Combined Transcriptome and Metabolome Analysis of Alfalfa Responses to Aphid Infestation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Material Preparation, Alfalfa Treatment, and Sample Collection
2.2. RNA Extraction, Library Construction, and Sequencing
2.3. Functional Annotation and Differentially Expressed Gene Identification
2.4. Extraction of Metabolites
2.5. LC-MS/MS Analysis
2.6. Metabolome Data Preprocessing, Annotation, and Analysis
2.7. Comprehensive Analysis of Transcriptome and Metabolome
2.8. Statistical Analysis
3. Result
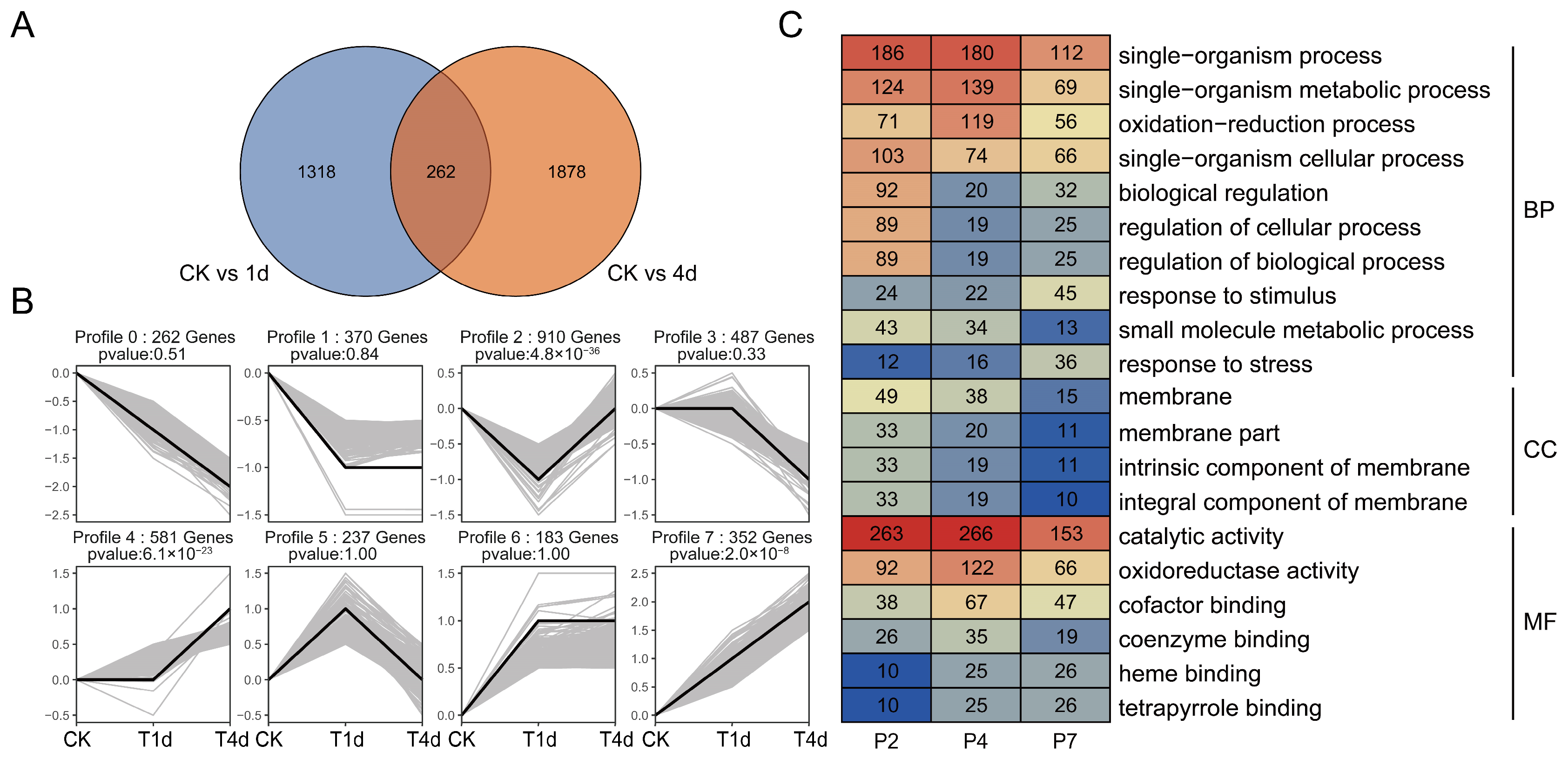
3.1. Analysis of Differentially Expressed Genes in Alfalfa Plants Infested with Aphids
3.2. Weighted Gene Co-Expression Network Analysis (WGCNA)
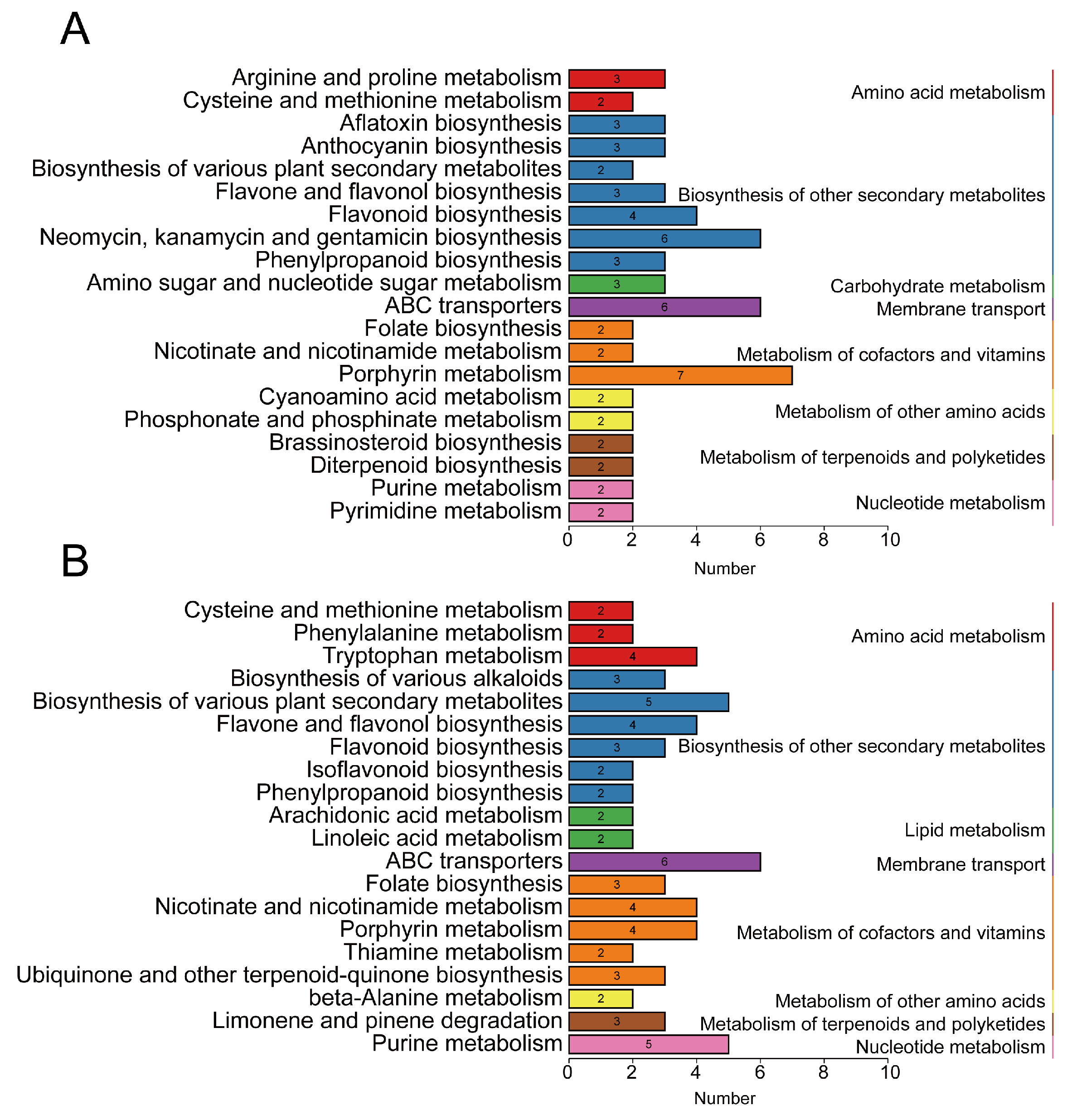
3.3. Analysis of Differentially Accumulated Metabolites in Alfalfa Infested with Aphids
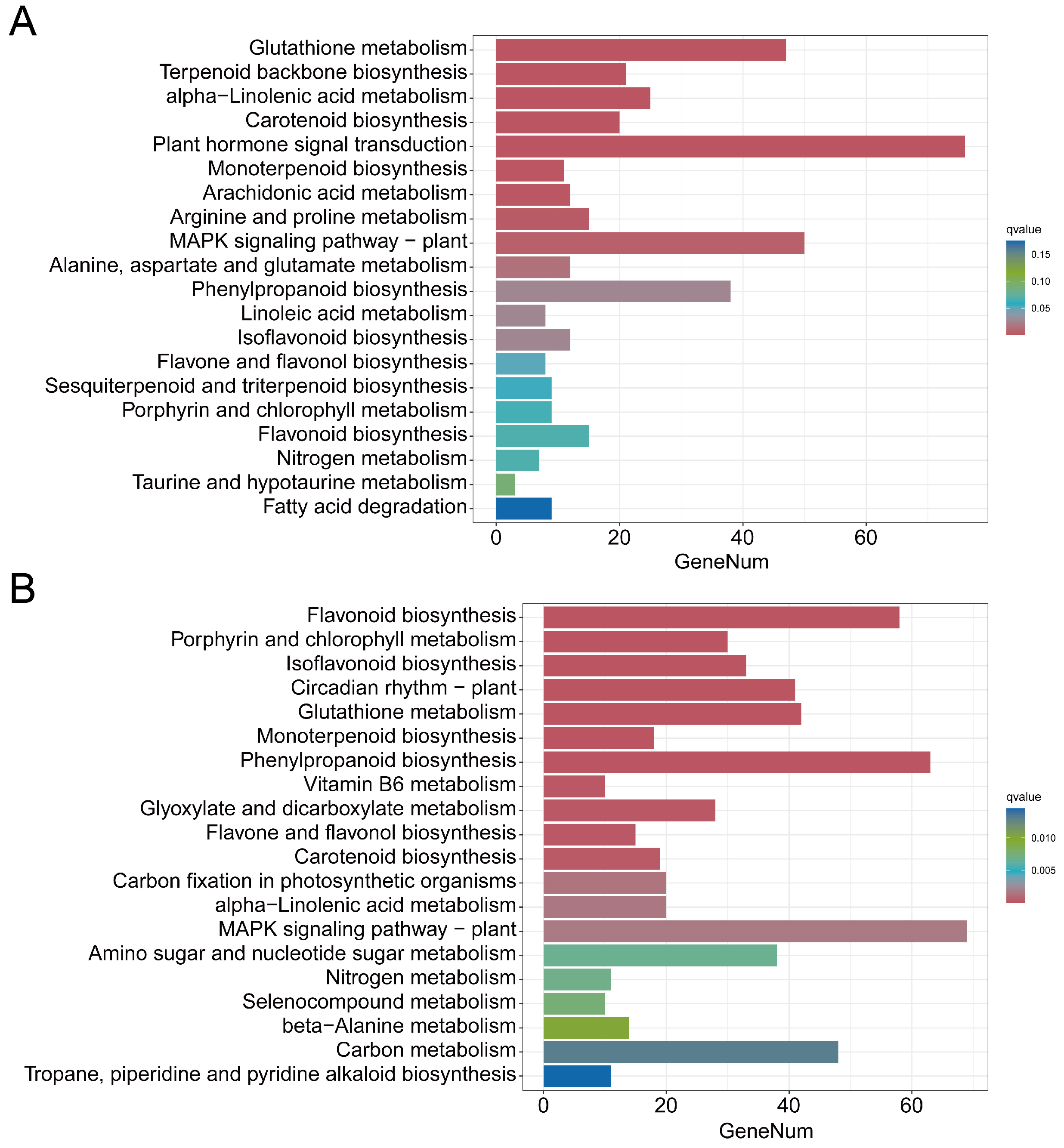
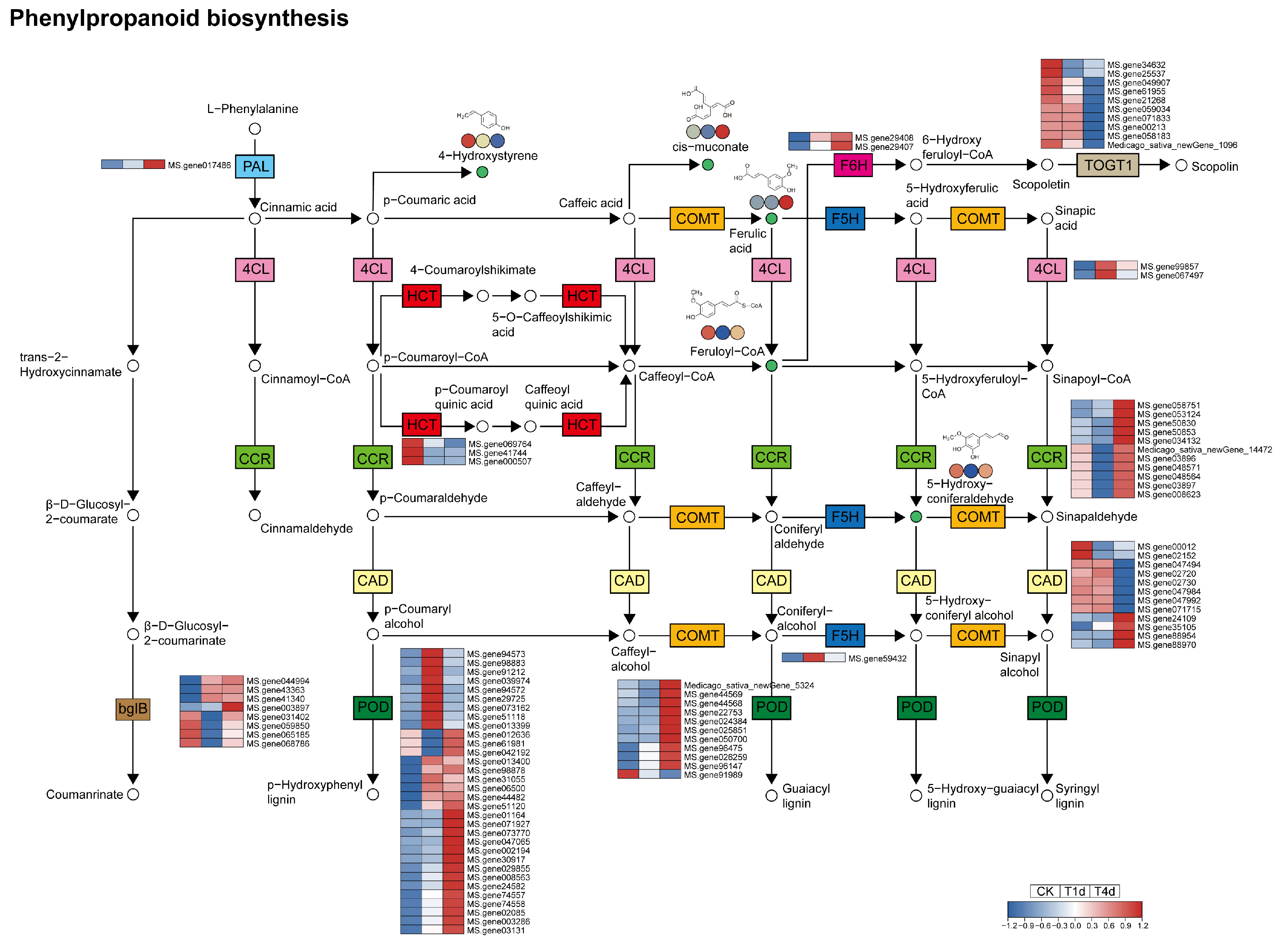
3.4. Analysis of DEGs and DAMs of the Phenylpropanoid Biosynthesis Pathway
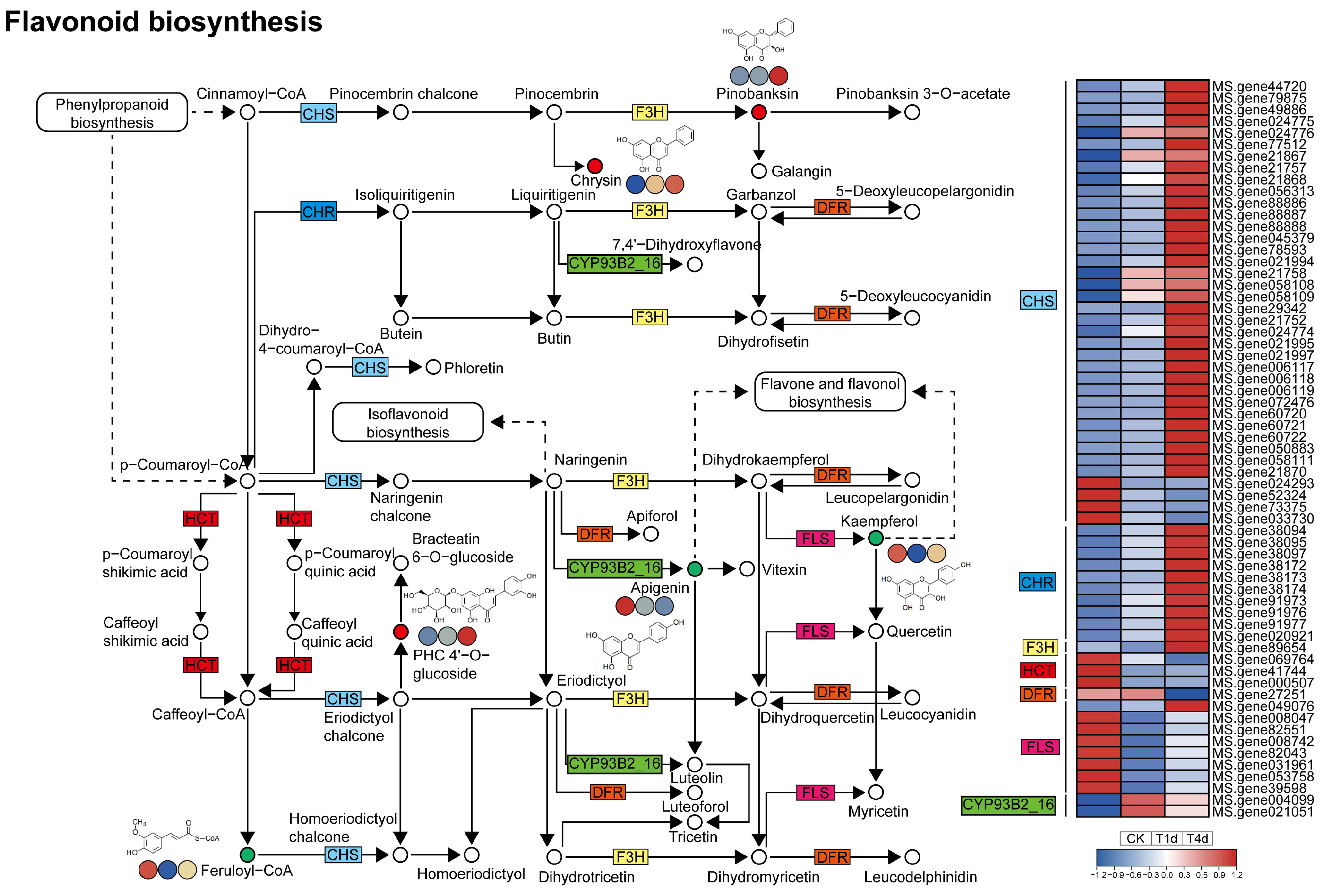
3.5. Analysis of DEGs and DAMs Involved in Flavonoid Biosynthesis
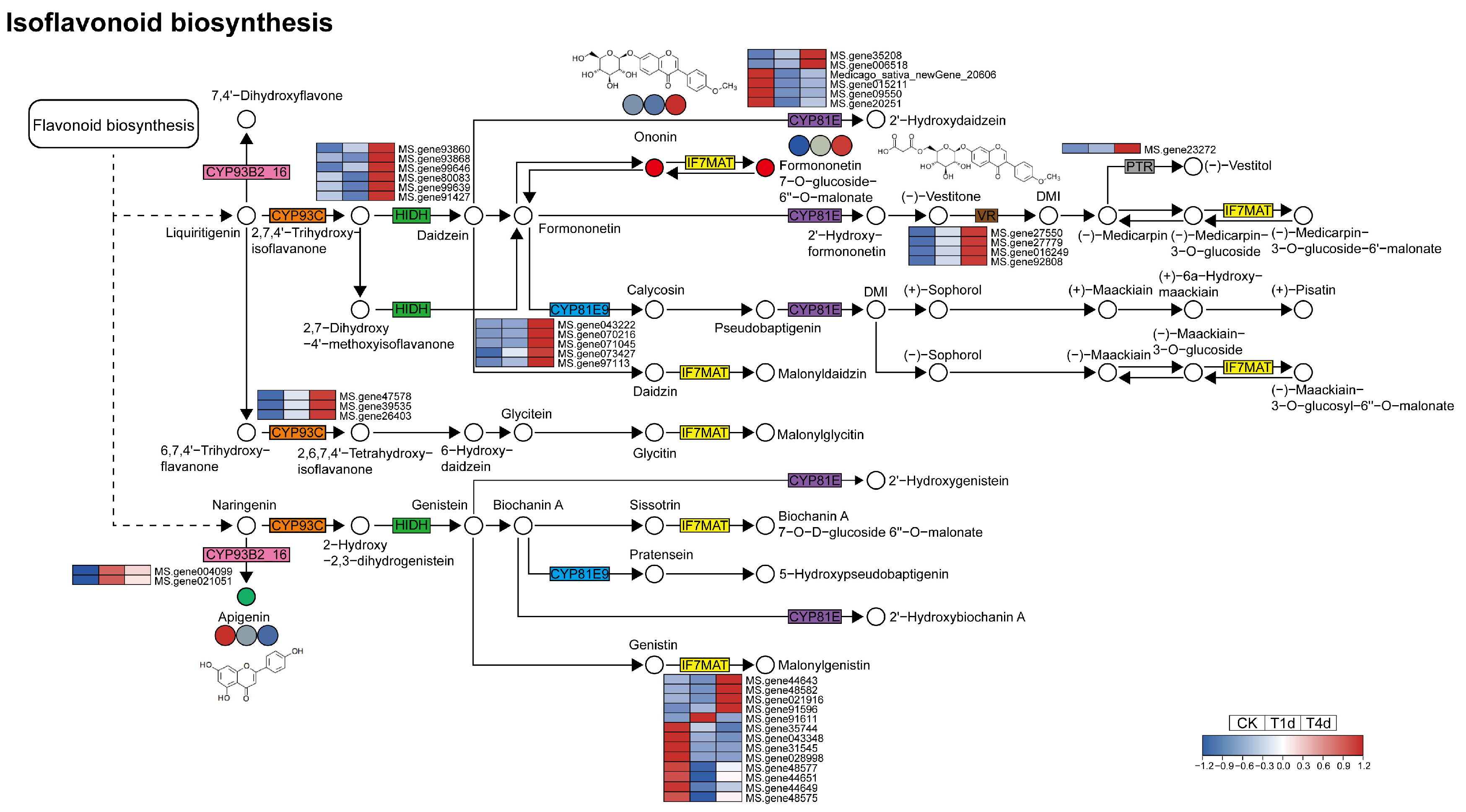
3.6. Analysis of DEGs and DAMs Involved in Isoflavonoid Biosynthesis
4. Discussion
4.1. Transcriptome Analysis
4.2. Metabolome Analysis
4.3. Phenylpropanoid Biosynthesis
4.4. Flavonoid Biosynthesis and Isoflavonoid Biosynthesis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Al-Farsi, S.M.; Nadaf, S.K.; Al-Sadi, A.M.; Ullah, A.; Farooq, M. Evaluation of indigenous Omani alfalfa landraces for morphology and forage yield under different levels of salt stress. Physiol. Mol. Biol. Plants 2020, 26, 1763–1772. [Google Scholar] [CrossRef]
- Ling, L.; An, Y.; Wang, D.; Lu, T.; Du, B.; Shu, Y.; Bai, Y.; Guo, C. Proteomic analysis reveals responsive mechanisms for saline–alkali stress in alfalfa. Plant Physiol. Biochem. 2022, 170, 146–159. [Google Scholar] [CrossRef]
- Zhang, C.; Shi, S.; Liu, Z.; Yang, F.; Yin, G. Drought tolerance in alfalfa (Medicago sativa L.) varieties is associated with enhanced antioxidative protection and declined lipid peroxidation. J. Plant Physiol. 2019, 232, 226–240. [Google Scholar] [CrossRef]
- Ma, Y.J.; Yang, G.Z.; Duan, R.J.; Li, X.A.; Zeng, S.H.; Yan, Y.J.; Zheng, C.; Hu, Y.M. Transcriptome analysis of alfalfa (Medicago sativa L.) roots reveals overwintering changes in different varieties. Czech J. Genet. Plant Breed. 2024, 60, 97–104. [Google Scholar] [CrossRef]
- De Matos, C.d.; Monteiro, L.C.P.; Gallo, S.A.D.; Costa, M.D.; da Silva, A.A. Changes in soil microbial communities modulate interactions between maize and weeds. Plant Soil 2019, 440, 249–264. [Google Scholar] [CrossRef]
