Identification and Fine Mapping of Quantitative Trait Loci for Tiller Angle Using Chromosome Segment Substitution Lines in Rice (Oryza Sativa L.)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Evaluation of the Tiller Angle
2.3. DNA Extraction, PCR, and Gene Sequencing
2.4. Identification of Quantitative Trait Loci
2.5. RNA Isolation and Quantitative Reverse Transcription PCR
2.6. Statistical Analysis
3. Results
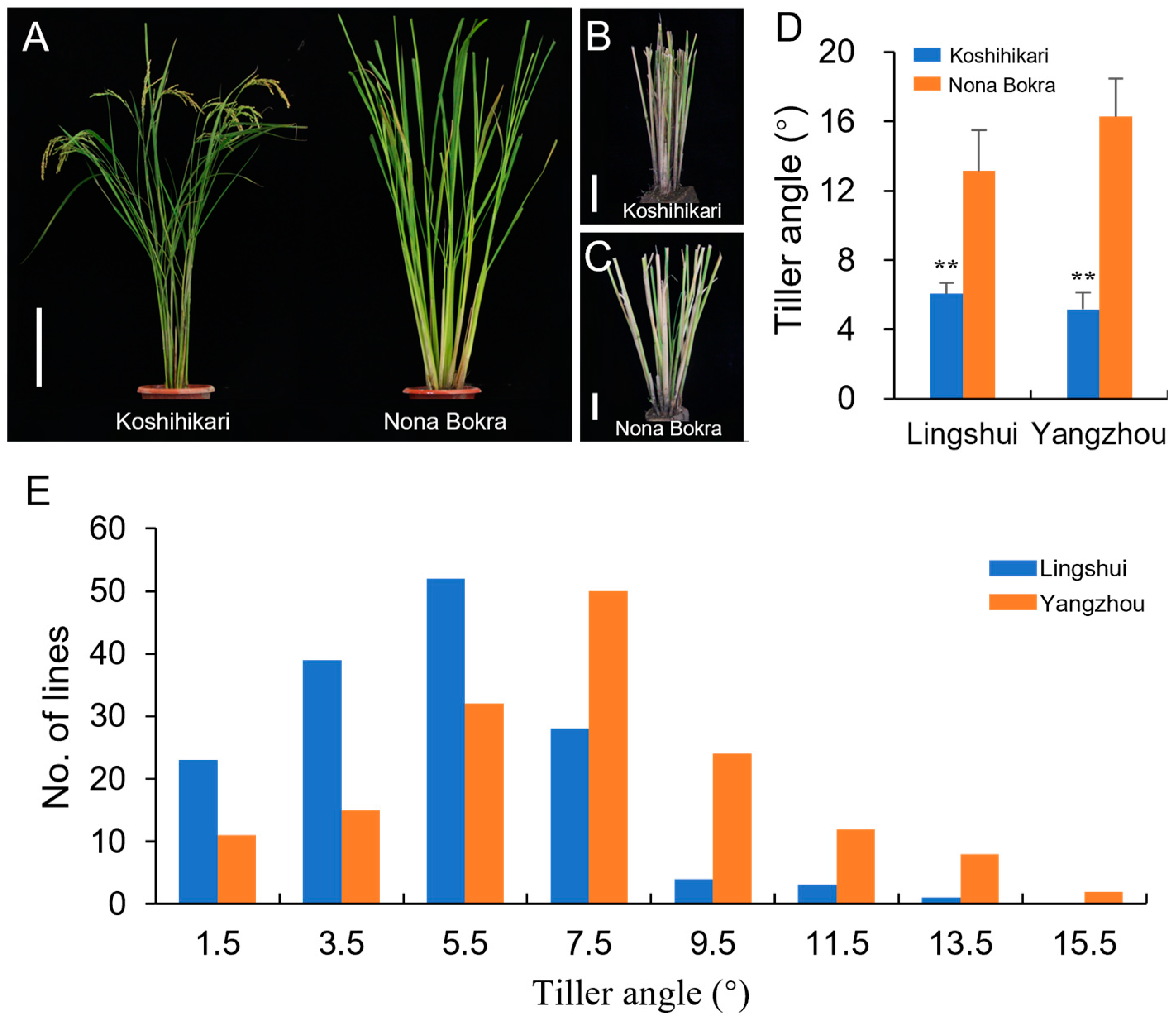
3.1. Phenotypic Variations in the Tiller Angle in the Chromosome Segment Substitution Lines
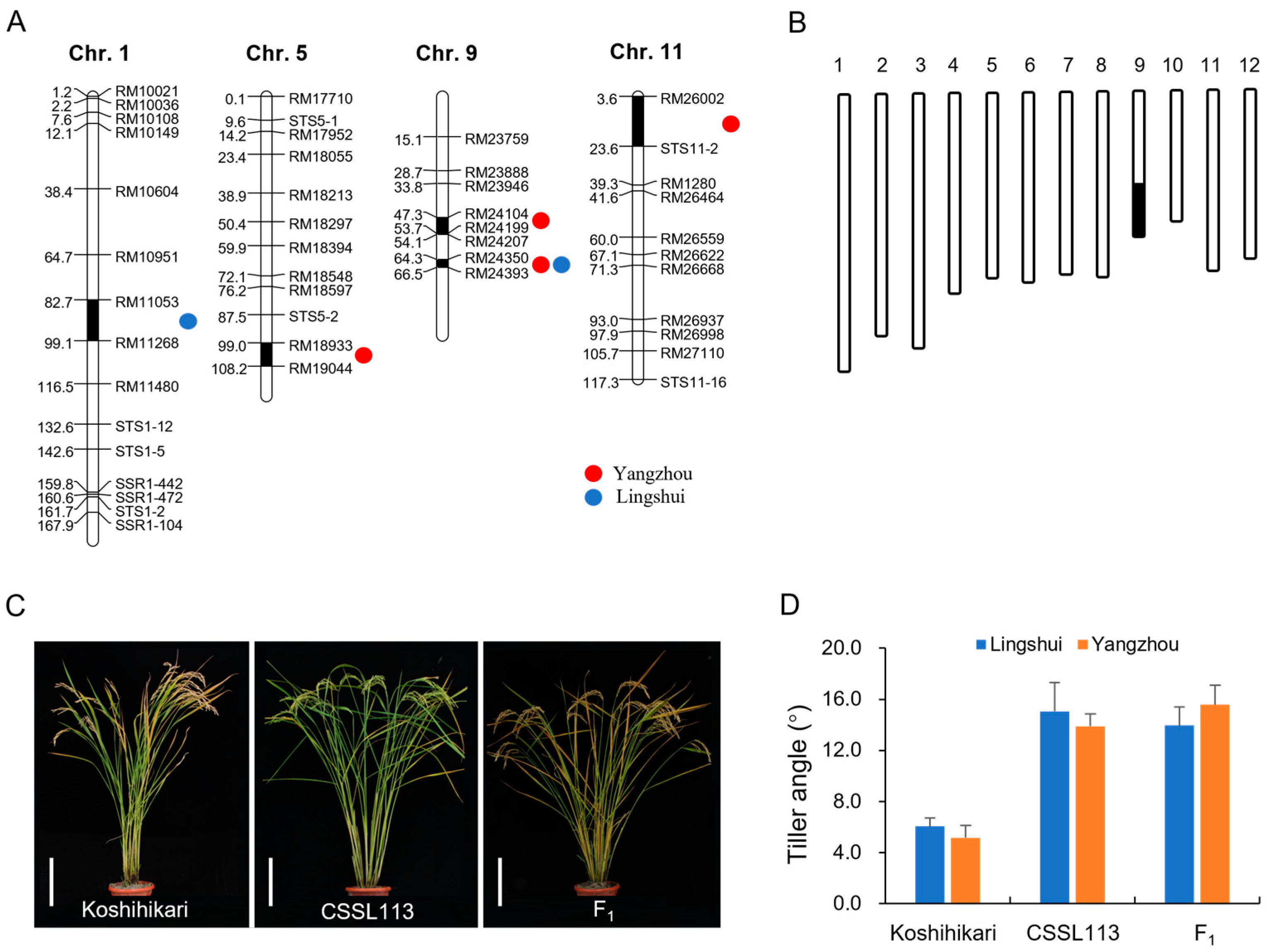
3.2. Quantitative Trait Locus Analysis for the Tiller Angle
3.3. Fine Mapping of qTA9-2 and Analysis of Candidate Genes
3.4. Potential Utilization of CSSLs to Improve Rice Architecture
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wang, Y.; Li, J. Branching in rice. Curr. Opin. Plant Biol. 2011, 14, 94–99. [Google Scholar] [CrossRef]
- Wang, Y.; Li, J. Molecular basis of plant architecture. Annu. Rev. Plant Biol. 2008, 59, 253–279. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, J. Rice, rising. Nat. Genet. 2008, 40, 1273–1275. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.B.; Susan, R.M.; Shen, Z.T. Transgressive segregation of tiller angle in rice caused by complementary gene action. Crop Sci. 1998, 38, 12–19. [Google Scholar] [CrossRef]
- Li, Z.K.; Paterson, A.H.; Pinson, S.R.M.; Stansel, J.W. RFLP facilitated analysis of tiller and leaf angles in rice (Oryza sativa L.). Euphytica 1999, 109, 79–84. [Google Scholar] [CrossRef]
- Qian, Q.; He, P.; Teng, S.; Zeng, D.L.; Zhu, L.H. QTLs analysis of tiller angle in rice (Oryza sativa L.). Acta Genet. Sin. 2001, 28, 29–32. [Google Scholar]
- Shen, S.Q.; Zhuang, J.Y.; Bao, J.S.; Zheng, K.L.; Xia, Y.W.; Shu, Q.Y. Analysis of QTLs with additive, epistasis and G×E interaction effects of the tillering angle trait in rice. J. Agric. Biotechnol. 2005, 13, 16–20. [Google Scholar]
- Yu, C.Y.; Liu, Y.Q.; Jiang, L.; Wang, C.M.; Zhai, H.Q.; Wan, J.M. QTLs mapping and genetic analysis of tiller angle in rice (Oryza sativa L.). Acta Genet. Sin. 2005, 32, 948–954. [Google Scholar]
- Bai, S.; Hong, J.; Su, S.; Li, Z.K.; Wang, W.S.; Shi, J.X.; Liang, W.; Zhang, D. Genetic basis underlying tiller angle in rice (Oryza sativa L.) by genome-wide association study. Plant Cell Rep. 2022, 41, 1707–1720. [Google Scholar] [CrossRef]
