The Genetic Characteristics and Carbapenem Resistance Mechanism of ST307 Klebsiella pneumoniae Coharbouring blaCMY-6, blaOXA-48, and a Truncated blaNDM-1
Abstract
1. Introduction
2. Results
2.1. Clonality Analysis and Population Structure of ST307
2.2. Antimicrobial Resistance Genes in the Isolates
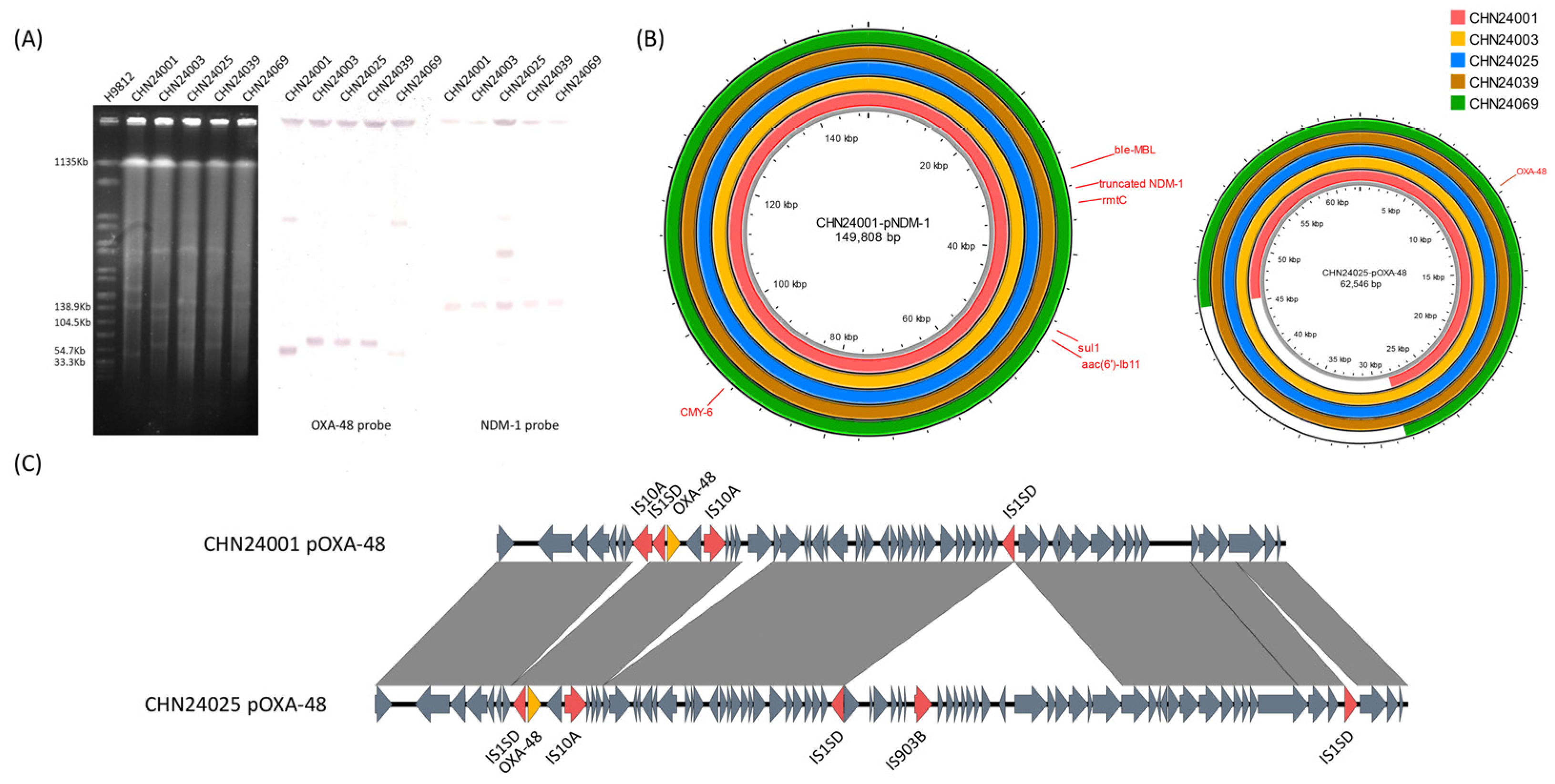
2.3. Difference in pNDM-1 and pOXA-48 among These Isolates
2.4. Conjugation Assay and Gene Cloning Experiments
3. Discussion
4. Materials and Methods
4.1. Collection of Isolates and Bacterial Identification
4.2. Antimicrobial Resistance Testing
4.3. PFGE, S1-PFGE, and Southern Blotting
4.4. Whole-Genome Sequencing and Subsequent Analysis
4.5. Conjugation Experiments
4.6. Plasmid Construction
4.7. Nucleotide Sequence Accession Numbers
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Paczosa, M.K.; Mecsas, J. Klebsiella pneumoniae: Going on the Offense with a Strong Defense. Microbiol. Mol. Biol. Rev. 2016, 80, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Sheu, C.C.; Chang, Y.T.; Lin, S.Y.; Chen, Y.H.; Hsueh, P.R. Infections Caused by Carbapenem-Resistant Enterobacteriaceae: An Update on Therapeutic Options. Front. Microbiol. 2019, 10, 80. [Google Scholar] [CrossRef] [PubMed]
- Hawser, S.P.; Bouchillon, S.K.; Lascols, C.; Hackel, M.; Hoban, D.J.; Badal, R.E.; Woodford, N.; Livermore, D.M. Susceptibility of Klebsiella pneumoniae isolates from intra-abdominal infections and molecular characterization of ertapenem-resistant isolates. Antimicrob. Agents Chemother. 2011, 55, 3917–3921. [Google Scholar] [CrossRef] [PubMed]
- Liao, W.; Liu, Y.; Zhang, W. Virulence evolution, molecular mechanisms of resistance and prevalence of ST11 carbapenem-resistant Klebsiella pneumoniae in China: A review over the last 10 years. J. Glob. Antimicrob. Resist. 2020, 23, 174–180. [Google Scholar] [CrossRef]
- Zhang, R.; Liu, L.; Zhou, H.; Chan, E.W.; Li, J.; Fang, Y.; Li, Y.; Liao, K.; Chen, S. Nationwide Surveillance of Clinical Carbapenem-resistant Enterobacteriaceae (CRE) Strains in China. EBioMedicine 2017, 19, 98–106. [Google Scholar] [CrossRef]
- Pitout, J.D.D.; Peirano, G.; Kock, M.M.; Strydom, K.A.; Matsumura, Y. The Global Ascendency of OXA-48-Type Carbapenemases. Clin. Microbiol. Rev. 2019, 33, e00102-19. [Google Scholar] [CrossRef]
