Whole-Genome Sequencing and Molecular Analysis of Ceftazidime–Avibactam-Resistant KPC-Producing Klebsiella pneumoniae from Intestinal Colonization in Elderly Patients
Abstract
:1. Introduction
2. Results
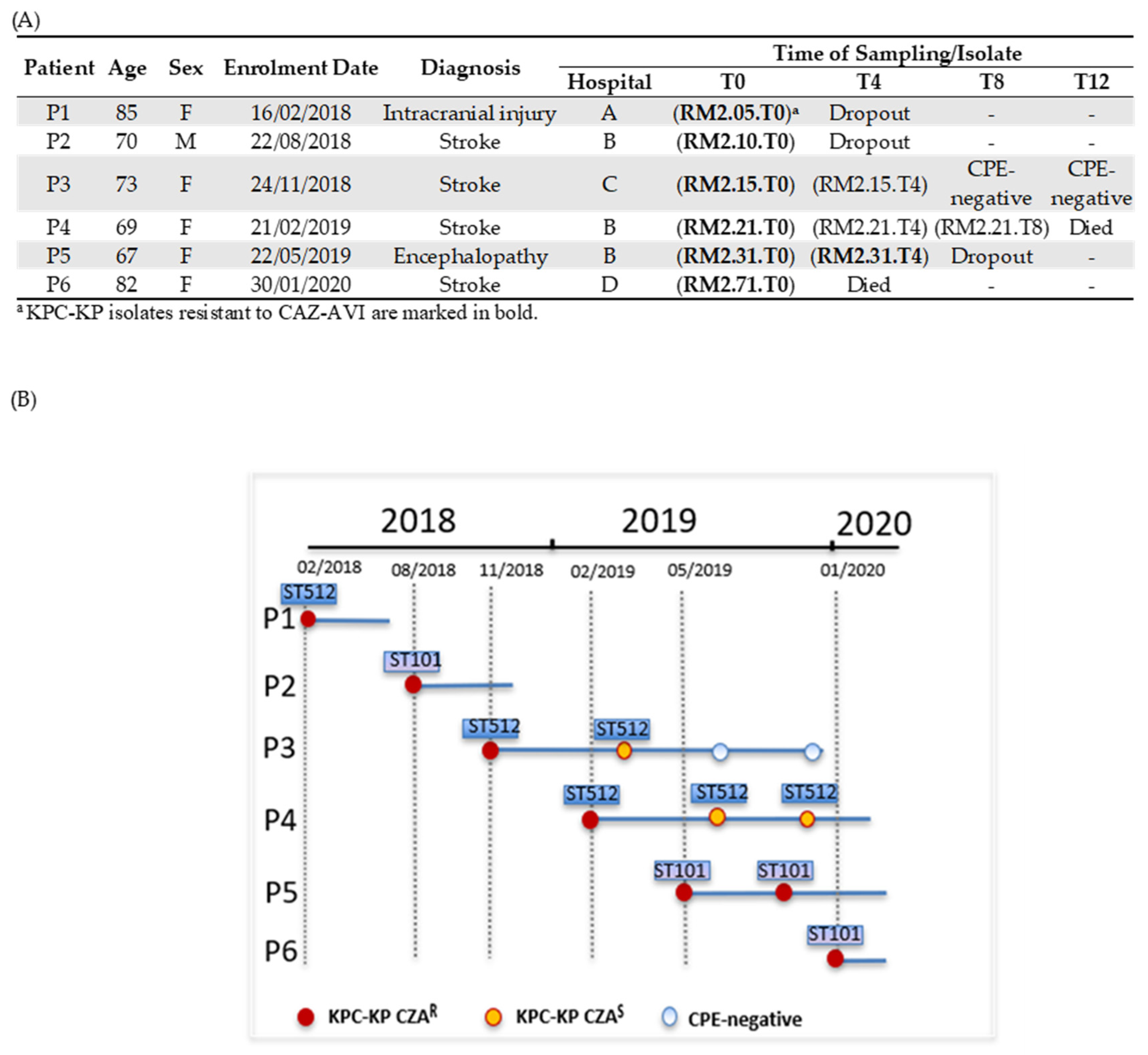
2.1. Patients Colonized by KPC-KP Resistant to CAZ-AVI and KPC-KP Isolates
2.2. Whole-Genome Sequencing and In Silico Analysis
3. Discussion
4. Materials and Methods
4.1. Bacterial Strain Identification and Antimicrobial Susceptibility Testing (AST)
4.2. Whole-Genome Sequencing and In Silico Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- David, S.; Reuter, S.; Harris, S.R.; Glasner, C.; Feltwell, T.; Argimon, S.; Abudahab, K.; Goater, R.; Giani, T.; Errico, G.; et al. Epidemic of carbapenem-resistant Klebsiella pneumoniae in Europe is driven by nosocomial spread. Nat. Microbiol. 2019, 4, 1919–1929. [Google Scholar] [CrossRef] [PubMed]
- Lerner, A.; Adler, A.; Abu-Hanna, J.; Percia, S.C.; Matalon, M.K.; Carmeli, Y. Spread of KPC-producing carbapenem-resistant Enterobacteriaceae: The importance of super-spreaders and rectal KPC concentration. Clin. Microbiol. Infect. 2015, 21, 470.e1–470.e7. [Google Scholar] [CrossRef] [Green Version]
- Tumbarello, M.; Viale, P.; Viscoli, C.; Trecarichi, E.M.; Tumietto, F.; Marchese, A.; Spanu, T.; Ambretti, S.; Ginocchio, F.; Cristini, F.; et al. Predictors of Mortality in Bloodstream Infections Caused by Klebsiella pneumoniae Carbapenemase-Producing K. pneumoniae: Importance of Combination Therapy. Clin. Infect. Dis. 2012, 55, 943–950. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Pilato, V.; Errico, G.; Monaco, M.; Giani, T.; Del Grosso, M.; Antonelli, A.; David, S.; Lindh, E.; Camilli, R.; Aanensen, D.M.; et al. The changing epidemiology of car-bapenemase-producing Klebsiella pneumoniae in Italy: Toward polyclonal evolution with emergence of high-risk lineages. J. Antimicrob. Chemother. 2021, 76, 355–361. [Google Scholar] [CrossRef] [PubMed]
- Porreca, A.M.; Sullivan, K.V.; Gallagher, J.C. The Epidemiology, Evolution, and Treatment of KPC-Producing Organisms. Curr. Infect. Dis. Rep. 2018, 20, 13. [Google Scholar] [CrossRef]
- Poulakou, G.; Bassetti, M.; Righi, E.; Dimopoulos, G. Current and future treatment options for infections caused by multidrug-resistant Gram-negative pathogens. Future Microbiol. 2014, 9, 1053–1069. [Google Scholar] [CrossRef]
- Liu, Y.-Y.; Wang, Y.; Walsh, T.R.; Yi, L.-X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Pogue, J.; Bonomo, R.A.; Kaye, K.S. Ceftazidime/Avibactam, Meropenem/Vaborbactam, or Both? Clinical and Formulary Considerations. Clin. Infect. Dis. 2019, 68, 519–524. [Google Scholar] [CrossRef] [Green Version]