- Dong, H.; Zhao, H.; Xie, W.; Han, Z.; Li, G.; Yao, W.; Bai, X.F.; Hu, Y.; Guo, Z.L.; Lu, K.; et al. A novel tiller angle gene, TAC3, together with TAC1 and D2 largely determine the natural variation of tiller angle in rice cultivars. PLoS Genet. 2016, 12, e1006412. [Google Scholar] [CrossRef]
- Yu, B.; Lin, Z.; Li, H.; Li, X.; Li, J.; Wang, Y.; Zhang, X.; Zhu, Z.F.; Zhai, W.X.; Wang, X.K.; et al. TAC1, a major quantitative trait locus controlling tiller angle in rice. Plant J. 2007, 52, 891–898. [Google Scholar] [CrossRef] [PubMed]
- Dardick, C.; Callahan, A.; Horn, R.; Ruiz, K.B.; Zhebentyayeva, T.; Hollender, C.; Whitaker, M.; Abbott, A.; Scorza, R. PpeTAC1 promotes the horizontal growth of branches in peach trees and is a member of a functionally conserved gene family found in diverse plants species. Plant J. 2013, 75, 618–630. [Google Scholar] [CrossRef] [PubMed]
- Waite, J.M.; Dardick, C. TILLER ANGLE CONTROL 1 modulates plant architecture in response to photosynthetic signals. J. Exp. Bot. 2018, 69, 4935–4944. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Huai, Z.X.; Xiao, Y.J.; Wang, X.H.; Yu, J.Y.; Ding, G.D.; Peng, J.H. Natural variation and genetic analysis of the tiller angle gene MsTAC1 in Miscanthus sinensis. Planta 2014, 240, 161–175. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Huang, W.; Gao, J.P.; Yang, J.; Shi, M.; Zhu, M.Z.; Luo, D.; Lin, H.X. Genetic control of rice plant architecture under domestication. Nat. Genet. 2008, 40, 1365–1369. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.B.; Li, X.R.; Liu, F.X.; Sun, X.Y.; Li, C.G.; Zhu, Z.F.; Fu, Y.C.; Cai, H.W.; Wang, X.K.; Xie, D.X.; et al. Control of a key transition from prostrate to erect growth in rice domestication. Nat. Genet. 2008, 40, 1360–1364. [Google Scholar] [CrossRef] [PubMed]
- Hu, M.; Lv, S.; Wu, W.; Fu, Y.; Liu, F.; Wang, B.; Li, W.; Gu, P.; Cai, H.; Sun, C.; et al. The domestication of plant architecture in African rice. Plant J. 2018, 94, 661–669. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhao, S.; Li, X.; Zhang, B.; Jiang, L.; Tang, Y.; Zhao, J.; Ma, X.; Cai, H.; Sun, C.; et al. Deletions linked to PROG1 gene participate in plant architecture domestication in Asian and African rice. Nat. Commun. 2018, 9, 4157. [Google Scholar] [CrossRef]
- Wang, W.; Gao, H.; Liang, Y.; Li, J.; Wang, Y. Molecular basis underlying rice tiller angle: Current progress and future perspectives. Mol. Plant 2022, 15, 125–137. [Google Scholar] [CrossRef]
