Abstract
The wildfire-like spread of COVID-19, caused by severe acute respiratory syndrome-associated coronavirus-2, has resulted in a pandemic that has put unprecedented stress on the world’s healthcare systems and caused varying severities of socio-economic damage. As there are no specific treatments to combat the virus, current approaches to overcome the crisis have mainly revolved around vaccination efforts, preventing human-to-human transmission through enforcement of lockdowns and repurposing of drugs. To efficiently facilitate the measures implemented by governments, rapid and accurate diagnosis of the disease is vital. Reverse-transcription polymerase chain reaction and computed tomography have been the standard procedures to diagnose and evaluate COVID-19. However, disadvantages, including the necessity of specialized equipment and trained personnel, the high financial cost of operation and the emergence of false negatives, have hindered their application in high-demand and resource-limited sites. Nanoparticle-based methods of diagnosis have been previously reported to provide precise results within short periods of time. Such methods have been studied in previous outbreaks of coronaviruses, including severe acute respiratory syndrome-associated coronavirus and middle east respiratory syndrome coronavirus. Given the need for rapid diagnostic techniques, this review discusses nanoparticle use in detecting the aforementioned coronaviruses and the recent severe acute respiratory syndrome-associated coronavirus-2 to highlight approaches that could potentially be used during the COVID-19 pandemic.
Keywords:
nanoparticles; coronaviruses; diagnostic techniques; COVID-19; SARS-CoV-2; MERS-CoV; SARS-CoV 1. Introduction
1.1. Global Prevalence
In December 2019, a new strain of coronavirus, called severe acute respiratory syndrome-associated coronavirus-2 (SARS-CoV-2), was identified in Wuhan, China [1]. The virus has led to the rapid outbreak of an infectious disease known as coronavirus disease 2019 (COVID-19). As of 2 August 2022, the disease has infected more than 580 million people and led to at least 6.4 million deaths globally [2].
A COVID-19 pandemic was declared by the WHO on March 11, 2020, causing governments and health agencies to take drastic measures in hopes of halting the spread of the virus, including restricting travel and nationwide lockdowns [3]. The current pandemic is considered a crisis, in which the world economy is expected to undergo severe negative and long-lived consequences [4].
1.2. Importance of Diagnostics in the COVID-19 Pandemic
With no specific treatments against COVID-19, the primary strategies adopted by governments have revolved around preventing the spread of the disease rather than relying on therapeutics or repurposed drugs. Such strategies include the enforcement of lockdowns (periodic shutdowns of most public amenities for 21–28 days), home quarantine and self-isolation (up to 14 days upon positive COVID-19 PCR result), social distancing, restricting social gatherings, developing low-cost detection methods with high specificity and selectivity (immunoassays and ‘test-at-home’ kits) and vaccination campaigns. Awareness campaigns on the use of personal protective equipment (PPE) and practicing vigilant personal hygiene have also been implemented [5,6].
The efficacy of such measures as home quarantine and self-isolation in slowing the spread of COVID-19 depends heavily on the rapid and accurate diagnosis of the disease. With healthcare systems worldwide being under unprecedented stress, the importance of efficient diagnostic techniques and preventive measures cannot be understated.
The current standard methods of diagnosis rely on reverse-transcription polymerase chain reaction (RT-PCR) and computed tomography (CT) [7,8]. RT-PCR entails the collection of viral RNA to detect the pathogen’s presence, while CT involves the stitching of several chest x-ray images to generate a 3-D image [9]. While accurate and effective, several disadvantages accompany RT-PCR, including the high cost of operation, lengthy sample processing, false positive or negative results and laborious handling [10,11]. On the other hand, besides exposing patients to radiation, standalone CT scanning is insufficient to provide a diagnosis as there are difficulties in precisely identifying a causative agent of pneumonia [11]. As such, CT has primarily been used as a confirmatory method for potential false-negative RT-PCR results [12,13].
With the continuous emergence of evolving variants of SARS-CoV-2, the need for economical, reliable and adaptive diagnostic methods is ever-increasing. The virus’s variants have introduced challenges in efficiently diagnosing patients, as the strain’s mutations result in increased rates of false-negatives and heterogeneity of clinical presentations [14]. Through consistent follow-up of the various SARS-CoV-2 sub-lineages, characteristic differences have been reported to influence transmission rates and disease severity [15]. For instance, the Omicron variant was seen to spread more easily amongst the general population while simultaneously having higher risk of re-infection [14]. Similarly, the Delta variant is also a significantly more contagious form of the virus [16]. Indeed, the effect of viral recombination was seen to create highly pathogenic strains with improved environmental survival fitness, leading to the reoccurring waves of COVID-19 infections [15]. Hence, adequate containment of the virus’s new variants depends heavily upon advancements of accurate, rapid, yet affordable means of diagnosis, especially in developing countries [16].
To this end, a quick, sensitive, specific and cost-effective diagnostic test for SARS-CoV-2 is urgently required. An option that would fit these criteria would be nanoparticles (NPs). The literature has documented that nanoparticles have rapidly been utilized to detect microbes, including coronaviruses. Similar to the repurposing of drugs to manage COVID-19, previous NP-based diagnostic approaches for coronaviruses have the potential to be adapted for use during the current pandemic.
With that said, this review will outline the relevant information for the detection of SARS-CoV-2 (the virus’s structure and pathogenesis) and delineate the major types of coronaviruses. Principally, this review aims to showcase and discuss NP-based endeavors for detecting coronaviruses, specifically those that caused past outbreaks like severe acute respiratory syndrome-associated coronavirus (SARS-CoV) and middle east respiratory syndrome coronavirus (MERS-CoV). Furthermore, recent developments in detecting SARS-CoV-2 using NPs will be explored.
1.3. Nanoparticles
Nanoparticles are defined as a group of small elemental units, collectively behaving as a singular unit, giving rise to novel properties not demonstrated or seen in the bulk material [17]. The size of a nanoparticle typically ranges between 1–100 nm [18]. At the scale of a nanometer, the unique arrangement of the atomic structure ultimately results in the portrayal of new chemical, physical and biological properties.
Overall, the unique properties of nanoparticles are largely due to their large surface areas and ability to self-assemble. The immensely small particle size and the increased proportion of surface atoms contribute to a significant increase in the surface area of active sites, leading to increased reactivity [17]. In addition, the process of self-assembly, where components are organized to produce an ordered pattern, is attributed to the individual properties of nanoparticles, including shape, charge and polarizability. Such processes would define the repulsive and attractive forces between individual nanoparticles, determining their unique sensitivity to external stimuli [17]. Furthermore, nanoparticles have been seen to contribute to lower toxicity rates while retaining greater efficacy, making them a promising tool for preventing, treating and diagnosing viral infections [19,20].
