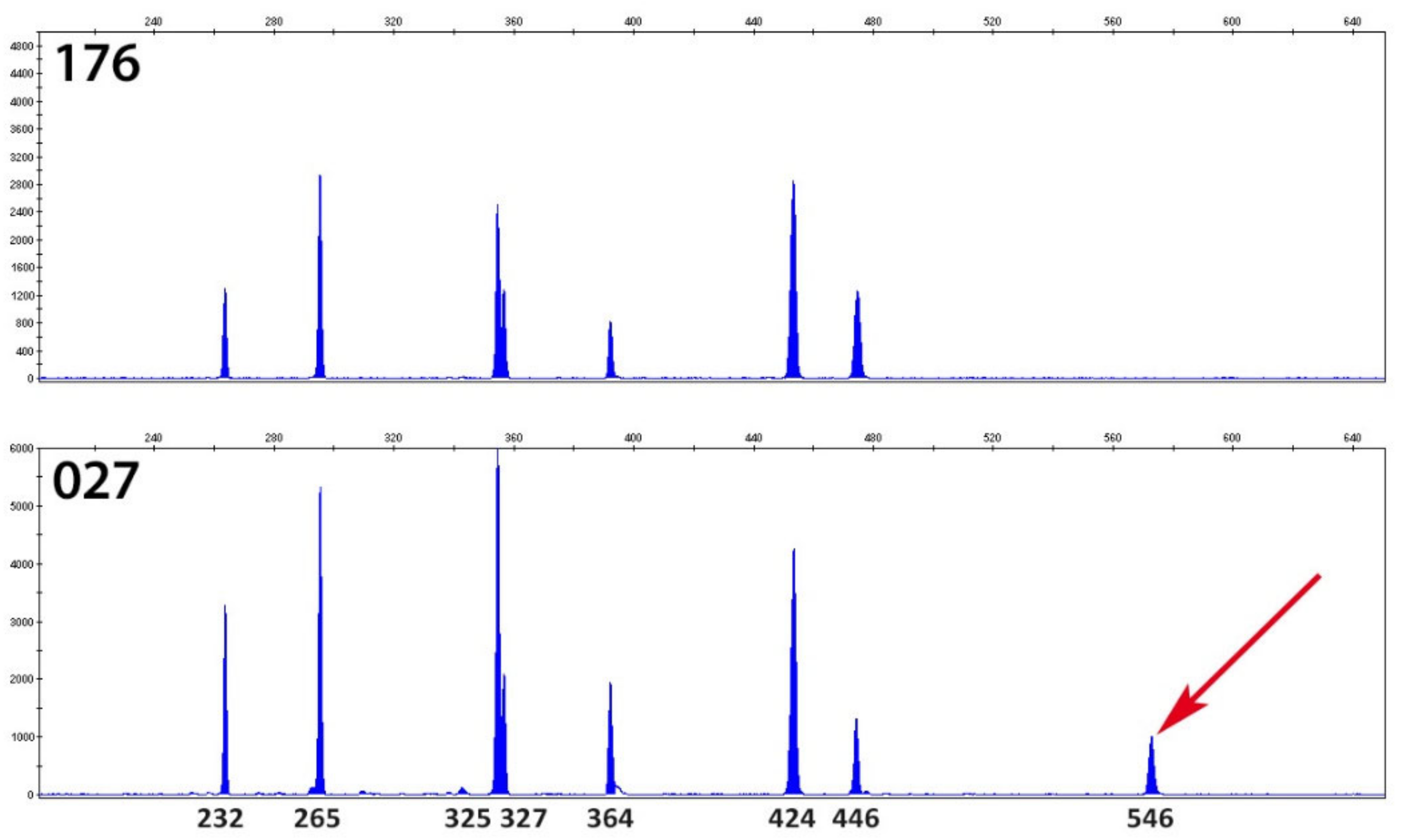
3.2. Suspected C. difficile RT027 in Stools Were Identified as C. difficile RT176 at a Strain Level
3.3. Moxifloxacin Resistance Due to Thr82Ile Was Identified in Majority of Isolates
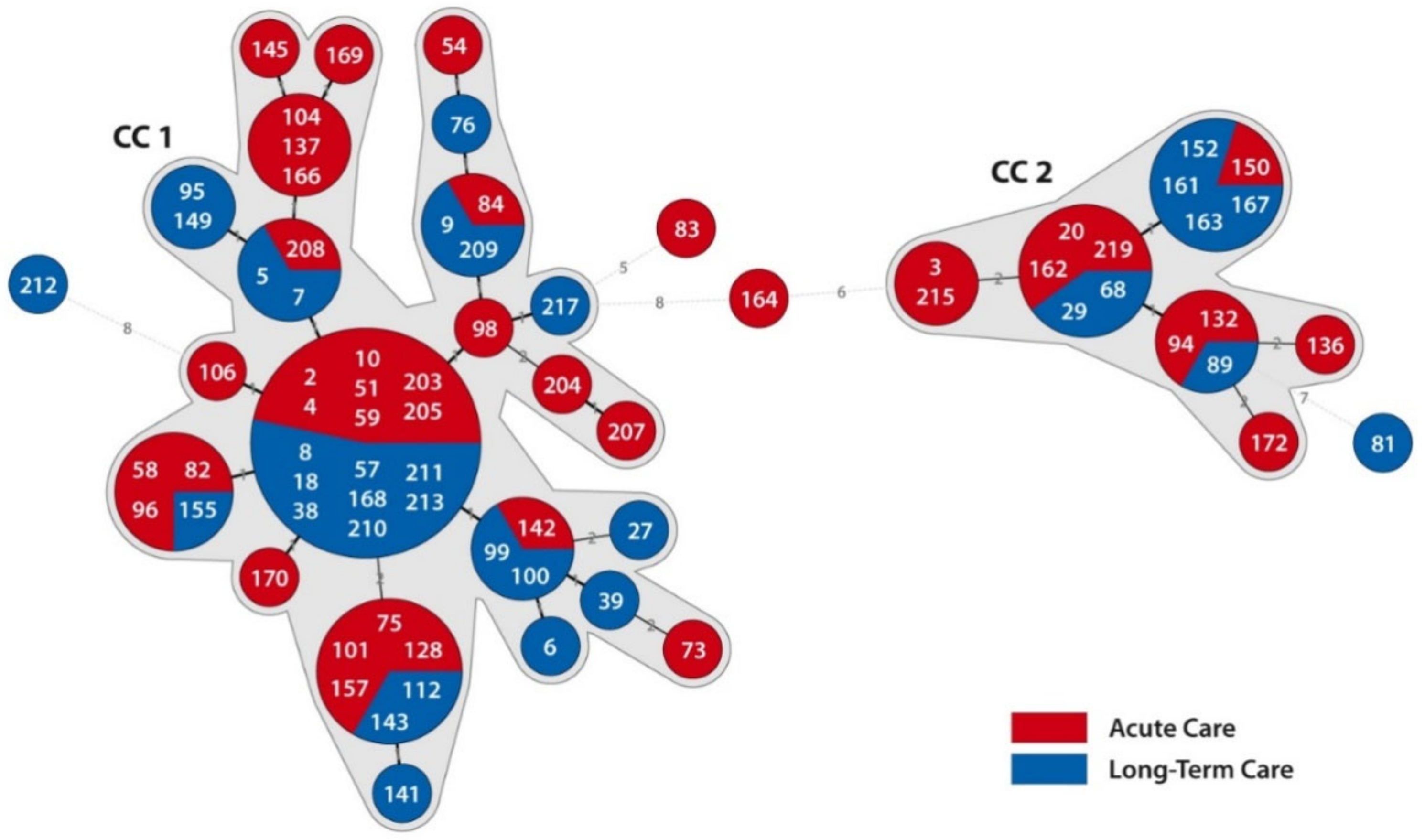
3.4. Clonal Relatedness of RT176 Isolates Were Identified from Samples within Acute and Long-Term Care Wards
3.5. RT176 Infection Is Not Associated with Fatal Outcome or Recurrence of CDI
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Lawson, P.A.; Citron, D.M.; Tyrrell, K.; Finegold, S.M. Reclassification of Clostridium difficile as Clostridioides difficile (Hall and O’Toole 1935) Prévot 1938. Anaerobe 2016, 40, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Cassini, A.; Plachouras, D.; Eckmanns, T.; Abu Sin, M.; Blank, H.-P.; Ducomble, T.; Haller, S.; Harder, T.; Klingeberg, A.; Sixtensson, M.; et al. Burden of Six Healthcare-Associated Infections on European Population Health: Estimating Incidence-Based Disability-Adjusted Life Years through a Population Prevalence-Based Modelling Study. PLoS Med. 2016, 13, e1002150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gerding, D.N.; Johnson, S.; Rupnik, M.; Aktories, K. Clostridium difficile binary toxin CDT. Gut Microbes 2013, 5, 15–27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin-Verstraete, I.; Peltier, J.; Dupuy, B. The Regulatory Networks That Control Clostridium difficile Toxin Synthesis. Toxins 2016, 8, 153. [Google Scholar] [CrossRef] [PubMed]
- Matamouros, S.; England, P.; Dupuy, B. Clostridium difficile toxin expression is inhibited by the novel regulator TcdC. Mol. Microbiol. 2007, 64, 1274–1288. [Google Scholar] [CrossRef]
- Krutova, M.; Kinross, P.; Barbut, F.; Hajdu, A.; Wilcox, M.; Kuijper, E.; Allerberger, F.; Delmée, M.; Van Broeck, J.; Vatcheva-Dobrevska, R.; et al. How to: Surveillance of Clostridium difficile infections. Clin. Microbiol. Infect. 2018, 24, 469–475. [Google Scholar] [CrossRef] [Green Version]
