1. Introduction
In the developed world, endometrial cancer (EC) is the most common neoplasm of the female genital tract, and its incidence is increasing [
1]. Although the prognosis of EC is generally good, many cases are diagnosed at advanced stages; these cases are usually high-grade carcinomas that are ultimately more likely to recur and are associated with high mortality. In addition, a non-negligible 2.5%–3% of low-risk patients with EC have recurrent disease [
2]. Overall, the prognosis for recurrent endometrial cancer is poor, especially for the 50% of advanced ECs with extra-pelvic disease recurrence [
3]. Much effort has been devoted to generate a consensus on EC risk classification to promote consistency for future clinical trial design, including EC molecular characterization and integrating it into clinicopathologic profiling to develop prognostic and predictive biomarkers [
4].
In parallel, the oncologic management of advanced disease has been revolutionized by the advent of liquid biopsy, i.e., the analysis of tumoral material shed from primary tumors and their metastatic sites into peripheral blood, such as circulating tumor cells (CTCs) and circulating tumor DNA (ctDNA). Major advantages of liquid biopsy analysis include its minimal invasiveness and ability to provide real-time information about the disease [
5]. To date, the utility of liquid biopsy in EC has been scarcely explored [
6]. The presence of CTCs has been shown to be limited in EC, and a small number of high-risk patients, mainly those with non-endometrioid high grade carcinomas, have been identified with Epithelial Cell Adhesion Molecule (EpCAM) positive CTCs in circulation at the time of diagnosis. Those studies—such as the one conducted by the European Network for Individualized Treatment in EC (ENITEC) Consortium, which found 22% CTC positivity in 32 high-risk patients with EC [
7]—have been performed in limited cohorts. Circulating free DNA (cfDNA) and ctDNA have been described to be promising in EC management. In fact, elevated cfDNA levels have been detected in patients with EC [
8]. Furthermore, targeted sequencing of commonly mutated genes in EC—such as
CTNNB1,
KRAS,
PTEN, or
PIK3CA—has also been performed on cfDNA at the time of surgery, thus leading to the identification of at least one mutation in more than 33% of patients with endometrioid-type endometrial carcinoma (EEC) [