- Bush, K.; Bradford, P.A. Interplay between β-lactamases and new β-lactamase inhibitors. Nat. Rev. Microbiol. 2019, 17, 295–306. [Google Scholar] [CrossRef]
- Thaden, J.T.; Pogue, J.M.; Kaye, K.S. Role of newer and re-emerging older agents in the treatment of infections caused by car-bapenem-resistant Enterobacteriaceae. Virulence 2017, 8, 403–416. [Google Scholar] [CrossRef] [Green Version]
- Shields, R.K.; Chen, L.; Cheng, S.; Chavda, K.D.; Press, E.G.; Snyder, A.; Pandey, R.; Doi, Y.; Kreiswirth, B.N.; Nguyen, M.H.; et al. Emergence of Ceftazidime-Avibactam Resistance Due to Plasmid-Borne bla KPC-3 Mutations during Treatment of Carbapenem-Resistant Klebsiella pneumoniae Infections. Antimicrob. Agents Chemother. 2016, 61, e02097-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- David, S.; Wong, J.L.C.; Sanchez-Garrido, J.; Kwong, H.-S.; Low, W.W.; Morecchiato, F.; Giani, T.; Rossolini, G.M.; Brett, S.J.; Clements, A.; et al. Widespread emergence of OmpK36 loop 3 insertions among multidrug-resistant clones of Klebsiella pneumoniae. PLoS Pathog. 2022, 18, e1010334. [Google Scholar] [CrossRef]
- Sun, D.; Rubio-Aparicio, D.; Nelson, K.; Dudley, M.N.; Lomovskaya, O. Meropenem-Vaborbactam Resistance Selection, Resistance Prevention, and Molecular Mechanisms in Mutants of KPC-Producing Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2017, 61, e01694-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gaibani, P.; Giani, T.; Bovo, F.; Lombardo, D.; Amadesi, S.; Lazzarotto, T.; Coppi, M.; Rossolini, G.M.; Ambretti, S. Resistance to Ceftazidime/Avibactam, Meropenem/Vaborbactam and Imipenem/Relebactam in Gram-Negative MDR Bacilli: Molecular Mechanisms and Susceptibility Testing. Antibiotics 2022, 11, 628. [Google Scholar] [CrossRef]
- Schechner, V.; Kotlovsky, T.; Kazma, M.; Mishali, H.; Schwartz, D.; Navon-Venezia, S.; Schwaber, M.; Carmeli, Y. Asymptomatic rectal carriage of blaKPC producing carbapenem-resistant Enterobacteriaceae: Who is prone to become clinically infected? Clin. Microbiol. Infect. 2013, 19, 451–456. [Google Scholar] [CrossRef] [Green Version]
- Tinelli, M.; Rossini, A.; Scudeller, L.; Zabzuni, D.; Errico, G.; Fogato, E.; D’Angelo, R.; Silverj, F.G.; Cesana, E.; Bergamaschini, L.C.; et al. Dynamics of carbapenemase-producing Enterobacterales intestinal colonisation in the elderly population after hospital discharge, Italy, 2018–2020. Int. J. Antimicrob. Agents 2022, 59, 106594. [Google Scholar] [CrossRef]
- Lim, C.J.; Cheng, A.C.; Kennon, J.; Spelman, D.; Hale, D.; Melican, G.; Sidjabat, H.E.; Paterson, D.L.; Kong, D.C.M.; Peleg, A.Y. Prevalence of multidrug-resistant organisms and risk factors for carriage in long-term care facilities: A nested case-control study. J. Antimicrob. Chemother. 2014, 69, 1972–1980. [Google Scholar] [CrossRef]