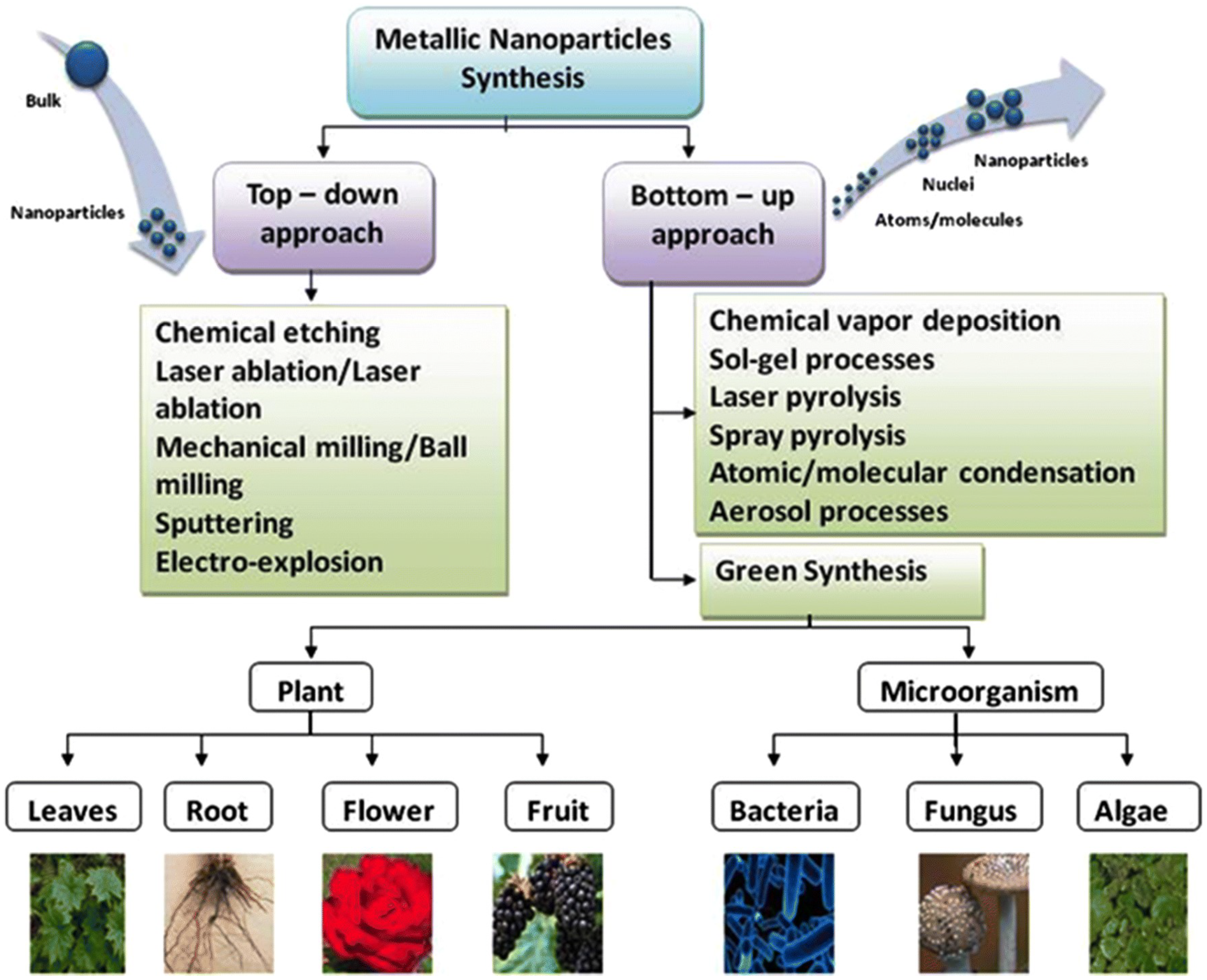
Many protocols have been developed to synthesize NPs using physical, chemical, or biological procedures, involving either top-down or bottom-up approaches [21] (illustrated in Figure 1). The top-down technique relies on breaking down material into nano-sized components through physicochemical means [21], such as thermolysis [22], nanolithography [23], laser ablation [24], irradiation [25,26] and sputter deposition [27]. The bottom-up technique combines and assembles atoms, molecules, or clusters to form nanoparticles using green and wet chemical synthesis methods.
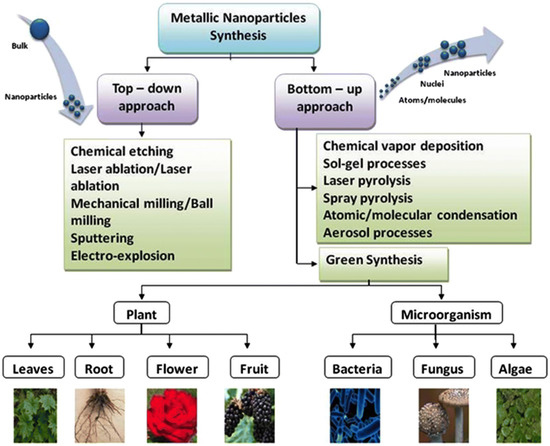
Figure 1.
Illustration of top-down and bottom-up nanoparticle synthesis Methods. Reproduced from [28] under the Creative Commons Attribution 4.0 International License.
However, the aforementioned physicochemical methods of NP synthesis present several disadvantages, including requirements for high amounts of temperature and pressure, production of toxic waste and the huge costs needed to initiate such processes [21]. On the other hand, biological methods use micro-organisms and plant-based compounds to produce intracellular or extracellular NPs [21]. They offer unique advantages such as being non-toxic, eco-friendly and considerably cheaper than their physicochemical counterpart [29]. Furthermore, green NPs boast rapid synthesis, high stability and increased drug conjugation potential while avoiding the need for high pressures or temperatures [30,31]. To this end, the green synthesis method has proved to be a desirable and reliable approach to generate NPs.
NPs of various compositions, such as metallic NPs, carbon-based materials NPs and quantum dots, have been widely used to diagnose viruses. More specifically, metallic NPs such as gold, silver and titanium have been seen to aid in diagnosing Ebola, HIV, influenzas and herpes simplex virus due to their encapsulation properties and optical characteristics [32,33]. Given their properties and the immense customizability of their physiological and physicochemical features, NPs are a promising tool that could be used to detect and diagnose coronaviruses [34]. Mechanisms of detection of coronaviruses using NPs vary greatly depending on the type of NP used, the desired molecular target and the apparatus that the NP is a part of. Potential mechanisms of action may include: (1) isolation of coronavirus RNA using superparamagnetic NPs; (2) aggregation-based columetric changes facilitated by gold NPs, detected through a biosensor or naked eye; (3) detection of electrochemical changes, facilitated by carbon nanostructures or gold NPs [35]. Further detail of detection mechanisms will be described in the following sections pertaining to nanoparticle application in the corresponding sections on the discussed coronaviruses.
3. Structure and Pathogenesis of SARS-CoV-2
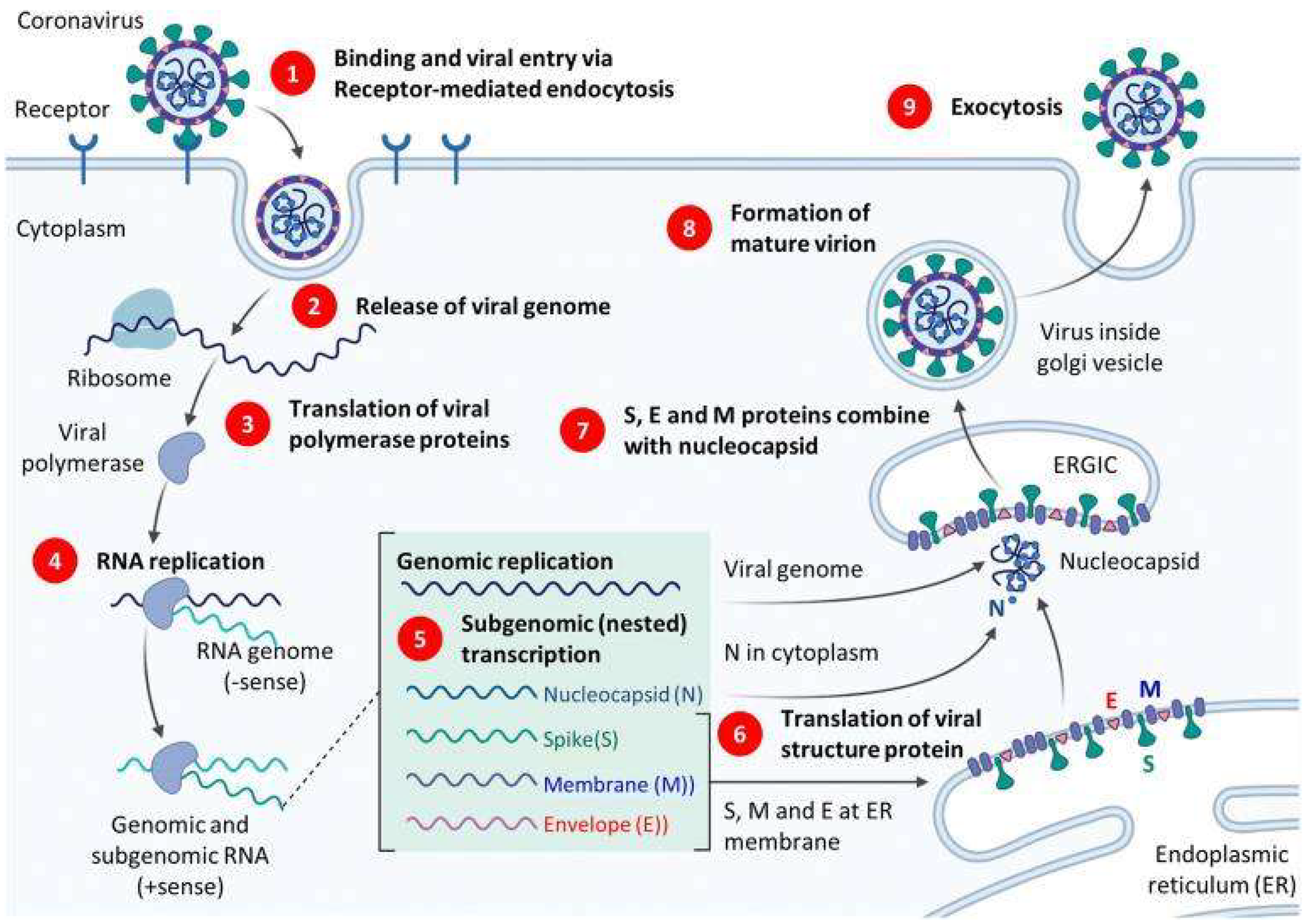
Our virus of interest, SARS-CoV-2, is a single-stranded RNA coronavirus made of approximately 26–32 kilobases and is covered with a helical-like nucleocapsid that protects its genome [36]. The main components that preserve its structural integrity include the spike (S) glycoprotein as well as the membrane (M), E (envelope) and the N (nucleocapsid) proteins [36]. The N and E proteins are expressed by the N and E RNA [42]; they are the nucleic acid targets of detection in RT-PCR [9]. An illustration of the replication cycle of SARS-CoV-2 and relevant RT-PCR targets can be seen in Figure 2.