9]. In addition to peripheral blood—the prototypical form of liquid biopsy and the main source of clinically useful tumor material—uterine aspirates (UAs) are a minimally invasive alternative form of liquid biopsy with high relevance in gynecologic malignancies. Notably, UA can be used to identify the tumor mutational landscape and importantly, this genetic analysis also captures the high intratumor heterogeneity associated with EC [
10]. In this regard, the recently developed PapSEEK test, applied to UA sampling, has shown that 81% of patients with EC have detectable mutations [
11].
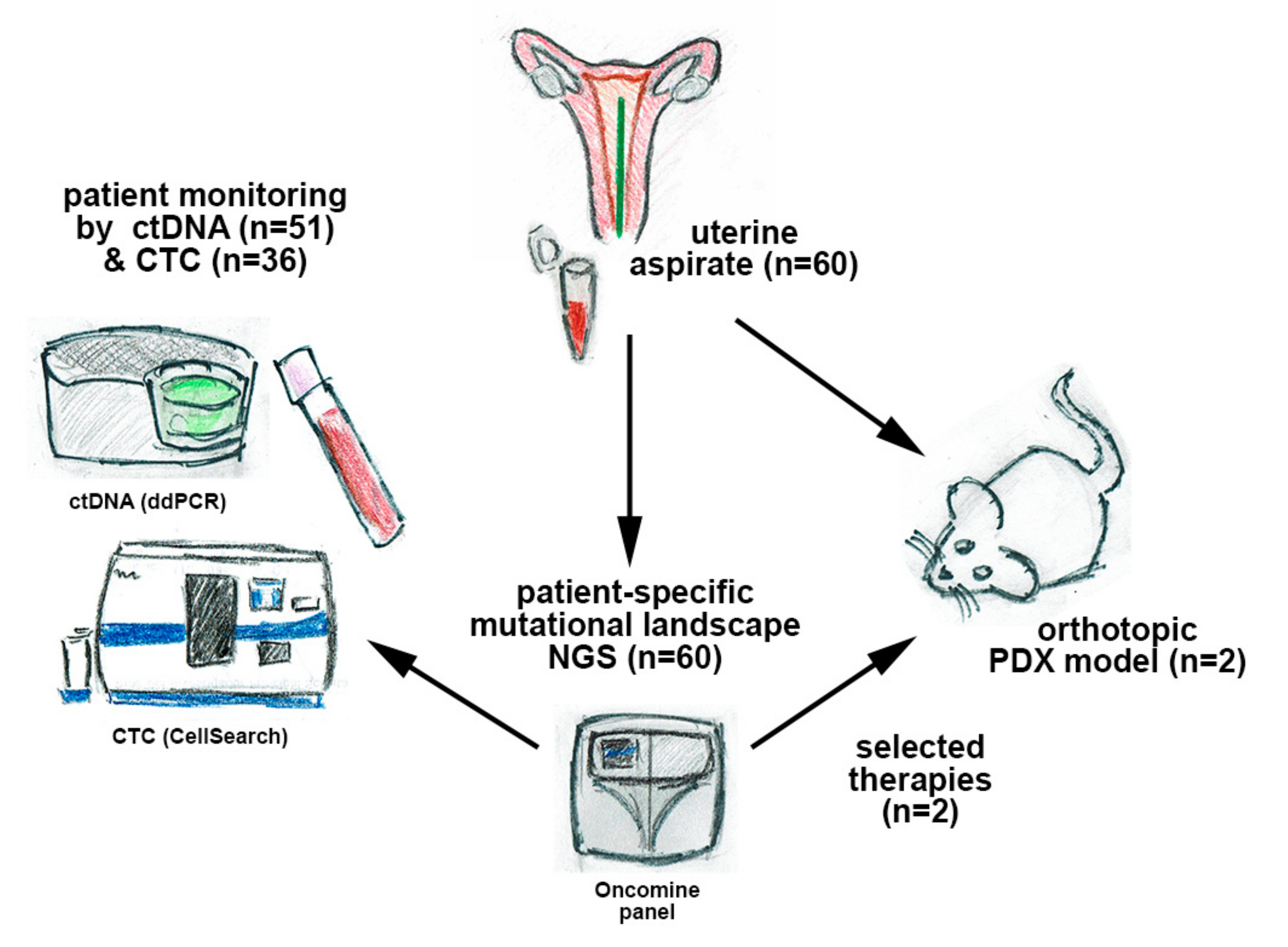
In this work, we developed an integrated strategy using different types of liquid biopsy in patients with EC to provide valuable and complementary information for the management of advanced disease. For this purpose, we performed targeted next-generation sequencing (NGS) on UA samples from a cohort of 60 patients with EC and monitored patient-specific mutations by droplet digital PCR (ddPCR) in cfDNA. In addition, we evaluated the clinical utility of cfDNA and CTCs in these patients; designed targeted therapies based on the molecular landscape; and finally, validated the therapies in patient-derived xenografts (PDXs) generated after orthotopic implantation of UA (scheme in
Figure 1).
2. Experimental Section
2.1. Patient Inclusion and Sample Collection
A total of 60 patients were recruited between January 2018 and July 2019 at the Gynecology Department of Vall d’Hebron University Hospital (Barcelona, Spain), the MD Anderson Cancer Center (Madrid, Spain), and the University Hospital of Santiago de Compostela (Santiago de Compostela, Spain). The EC cohort included low- to high-risk, grade 1–3, and stage I–IV cases, at first diagnosis or recurrence (
Table 1). The study was carried out according to the rules of the Declaration of Helsinki of 1975, revised in 2013, and according to the standards for good clinical practice and other local ethical laws and regulations. Informed consent forms, approved by the pertinent ethical committees, were signed by all patients (Autonomic Galician Ethical Committee Code 2017/430, approval number PRAMI276–2018 of the Vall d’Hebron Ethical Committee).