- Peirano, G.; Chen, L.; Kreiswirth, B.N.; Pitout, J.D.D. Emerging Antimicrobial-Resistant High-Risk Klebsiella pneumoniae Clones ST307 and ST147. Antimicrob. Agents Chemother. 2020, 64, e01148-20. [Google Scholar] [CrossRef]
- Chen, J.; Hu, C.; Wang, R.; Li, F.; Sun, G.; Yang, M.; Chu, Y. Shift in the Dominant Sequence Type of Carbapenem-Resistant Klebsiella pneumoniae Bloodstream Infection from ST11 to ST15 at a Medical Center in Northeast China, 2015–2020. Infect Drug Resist. 2021, 14, 1855–1863. [Google Scholar] [CrossRef]
- Shropshire, W.C.; Dinh, A.Q.; Earley, M.; Komarow, L.; Panesso, D.; Rydell, K.; Gomez-Villegas, S.I.; Miao, H.; Hill, C.; Chen, L.; et al. Accessory Genomes Drive Independent Spread of Carbapenem-Resistant Klebsiella pneumoniae Clonal Groups 258 and 307 in Houston, TX. mBio 2022, 13, e0049722. [Google Scholar] [CrossRef]
- Castanheira, M.; Farrell, S.E.; Wanger, A.; Rolston, K.V.; Jones, R.N.; Mendes, R.E. Rapid expansion of KPC-2-producing Klebsiella pneumoniae isolates in two Texas hospitals due to clonal spread of ST258 and ST307 lineages. Microb. Drug Resist. 2013, 19, 295–297. [Google Scholar] [CrossRef]
- Wyres, K.L.; Hawkey, J.; Hetland, M.A.K.; Fostervold, A.; Wick, R.R.; Judd, L.M.; Hamidian, M.; Howden, B.P.; Lohr, I.H.; Holt, K.E. Emergence and rapid global dissemination of CTX-M-15-associated Klebsiella pneumoniae strain ST307. J. Antimicrob. Chemother. 2019, 74, 577–581. [Google Scholar] [CrossRef] [PubMed]
- Novovic, K.; Trudic, A.; Brkic, S.; Vasiljevic, Z.; Kojic, M.; Medic, D.; Cirkovic, I.; Jovcic, B. Molecular Epidemiology of Colistin-Resistant, Carbapenemase-Producing Klebsiella pneumoniae in Serbia from 2013 to 2016. Antimicrob. Agents Chemother. 2017, 61, e02550-16. [Google Scholar] [CrossRef] [PubMed]
- Lowe, M.; Kock, M.M.; Coetzee, J.; Hoosien, E.; Peirano, G.; Strydom, K.A.; Ehlers, M.M.; Mbelle, N.M.; Shashkina, E.; Haslam, D.B.; et al. Klebsiella pneumoniae ST307 with blaOXA-181, South Africa, 2014–2016. Emerg. Infect Dis. 2019, 25, 739–747. [Google Scholar] [CrossRef] [PubMed]
- Bocanegra-Ibarias, P.; Garza-Gonzalez, E.; Morfin-Otero, R.; Barrios, H.; Villarreal-Trevino, L.; Rodriguez-Noriega, E.; Garza-Ramos, U.; Petersen-Morfin, S.; Silva-Sanchez, J. Molecular and microbiological report of a hospital outbreak of NDM-1-carrying Enterobacteriaceae in Mexico. PLoS ONE 2017, 12, e0179651. [Google Scholar] [CrossRef]