- Li, P.; Wang, Y.; Qian, Q.; Fu, Z.; Wang, M.; Zeng, D.; Li, B.; Wang, X.J.; Li, J.Y. LAZY1 controls rice shoot gravitropism through regulating polar auxin transport. Cell Res. 2007, 17, 402–410. [Google Scholar] [CrossRef]
- Yoshihara, T.; Iino, M. Identification of the gravitropism-related rice gene LAZY1 and elucidation of LAZY1-dependent and -independent gravity signaling pathways. Plant Cell Physiol. 2007, 48, 678–688. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Liang, Y.; Yuan, Y.; Wang, L.; Meng, X.; Xiong, G.; Zhou, J.; Cai, Y.; Han, N.; Hua, L.; et al. OsBRXL4 regulates shoot gravitropism and rice tiller angle through affecting LAZY1 nuclear localization. Mol. Plant 2019, 12, 1143–1156. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Yu, H.; Yu, H.; Cai, Y.; Huang, L.; Xu, C.; Xiong, G.; Meng, X.; Wang, J.; Chen, H.; et al. A core regulatory pathway controlling rice tiller angle mediated by the LAZY1-dependent asymmetric distribution of auxin. Plant Cell 2018, 30, 1461–1475. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Li, S.L.; Fan, X.; Song, S.; Zhou, X.; Weng, X.; Xiao, J.; Li, X.; Xiong, L.; You, A.; et al. OsHOX1 and OsHOX28 redundantly shape rice tiller angle by reducing HSFA2D expression and auxin content. Plant Physiol. 2020, 184, 1424–1437. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.Z.; Wang, W.G.; Zhang, N.; Cai, Y.Y.; Liang, Y.; Meng, X.B.; Yuan, Y.D.; Li, J.Y.; Wu, D.X.; Wang, Y.H. LAZY2 controls rice tiller angle through regulating starch biosynthesis in gravity-sensing cells. New Phytol. 2021, 231, 1073–1087. [Google Scholar] [CrossRef]
- Li, H.; Sun, H.Y.; Jiang, J.H.; Sun, X.Y.; Tan, L.B.; Sun, C.Q. TAC4 controls tiller angle by regulating the endogenous auxin content and distribution in rice. Plant Biotechnol. J. 2021, 19, 64–73. [Google Scholar] [CrossRef]
- Zhao, L.; Tan, L.B.; Zhu, Z.F.; Xiao, L.T.; Xie, D.X.; Sun, C.Q. PAY1 improves plant architecture and enhances grain yield in rice. Plant J. 2015, 83, 528–536. [Google Scholar] [CrossRef]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Ikeda, M.; Matsubara, A.; Song, X.J.; Ito, M.; Asano, K.; Matsuoka, M.; Kitano, H.; Ashikari, M. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 2010, 42, 545–549. [Google Scholar] [CrossRef]