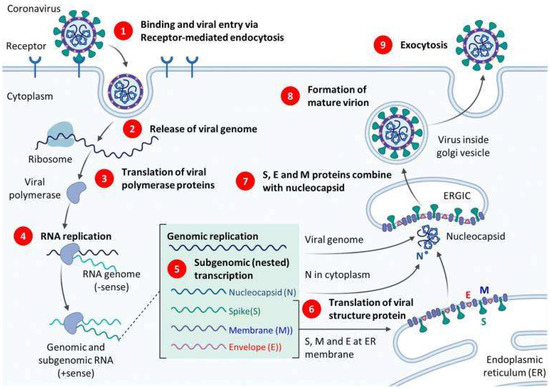
Figure 2.
Illustration of SARS-CoV-2 replication cycle. Reproduced from [43] under the Creative Commons Attribution 4.0 International License.
It is noteworthy to mention that SARS-CoV-2′s genome is approximately 82% identical to the original 2002 SARS-CoV and the 2012 MERS-CoV, with >90% similarity in sequences coding for essential enzymes and structural proteins [44]. Therefore, it is essential to examine the previous diagnostic approaches for both viruses and attempt to translate them to a SARS-CoV-2 diagnostic approach.
The Spike S protein is the most extensively studied antigen. This protein binds to the angiotensin-converting enzyme 2 (ACE2) receptor, a receptor generally expressed in epithelial linings, such as respiratory and gastrointestinal tracts and organs such as the heart, liver and kidneys [39]. The S protein has two subunits, S1 and S2. Within the S1 subunit is a domain named the receptor-binding domain (RBD), responsible for binding to the ACE-2 binding domain [45]. The S2 Subunit is essential for membrane fusion [45]. The viral RNA then translocates and integrates within the host cell’s DNA, utilizing its replication machinery to transcribe the viral mRNA. The mRNA is then translated to proteins that become2 assembled to new replicated SARS-CoV-2 and subsequently released [45]. One of the important genes involved in replication and transcribing of the viral mRNA is the RNA-dependent RNA polymerase (RdRp) gene. This gene, along with the N and E genes, is also used to detect SARS-CoV-2 [9].
ACE-2 receptors are highly expressed in the epithelium of lungs, including bronchial and bronchial branches’ epithelia [46]. As a result, the symptoms of SARS-CoV-2 invasion are clinically presented as pneumonia and acute respiratory distress syndrome [47]. ACE-2 receptors are also expressed in the kidneys, heart and liver [48]. Therefore, SARS-CoV-2 infection can systematically spread and, in severe cases, result in multiple organ failures [47]. In the cardiovascular system, the binding of the virus to the ACE-2 receptor activates signal transduction pathways, including the as-ERK-AP-1 pathway that activates pro-fibrosis factor, resulting in the development of cardiac inflammation and fibrosis [49].
Mutations to the aforementioned molecular targets have3333 led to increased pathogenicity and detection alterations of the various SARS-CoV-2 variants. Regarding the Omicron variant, a deficiency in spike cleavage, resulting in inefficient transmembrane protease-serine 2 (TMPRSS2) usage, allows for a dramatic increase in replication within the lungs [50]. As a result, RT-PCR can be set to search for a S gene target dropout, facilitating faster detection rates [51]. Conversely, the Delta variant’s increased contagiousness is attributed to an increased propensity for more RBD-up states and an affinity-enhancing T478K substitution, both of which increase ACE2 receptor binding [52]. The subsequent enhances replication allows for earlier detection, where the Delta variant has an average detection time of 3.2 days from exposure compared to the 4.5 days of the early alpha variant virus [53].
4. Current Methods of COVID-19 Diagnosis
Numerous methodologies have been applied to detect COVID-19. Currently, COVID-19 is primarily diagnosed using three techniques: (1) reverse-transcription polymerase chain reaction (RT-PCR) and gene sequencing, (2) chest computed tomography (CT) and (3) lateral flow immunoassay [7,8].
4.1. Reverse-Transcription Polymerase Chain Reaction (RT-PCR)
The current gold standard for diagnosing COVID-19 is RT-PCR due to its high selectivity and sensitivity [9,54,55,56,57], providing ~95% accuracy under ideal conditions [58]. In order to perform RT-PCR, a biological fluid sample containing SARS-CoV-2 strains is collected. A sample would usually entail upper and lower respiratory fluid collected using nasopharyngeal and oropharyngeal swabs [9]. Subsequently, the collected fluid would undergo filtration and separation steps to isolate the viral RNA, from which complementary viral DNA (cDNA) is created [59]. A large number of cDNA (proportional to the concentration of the virus) would generate a sizeable fluorescent signal following several rounds of polymerase reaction [59]. If the system is well-calibrated, fluorescence intensity would directly reflect the concentration of the virus within an infected patient [59]. Until now, three regions of the cDNA have been identified for detection of SARS-CoV-2: E, N and RdRP genes [9].
Despite its high selectivity and sensitivity, RT-PCR is accompanied by numerous challenges and disadvantages. The technique requires complex and expensive equipment and a laboratory with biosafety level 2 or above [60]. In addition, further prerequisites, including technically skilled personnel and a stable power supply (9), hinder the use of RT-PCR in several COVID-19 outbreak regions where there is a lack of infrastructure. The method takes 3–6 h from sample collection to result generation [10]. However, due to the sheer number of requested tests, the time required to obtain the results can be up to 2–3 days [60]. The massive amounts of reagents needed to perform the testing have also become a bottleneck [61]. Such inefficient systems can be detrimental to public health safety, especially since we live during a COVID-19 crisis. Furthermore, the wide variability of viruses in different patient samples has given rise to false-negative results [62]. The relatively low sensitivity of RT-PCR if patient samples are not of high purity (blood samples or sputum) can also explain the emergence of false-negative results [63,64]. Lastly, isolated RNA degrades rapidly and requires immediate freezing [59]; poor handling of samples may be another reason for false-negative results. All the factors mentioned above contributing to lower sensitivities can be exemplified by the sensitivity of the first RT-PCR test for SARS-CoV-2 being only 70% [65]. Consequently, CT is being used in combination with RT-PCR as a confirmatory diagnostic measure for patients with clinical suspicion who gave a negative RT-PCR result [12,13].
4.2. Computer Tomography (CT) Scan
CT scan is currently being used as either a standalone diagnostic tool or alongside RT-PCR (to ascertain negative results in symptomatic patients) in diagnosing COVID-19 [12,13]. In the context of COVID-19, CT revolves around taking many chest X-ray measurements from different angles. These measurements would then be compiled to create a three-dimensional (3D) image with contrast; such images are investigated by radiologists. Characteristic COVID-19 presentations that appear in CT include “areas of subpleural regions of ground glass opacification affecting the lower parts of either a single lobe or both lobes” [59]. CT scans can further be used to assess the prognosis of a patient with COVID-19. For instance, within the first few days of the infection, the image mimics a regular chest CT. However, as the disease progresses, the ground-glass-like opacity becomes more pronounced with bilateral peripheral predominant consolidation [66].
One of the main challenges that radiologists face is attempting to differentiate the symptoms of COVID-19 from other lung conditions, especially other pneumonia-causing pathologies [59]. Indeed, CT scans are reported to have a specificity of detection of approximately 25%, mainly due to the imaging characteristics and signs sharing significant similarities with other viral pneumonia [67]. Nonetheless, as CT scans are expensive and require knowledgeable personnel with high technical skills to operate and interpret, it has been chiefly used as a complementary method in SARS-CoV-2 detection [12,13].