- Yang, B.; Zhao, Y.; Guo, Z. Research progress and prospect of alfalfa resistance to pathogens and pests. Plants 2022, 11, 2008. [Google Scholar] [CrossRef] [PubMed]
- Dan, Z.; Xu, M.; Zhang, L.; Li, H.; Wang, Z.; Guo, Y.; Yang, G.; Li, Z.; Cong, L. First report of alfalfa root rot caused by Trichothecium roseum in China. Plant Dis. 2023, 107, 1941. [Google Scholar] [CrossRef]
- Ryalls, J.M.W.; Riegler, M.; Moore, B.D.; Johnson, S.N. Biology and trophic interactions of lucerne aphids. Agric. For. Entomol. 2013, 15, 335–350. [Google Scholar] [CrossRef]
- Züst, T.; Agrawal, A.A. Mechanisms and evolution of plant resistance to aphids. Nat. Plants 2012, 2, 15206. [Google Scholar] [CrossRef]
- Dedryver, C.A.; Le Ralec, A.; Fabre, F. The conflicting relationships between aphids and men: A review of aphid damage and control strategies. C. R. Biol. 2010, 333, 539–553. [Google Scholar] [CrossRef]
- Michaud, J.P. The ecological significance of aphid cornicles and their secretions. Annu. Rev. Entomol. 2022, 67, 65–81. [Google Scholar] [CrossRef]
- Wang, X.W.; Blanc, S. Insect transmission of plant single-stranded DNA viruses. Annu. Rev. Entomol. 2021, 66, 389–405. [Google Scholar] [CrossRef] [PubMed]
- Gallet, R.; Michalakis, Y.; Blanc, S. Vector-transmission of plant viruses and constraints imposed by virus-vector interactions. Curr. Opin. Virol. 2018, 33, 144–150. [Google Scholar] [CrossRef]
- Carr, J.P.; Tungadi, T.; Donnelly, R.; Bravo-Cazar, A.; Rhee, S.J.; Watt, L.G.; Mutuku, J.M.; Wamonje, F.O.; Murphy, A.M.; Arinaitwe, W.; et al. Modelling and manipulation of aphid-mediated spread of non-persistently transmitted viruses. Virus Res. 2020, 277, 197845. [Google Scholar] [CrossRef] [PubMed]
- Mondal, S.; Wenninger, E.J.; Hutchinson, P.J.S.; Whitworth, J.L.; Shrestha, D.; Eigenbrode, S.D.; Bosque-Pérez, N.A.; Snyder, W.E. Responses of aphid vectors of potato leaf roll virus to potato varieties. Plant Dis. 2017, 101, 1812–1818. [Google Scholar] [CrossRef] [PubMed]
- Lojek, J.S.; Orlob, G.B. Aphid transmission of tobacco mosaic virus. Science 1969, 164, 1407–1408. [Google Scholar] [CrossRef]
- Irwin, M.E.; Kampmeier, G.; Weisser, W. Aphid movement: Process and consequences. In Aphids as Crop Pests; Van Emden, H.F., Harrington, R., Eds.; CABI: Wallingford, UK, 2007; pp. 153–186. [Google Scholar]
- Marshall, S.A. Insects: Their Natural History and Diversity: With a Photographic Guide to Insects of Eastern North America; Firefly Books Inc.: Buffalo, NY, USA, 2007; p. 736. [Google Scholar]