- Piazza, A.; Comandatore, F.; Romeri, F.; Brilli, M.; Dichirico, B.; Ridolfo, A.; Antona, C.; Bandi, C.; Gismondo, M.R.; Rimoldi, S.G. Identification of blaVIM-1 Gene in ST307 and ST661 Klebsiella pneumoniae Clones in Italy: Old Acquaintances for New Combinations. Microb. Drug Resist. 2019, 25, 787–790. [Google Scholar] [CrossRef]
- Carattoli, A.; Seiffert, S.N.; Schwendener, S.; Perreten, V.; Endimiani, A. Differentiation of IncL and IncM Plasmids Associated with the Spread of Clinically Relevant Antimicrobial Resistance. PLoS ONE 2015, 10, e0123063. [Google Scholar] [CrossRef]
- Boyd, S.E.; Livermore, D.M.; Hooper, D.C.; Hope, W.W. Metallo-beta-Lactamases: Structure, Function, Epidemiology, Treatment Options, and the Development Pipeline. Antimicrob. Agents Chemother. 2020, 64, e00397-20. [Google Scholar] [CrossRef]
- Villa, L.; Feudi, C.; Fortini, D.; Brisse, S.; Passet, V.; Bonura, C.; Endimiani, A.; Mammina, C.; Ocampo, A.M.; Jimenez, J.N.; et al. Diversity, virulence, and antimicrobial resistance of the KPC-producing Klebsiella pneumoniae ST307 clone. Microb. Genom. 2017, 3, e000110. [Google Scholar] [CrossRef]
- Gonzalez, L.J.; Bahr, G.; Nakashige, T.G.; Nolan, E.M.; Bonomo, R.A.; Vila, A.J. Membrane anchoring stabilizes and favors secretion of New Delhi metallo-beta-lactamase. Nat. Chem. Biol. 2016, 12, 516–522. [Google Scholar] [CrossRef]
- Yan, J.; Pu, S.; Jia, X.; Xu, X.; Yang, S.; Shi, J.; Sun, S.; Zhang, L. Multidrug Resistance Mechanisms of Carbapenem Resistant Klebsiella pneumoniae Strains Isolated in Chongqing, China. Ann. Lab. Med. 2017, 37, 398–407. [Google Scholar] [CrossRef][Green Version]
- Stapleton, P.D.; Shannon, K.P.; French, G.L. Carbapenem resistance in Escherichia coli associated with plasmid-determined CMY-4 beta-lactamase production and loss of an outer membrane protein. Antimicrob. Agents Chemother. 1999, 43, 1206–1210. [Google Scholar] [CrossRef] [PubMed]
- van Boxtel, R.; Wattel, A.A.; Arenas, J.; Goessens, W.H.; Tommassen, J. Acquisition of Carbapenem Resistance by Plasmid-Encoded-AmpC-Expressing Escherichia coli. Antimicrob. Agents Chemother. 2017, 61, e01413-16. [Google Scholar] [CrossRef] [PubMed]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing, CLSI Supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2019. [Google Scholar]