- Tinelli, M.; Tiseo, G.; Falcone, M. ESCMID Study Group for Infections in the Elderly. Prevention of the spread of multi-drug-resistant organisms in nursing homes. Aging Clin. Exp. Res. 2021, 33, 679–687. [Google Scholar] [CrossRef]
- WHO Regional Office for Europe/European Centre for Disease Prevention and Control. Antimicrobial Resistance Surveillance in Europe 2022–2020 Data; WHO Regional Office for Europe: Copenhagen, Denmark, 2022. [Google Scholar]
- Roe, C.C.; Vazquez, A.J.; Esposito, E.P.; Zarrilli, R.; Sahl, J.W. Diversity, Virulence, and Antimicrobial Resistance in Isolates From the Newly Emerging Klebsiella pneumoniae ST101 Lineage. Front Microbiol. 2019, 10, 542. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oteo, J.; Pérez-Vázquez, M.; Bautista, V.; Ortega, A.; Zamarrón, P.; Saez, D.; Fernández-Romero, S.; Lara, N.; Ramiro, R.; Aracil, B.; et al. The spread of KPC-producing Enterobacteriaceae in Spain: WGS analysis of the emerging high-risk clones of Klebsiella pneumoniae ST11/KPC-2, ST101/KPC-2 and ST512/KPC-3. J. Antimicrob. Chemother. 2016, 71, 3392–3399. [Google Scholar] [CrossRef] [Green Version]
- Kiratisin, P.; Kazmierczak, K.; Stone, G.G. In vitro activity of ceftazidime/avibactam and comparators against carbapenemase-producing Enterobacterales and Pseudomonas aeruginosa isolates collected globally between 2016 and 2018. J. Glob. Antimicrob. Resist. 2021, 27, 132–141. [Google Scholar] [CrossRef]
- Winkler, M.L.; Papp-Wallace, K.M.; Bonomo, R.A. Activity of ceftazidime/avibactam against isogenic strains of Escherichia coli containing KPC and SHV b-lactamases with single amino acid substitutions in the X-loop. J. Antimicrob. Chemother. 2015, 70, 2279–2286. [Google Scholar] [CrossRef] [Green Version]
- Moreira, N.K.; Caierão, J. Ceftazidime-avibactam: Are we safe from class A carbapenemase producers’ infections? Folia Microbiol. 2021, 66, 879–896. [Google Scholar] [CrossRef] [PubMed]
- Nicola, F.; Cejas, D.; González-Espinosa, F.; Relloso, S.; Herrera, F.; Bonvehí, P.; Smayevsky, J.; Figueroa-Espinosa, R.; Gutkind, G.; Radice, M. Outbreak of Klebsiella pneumoniae ST11 Resistant to Ceftazidime-Avibactam Producing KPC-31 and the Novel Variant KPC-115 during COVID-19 Pandemic in Argentina. Microbiol. Spectr. 2022, 10, e0373322. [Google Scholar] [CrossRef] [PubMed]