4.3. Rapid Antigen Testing (RAT)
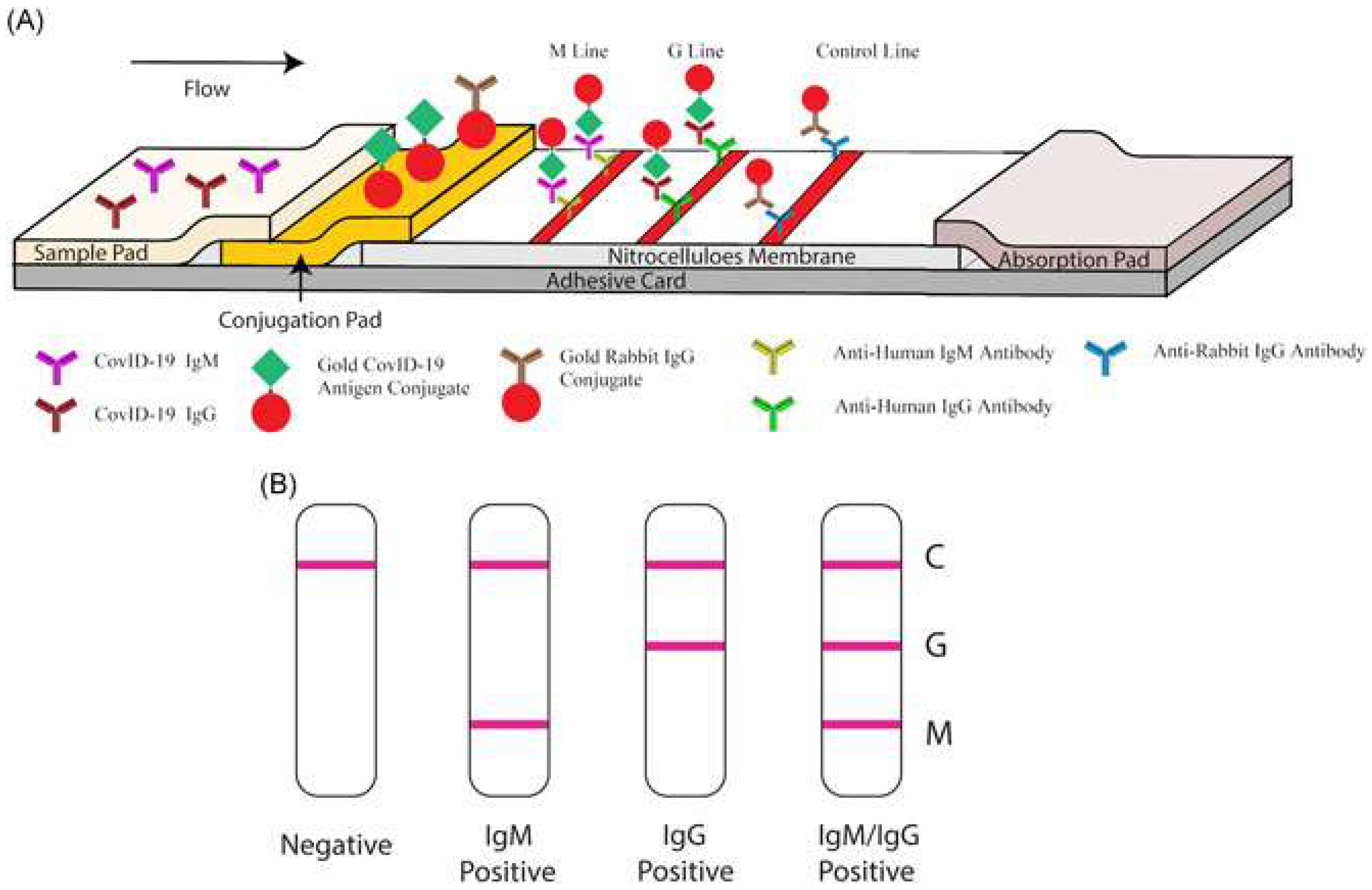
As an alternative to RT-PCR and radiological imaging, healthcare centers, particularly those in developing nations or rural regions, have been employing RAT kits to quickly determine the clinical management of symptomatic patients [68]. Using immunochromatography, the method revolves around the interaction between desired antigens (typically SARS-CoV-2 nucleocapsid protein) and antibodies implanted onto nitrocellulose membranes [69]. Results can be interpreted either through immunofluorescence or with the naked eye, through the presence of a colored band. RAT offers unique advantages including being an inexpensive, rapid and intuitive test that could be performed in a point-of-care setting or by patients themselves. Yet its main drawback is relatively poorer sensitivity and selectivity when compared to RT-PCR [68]. Indeed, the WHO recommends that a RAT kit should have a sensitivity of 80% and a specificity of 97% (compared to RT-PCR) to be clinically appropriate [69]. While such performance markers vary between RAT kit types and manufacturers as well as the included patient selection, a meta-analysis conducted by Khandker et al. involving 17,171 suspected COVID-19 patients reported a lack-luster sensitivity of 68.4% (95% CI: 60.8–75.9) and specificity of 99.4% (95% CI: 99.1–99.8). Further, the reported sensitivity of RAT kits within asymptomatic patients was 54.5% [69]. To this end, negative RAT results are not enough to rule out a COVID-19 infection and further molecular investigation may be indicated, especially for symptomatic patients.
4.4. Immunoassays and Enzyme-Linked Immunosorbent Assays (ELISA)
COVID-19 has been diagnosed through the detection of anti-SARS-CoV-2 IgG in serum. A prominent type of this technique is enzyme-linked immunosorbent assay (ELISA). Through the use of microtiter plates, such as the 96-well, antibodies are detected through protein–protein interactions [59]. A signal is then detected via fluorescence, luminescence, or colorimetric techniques [59]. The applications of this procedure can go beyond diagnosis. Changes in serum SARS-CoV-2 IgG levels can be monitored, providing a way to evaluate treatment response and prognosis [70]. Furthermore, the presence of antibodies can confidently confirm whether vaccines are effective [59]. It can also be an invaluable procedure in aiding intervention policymakers on the number of asymptomatic patients within a population [59].
Within diagnosis, immunoassays have the potential to provide a result very rapidly. A recent report has described the performance of a rapid test based on IgM and IgG; compared to RT-PCR, it showcased a lower sensitivity of 86.66% [60]. Nonetheless, the main advantage of using immunoassays remains in their remarkable speed in providing a result.
Challenges associated with immunoassays revolve around their sensitivity, technical issues and the time required to produce antibodies since the onset of infection. To begin, there are potential difficulties with creating an accurate serological test that can precisely differentiate between SARS-CoV-2 antibodies and antibodies generated against other coronaviruses [59]. As such, false-positive results may emerge due to a lack of selectivity between different coronaviruses’ antibodies. Interestingly, current antibody tests have been reported to present false-negative results. The majority of false-negative results are said to be caused by the following technical issues: “(1) a low concentration of antibodies typically present in fluidic samples; (2) presence of homologous proteins; and (3) lack of sensitivity from the detection instrument” [59]. Perhaps the most critical disadvantage of immunoassay lies in the fact that IgM and IgG are typically detectable two weeks after the onset of the infection (8). As a result, early detection of SARS-CoV-2 using immunoassays is difficult, forcing health workers to rely on other diagnostic methods such as RT-PCR and CT. Other issues pertaining to immunoassays, specifically ELISA, include labor-intensive sample collection and long incubation times [71].
4.5. Loop-Mediated Isothermal Amplification (LAMP)
Developed by Notomi et al. in 2000, LAMP has garnered interest in diagnostics due to its quick, sensitive and ultimately effective amplification method using nucleic acids [72]. Detection can be seen visually using intercalating fluorescent dyes [72] or colorimetric techniques [73]. As it is notably cheaper to operate than RT-PCR and still able to detect nucleic acids, isothermal thermal amplification methods are being developed to overcome RT-PCR’s cost and resource [74]; the method presents advantages that can be utilized in POC testing [74]. In some instances, LAMP has been reported to be faster and marginally more sensitive than conventional PCR methods [62]. Consequently, the technique has been deemed appropriate for detecting viruses such as MERS-CoV, SARS-CoV and influenza A [62].
LAMP has been implemented alongside RT (RT-LAMP) to provide a simple and high-throughput method of detecting SARS-CoV-2. The detection time was reported to be 30 min, highlighting its potential in POC and screening tests [74]. Furthermore, LAMP’s efficiency in mass detection can be further amplified as it is possible to use unpurified samples in conjunction with colorimetric detection [74].