UAs were obtained at surgery with a Cornier cannula and kept on ice until they were processed, always within 1 hour. Each UA was then homogenized with phosphate-buffered saline (PBS) at a 1:1 ratio and centrifuged at 4 °C for 20 min at 2500 × g. The supernatant and pellet were stored at −80 °C until use. Peripheral blood samples were also collected at surgery into CellSave Preservative tubes (Silicon Biosystems Inc, Huntington Valley, USA). A two-step centrifugation was performed to isolate the plasma. First, blood samples were centrifuged for 10 min at 1600 × g at room temperature. Supernatant was collected, avoiding the buffy coat, and then centrifuged again for 15 min at room temperature and 6000 × g to remove remaining cells. Plasma supernatants were stored at −80 °C until use.
The fraction containing the mononuclear cells obtained after the first centrifugation was used for CTC isolation with the CellSearch system (Menarini, Sylicon Biosystems, Bologna, Italy). This system allows for the isolation and enumeration of EpCAM-positive CTCs. After CTC isolation with the CellSearch Epithelial Circulating Tumor Cell Kit (Menarini, Silicon Biosystems Inc), cells were labeled with phycoerythrin-conjugated anti-cytokeratin (CK) antibodies, allophycocyanin-conjugated anti-CD45 antibodies, and 4,6-diamino-2-phenylindole (DAPI) to stain the nuclei (
Figure S1). The CellTracks Analyzer (Menarini, Silicon Biosystems, Bologna, Italy) was used to acquire digital images of the three different fluorescent dyes with a 12-bit camera; the images were reviewed by trained operators to determine the CTC count.
2.2. DNA Extraction
DNA extraction from cells present in the pellet, obtained after UA processing, was performed with the MagMAX
TM Total Nucleic Acid Isolation Kit (Applied Biosystems, Foster City, California, USA), according to the manufacturer’s specifications. DNA from plasma samples was extracted with the QIAamp DNA Circulating Nucleic Acid Kit (Qiagen, Venlo, Netherlands), according to the manufacturer’s instructions. DNA samples were stored at −20 °C until use. The quantification of DNA from all samples was performed with the Qubit Fluorometer (Thermo Fisher Scientific, Waltham, MA, USA) and the Qubit DNA High-sensitivity Assay (Thermo Fisher Scientific, Waltham, MA, USA). Agilent’s TapeStation 2200 (Agilent Technologies, Santa Clara, CA, USA) was used to assess the fragment distribution of the extracted DNA (
Figure S2A).
2.3. Targeted Sequencing of UA, Personalized Therapy Selection, and ddPCR Assays
Targeted sequencing of UA was performed with the Oncomine Comprehensive Panel v3 (Thermo Fisher, Pleasanton, CA, USA), and personalized therapies identified through an in silico study using various compound databases are detailed below.
To prepare amplicon libraries, we performed targeted sequencing of uterine multiplex PCR with the Ion AmpliSeq Library Kit 2.0 and Oncomine Comprehensive Panel v3 (Thermo Fisher, Pleasanton, CA, USA). For PCR, a total of 17 and 20 cycles were performed. The PCR template preparation and enrichment were performed with the Ion PGM Template OT2 200 Kit and Ion OneTouch 2 System. Finally, the Ion PGM Sequencing 200 Kit v2 and Ion PGM System (Life Technologies, Santa Clara, CA, USA) were used for DNA sequencing, according to the manufacturer’s protocols. Duplicates were analyzed for 10% of the samples and found to yield equivalent results.