- Hernández-García, M.; Castillo-Polo, J.A.; Cordero, D.G.; Pérez-Viso, B.; García-Castillo, M.; de la Fuente, J.S.; Morosini, M.I.; Cantón, R.; Ruiz-Garbajosa, P. Impact of Ceftazidime-Avibactam Treatment in the Emergence of Novel KPC Variants in the ST307-Klebsiella pneumoniae High-Risk Clone and Consequences for Their Routine Detection. J. Clin. Microbiol. 2022, 60, e0224521. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Ding, B.; Ye, M.; Wang, P.; Bi, Y.; Wu, S.; Xu, X.; Guo, Q.; Wang, M. High ceftazidime hydrolysis activity and porin OmpK35 deficiency contribute to the decreased susceptibility to ceftazidime/avibactam in KPC-producing Klebsiella pneu-moniae. J. Antimicrob. Chemother. 2017, 72, 1930–1936. [Google Scholar] [CrossRef] [Green Version]
- Castanheira, M.; Doyle, T.B.; Hubler, C.; Sader, H.S.; Mendes, R.E. Ceftazidime-avibactam activity against a challenge set of carbapenem-resistant Enterobacterales: Ompk36 L3 alterations and β-lactamases with ceftazidime hydrolytic activity lead to elevated MIC values. Int. J. Antimicrob. Agents 2020, 56, 106011. [Google Scholar] [CrossRef] [PubMed]
- Findlay, J.; Poirel, L.; Juhas, M.; Nordmann, P. KPC-Mediated Resistance to Ceftazidime-Avibactam and Collateral Effects in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2021, 65, e0089021. [Google Scholar] [CrossRef]
- Di Pilato, V.; Principe, L.; Andriani, L.; Aiezza, N.; Coppi, M.; Ricci, S.; Giani, T.; Luzzaro, F.; Rossolini, G.M. Deciphering variable resistance to novel carbapenem-based β-lactamase inhibitor combinations in a multi-clonal outbreak caused by Klebsiella pneumoniae carbapenemase (KPC)-producing Klebsiella pneumoniae resistant to ceftazidime/avibactam. Clin. Microbiol. Infect. 2023, 29, 537.e1–537.e8. [Google Scholar] [CrossRef]
- The European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters. Version 13.0. 2023. Available online: http://www.eucast.org (accessed on 18 February 2023).
- Kaas, R.S.; Leekitcharoenphon, P.; Aarestrup, F.M.; Lund, O. Solving the Problem of Comparing Whole Bacterial Genomes across Different Sequencing Platforms. PLoS ONE 2014, 9, e104984. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows—Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Porins | MIC (mg/L) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Patient | Isolate | ST | Carba | OmpK35 * | OmpK36 | CAZ-AVI | MEM | MVB | IMI | IMR | |||||
| P1 | RM2.05.T0 | 512 | KPC-3 | AA89 stop | GD ins | 16/4 | R | >64 | R | 1/8 | S | >8 | R | 0.5/4 | S |
| P2 | RM2.10.T0 | 101 | KPC-31 | AA63 stop | TD ins | >64/4 | R | 2 | S | 0.25/8 | S | ≤1 | S | 0.12/4 | S |
| P3 | RM2.15.T0 | 512 | KPC-31 | AA89 stop | GD ins | 64/4 | R | 2 | S | 1/8 | S | ≤1 | S | 0.12/4 | S |
| RM2.15.T4 | 512 | KPC-3 | AA89 stop | GD ins | 8/4 | S | >64 | R | 0.25/8 | S | >8 | R | 0.25/4 | S | |
| P4 | RM2.21.T0 | 512 | KPC-3 | AA89 stop | GD ins | 16/4 | R | >64 | R | 8/8 | S | >8 | R | 0.25/4 | S |
| RM2.21.T4 | 512 | KPC-3 | AA89 stop | GD ins | 4/4 | S | 32 | R | ≤0.06/8 | S | 4 | I | 0.12/4 | S | |
| RM2.21.T8 | 512 | KPC-3 | AA89 stop | GD ins | 2/4 | S | >64 | R | 0.5/8 | S | >8 | R | 0.12/4 | S | |