- Bass, C.; Nauen, R. The molecular mechanisms of insecticide resistance in aphid crop pests. Insect Biochem. Mol. Biol. 2023, 156, 103937. [Google Scholar] [CrossRef] [PubMed]
- Loxdale, H.D.; Balog, A. Aphid specialism as an example of ecological-evolutionary divergence. Biol. Rev. Camb. Philos. Soc. 2018, 93, 642–657. [Google Scholar] [CrossRef]
- Howe, G.A.; Jander, G. Plant immunity to insect herbivores. Annu. Rev. Plant Biol. 2008, 59, 41–66. [Google Scholar] [CrossRef]
- Fürstenberg-Hägg, J.; Zagrobelny, M.; Bak, S. Plant defense against insect herbivores. Int. J. Mol. Sci. 2013, 14, 10242–10297. [Google Scholar] [CrossRef]
- Sidhu, J.K.; Stout, M.J.; Blouin, D.C.; Datnoff, L.E. Effect of silicon soil amendment on performance of sugarcane borer, Diatraea saccharalis (Lepidoptera: Crambidae) on rice. Bull. Entomol. Res. 2013, 103, 656–664. [Google Scholar] [CrossRef] [PubMed]
- Aljbory, Z.; Chen, M.S. Indirect plant defense against insect herbivores: A review. Insect Sci. 2018, 25, 2–23. [Google Scholar] [CrossRef]
- Plett, J.M.; Wilkins, O.; Campbell, M.M.; Ralph, S.G.; Regan, S. Endogenous overexpression of Populus MYB186 increases trichome density, improves insect pest resistance, and impacts plant growth. Plant J. 2010, 64, 419–432. [Google Scholar] [CrossRef] [PubMed]
- War, A.R.; Paulraj, M.G.; Ahmad, T.; Buhroo, A.A.; Hussain, B.; Ignacimuthu, S.; Sharma, H.C. Mechanisms of plant defense against insect herbivores. Plant Signal. Behav. 2012, 7, 1306–1320. [Google Scholar] [CrossRef]
- Lou, Y.; Baldwin, I.T. Manduca sexta recognition and resistance among allopolyploid Nicotiana host plants. Proc. Natl. Acad. Sci. USA 2003, 100, 14581–14586. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Wilkerson, C.G.; Kuchar, J.A.; Phinney, B.S.; Howe, G.A. Jasmonate-inducible plant enzymes degrade essential amino acids in the herbivore midgut. Proc. Natl. Acad. Sci. USA 2005, 102, 19237–19242. [Google Scholar] [CrossRef]
- Divekar, P.A.; Narayana, S.; Divekar, B.A.; Kumar, R.; Gadratagi, B.G.; Ray, A.; Singh, A.K.; Rani, V.; Singh, V.; Singh, A.K.; et al. Plant secondary metabolites as defense tools against herbivores for sustainable crop protection. Int. J. Mol. Sci. 2022, 23, 2690. [Google Scholar] [CrossRef]
- Dicke, M.; Baldwin, I.T. The evolutionary context for herbivore-induced plant volatiles: Beyond the ‘cry for help’. Trends Plant Sci. 2010, 15, 167–175. [Google Scholar] [CrossRef]
- Chen, L.; He, F.; Long, R.; Zhang, F.; Li, M.; Wang, Z.; Kang, J.; Yang, Q. A global alfalfa diversity panel reveals genomic selection signatures in Chinese varieties and genomic associations with root development. J. Integr. Plant Biol. 2021, 63, 1937–1951. [Google Scholar] [CrossRef]