- Du, X.; He, F.; Shi, Q.; Zhao, F.; Xu, J.; Fu, Y.; Yu, Y. The Rapid Emergence of Tigecycline Resistance in blaKPC-2 Harboring Klebsiella pneumoniae, as Mediated in Vivo by Mutation in tetA During Tigecycline Treatment. Front. Microbiol. 2018, 9, 648. [Google Scholar] [CrossRef] [PubMed]
- Shi, Q.; Ye, Y.; Lan, P.; Han, X.; Quan, J.; Zhou, M.; Yu, Y.; Jiang, Y. Prevalence and Characteristics of Ceftriaxone-Resistant Salmonella in Children’s Hospital in Hangzhou, China. Front. Microbiol. 2021, 12, 764787. [Google Scholar] [CrossRef]
- Wick, R.R.; Holt, K.E. Polypolish: Short-read polishing of long-read bacterial genome assemblies. PLoS Comput. Biol. 2022, 18, e1009802. [Google Scholar] [CrossRef]
- Feldgarden, M.; Brover, V.; Haft, D.H.; Prasad, A.B.; Slotta, D.J.; Tolstoy, I.; Tyson, G.H.; Zhao, S.; Hsu, C.H.; McDermott, P.F.; et al. Validating the AMRFinder Tool and Resistance Gene Database by Using Antimicrobial Resistance Genotype-Phenotype Correlations in a Collection of Isolates. Antimicrob. Agents Chemother. 2019, 63, e00483-19. [Google Scholar] [CrossRef]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Hua, X.; Liang, Q.; Deng, M.; He, J.; Wang, M.; Hong, W.; Wu, J.; Lu, B.; Leptihn, S.; Yu, Y.; et al. BacAnt: A Combination Annotation Server for Bacterial DNA Sequences to Identify Antibiotic Resistance Genes, Integrons, and Transposable Elements. Front. Microbiol. 2021, 12, 649969. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Shi, Q.; Quan, J.; Lan, P.; Huang, D.; Zhou, J.; Jiang, Y.; Yu, Y. Prevalence and characteristics of pks gene cluster harbouring Klebsiella pneumoniae from bloodstream infection in China. Epidemiol. Infect. 2020, 148, e69. [Google Scholar] [CrossRef]

| Isolates | Department | Specimen Type | ST | MIC (mg/L) 1 | Beta-Lactamases | ||||
|---|---|---|---|---|---|---|---|---|---|
| ATM | IPM | MEM | ETP | CAZ | |||||
| CHN24001 | Department of Burns | Blood | 307 | >64 | 64 | 64 | >64 | >64 | blaNDM-12, blaOXA-48, blaCMY-6, blaCTX-M-15, blaSHV-106, blaTEM-1, blaOXA-1 |
| CHN24003 | Department of Burns | Secretion | 307 | >64 | 16 | 16 | >64 | >64 | blaNDM-12, blaOXA-48, blaCMY-6, blaSHV-106, blaTEM-1 |
| CHN24025 | Department of Burns | Secretion | 307 | >64 | 32 | 32 | >64 | >64 | blaNDM-12, blaOXA-48, blaCMY-6, blaSHV-106, blaTEM-1 |
| CHN24039 | Department of Burns | Secretion | 307 | >64 | 16 | 16 | >64 | >64 | blaNDM-12, blaOXA-48, blaCMY-6, blaSHV-106, blaTEM-1 |