For the bioinformatics analysis, alignment to the Hg19 human reference genome and variant calling were performed with Torrent Suite™ Software v.4.2.1 (Life Technologies, Santa Clara, CA, USA). All samples were sequenced and analyzed in comparable conditions. The mean coverage per sequenced sample was approximately 1500 reads per base. Variants with a Phred quality score field value less than 100 were considered as low-quality variants. The prediction of genomic variant effects on protein function was performed with the PROVEAN Genome Variants tool (
http://provean.jcvi.org/index.php). Variants with possibly damaging or deleterious consequences, as predicted by at least one of the PROVEAN predictors, were considered to be of interest and were visually checked with Integrative Genomics Viewer (IGV) v.2.3.40, Broad Institute. Variants with a global minor allele frequency above 0.05 were considered as single nucleotide polymorphisms and were rejected (data from dbSNP,
http://www.ncbi.nlm.nih.gov/SNP/).
2.4. Personalized Therapy Selection
To identify potential effective drugs on the basis of the mutational profile obtained from the targeted sequencing analysis, we performed an in silico study using the CTD (
http://ctdbase.org/) and STITCH (
http://stitch.embl.de/) compound databases. These databases contain peer-reviewed information about the effect of particular drugs on specific genes or signaling pathways, thus indicating the relationship between chemicals and proteins in the human context. In this analysis, we included the mutational status of each identified gene per case. The identified drugs were then used to treat PDX models, and standard protocol treatment was included as a control.
2.5. Detection of ctDNA with ddPCR
For each patient, specific ddPCR assays were designed and run on a QX-200 dPCR system (Bio-Rad, California, USA) using TaqMan chemistry, with primers and probes at final concentrations of 900 nM and 250 nM, respectively. For each patient-specific assay, one non-target control, one wild-type control (fragmented genomic DNA from a healthy donor), and one positive control (patient UA DNA) were used. Custom TaqMan assays were used (Bio-Rad, CA, USA) according to the variants discovered from tumor tissue sequencing. The linearity and the lower limit of detection (LOD) for all ddPCR assays used in the study were tested with dilutions of synthetic targeted sequences in a background of 20,000 copies of wild type (WT) DNA (mutant copies ranging from 0 to 40). All assays were run in triplicates and to assure their performance, the Pearson correlation between the estimated and experimental mutant copies detected was calculated (Pearson r = 0.995) (
Figure S2B–C).
PCR was performed with the ddPCR Supermix for probes (Bio-Rad, Santa Rosa, CA, USA) and partitioned into a median of 50,000 droplets per sample (run in triplicates) in an automated droplet generator (Bio-Rad, CA, USA), according to the manufacturer’s instructions. Emulsified PCR reactions were run on 96-well plates on a C1000 TouchTM thermal cycler (Bio-Rad, CA, USA) by incubating the plates at 95 °C for 10 min followed by 40 cycles of 95 °C for 15 sec; the specific assay extension temperature for 60 sec; and 98 °C for 10 min. The temperature ramp increment was 2.5 °C/sec for all steps. Plates were read on a Bio-Rad QX-200 droplet reader with Bio-Rad’s QuantaSoft v1.7.4 software to quantify the number of droplets positive for mutant DNA, wild-type DNA, both, or neither. A minimum of 30,000 positive droplets across wells were required for a valid assay, and a minimum of five, single FAM-positive droplets with no positive events were required for the WT control.
2.6. PDX Generation and Therapy Testing
The UA was mechanically disaggregated with the VWR Pellet Mixer (VWR International, Radnor, PA, USA) and through a 30 G needle. Fetal bovine serum (Invitrogen, Carlsbad, CA, USA) was added to facilitate disaggregation. A Matrigel matrix (BD Biosciences, Franklin Lakes, NJ, USA) was added in a 1:2 ratio and kept on ice until use. Then, 50 µL of this mix was injected into the uteri of 8-week-old female SCID-beige mice (Janvier Labs; Le Genest Saint-Isle, France).