- Wu, X.R.; Tang, D.; Li, M.; Wang, K.J.; Cheng, Z.K. Loose plant architecture1, an indeterminate domain protein involved in shoot gravitropism, regulates plant architecture in rice. Plant Physiol. 2013, 161, 317–329. [Google Scholar] [CrossRef]
- Zhang, W.F.; Tan, L.B.; Sun, H.Y.; Zhao, X.H.; Liu, F.X.; Cai, H.W.; Fu, Y.C.; Sun, X.Y.; Gu, P.; Zhu, Z.F.; et al. Natural variations at TIG1 encoding a TCP transcription factor contribute to plant architecture domestication in rice. Mol. Plant 2019, 12, 1075–1089. [Google Scholar] [CrossRef] [PubMed]
- Balakrishnan, D.; Surapaneni, M.; Mesapogu, S.; Neelamraju, S. Development and use of chromosome segment substitution lines as a genetic resource for crop improvement. Theor. Appl. Genet. 2019, 132, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Hua, H.; Luo, Z.; Zhang, Q.; Chen, M.; Gong, J.; Wei, X.; Huang, Z.; Huang, X.; Wang, Q. Whole-genome sequencing of 117 chromosome segment substitution lines for genetic analyses of complex traits in rice. Rice 2022, 15, 5. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhang, J.Y.; Naz, F.; Li, J.; Sun, S.F.; He, G.H.; Zhang, T.; Ling, Y.H.; Zhao, F.M. Identification of rice QTLs for important agronomic traits with long-kernel CSSL-Z741 and three SSSLs. Rice Sci. 2020, 27, 414–422. [Google Scholar]
- Liang, P.Y.; Wang, H.; Zhang, Q.L.; Zhou, K.; Li, M.M.; Li, R.X.; Xiang, S.Q.; Zhang, T.; Ling, Y.H.; Yang, Z.L.; et al. Identification and pyramiding of QTLs for rice grain size based on short-wide grain CSSL-Z563 and fine-mapping of qGL3–2. Rice 2021, 14, 35. [Google Scholar] [CrossRef] [PubMed]
- Bian, J.M.; Jiang, L.; Liu, L.L.; Wei, X.J.; Xiao, Y.H.; Zhang, L.J.; Zhao, Z.G.; Zhai, H.Q.; Wan, J.M. Construction of a new set of rice chromosome segment substitution lines and identification of grain weight and related traits QTLs. Breed. Sci. 2010, 60, 305–313. [Google Scholar] [CrossRef]
- Kobayashi, A.; Hori, K.; Yamamoto, T.; Yano, M. Koshihikari: A premium short-grain rice cultivar—Its expansion and breeding in Japan. Rice 2018, 11, 15. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.K.; Xu, L.N.; Yang, Q.Q.; Zhang, M.Q.; Wang, S.L.; Wang, R.A.; Tao, T.; Hong, L.M.; Guo, Q.Q.; Jia, S.W.; et al. High-resolution quantitative trait locus mapping for rice grain quality traits using genotyping by sequencing. Front. Plant Sci. 2023, 13, 1050882. [Google Scholar] [CrossRef] [PubMed]