- Sun, Q.Z.; Xu, L.J.; Tang, X.J.; Ma, J.T.; Wang, D.; Li, D.; Liu, Q.; Tao, Y.; Li, F. Investigating the origin of the Chinese name for alfalfa. IOP Conf. Ser. Earth Environ. Sci. 2017, 57, 012053. [Google Scholar] [CrossRef]
- Mei, L.; Zhang, N.; Wei, Q.; Cao, Y.; Li, D.; Cui, G. Alfalfa modified the effects of degraded black soil cultivated land on the soil microbial community. Front Plant Sci. 2022, 13, 938187. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.L.; Gao, Z.W.; Gao, Y.; Liu, G.F.; Sheng, L.X.; Wang, D.L. Eco-physiological characteristics of alfalfa seedlings in response to various mixed salt-alkaline stresses. J. Integr. Plant Biol. 2008, 50, 29–39. [Google Scholar] [CrossRef]
- Brogdon, W.G.; McAllister, J.C. Insecticide resistance and vector control. Emerg. Infect. Dis. 1998, 4, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Guo, K.; Yang, P.; Chen, J.; Lu, H.; Cui, F. Transcriptomic responses of three aphid species to chemical insecticide stress. Sci. China Life Sci. 2017, 60, 931–934. [Google Scholar] [CrossRef]
- Liu, C.; Pan, J.; Yin, Z.G.; Feng, T.; Zhao, J.; Dong, X.; Zhou, Y. Integrated transcriptome and metabolome analyses revealed regulatory mechanisms of flavonoid biosynthesis in Radix Ardisia. PeerJ 2022, 10, e13670. [Google Scholar] [CrossRef]
- Modi, A.; Vai, S.; Caramelli, D.; Lari, M. The illumina sequencing protocol and the NovaSeq 6000 system. Methods Mol. Biol. 2021, 2242, 15–42. [Google Scholar]
- Chen, H.; Zeng, Y.; Yang, Y.; Huang, L.; Tang, B.; Zhang, H.; Hao, F.; Liu, W.; Li, Y.; Liu, Y.; et al. Allele-aware chromosome-level genome assembly and efficient transgene-free genome editing for the autotetraploid cultivated alfalfa. Nat. Commun. 2020, 11, 2494. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Wang, P.; Chen, J. 2dGBH: Two-dimensional group Benjamini-Hochberg procedure for false discovery rate control in two-way multiple testing of genomic data. Bioinformatics 2024, 40, btae035. [Google Scholar] [CrossRef]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef]
- Mao, X.; Cai, T.; Olyarchuk, J.G.; Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 2005, 21, 3787–3793. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; An, M.; Hong, D.; Chang, D.; Wang, K.; Fan, H. Transcriptomic and metabolomic analyses reveal the differential regulatory mechanisms of compound material on the responses of Brassica campestris to saline and alkaline stresses. Front. Plant Sci. 2022, 13, 820540. [Google Scholar] [CrossRef]