For implantation of the processed UA, mice were housed and maintained under specific-pathogen-free conditions, and procedures were performed in accordance with institutional guidelines and approved by the Use Committee for Animal Care from the Universidad de Santiago de Compostela. Aseptic procedures were followed for all surgeries. Each 8-week-old female SCID-beige mouse (Janvier Labs, Le Genest Saint-Isle, France) was anesthetized with 2% isoflurane/air (Isoflo, Esteve Farma, Carnaxide, Portugal) and kept under anesthesia for the entire procedure. The area was prepared for sterile surgery by shaving off the fur and scrubbing with a betadine solution and sterile 4×4 gauze. A midline ventral incision was made in the skin and abdominal wall. Once open, the uterus was located and 50 µL of cell suspension (UA cell suspension in the Matrigel matrix (BD Biosciences) in a 1:2 ratio) was injected with a 30 G syringe. After the Matrigel was allowed to solidify, the syringe was removed and the uterus was returned to the abdominal cavity. The wound was closed with Ethicon VICRIL (Johnson & Johnson International, Diegem, Belgium) sutures. Buprenodale (Dechra, Dales Pharmaceuticals, Keighley Road, Skipton, UK) was used for postoperative analgesia.
The orthotopic tumors that developed in the uteri were minced into small pieces and placed subcutaneously in the flanks of 8-week-old female SCID-beige mice (
n = 11). For the preclinical studies, once the tumors reached a suitable size of 150–200 mm, they were randomly assigned to one of three groups and treated with placebo (methylcellulose 0.5%; Sigma;
n = 4); CarboTaxol (carboplatin (50 mg/kg) or paclitaxel (20 mg/kg) combined therapy was administered intraperitoneally once every week for 4 weeks (
n = 3); or BYL719, (50 mg/kg; Achemblock, Burlingame, CA; on the basis of the molecular alterations found in the UAs and a molecular tumor board consensus, as described in
Section 2.4) previously dissolved in dimethylsulfoxide, was administered through oral gavage in methylcellulose 0.5% (Sigma) for 5 days on/2 days off until sacrifice (
n = 4). All mice were sacrificed after the tumor diameter reached 15 mm, the tumor volume exceeded 1200 mm
3, or the human endpoint criteria were met, according to the guidelines of Directive 2010/63/EU for the protection of animals used for scientific purposes, including assessment of appearance, body function, environment, and behavior. Tumor size was measured with calipers (Rohs) twice per week. Tumor volume was calculated according to the following formula: V = (W
2 × L)/2 where V is the tumor volume, W is the tumor width, and L is the tumor length.
2.7. Statistical Analysis
Statistical analyses were performed in IBM SPSS Statistics 20, and graphs were generated in GraphPad Prism 5.0 (GraphPad Software, Inc., San Diego, CA, USA). Two-tailed Mann–Whitney U test or Kruskal–Wallis test was used to evaluate the differential cfDNA, ctDNA, and CTC levels among clinical groups. The Pearson correlation test was performed to determine the relationship between quantitative experimental and clinical variables. Associations between clinicopathologic features and the presence of ctDNA or CTCs were examined with the chi-square test (Fisher’s exact test). A p-value < 0.05 was set as the level of statistical significance.
4. Discussion
Patients normally have good prognosis if diagnosis occurs in early stages of EC. However, some patients experience recurrence after surgery, and this recurrence is not predictable with the current risk classification systems. Clinical management of the risk of recurrence remains an unsolved issue that is probably associated with tumor heterogeneity and early tumor dissemination [
15]. Therefore, new strategies must be developed to improve risk stratification, therapy selection, and monitoring of this disease. In this sense, studies based on liquid biopsy may be essential to achieve more precise clinical management in EC. The advantages of using liquid biopsies rather than tissue samples are clear: the samples are easy to obtain, provide information in real time, and improve the understanding of tumor heterogeneity [
16]. Nevertheless, despite the many technical improvements in liquid biopsies in other tumors during the past two decades, information regarding the clinical benefit of using these alternative biopsies to manage patients with EC is quite limited [
6].
Our study performed the first combined characterization of UAs, ctDNA, and CTCs to comprehensively explore the value of liquid biopsies for personalizing EC treatment. To do so, we determined the genomic landscape of UA samples obtained at surgery in a relevant cohort of patients. The samples recapitulated the mutational patterns found in tissue samples from ECs and had the additional advantages of being minimally invasive and representative of the molecular heterogeneity of primary carcinomas. We were able to detect genetic alterations in 93% of the UA samples analyzed with targeted sequencing.