However, LAMP still has its share of disadvantages. It has been reported that LAMP is less sensitive than PCR in cases of complicated samples, such as blood. The presumed reason for this is its use of Bst DNA polymerase, while PCR employs Taq polymerase [75]. In addition, the suitable development of primers is a limitation associated with LAMP [75]. Additional research on the stability and consistency of LAMP is required for its widespread application in clinical diagnosis [76].
6. Conclusions
The spread of COVID-19 has caused massive repercussions to the world’s healthcare system. Without any specific therapeutic procedures for COVID-19, proper knowledge of the virus’s structural and genomic components is required to develop alternative methods of combating the virus. One such method is using efficient diagnostic techniques as a preventative approach for the outbreak. RT-PCR and CT scan are used concomitantly as the standard procedure for diagnosing COVID-19 due to their accuracy and efficacy; however, several disadvantages have been noted for the aforementioned techniques, including false-negative results, high cost of operation and requirement of trained personnel. With the rise of SARS-CoV-2 variants and mutations that both worsen prognosis and complicate virus detection, challenges obstructing rapid, reliable and inexpensive COVID-19 diagnosis continue to emerge. As such, NPs have been identified in this review as possible candidates to cover the fallacies of RT-PCR and CT in the diagnosis of COVID-19.
This review has showcased the main studies revolving around the potential diagnostic utility of NPs in the COVID-19 pandemic. Considering the flexibility of NPs, they can be utilized either as standalone or supporting devices to current methods such as RT-PCR. Particularly, AuNP, AgNP, carbon nanotubes and graphene sheets have displayed promising capability to be used as a basis of a biosensor. AuNP has also portrayed great results when applied in lateral flow immunoassays. Moreover, superparamagnetic NPs can aid in RT-PCR by efficiently extracting RNA for the desired application. Given the ability of NPs to produce precise results within a relatively short period of time, NP-based diagnostic studies for SARS-CoV, MERS-CoV and SARS-CoV-2 can be utilized to better the available strategies currently used in the COVID-19 pandemic.
Author Contributions
Conceptualization, A.A.-H.; project administration A.A.-H.; literature review and investigation, A.A.-H., U.A., Y.M.W., O.S.M.M.S.E., M.H.H. and M.S.; writing—original draft preparation, A.A.-H., U.A. and O.S.M.M.S.E.; writing—review and editing, A.A.-H., Y.M.W., M.H.H. and M.S.; supervision, G.R.D. and F.Z.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- John Hopkins University. Coronavirus COVID-19 Global Cases by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University. Available online: https://coronavirus.jhu.edu/map.html (accessed on 2 August 2022).
- World Health Organization. Coronavirus Disease (COVID-19) Pandemic. Available online: https://www.who.int/europe/emergencies/situations/covid-19 (accessed on 28 September 2022).
- McKibbin, W.; Fernando, R. The Global Macroeconomic Impacts of COVID-19- Seven Scenarios. SSRN Electron. J. 2020, 20, 1–30. [Google Scholar] [CrossRef]
- Talic, S.; Shah, S.; Wild, H.; Gasevic, D.; Maharaj, A.; Ademi, Z.; Li, X.; Xu, W.; Mesa-Eguiagaray, I.; Rostron, J.; et al. Effectiveness of public health measures in reducing the incidence of COVID-19, SARS-CoV-2 transmission, and covid-19 mortality: Systematic review and meta-analysis. BMJ 2021, 375, e068302. [Google Scholar] [CrossRef]
- World Health Organization. Coronavirus Disease (COVID-19) Advice for the Public. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/advice-for-public (accessed on 20 September 2020).
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA 2020, 323, 1843–1844. [Google Scholar] [CrossRef] [PubMed]
- Kim, H. Outbreak of novel coronavirus (COVID-19): What is the role of radiologists? Eur. Radiol. 2020, 30, 3266–3267. [Google Scholar] [CrossRef] [PubMed]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.C.; Chen, H.; Mubareka, S.; Gubbay, J.B.; Chan, W.C.W. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822–3835. [Google Scholar] [CrossRef] [PubMed]
- Cui, F.; Zhou, H.S. Diagnostic methods and potential portable biosensors for coronavirus disease 2019. Biosens. Bioelectron. 2020, 165, 112349. [Google Scholar] [CrossRef]
- Zhu, X.; Wang, X.; Han, L.; Chen, T.; Wang, L.; Li, H.; Li, S.; He, L.; Fu, X.; Chen, S.; et al. Multiplex reverse transcription loop-mediated isothermal amplification combined with nanoparticle-based lateral flow biosensor for the diagnosis of COVID-19. Biosens. Bioelectron. 2020, 166, 112437. [Google Scholar] [CrossRef]
- Li, D.; Wang, D.; Dong, J.; Wang, N.; Huang, H.; Xu, H.; Xia, C. False-Negative Results of Real-Time Reverse-Transcriptase Polymerase Chain Reaction for Severe Acute Respiratory Syndrome Coronavirus 2: Role of Deep-Learning-Based CT Diagnosis and Insights from Two Cases. Korean J. Radiol. 2020, 21, 505–508. [Google Scholar] [CrossRef]
- Zu, Z.Y.; Jiang, M.D.; Xu, P.P.; Chen, W.; Ni, Q.Q.; Lu, G.M.; Zhang, L.J. Coronavirus Disease 2019 (COVID-19)- A Perspective from China. Radiology 2020, 296, E15–E25. [Google Scholar] [CrossRef]
- Fernandes, Q.; Inchakalody, V.P.; Merhi, M.; Mestiri, S.; Taib, N.; Moustafa Abo El-Ella, D.; Bedhiafi, T.; Raza, A.; Al-Zaidan, L.; Mohsen, M.O.; et al. Emerging COVID-19 variants and their impact on SARS-CoV-2 diagnosis, therapeutics and vaccines. Ann. Med. 2022, 54, 524–540. [Google Scholar] [CrossRef]
- Thakur, P.; Thakur, V.; Kumar, P.; Singh Patel, S.K. Emergence of novel omicron hybrid variants: BA(x), XE, XD, XF more than just alphabets. Int. J. Surg. 2022, 104, 106727. [Google Scholar] [CrossRef] [PubMed]
- Thakur, V.; Bhola, S.; Thakur, P.; Patel, S.; Kulshrestha, S.; Ratho, R.K.; Kumar, P. Waves and variants of SARS-CoV-2: Understanding the causes and effect of the COVID-19 catastrophe. Infection 2022, 50, 309–325. [Google Scholar] [CrossRef]
- Kumar, N.; Kumbhat, S. Unique Properties. In Essentials in Nanoscience and Nanotechnology; John Wiley & Sons: Hoboken, NJ, USA, 2016. [Google Scholar]
- Tiloke, C.; Phulukdaree, A.; Chuturgoon, A.A. The Chemotherapeutic Potential of Gold Nanoparticles against Human Carcinomas: A Review. In Nanoarchitectonics for Smart Delivery and Drug Targeting, 1st ed.; Holban, A.M., Grumezescu, A.M., Eds.; Elsevier: Amsterdam, The Netherlands, 2016; pp. 783–811. [Google Scholar]
- Thakur, N.; Kumar, A.K.; Kumar, A. Effect of (Ag, Zn) co-doping on structural, optical and bactericidal properties of CuO nanoparticles synthesized by a microwave-assisted method. Dalton Trans. 2021, 50, 6188–6203. [Google Scholar] [CrossRef]