- Wang, J.; Zhang, T.; Shen, X.; Liu, J.; Zhao, D.; Sun, Y.; Wang, L.; Liu, Y.; Gong, X.; Liu, Y.; et al. Serum metabolomics for early diagnosis of esophageal squamous cell carcinoma by UHPLC-QTOF/MS. Metabolomics 2016, 12, 116. [Google Scholar] [CrossRef]
- Yang, C.; Wu, P.; Yao, X.; Sheng, Y.; Zhang, C.; Lin, P.; Wang, K. Integrated transcriptome and metabolome analysis reveals key metabolites involved in Camellia oleifera defense against anthracnose. Int. J. Mol. Sci. 2022, 23, 536. [Google Scholar] [CrossRef]
- Zhang, Z.; Chen, Q.; Tan, Y.; Shuang, S.; Dai, R.; Jiang, X.; Temuer, B. Combined transcriptome and metabolome analysis of alfalfa response to thrips infection. Genes 2021, 12, 1967. [Google Scholar] [CrossRef] [PubMed]
- Łukasik, I.; Wołoch, A.; Sytykiewicz, H.; Sprawka, I.; Goławska, S. Changes in the content of thiol compounds and the activity of glutathione s-transferase in maize seedlings in response to a rose-grass aphid infestation. PLoS ONE 2019, 14, e0221160. [Google Scholar] [CrossRef]
- Dampc, J.; Kula-Maximenko, M.; Molon, M.; Durak, R. Enzymatic defense response of apple aphid aphis pomi to increased temperature. Insects 2020, 11, 436. [Google Scholar] [CrossRef]
- Gao, P.; Zhang, H.; Yan, H.; Zhou, N.; Yan, B.; Fan, Y.; Tang, K.; Qiu, X. Transcriptomic and metabolomic changes triggered by Macrosiphum rosivorum in rose (Rosa longicuspis). BMC Genom. 2021, 22, 885. [Google Scholar] [CrossRef]
- Sytykiewicz, H. Expression patterns of genes involved in ascorbate-glutathione cycle in aphid-infested maize (Zea mays L.) Seedlings. Int. J. Mol. Sci. 2016, 17, 268. [Google Scholar] [CrossRef]
- Verma, V.; Ravindran, P.; Kumar, P.P. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016, 16, 86. [Google Scholar] [CrossRef]
- Bari, R.; Jones, J.D. Role of plant hormones in plant defence responses. Plant Mol. Biol. 2009, 69, 473–488. [Google Scholar] [CrossRef] [PubMed]
- Cavaiuolo, M.; Cocetta, G.; Ferrante, A. The antioxidants changes in ornamental flowers during development and senescence. Antioxidants 2013, 2, 132–155. [Google Scholar] [CrossRef]
- Iwashina, T. Flavonoid function and activity to plants and other organisms. Biol. Sci. Space 2003, 17, 24–44. [Google Scholar] [CrossRef]
- Danquah, A.; de Zelicourt, A.; Colcombet, J.; Hirt, H. The role of ABA and MAPK signaling pathways in plant abiotic stress responses. Biotechnol. Adv. 2014, 32, 40–52. [Google Scholar] [CrossRef] [PubMed]
- Hettenhausen, C.; Schuman, M.C.; Wu, J. MAPK signaling: A key element in plant defense response to insects. Insect Sci. 2015, 22, 157–164. [Google Scholar] [CrossRef]
- Rodriguez, M.C.; Petersen, M.; Mundy, J. Mitogen-activated protein kinase signaling in plants. Annu. Rev. Plant Biol. 2010, 61, 621–649. [Google Scholar]