- Hao, W.; Zhu, M.Z.; Gao, J.P.; Sun, S.Y.; Lin, H.X. Identification of quantitative trait loci for rice quality in a population of chromosome segment substitution lines. J. Integr. Plant Biol. 2009, 51, 500–512. [Google Scholar] [CrossRef]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucl. Acids Res. 1980, 8, 4321–4326. [Google Scholar] [CrossRef]
- Leng, Y.; Hong, L.; Tao, T.; Guo, Q.; Yang, Q.; Zhang, M.; Ren, X.; Jin, S.; Cai, X.; Gao, J. Mapping of QTLs for brown rice traits based on chromosome segment substitution line in rice (Oryza sativa L.). Agriculture 2023, 13, 928. [Google Scholar] [CrossRef]
- Li, H.H.; Ye, G.Y.; Wang, J.K. A modifed algorithm for the improvement of composite interval mapping. Genetics 2007, 175, 361–374. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Li, H.H.; Zhang, L.Y.; Wang, J.K. QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J. 2015, 3, 269–283. [Google Scholar]
- McCouch, S.R.; Cho, Y.G.; Yano, M.; Paul, E.; Blinstrub, M.; Morishima, H.; Kinosita, T. Report on QTL nomenclature. Rice Genet. Newsl. 1997, 14, 11–13. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Thomson, M.J.; Tai, T.H.; McClung, A.M.; Lai, X.H.; Hinga, M.E.; Lobos, K.B.; Xu, Y.; Martinez, C.P.; McCouch, S.R. Mapping quantitative trait loci for yield, yield components and morphological traits in an advanced backcross population between Oryza rufipogon and the Oryza sativa cultivar Jefferson. Theor. Appl. Genet. 2003, 107, 479–493. [Google Scholar] [CrossRef]
- Yan, J.; Zhu, J.; He, C.; Benmoussa, M.; Wu, P. Molecular marker-assisted dissection of genotype × environment interaction for plant type traits in rice (Oryza sativa L.). Crop Sci. 1999, 39, 538–544. [Google Scholar] [CrossRef]
- Li, H.; Zhou, A.; Sang, T. Genetic analysis of rice domestication syndrome with the wild annual species, Oryza nivara. New Phytol. 2006, 170, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Kennard, C.; Phillips, L.; Porter, A. Genetic dissection of seed shattering, agronomic, and color traits in American wildrice (Zizania palustris var. interior L.) with a comparative map. Theor. Appl. Genet. 2002, 105, 1075–1086. [Google Scholar] [CrossRef]
- Gao, J.; Liang, H.; Huang, J.; Qing, D.; Wu, H.; Zhou, W.; Chen, W.; Pan, Y.; Dai, G.; Gao, L.; et al. Development of the PARMS marker of the TAC1 gene and its utilization in rice plant architecture breeding. Euphytica 2021, 217, 49. [Google Scholar] [CrossRef]