- He, M.; Miyajima, F.; Roberts, P.; Ellison, L.; Pickard, D.J.; Martin, M.J.; Connor, T.R.; Harris, S.R.; Fairley, D.; Bamford, K.B.; et al. Emergence and global spread of epidemic healthcare-associated Clostridium difficile. Nat. Genet. 2013, 45, 109–113. [Google Scholar] [CrossRef]
- Curry, S.R.; Marsh, J.W.; Muto, C.A.; O’Leary, M.M.; Pasculle, A.W.; Harrison, L.H. tcdC Genotypes Associated with Severe TcdC Truncation in an Epidemic Clone and Other Strains of Clostridium difficile. J. Clin. Microbiol. 2006, 45, 215–221. [Google Scholar] [CrossRef] [Green Version]
- Gateau, C.; Couturier, J.; Coia, J.; Barbut, F. How to: Diagnose infection caused by Clostridium difficile. Clin. Microbiol. Infect. 2018, 24, 463–468. [Google Scholar] [CrossRef] [Green Version]
- Fawley, W.N.; Knetsch, C.W.; MacCannell, D.R.; Harmanus, C.; Du, T.; Mulvey, M.R.; Paulick, A.; Anderson, L.; Kuijper, E.J.; Wilcox, M.H. Development and Validation of an Internationally-Standardized, High-Resolution Capillary Gel-Based Electrophoresis PCR-Ribotyping Protocol for Clostridium difficile. PLoS ONE 2015, 10, e0118150. [Google Scholar] [CrossRef] [Green Version]
- Persson, S.; And, M.T.; Olsen, K. New multiplex PCR method for the detection of Clostridium difficile toxin A (tcdA) and toxin B (tcdB) and the binary toxin (cdtA/cdtB) genes applied to a Danish strain collection. Clin. Microbiol. Infect. 2008, 14, 1057–1064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Spigaglia, P.; Mastrantonio, P. Molecular Analysis of the Pathogenicity Locus and Polymorphism in the Putative Negative Regulator of Toxin Production (TcdC) among Clostridium difficile Clinical Isolates. J. Clin. Microbiol. 2002, 40, 3470–3475. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berg, R.J.V.D.; Schaap, I.; Templeton, K.E.; Klaassen, C.H.W.; Kuijper, E.J. Typing and Subtyping of Clostridium difficile Isolates by Using Multiple-Locus Variable-Number Tandem-Repeat Analysis. J. Clin. Microbiol. 2006, 45, 1024–1028. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Griffiths, D.; Fawley, W.; Kachrimanidou, M.; Bowden, R.; Crook, D.W.; Fung, R.; Golubchik, T.; Harding, R.M.; Jeffery, K.J.M.; Jolley, K.A.; et al. Multilocus Sequence Typing of Clostridium difficile. J. Clin. Microbiol. 2010, 48, 770–778. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dridi, L.; Tankovic, J.; Burghoffer, B.; Barbut, F.; Petit, J.-C. gyrA and gyrB Mutations Are Implicated in Cross-Resistance to Ciprofloxacin and Moxifloxacin in Clostridium difficile. Antimicrob. Agents Chemother. 2002, 46, 3418–3421. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Novakova, E.; Stefkovicova, M.; Kopilec, M.G.; Novak, M.; Kotlebova, N.; Kuijper, E.; Krutova, M.; Garabasova, M.K. The emergence of Clostridium difficile ribotypes 027 and 176 with a predominance of the Clostridium difficile ribotype 001 recognized in Slovakia following the European standardized Clostridium difficile infection surveillance of 2016. Int. J. Infect. Dis. 2019, 90, 111–115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krutova, M.; Wilcox, M.; Kuijper, E. A two-step approach for the investigation of a Clostridium difficile outbreak by molecular methods. Clin. Microbiol. Infect. 2019, 25, 1300–1301. [Google Scholar] [CrossRef] [Green Version]
- Pituch, H.; Obuch-Woszczatyński, P.; Lachowicz, D.; Wultańska, D.; Karpiński, P.; Młynarczyk, G.; Van Dorp, S.M.; Kuijper, E.J. Hospital-based Clostridium difficile infection surveillance reveals high proportions of PCR ribotypes 027 and 176 in different areas of Poland, 2011 to 2013. Eurosurveillance 2015, 20, 30025. [Google Scholar] [CrossRef] [Green Version]
- Krutova, M.; Matejkova, J.; Kuijper, E.J.; Drevinek, P.; Nyc, O.; Czech Clostridium difficile study group. Clostridium difficile PCR ribotypes 001 and 176—The common denominator of C. difficile infection epidemiology in the Czech Republic, 2014. Eurosurveillance 2016, 21. [Google Scholar] [CrossRef]
- Emele, M.F.; Joppe, F.M.; Riedel, T.; Overmann, J.; Rupnik, M.; Cooper, P.; Kusumawati, R.L.; Berger, F.K.; Laukien, F.; Zimmermann, O.; et al. Proteotyping of Clostridioides difficile as Alternate Typing Method to Ribotyping Is Able to Distinguish the Ribotypes RT027 and RT176 From Other Ribotypes. Front. Microbiol. 2019, 10, 2087. [Google Scholar] [CrossRef]