- Zhang, B.; Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 2005, 4, 17. [Google Scholar] [CrossRef] [PubMed]
- El-Sharkawy, I.; Liang, D.; Xu, K. Transcriptome analysis of an apple (Malus × domestica) yellow fruit somatic mutation identifies a gene network module highly associated with anthocyanin and epigenetic regulation. J. Exp. Bot. 2015, 66, 7359–7376. [Google Scholar] [CrossRef]
- Wang, S.; Li, Y.; He, L.; Yang, J.; Fernie, A.R.; Luo, J. Natural variance at the interface of plant primary and specialized metabolism. Curr. Opin. Plant Biol. 2022, 67, 102201. [Google Scholar] [CrossRef]
- Fernie, A.R.; Pichersky, E. Focus issue on metabolism: Metabolites, metabolites everywhere. Plant Physiol. 2015, 169, 1421–1423. [Google Scholar] [CrossRef]
- Zhou, S.; Lou, Y.R.; Tzin, V.; Jander, G. Alteration of plant primary metabolism in response to insect herbivory. Plant Physiol. 2015, 169, 1488–1498. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, T.C. ‘Trophic’ and ‘source’ amino acids in trophic estimation: A likely metabolic explanation. Oecologia 2017, 184, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Tegeder, M. Transporters for amino acids in plant cells: Some functions and many unknowns. Curr. Opin. Plant Biol. 2012, 15, 315–321. [Google Scholar] [CrossRef]
- Erb, M.; Kliebenstein, D.J. Plant secondary metabolites as defenses, regulators, and primary metabolites: The blurred functional trichotomy. Plant Physiol. 2020, 184, 39–52. [Google Scholar] [CrossRef]
- Jamwal, K.; Bhattacharya, S.; Puri, S. Plant growth regulator mediated consequences of secondary metabolites in medicinal plants. J. Appl. Res. Med. Aromat. Plants 2018, 9, 26–38. [Google Scholar] [CrossRef]
- Al-Khayri, J.M.; Rashmi, R.; Toppo, V.; Chole, P.B.; Banadka, A.; Sudheer, W.N.; Nagella, P.; Shehata, W.F.; Al-Mssallem, M.Q.; Alessa, F.M.; et al. Plant secondary metabolites: The weapons for biotic stress management. Metabolites 2023, 13, 716. [Google Scholar] [CrossRef]
- Li, X.; Ouyang, X.; Zhang, Z.; He, L.; Wang, Y.; Li, Y.; Zhao, J.; Chen, Z.; Wang, C.; Ding, L.; et al. Over-expression of the red plant gene R1 enhances anthocyanin production and resistance to bollworm and spider mite in cotton. Mol. Genet. Genom. 2019, 294, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Sasaki, N.; Ohmiya, A. Biosynthesis of plant pigments: Anthocyanins, betalains and carotenoids. Plant J. 2008, 54, 733–749. [Google Scholar] [CrossRef]