- Jiang, J.; Tan, L.; Zhu, Z.; Fu, Y.; Liu, F.; Cai, H.; Sun, C. Molecular evolution of the TAC1 gene from rice (Oryza sativa L.). J. Genet. Genom. 2012, 39, 551–560. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Li, L.; Jiang, D. Understanding the regulatory mechanisms of rice tiller angle, then and now. Plant Mol. Biol. Rep. 2021, 39, 640–647. [Google Scholar] [CrossRef]

| Location | Parents (Mean ± SD) | CSSLs | ||||||
|---|---|---|---|---|---|---|---|---|
| Koshihikari | Nona Bokra | p Value | Mean ± SD | Range | Skewness | Kurtosis | p Value a | |
| Lingshui | 6.1 ± 0.64 | 13.1 ± 2.37 | <0.0001 | 6.3 ± 2.68 | 1.49~15.05 | 1.09 | 1.80 | 0.065 |
| Yangzhou | 5.2 ± 0.98 | 16.3 ± 2.2 | <0.0001 | 8.3 ± 3.05 | 1.85~18.38 | 0.41 | 0.51 | 0.333 |
| QTL | Chr. | Marker Interval | Position (cM) | LOD | Additive Effect | PVE (%) | |||
|---|---|---|---|---|---|---|---|---|---|
| Lingshui | Yangzhou | Lingshui | Yangzhou | Lingshui | Yangzhou | ||||
| qTA1 | 1 | RM11053–RM11268 | 82.7~99.1 | 2.64 | — | 0.06 | — | 4.12 | — |
| qTA5 | 5 | RM18933–RM19044 | 99.0~108.2 | — | 2.97 | — | −1.34 | — | 3.78 |
| qTA9-1 | 9 | RM24104–RM24199 | 47.3~53.7 | — | 5.52 | — | 1.93 | — | 7.12 |
| qTA9-2 | 9 | RM24350–RM24393 | 64.3~66.5 | 7.56 | 3.04 | 2.32 | 0.62 | 8.06 | 5.82 |
| qTA11 | 11 | RM26002–STS11-2 | 3.6~23.6 | — | 2.71 | — | −0.78 | — | 8.22 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leng, Y.; Tao, T.; Lu, S.; Liu, R.; Yang, Q.; Zhang, M.; Hong, L.; Guo, Q.; Ren, X.; Yang, Z.; et al. Identification and Fine Mapping of Quantitative Trait Loci for Tiller Angle Using Chromosome Segment Substitution Lines in Rice (Oryza Sativa L.). Agriculture 2024, 14, 1002. https://doi.org/10.3390/agriculture14071002
Leng Y, Tao T, Lu S, Liu R, Yang Q, Zhang M, Hong L, Guo Q, Ren X, Yang Z, et al. Identification and Fine Mapping of Quantitative Trait Loci for Tiller Angle Using Chromosome Segment Substitution Lines in Rice (Oryza Sativa L.). Agriculture. 2024; 14(7):1002. https://doi.org/10.3390/agriculture14071002
Chicago/Turabian StyleLeng, Yujia, Tao Tao, Shuai Lu, Ran Liu, Qingqing Yang, Mingqiu Zhang, Lianmin Hong, Qianqian Guo, Xinzhe Ren, Zhidi Yang, and et al. 2024. "Identification and Fine Mapping of Quantitative Trait Loci for Tiller Angle Using Chromosome Segment Substitution Lines in Rice (Oryza Sativa L.)" Agriculture 14, no. 7: 1002. https://doi.org/10.3390/agriculture14071002