- Pelaz, B.; Alexiou, C.; Alvarez-Puebla, R.A.; Alves, F.; Andrews, A.M.; Ashraf, S.; Balogh, L.P.; Ballerini, L.; Bestetti, A.; Brendel, C.; et al. Diverse Applications of Nanomedicine. ACS Nano 2017, 11, 2313–2381. [Google Scholar] [CrossRef] [PubMed]
- Ielo, I.; Rando, G.; Giacobello, F.; Sfameni, S.; Castellano, A.; Galletta, M.; Drommi, D.; Rosace, G.; Plutino, M.R. Synthesis, Chemical-Physical Characterization, and Biomedical Applications of Functional Gold Nanoparticles: A Review. Molecules 2021, 26, 5823. [Google Scholar] [CrossRef]
- Davies, G.L.; O’Brien, J.; Gun’ko, Y.K. Rare Earth Doped Silica Nanoparticles via Thermolysis of a Single Source Metallasilsesquioxane Precursor. Sci. Rep. 2017, 7, 45862. [Google Scholar] [CrossRef]
- Yu, X.; Pham, J.T.; Subramani, C.; Creran, B.; Yeh, Y.C.; Du, K.; Patra, D.; Miranda, O.R.; Crosby, A.J.; Rotello, V.M. Direct patterning of engineered ionic gold nanoparticles via nanoimprint lithography. Adv. Mater. 2012, 24, 6330–6334. [Google Scholar] [CrossRef]
- Sportelli, M.C.; Izzi, M.; Volpe, A.; Clemente, M.; Picca, R.A.; Ancona, A.; Lugarà, P.M.; Palazzo, G.; Cioffi, N. The Pros and Cons of the Use of Laser Ablation Synthesis for the Production of Silver Nano-Antimicrobials. Antibiotics 2018, 7, 67. [Google Scholar] [CrossRef]
- Mirzaei, A.; Neri, G. Microwave-assisted synthesis of metal oxide nanostructures for gas sensing application: A review. Sens. Actuators B Chem. 2016, 237, 749–775. [Google Scholar] [CrossRef]
- Flores-Rojas, G.; López-Saucedo, F.; Bucio, E. Gamma-irradiation applied in the synthesis of metallic and organic nanoparticles: A short review. Radiat. Phys. Chem. 2020, 169, 107962. [Google Scholar] [CrossRef]
- Hatakeyama, Y.; Onishi, K.; Nishikawa, K. Effects of sputtering conditions on formation of gold nanoparticles in sputter deposition technique. RSC Adv. 2011, 1, 1815–1821. [Google Scholar] [CrossRef]
- Singh, J.; Dutta, T.; Kim, K.H.; Rawat, M.; Samddar, P.; Kumar, P. ‘Green’ synthesis of metals and their oxide nanoparticles: Applications for environmental remediation. J. Nanobiotechnol. 2018, 16, 84. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.; Annu, I.S.; Yudha, S.S. Biosynthesis of gold nanoparticles: A green approach. J. Photochem. Photobiol. B 2016, 161, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Salem, S.S.; Fouda, A. Synthesis of Metallic Nanoparticles and Their Prospective Biotechnological Applications: An Overview. Biol. Trace Elem. Res. 2021, 199, 344–370. [Google Scholar] [CrossRef]
- Kumar, S.; Lather, V.; Pandita, D. Green synthesis of therapeutic nanoparticles: An expanding horizon. Nanomedicine 2015, 10, 2451–2471. [Google Scholar] [CrossRef] [PubMed]
- Caires, A.J.; Mansur, H.S.; Mansur, A.; Carvalho, S.M.; Lobato, Z.; Dos Reis, J. Gold nanoparticle-carboxymethyl cellulose nanocolloids for detection of human immunodeficiency virus type-1 (HIV-1) using laser light scattering immunoassay. Colloids. Surf. B Biointerfaces 2019, 177, 377–388. [Google Scholar] [CrossRef]
- Liu, Z.; Shang, C.; Ma, H.; You, M. An upconversion nanoparticle-based photostable FRET system for long-chain DNA sequence detection. Nanotechnology 2020, 31, 235501. [Google Scholar] [CrossRef]
- Medhi, R.; Srinoi, P.; Ngo, N.; Tran, H.V.; Lee, T.R. Nanoparticle-Based Strategies to Combat COVID-19. ACS Appl. Nano Mater. 2020, 3, 8557–8580. [Google Scholar] [CrossRef]
- Asdaq, S.; Ikbal, A.; Sahu, R.K.; Bhattacharjee, B.; Paul, T.; Deka, B.; Fattepur, S.; Widyowati, R.; Vijaya, J.; Al Mohaini, M.; et al. Nanotechnology Integration for SARS-CoV-2 Diagnosis and Treatment: An Approach to Preventing Pandemic. Nanomaterials 2021, 11, 1841. [Google Scholar] [CrossRef]
- Rabaan, A.A.; Al-Ahmed, S.H.; Haque, S.; Sah, R.; Tiwari, R.; Malik, Y.S.; Dhama, K.; Yatoo, M.I.; Bonilla-Aldana, D.K.; Rodriguez-Morales, A.J. SARS-CoV-2, SARS-CoV, and MERS-CoV- a comparative overview. Le Infez. Med. 2020, 28, 174–184. [Google Scholar]
- Demmler, G.J.; Ligon, B.L. Severe Acute Respiratory Syndrome (SARS)- A Review of the History, Epidemiology, Prevention, and Concerns for the Future. Semin. Pediatr. Infect. Dis. 2003, 14, 240–244. [Google Scholar] [CrossRef]
- Coleman, C.M.; Frieman, M.B. Coronaviruses: Important emerging human pathogens. J. Virol. 2014, 88, 5209–5212. [Google Scholar] [CrossRef] [PubMed]
- Skowronski, D.M.; Astell, C.; Brunham, R.C.; Low, D.E.; Petric, M.; Roper, R.L.; Talbot, P.J.; Tam, T.; Babiuk, L. Severe acute respiratory syndrome (SARS): A year in review. Annu. Rev. Med. 2005, 56, 357–381. [Google Scholar] [CrossRef]
- Baharoon, S.; Memish, Z.A. MERS-CoV as an emerging respiratory illness: A review of prevention methods. Travel Med. Infect. Dis. 2019, 32, 101520. [Google Scholar] [CrossRef]
- Mackay, I.M.; Arden, K.E. MERS coronavirus: Diagnostics, epidemiology and transmission. Virol. J. 2015, 12, 222. [Google Scholar] [CrossRef]
- Khailany, R.A.; Safdar, M.; Ozaslan, M. Genomic characterization of a novel SARS-CoV-2. Gene Rep. 2020, 19, 100682. [Google Scholar] [CrossRef]
- Majumder, J.; Minko, T. Recent Developments on Therapeutic and Diagnostic Approaches for COVID-19. AAPS J. 2021, 23, 14. [Google Scholar] [CrossRef]
- Naqvi, A.A.T.; Fatima, K.; Mohammad, T.; Fatima, U.; Singh, I.K.; Singh, A.; Atif, S.M.; Hariprasad, G.; Hasan, G.M.; Hassan, M.I. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165878. [Google Scholar] [CrossRef]
- Yuki, K.; Fujiogi, M.; Koutsogiannaki, S. COVID-19 pathophysiology: A review. Clin. Immunol. 2020, 215, 108427. [Google Scholar] [CrossRef]
- Turner, A.J.; Hiscox, J.A.; Hooper, N.M. ACE2: From vasopeptidase to SARS virus receptor. Trends Pharm. Sci 2004, 25, 291–294. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Tipnis, S.R.; Hooper, N.M.; Hyde, R.; Karran, E.; Christie, G.; Turner, A.J. A human homolog of angiotensin-converting enzyme. Cloning and functional expression as a captopril-insensitive carboxypeptidase. J. Biol. Chem. 2000, 275, 33238–33243. [Google Scholar] [CrossRef] [PubMed]
- Chen, I.Y.; Chang, S.C.; Wu, H.Y.; Yu, T.C.; Wei, W.C.; Lin, S.; Chien, C.L.; Chang, M.F. Upregulation of the chemokine (C-C motif) ligand 2 via a severe acute respiratory syndrome coronavirus spike-ACE2 signaling pathway. J. Virol. 2010, 84, 7703–7712. [Google Scholar] [CrossRef]
- Chu, H.; Yuen, K.Y. Pathogenicity of SARS-CoV-2 Omicron. Clin. Transl. Med. 2022, 12, e880. [Google Scholar] [CrossRef]
- World Health Organization. Classification of Omicron (B.1.1.529): SARS-CoV-2 Variant of Concern. Available online: https://www.who.int/news/item/26-11-2021-classification-of-omicron-(b.1.1.529)-sars-cov-2-variant-of-concern (accessed on 9 September 2022).