- Deng, Z.; Zhang, Y.; Fang, L.; Zhang, M.; Wang, L.; Ni, X.; Li, X. Identification of the flavone-inducible counter-defense genes and their cis-elements in Helicoverpa armigera. Toxins 2023, 15, 365. [Google Scholar] [CrossRef]
- Lane, G.A.; Sutherland, O.R.; Skipp, R.A. Isoflavonoids as insect feeding deterrents and antifungal components from root of Lupinus angustifolius. J. Chem. Ecol. 1987, 13, 771–783. [Google Scholar] [CrossRef]
- Vogt, T. Phenylpropanoid biosynthesis. Mol. Plant 2010, 3, 2–20. [Google Scholar] [CrossRef]
- La Camera, S.; Gouzerh, G.; Dhondt, S.; Hoffmann, L.; Fritig, B.; Legrand, M.; Heitz, T. Metabolic reprogramming in plant innate immunity: The contributions of phenylpropanoid and oxylipin pathways. Immunol. Rev. 2004, 198, 267–284. [Google Scholar] [CrossRef] [PubMed]
- Chaman, M.E.; Copaja, S.V.; Argandoña, V.H. Relationships between salicylic acid content, phenylalanine ammonia-lyase (PAL) activity, and resistance of barley to aphid infestation. J. Agric. Food Chem. 2003, 51, 2227–2231. [Google Scholar] [CrossRef] [PubMed]
- Lavhale, S.G.; Kalunke, R.M.; Giri, A.P. Structural, functional and evolutionary diversity of 4-coumarate-CoA ligase in plants. Planta 2018, 248, 1063–1078. [Google Scholar] [CrossRef]
- Lee, D.; Douglas, C.J. Two divergent members of a tobacco 4-coumarate: Coenzyme A ligase (4CL) gene family. cDNA structure, gene inheritance and expression, and properties of recombinant proteins. Plant Physiol. 1996, 112, 193–205. [Google Scholar] [CrossRef]
- Ehlting, J.; Büttner, D.; Wang, Q.; Douglas, C.J.; Somssich, I.E.; Kombrink, E. Three 4-coumarate: Coenzyme A ligases in Arabidopsis thaliana represent two evolutionarily divergent classes in angiosperms. Plant J. 1999, 19, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Humphreys, J.M.; Chapple, C. Rewriting the lignin roadmap. Curr. Opin. Plant Biol. 2002, 5, 224–229. [Google Scholar] [CrossRef]
- Wang, M.; Wang, Y.; Li, X.; Zhang, Y.; Chen, X.; Liu, J.; Qiua, Y.; Wang, A. Integration of metabolomics and transcriptomics reveals the regulation mechanism of the phenylpropanoid biosynthesis pathway in insect resistance traits in Solanum habrochaites. Hortic. Res. 2024, 11, 277. [Google Scholar] [CrossRef]
- Kaur, S.; Tiwar, V.; Kumari, A.; Chaudhary, E.; Sharma, A.; Ali, U.; Garg, M. Protective and defensive role of anthocyanins under plant abiotic and biotic stresses: An emerging application in sustainable agriculture. J. Biotechnol. 2023, 361, 12–29. [Google Scholar] [CrossRef]