- Rupnik, M.; Andrasevic, A.T.; Dokic, E.T.; Matas, I.; Jovanovic, M.; Pasic, S.; Kocuvan, A.; Janezic, S. Distribution of Clostridium difficile PCR ribotypes and high proportion of 027 and 176 in some hospitals in four South Eastern European countries. Anaerobe 2016, 42, 142–144. [Google Scholar] [CrossRef] [PubMed]
- Krutova, M.; Nyc, O.; Matejkova, J.; Kuijper, E.J.; Jalava, J.; Mentula, S. The recognition and characterisation of Finnish Clostridium difficile isolates resembling PCR-ribotype 027. J. Microbiol. Immunol. Infect. 2018, 51, 344–351. [Google Scholar] [CrossRef] [PubMed]
- Tóth, J.; Urbán, E.; Osztie, H.; Benczik, M.; Indra, A.; Nagy, E.; Allerberger, F. Distribution of PCR ribotypes among recent Clostridium difficile isolates collected in two districts of Hungary using capillary gel electrophoresis and review of changes in the circulating ribotypes over time. J. Med. Microbiol. 2016, 65, 1158–1163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Couturier, J.; Eckert, C.; Barbut, F. Spatio-temporal variability of the epidemic 027 Clostridium difficile strains in France based on MLVA typing. Anaerobe 2017, 48, 179–183. [Google Scholar] [CrossRef]
- Krůtová, M.; Matejkova, J.; Nyc, O.C. C. difficile ribotype 027 or 176? Folia Microbiol. 2014, 59, 523–526. [Google Scholar] [CrossRef]
- Lim, S.K.; Stuart, R.L.; Mackin, K.E.; Carter, G.P.; Kotsanas, D.; Francis, M.J.; Easton, M.; Dimovski, K.; Elliott, B.; Riley, T.V.; et al. Emergence of a Ribotype 244 Strain of Clostridium difficile Associated with Severe Disease and Related to the Epidemic Ribotype 027 Strain. Clin. Infect. Dis. 2014, 58, 1723–1730. [Google Scholar] [CrossRef] [Green Version]
- Skinner, A.M.; Petrella, L.; Siddiqui, F.; Sambol, S.P.; Gulvik, C.A.; Gerding, D.N.; Donskey, C.J.; Johnson, S. Unique Clindamycin-Resistant Clostridioides difficile Strain Related to Fluoroquinolone-Resistant Epidemic BI/RT027 Strain. Emerg. Infect. Dis. 2020, 26, 247–254. [Google Scholar] [CrossRef]