- Wang, Y.; Liu, C.; Zhang, C.; Wang, Y.; Hong, Q.; Xu, S.; Li, Z.; Yang, Y.; Huang, Z.; Cong, Y. Structural basis for SARS-CoV-2 Delta variant recognition of ACE2 receptor and broadly neutralizing antibodies. Nat. Commun. 2022, 13, 871. [Google Scholar] [CrossRef]
- Hart, W.S.; Miller, E.; Andrews, N.J.; Waight, P.; Maini, P.K.; Funk, S.; Thompson, R.N. Generation time of the alpha and delta SARS-CoV-2 variants: An epidemiological analysis. Lancet Infect. Dis. 2022, 25, 603–610. [Google Scholar] [CrossRef]
- Carter, L.J.; Garner, L.V.; Smoot, J.W.; Li, Y.; Zhou, Q.; Saveson, C.J.; Sasso, J.M.; Gregg, A.C.; Soares, D.J.; Beskid, T.R.; et al. Assay Techniques and Test Development for COVID-19 Diagnosis. ACS Cent. Sci. 2020, 6, 591–605. [Google Scholar] [CrossRef]
- Yu, L.; Wu, S.; Hao, X.; Dong, X.; Mao, L.; Pelechano, V.; Chen, W.; Yin, X. Rapid detection of COVID-19 coronavirus using a reverse transcriptional loop-mediated isothermal amplification (RT-LAMP) diagnostic platform. Clin. Chem. 2020, 66, 975–977. [Google Scholar] [CrossRef]
- Haijema, B.J.; Volders, H.; Rottier, P.J. Live, attenuated coronavirus vaccines through the directed deletion of group-specific genes provide protection against feline infectious peritonitis. J. Virol. 2004, 78, 3863–3871. [Google Scholar] [CrossRef]
- Fukushi, S.; Fukuma, A.; Kurosu, T.; Watanabe, S.; Shimojima, M.; Shirato, K.; Iwata-Yoshikawa, N.; Nagata, N.; Ohnishi, K.; Ato, M.; et al. Characterization of novel monoclonal antibodies against the MERS-coronavirus spike protein and their application in species-independent antibody detection by competitive ELISA. J. Virol. Methods 2018, 251, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Tahamtan, A.; Ardebili, A. Real-time RT-PCR in COVID-19 detection: Issues affecting the results. Expert Rev. Mol. Diagn. 2020, 20, 453–454. [Google Scholar] [CrossRef] [PubMed]
- Bhalla, N.; Pan, Y.; Yang, Z.; Payam, A.F. Opportunities and Challenges for Biosensors and Nanoscale Analytical Tools for Pandemics: COVID-19. ACS Nano 2020, 14, 7783–7807. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.; Duong Bang, D.; Wolff, A. 2019 Novel Coronavirus Disease (COVID-19): Paving the Road for Rapid Detection and Point-of-Care Diagnostics. Micromachines 2020, 11, 306. [Google Scholar] [CrossRef] [PubMed]
- Palestino, G.; Garcia-Silva, I.; Gonzalez-Ortega, O.; Rosales-Mendoza, S. Can nanotechnology help in the fight against COVID-19? Expert Rev. Anti. Infect. 2020, 18, 849–864. [Google Scholar] [CrossRef] [PubMed]
- Rabiee, N.; Bagherzadeh, M.; Ghasemi, A.; Zare, H.; Ahmadi, S.; Fatahi, Y.; Dinarvand, R.; Rabiee, M.; Ramakrishna, S.; Shokouhimehr, M.; et al. Point-of-Use Rapid Detection of SARS-CoV-2: Nanotechnology-Enabled Solutions for the COVID-19 Pandemic. Int. J. Mol. Sci. 2020, 21, 5126. [Google Scholar] [CrossRef]
- Li, Y.; Yao, L.; Li, J.; Chen, L.; Song, Y.; Cai, Z.; Yang, C. Stability issues of RT-PCR testing of SARS-CoV-2 for hospitalized patients clinically diagnosed with COVID-19. J. Med. Virol. 2020, 92, 903–908. [Google Scholar] [CrossRef]
- Tang, Y.; Schmitz, J.; Persing, D.; CW, S. Laboratory Diagnosis of COVID-19- Current Issues and Challenges. J Clin Microbiol 2020, 58, e00512-20. [Google Scholar] [CrossRef]
- Al-Tawfiq, J.A.; Memish, Z.A. Diagnosis of SARS-CoV-2 infection based on CT scan vs RT-PCR: Reflecting on experience from MERS-CoV. J. Hosp. Infect. 2020, 105, 154–155. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Han, X.; Jiang, N.; Cao, Y.; Alwalid, O.; Gu, J.; Fan, Y.; Zheng, C. Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: A descriptive study. Lancet Infect. Dis. 2020, 20, 425–434. [Google Scholar] [CrossRef]
- Ai, T.; Yang, Z.; Hou, H.; Zhan, C.; Chen, C.; Lv, W.; Tao, Q.; Sun, Z.; Xia, L. Correlation of Chest CT and RT-PCR Testing in Coronavirus Disease 2019 (COVID-19) in Chin—A Report of 1014 Cases. Radiology 2020, 296, E32–E40. [Google Scholar] [CrossRef]
- Peeling, R.W.; Olliaro, P.L.; Boeras, D.I.; Fongwen, N. Scaling up COVID-19 rapid antigen tests: Promises and challenges. Lancet Infect. Dis. 2021, 21, e290–e295. [Google Scholar] [CrossRef]
- Khandker, S.S.; Nik Hashim, N.; Deris, Z.Z.; Shueb, R.H.; Islam, M.A. Diagnostic Accuracy of Rapid Antigen Test Kits for Detecting SARS-CoV-2: A Systematic Review and Meta-Analysis of 17,171 Suspected COVID-19 Patients. J. Clin. Med. 2021, 10, 3493. [Google Scholar] [CrossRef] [PubMed]
- Xie, L.; Liu, Y.; Fan, B.; Xiao, Y.; Tian, Q.; Chen, L.; Zhao, H.; Chen, W. Dynamic changes of serum SARS-coronavirus IgG, pulmonary function and radiography in patients recovering from SARS after hospital discharge. Respir. Res. 2005, 6, 5. [Google Scholar] [CrossRef] [PubMed]
- Wen, T.; Huang, C.; Shi, F.J.; Zeng, X.Y.; Lu, T.; Ding, S.N.; Jiao, Y.J. Development of a lateral flow immunoassay strip for rapid detection of IgG antibody against SARS-CoV-2 virus. Analyst 2020, 145, 5345–5352. [Google Scholar] [CrossRef] [PubMed]
- Oscorbin, I.P.; Belousova, E.A.; Zakabunin, A.I.; Boyarskikh, U.A.; Filipenko, M.L. Comparison of fluorescent intercalating dyes for quantitative loop-mediated isothermal amplification (qLAMP). Biotechniques 2016, 61, 20–25. [Google Scholar] [CrossRef]
- Goto, M.; Honda, E.; Ogura, A.; Nomoto, A.; Hanaki, K. Colorimetric detection of loop-mediated isothermal amplification reaction by using hydroxy naphthol blue. Biotechniques 2009, 46, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Park, G.S.; Ku, K.; Baek, S.H.; Kim, S.J.; Kim, S.I.; Kim, B.T.; Maeng, J.S. Development of Reverse Transcription Loop-Mediated Isothermal Amplification Assays Targeting Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). J. Mol. Diagn. 2020, 22, 729–735. [Google Scholar] [CrossRef]
- Sahoo, P.R.; Sethy, K.; Mohapatra, S.; Panda, D. Loop mediated isothermal amplification: An innovative gene amplification technique for animal diseases. Vet. World 2016, 9, 465–469. [Google Scholar] [CrossRef]
- Deguo, W.; Guicheng, H.; Fugui, W.; Yonggang, L.; Daxi, R. Drawback of loop-mediated isothermal amplification. Afr. J. Food Sci. 2008, 2, 83–86. [Google Scholar]
- Moitra, P.; Alafeef, M.; Dighe, K.; Frieman, M.B.; Pan, D. Selective Naked-Eye Detection of SARS-CoV-2 Mediated by N Gene Targeted Antisense Oligonucleotide Capped Plasmonic Nanoparticles. ACS Nano 2020, 14, 7617–7627. [Google Scholar] [CrossRef]
- Nikaeen, G.; Abbaszadeh, S.; Yousefinejad, S. Application of nanomaterials in treatment, anti-infection and detection of coronaviruses. Nanomedicine 2020, 15, 1501–1512. [Google Scholar] [CrossRef] [PubMed]