PI3KCA,
PTEN,
TP53,
CTNB1, and
KRAS were the genes most frequently mutated in our cohort, in accordance with the genomic pattern previously described in primary carcinomas [
10,
14,
17,
18]. Moreover, the genomic signatures reliably reproduced the molecular classification of the histology subtypes: serous UA was mainly characterized by
TP53 point mutations and CNVs, whereas endometrioid UA showed a broader spectrum of mutations, primarily
PI3KCA and
PTEN alterations [
17]. Importantly, 23.3% of UAs showed intratumor genetic heterogeneity (ITH), considered as the presence of more than one point mutation in the same genes, mostly in
PTEN and
PIK3CA. These findings reinforced our previous observations [
10] regarding the use of sequencing analysis of UAs to capture ITH as an alternative diagnostic biopsy to aid in the selection of more specific treatments.
In addition to providing a valuable tool to better understand the biology of each endometrial tumor, the characterization of UAs also facilitated the translation of a personalized approach to relevant clinical blood samples through the analysis of patient-specific mutations with cfDNA. The levels of cfDNA in EC have been studied [
19,
20]; higher cfDNA levels have been reported in serum samples from patients with EC than in healthy control individuals and patients with benign gynecologic disorders, although the cfDNA levels are independent of tumor stage or grade [
19]. In our study, including both endometrioid and non-endometrioid tumors, cfDNA levels at surgery were significantly higher in high-risk tumors. Therefore, cfDNA levels have potential as a prognostic factor, although long-term follow-up of patients will be required to definitively demonstrate the clinical value.
Importantly, the rate of positive cases for ctDNA at surgery was 41.2%, and this rate was higher in high-risk tumors. Even though these tumors were not characterized for their DNA release and not all mutations found in UA sequencing were followed by ddPCR, the detection rate presented in this article with our targeted approach was higher than that previously described with other technologies, such as NGS [
9]. Moreover, the rate of ctDNA-positive cases found correlated with myometrial and lymphovascular infiltration and with histology grade, in line with recent data [
9]. In addition, our results also demonstrated the feasibility of ctDNA monitoring in patients with EC to identify the presence of a more aggressive disease, as well as the utility of assessment of ctDNA in combination with cfDNA as potential risk factors for identifying patients with higher risk of recurrence. In fact, the levels of ctDNA at surgery and during follow-up were associated with the recurrence of the disease in three of the patients included in the study. As mentioned above, long-term follow-up of the patients included in the study will be necessary to confirm the clinical value of ctDNA determination in improving the risk classification of patients with EC. In this regard, Pereira et al., by analyzing a retrospective cohort of gynecologic tumors (ovary and endometrial tumors), have found lower survival rates in patients with detectable ctDNA levels at surgery [
21].
CTC enumeration showed similar, but slightly higher, rates of positivity than that reported in previous studies in EC analyzed with the CellSearch system (probably associated with the specific characteristics of the different cohorts) [
7,
22]. Notably, we found that in patients with detectable ctDNA and CTCs, the levels of both markers were correlated, and by considering the presence of CTCs and/or ctDNA, most high-risk tumors (80%) could be identified. In line with our results, previous studies have also indicated discrepancies between CTC presence and the release of ctDNA into the bloodstream, owing to the different natures of both circulating tumor markers, thus suggesting that this method provides complementary, clinically valuable information in EC [
23,
24].
Finally, our study also pioneered the use of UAs in selecting targeted therapies based on the patient-specific genomic landscape, and generating PDX models as an individualized approach to evaluate their efficacy. We were able to generate PDX models closely resembling the patient tumor characteristics and reproducing the molecular and histologic features. The preclinical study targeting the identified PI3KCA-activating alteration confirmed the activity of BYL719 (a specific PI3KCA inhibitor) and demonstrated the clinical value of our personalized pipeline based on liquid biopsies to address therapeutic alternatives.