- Rao, K.; Micic, D.; Natarajan, M.; Winters, S.; Kiel, M.J.; Walk, S.T.; Santhosh, K.; Mogle, J.A.; Galecki, A.T.; Lebar, W.; et al. Clostridium difficile Ribotype 027: Relationship to Age, Detectability of Toxins A or B in Stool With Rapid Testing, Severe Infection, and Mortality. Clin. Infect. Dis. 2015, 61, 233–241. [Google Scholar] [CrossRef] [Green Version]
- Dingle, K.E.; Didelot, X.; Quan, T.P.; Eyre, D.W.; Stoesser, N.; Golubchik, T.; Harding, R.M.; Wilson, D.J.; Griffiths, D.; Vaughan, A.; et al. Effects of control interventions on Clostridium difficile infection in England: An observational study. Lancet Infect. Dis. 2017, 17, 411–421. [Google Scholar] [CrossRef] [Green Version]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Novakova, E.; Kotlebova, N.; Gryndlerova, A.; Novak, M.; Vladarova, M.; Wilcox, M.; Kuijper, E.; Krutova, M. An Outbreak of Clostridium (Clostridioides) difficile Infections within an Acute and Long-Term Care Wards Due to Moxifloxacin-Resistant PCR Ribotype 176 Genotyped as PCR Ribotype 027 by a Commercial Assay. J. Clin. Med. 2020, 9, 3738. https://doi.org/10.3390/jcm9113738
Novakova E, Kotlebova N, Gryndlerova A, Novak M, Vladarova M, Wilcox M, Kuijper E, Krutova M. An Outbreak of Clostridium (Clostridioides) difficile Infections within an Acute and Long-Term Care Wards Due to Moxifloxacin-Resistant PCR Ribotype 176 Genotyped as PCR Ribotype 027 by a Commercial Assay. Journal of Clinical Medicine. 2020; 9(11):3738. https://doi.org/10.3390/jcm9113738
Chicago/Turabian StyleNovakova, Elena, Nina Kotlebova, Anezka Gryndlerova, Martin Novak, Michala Vladarova, Mark Wilcox, Ed Kuijper, and Marcela Krutova. 2020. "An Outbreak of Clostridium (Clostridioides) difficile Infections within an Acute and Long-Term Care Wards Due to Moxifloxacin-Resistant PCR Ribotype 176 Genotyped as PCR Ribotype 027 by a Commercial Assay" Journal of Clinical Medicine 9, no. 11: 3738. https://doi.org/10.3390/jcm9113738