- Layqah, L.A.; Eissa, S. An electrochemical immunosensor for the corona virus associated with the Middle East respiratory syndrome using an array of gold nanoparticle-modified carbon electrodes. Mikrochim. Acta 2019, 186, 224. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Park, M.; Hwang, J.; Kim, J.H.; Chung, D.R.; Lee, K.S.; Kang, M. Development of Label-Free Colorimetric Assay for MERS-CoV Using Gold Nanoparticles. ACS Sens. 2019, 4, 1306–1312. [Google Scholar] [CrossRef]
- Huang, P.; Wang, H.; Cao, Z.; Jin, H.; Chi, H.; Zhao, J.; Yu, B.; Yan, F.; Hu, X.; Wu, F.; et al. A Rapid and Specific Assay for the Detection of MERS-CoV. Front. Microbiol. 2018, 9, 1101. [Google Scholar] [CrossRef] [PubMed]
- Teengam, P.; Siangproh, W.; Tuantranont, A.; Vilaivan, T.; Chailapakul, O.; Henry, C.S. Multiplex Paper-Based Colorimetric DNA Sensor Using Pyrrolidinyl Peptide Nucleic Acid-Induced AgNPs Aggregation for Detecting MERS-CoV, MTB, and HPV Oligonucleotides. Anal. Chem. 2017, 89, 5428–5435. [Google Scholar] [CrossRef]
- Qiao, J.; Li, Y.; Wei, C.; Yang, H.; Yu, J.; Wei, H. Rapid detection of viral antibodies based on multifunctional Staphylococcus aureus nanobioprobes. Enzym. Microb. Technol. 2016, 95, 94–99. [Google Scholar] [CrossRef]
- Park, T.J.; Lee, S.Y.; Lee, S.J.; Park, J.P.; Yang, S.K.; Lee, K.B.; Ko, S.; Park, J.B.; Kim, T.; Kim, S.K.; et al. Protein Nanopatterns and Biosensors Using Gold Binding Polypeptide as a Fusion Partner. Anal. Chem. 2006, 78, 7197–7205. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.C.; Chang, Y.F.; Chen, K.H.; Su, L.C.; Lee, C.W.; Chen, C.C.; Chen, Y.M.; Chou, C. Detection of severe acute respiratory syndrome (SARS) coronavirus nucleocapsid protein in human serum using a localized surface plasmon coupled fluorescence fiber-optic biosensor. Biosens. Bioelectron. 2009, 25, 320–325. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Paredes, G.; González-García, M.B.; Costa-García, A. Genosensor for SARS Virus Detection Based on Gold Nanostructured Screen-Printed Carbon Electrodes. Electroanalysis 2009, 21, 379–385. [Google Scholar] [CrossRef]
- Ishikawa, F.N.; Curreli, M.; Olson, C.A.; Liao, H.I.; Sun, R.; Roberts, R.W.; Cote, R.J.; Thompson, M.E.; Zhou, C. Importance of Controlling Nanotube Density for Highly Sensitive and Reliable Biosensors Functional in Physiological Conditions. ACS Nano 2010, 4, 6914–6922. [Google Scholar] [CrossRef] [PubMed]
- Gong, P.; He, X.; Wang, K.; Tan, W.; Xie, W.; Wu, P.; Li, H. Combination of functionalized nanoparticles and polymerase chain reaction-based method for SARS-CoV gene detection. Nanosci. Nanotechnol. 2008, 8, 293–300. [Google Scholar] [CrossRef]
- Kumar, V.; Mishra, S.; Sharma, R.; Agarwal, J.; Ghoshal, U.; Khanna, T.; Sharma, L.K.; Verma, S.K.; Mishra, P.; Tiwari, S. Development of RNA-based assay for rapid detection of SARS-CoV-2 in clinical samples. Intervirology 2022, 1–7. [Google Scholar] [CrossRef]
- Li, J.; Wu, D.; Yu, Y.; Li, T.; Li, K.; Xiao, M.M.; Li, Y.; Zhang, Z.Y.; Zhang, G.J. Rapid and unamplified identification of COVID-19 with morpholino-modified graphene field-effect transistor nanosensor. Biosens. Bioelectron. 2021, 183, 113206. [Google Scholar] [CrossRef] [PubMed]
- Qiu, G.; Gai, Z.; Tao, Y.; Schmitt, J.; Kullak-Ublick, G.A.; Wang, J. Dual-Functional Plasmonic Photothermal Biosensors for Highly Accurate Severe Acute Respiratory Syndrome Coronavirus 2 Detection. ACS Nano 2020, 14, 5268–5277. [Google Scholar] [CrossRef]
- Chen, Z.; Zhang, Z.; Zhai, X.; Li, Y.; Lin, L.; Zhao, H.; Bian, L.; Li, P.; Yu, L.; Wu, Y.; et al. Rapid and Sensitive Detection of anti-SARS-CoV-2 IgG, Using Lanthanide-Doped Nanoparticles-Based Lateral Flow Immunoassay. Anal. Chem. 2020, 92, 7226–7231. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wen, T.; Shi, F.J.; Zeng, X.Y.; Jiao, Y.J. Rapid Detection of IgM Antibodies against the SARS-CoV-2 Virus via Colloidal Gold Nanoparticle-Based Lateral-Flow Assay. ACS Omega 2020, 5, 12550–12556. [Google Scholar] [CrossRef]
- Li, Z.; Yi, Y.; Luo, X.; Xiong, N.; Liu, Y.; Li, S.; Sun, R.; Wang, Y.; Hu, B.; Chen, W.; et al. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J. Med. Virol. 2021, 92, 1518–1524. [Google Scholar] [CrossRef]
- Seo, G.; Lee, G.; Kim, M.J.; Baek, S.H.; Choi, M.; Ku, K.B.; Lee, C.S.; Jun, S.; Park, D.; Kim, H.G.; et al. Rapid Detection of COVID-19 Causative Virus (SARS-CoV-2) in Human Nasopharyngeal Swab Specimens Using Field-Effect Transistor-Based Biosensor. ACS Nano 2020, 14, 5135–5142. [Google Scholar] [CrossRef] [PubMed]
- Zhong, J.; Rösch, E.L.; Viereck, T.; Schilling, M.; Ludwig, F. Toward Rapid and Sensitive Detection of SARS-CoV-2 with Functionalized Magnetic Nanoparticles. ACS Sens. 2021, 6, 976–984. [Google Scholar] [CrossRef] [PubMed]
- Hryniewicz, B.M.; Volpe, J.; Bach-Toledo, L.; Kurpel, K.C.; Deller, A.E.; Soares, A.L.; Nardin, J.M.; Marchesi, L.F.; Simas, F.F.; Oliveira, C.C.; et al. Development of polypyrrole (nano)structures decorated with gold nanoparticles toward immunosensing for COVID-19 serological diagnosis. Mater. Today Chem. 2022, 24, 100817. [Google Scholar] [CrossRef] [PubMed]
- Durmus, C.; Balaban Hanoglu, S.; Harmanci, D.; Moulahoum, H.; Tok, K.; Ghorbanizamani, F.; Sanli, S.; Zihnioglu, F.; Evran, S.; Cicek, C.; et al. Indiscriminate SARS-CoV-2 multivariant detection using magnetic nanoparticle-based electrochemical immunosensing. Talanta 2022, 243, 123356. [Google Scholar] [CrossRef] [PubMed]
- Ellipilli, S.; Wang, H.; Lee, W.J.; Shu, D.; Guo, P. Proof-of-concept for speedy development of rapid and simple at-home method for potential diagnosis of early COVID-19 mutant infections using nanogold and aptamer. Nanomedicine 2022, 45, 102590. [Google Scholar] [CrossRef]
- Blumenfeld, N.R.; Bolene, M.; Jaspan, M.; Ayers, A.G.; Zarrandikoetxea, S.; Freudman, J.; Shah, N.; Tolwani, A.M.; Hu, Y.; Chern, T.L.; et al. Multiplexed reverse-transcriptase quantitative polymerase chain reaction using plasmonic nanoparticles for point-of-care COVID-19 diagnosis. Nat. Nanotechnol. 2022, 17, 984–992. [Google Scholar] [CrossRef] [PubMed]
- Miripour, Z.S.; Sarrami-Forooshani, R.; Sanati, H.; Makarem, J.; Taheri, M.S.; Shojaeian, F.; Eskafi, A.H.; Abbasvandi, F.; Namdar, N.; Ghafari, H.; et al. Real-time diagnosis of reactive oxygen species (ROS) in fresh sputum by electrochemical tracing; correlation between COVID-19 and viral-induced ROS in lung/respiratory epithelium during this pandemic. Biosens. Bioelectron. 2020, 165, 112435. [Google Scholar] [CrossRef]
- Shan, B.; Broza, Y.Y.; Li, W.; Wang, Y.; Wu, S.; Liu, Z.; Wang, J.; Gui, S.; Wang, L.; Zhang, Z.; et al. Multiplexed Nanomaterial-Based Sensor Array for Detection of COVID-19 in Exhaled Breath. ACS Nano 2020, 14, 12125–12132. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).