- Zhuang, W.B.; Li, Y.H.; Shu, X.C.; Pu, Y.T.; Wang, X.J.; Wang, T.; Wang, Z. The classification, molecular structure and biological biosynthesis of flavonoids, and their roles in biotic and abiotic stresses. Molecules 2023, 28, 3599. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Arif, Y.; Bajguz, A.; Hayat, S. The role of quercetin in plants. Plant Physiol. Biochem. 2021, 166, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Lewinsohn, E.; Britsch, L.; Mazu, Y.; Gressel, J. Flavanone glycoside biosynthesis in citrus: Chalcone synthase, UDP-Glucose: Flavanone-7-o-glucosyl-transferase and -rhamnosyl-transferase activities in cell-free extracts. Plant Physiol. 1989, 91, 1323–1328. [Google Scholar] [CrossRef] [PubMed]
- Richard, S.; Lapointe, G.; Rutledge, R.G.; Séguin, A. Induction of chalcone synthase expression in white spruce by wounding and jasmonate. Plant Cell Physiol. 2000, 41, 982–987. [Google Scholar] [CrossRef] [PubMed]
- Fliegmann, J.; Furtwängler, K.; Malterer, G.; Cantarello, C.; Schüler, G.; Ebel, J.; Mithöfer, A. Flavone synthase II (CYP93B16) from soybean (Glycine max L.). Phytochemistry 2010, 71, 508–514. [Google Scholar] [CrossRef]
- Schuler, M.A. The role of cytochrome P450 monooxygenases in plant-insect interactions. Plant Physiol. 1996, 112, 1411–1419. [Google Scholar] [CrossRef]
- Gu, L.; Chen, P.; Yu, S. The cytochrome P450 gene GhCYP94C1 is involved in drought stress in upland cotton (Gossypium hirsutum L.). Czech J. Genet. Plant Breed. 2023, 59, 189–195. [Google Scholar] [CrossRef]
- Diharce, J.; Bignon, E.; Fiorucci, S.; Antonczak, S. Exploring dihydroflavonol-4-reductase reactivity and selectivity by QM/MM-MD simulations. Chembiochem 2022, 23, e202100553. [Google Scholar] [CrossRef]
- Schilbert, H.M.; Schöne, M.; Baier, T.; Busche, M.; Viehöver, P.; Weisshaar, B.; Holtgräwe, D. Characterization of the brassica napus flavonol synthase gene family reveals bifunctional flavonol synthases. Front. Plant Sci. 2021, 12, 733762. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, H.; Xu, M.; Guo, Y.; Dan, Z.; Liu, X.; Zhang, J.; Li, C.; Jia, S.; Jia, L.; Yu, A.; et al. Combined Transcriptome and Metabolome Analysis of Alfalfa Responses to Aphid Infestation. Agriculture 2024, 14, 1545. https://doi.org/10.3390/agriculture14091545
Liu H, Xu M, Guo Y, Dan Z, Liu X, Zhang J, Li C, Jia S, Jia L, Yu A, et al. Combined Transcriptome and Metabolome Analysis of Alfalfa Responses to Aphid Infestation. Agriculture. 2024; 14(9):1545. https://doi.org/10.3390/agriculture14091545
Chicago/Turabian StyleLiu, Hao, Ming Xu, Yuhan Guo, Zhencuo Dan, Xin Liu, Jiayi Zhang, Cong Li, Shizhen Jia, Lei Jia, Ailing Yu, and et al. 2024. "Combined Transcriptome and Metabolome Analysis of Alfalfa Responses to Aphid Infestation" Agriculture 14, no. 9: 1545. https://doi.org/10.3390/agriculture14091545