| CHN24069 | Department of Hematology | Blood | 307 | >64 | 64 | 64 | >64 | >64 | blaNDM-12, blaOXA-48, blaCMY-6, blaCTX-M-15, blaSHV-106, blaTEM-1, blaOXA-1 |
| Isolates | MIC (mg/L) 1 | ||||
|---|---|---|---|---|---|
| ATM | IPM | MEM | ETP | CAZ | |
| Conjugation | |||||
| EC600 | 0.125 | 0.25 | <0.03 | <0.03 | 0.5 |
| E24039J-pOXA-48 | 0.25 | 2 | 1 | 16 | 0.25 |
| E24001J-pNDM-1 | >64 | 1 | 0.25 | 2 | >64 |
| E24025J-pNDM-1 | >64 | 1 | 0.25 | 4 | >64 |
| E24039J-pNDM-1 | >64 | 1 | 0.25 | 8 | >64 |
| Gene cloning | |||||
| DH5α | <0.03 | 0.25 | <0.03 | <0.03 | 0.125 |
| DH5α-pCR2.1K | 0.0625 | 0.25 | <0.03 | <0.03 | 0.25 |
| DH5α-pCR2.1K::NDM-1 | 0.0625 | 32 | 64 | 64 | >64 |
| DH5α-pCR2.1K::NDM-1(T) 2 | 0.125 | 0.25 | <0.03 | <0.03 | 0.5 |
| DH5α-pCR2.1K::CMY-6 | >64 | 1 | 0.25 | 2 | >64 |
| DH5α-pCR2.1K::OXA-48 | 0.25 | 2 | 0.5 | 16 | 0.5 |
| DH5α-pCR2.1K::NDM-1(T) 2::CMY-6 | >64 | 0.5 | 0.25 | 2 | >64 |
| DH5α-pCR2.1K::NDM-1(T) 2::OXA-48 | 0.25 | 2 | 1 | 8 | 0.5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shi, Q.; Han, X.; Huang, Q.; Meng, Y.; Zhang, P.; Wang, Z.; Hu, H.; Jiang, Y.; Du, X.; Yu, Y. The Genetic Characteristics and Carbapenem Resistance Mechanism of ST307 Klebsiella pneumoniae Coharbouring blaCMY-6, blaOXA-48, and a Truncated blaNDM-1. Antibiotics 2022, 11, 1616. https://doi.org/10.3390/antibiotics11111616
Shi Q, Han X, Huang Q, Meng Y, Zhang P, Wang Z, Hu H, Jiang Y, Du X, Yu Y. The Genetic Characteristics and Carbapenem Resistance Mechanism of ST307 Klebsiella pneumoniae Coharbouring blaCMY-6, blaOXA-48, and a Truncated blaNDM-1. Antibiotics. 2022; 11(11):1616. https://doi.org/10.3390/antibiotics11111616
Chicago/Turabian StyleShi, Qiucheng, Xinhong Han, Qin Huang, Yan Meng, Ping Zhang, Zhengan Wang, Huangdu Hu, Yan Jiang, Xiaoxing Du, and Yunsong Yu. 2022. "The Genetic Characteristics and Carbapenem Resistance Mechanism of ST307 Klebsiella pneumoniae Coharbouring blaCMY-6, blaOXA-48, and a Truncated blaNDM-1" Antibiotics 11, no. 11: 1616. https://doi.org/10.3390/antibiotics11111616
APA StyleShi, Q., Han, X., Huang, Q., Meng, Y., Zhang, P., Wang, Z., Hu, H., Jiang, Y., Du, X., & Yu, Y. (2022). The Genetic Characteristics and Carbapenem Resistance Mechanism of ST307 Klebsiella pneumoniae Coharbouring blaCMY-6, blaOXA-48, and a Truncated blaNDM-1. Antibiotics, 11(11), 1616. https://doi.org/10.3390/antibiotics11111616