| P5 | RM2.31.T0 | 101 | KPC-46 | AA63 stop | TD ins | 32/4 | R | 1 | S | 0.25/8 | S | ≤1 | S | 0.12/4 | S |
| RM2.31.T4 | 101 | KPC-46 | AA63 stop | TD ins | 64/4 | R | 1 | S | 0.25/8 | S | ≤1 | S | 0.12/4 | S | |
| P6 | RM2.71.T0 | 101 | KPC-46 | AA63 stop | TD ins | 32/4 | R | 1 | S | 0.25/8 | S | ≤1 | S | 0.12/4 | S |
| Patient | Isolate | ST | β-Lactamases | Aminoglycoside Modifying Enzyme | Fluoroquinolones | MLS | Phenicols | RIF | SUL | TET | TMT |
|---|---|---|---|---|---|---|---|---|---|---|---|
| P1 | RM2.05.T0 | 512 | KPC-3, TEM-1D, SHV-11 | aac(6′)-Ib, aadA2, aph(3′)-Ia | GyrA-S83I, ParC-S80I | mphA | catA1 | - | sul1 | - | dfrA12 |
| P2 | RM2.10.T0 | 101 | KPC-31, SHV-1 | armA | GyrA-S83Y/D87N ParC-S80I | mphE, msrE | - | - | - | - | - |
| P3 | RM2.15.T0 | 512 | KPC-31, SHV-11 | aac(6′)-Ib’,aadA2, aph(3′)-Ia | GyrA-S83I, ParC-S80I | mphA | catA1 | - | sul1 | - | dfrA12 |
| RM2.15.T4 | 512 | KPC-3, SHV-11 | aac(6′)-Ib’ | GyrA-S83I, ParC-S80I | - | - | - | - | - | - | |
| P4 | RM2.21.T0 | 512 | KPC-3, CMY-16, OXA-10, SHV11 | aadA1, aadA2, aac(6′)-Ib3, strA | GyrA-S83I, ParC-S80I | mphA | catA1, cmlA5, floR | arr2 | sul1, sul2 | tetA | dfrA12, dfrA14 |
| RM2.21.T4 | 512 | KPC-3, CMY-16, OXA-10, SHV-11, TEM-1D | aadA1, aadA2, aac(6′)-Ib3,strA | GyrA-83I, ParC-80I | - | cmlA5, floR | arr2 | sul1, sul2 | tetA | dfrA14 | |
| RM2.21.T8 | 512 | KPC-3, CMY-16, OXA-10, SHV-11, TEM-1D | aadA1, aadA2, aac(6′)-Ib3, strA | GyrA-83I, ParC-80I | - | cmlA5, floR | arr2 | sul1, sul2 | tetA | dfrA14 | |
| P5 | RM2.31.T0 | 101 | KPC-46, SHV-1 | armA | GyrA-S83Y/D87N ParC-S80I | mphE, msrE | - | - | - | - | - |
| RM2.31.T4 | 101 | KPC-46, SHV-1 | armA | GyrA-S83Y/D87N ParC-S80I | mphE, msrE | - | - | - | - | - | |
| P6 | RM2.71.T0 | 101 | KPC-46, SHV-1 | - | GyrA-S83Y/D87NParC-S80I | mphE, msrE | - | - | - | - | - |
| Patient | Isolate | ST | KLocus | Virulence Factor | Plasmid | pMLST |
|---|---|---|---|---|---|---|
| P1 | RM2.05.T0 | 512 | KL107 (wzi-154) | - | IncFIB, IncFII, IncX3,ColRNAI | IncF [K1:A-:B-] |
| P2 | RM2.10.T0 | 101 | KL17 (wzi-137) | ybt9/ ICEKp3 | IncFIA, IncFIB, IncFII, IncR,Col440,ColRNAI | IncF [K1:A13:B-] |
| P3 | RM2.15.T0 | 512 | KL107 (wzi-154) | - | IncFIB, IncFII, IncX3 | IncF [K2:A-:B-] |
| RM2.15.T4 | 512 | KL107 (wzi-154) | - | IncFIB, IncFII, IncX3, ColRNAI | IncF [K2:A-:B-] | |
| P4 | RM2.21.T0 | 512 | KL107 (wzi-154) | - | IncFIB, IncFII, IncC IncX3 | IncF [K2:A-:B-] IncC [C3] |
| RM2.21.T4 | 512 | KL107 (wzi-154) | - | IncFIB, IncFII, IncC IncX3 | IncF [K2:A-:B-] IncC [C3] | |
| RM2.21.T8 | 512 | KL107 (wzi-154) | - | IncFIB, IncFII, IncC IncX3 | IncF [K2:A-:B-] IncC [C3] | |
| P5 | RM2.31.T0 | 101 | KL17 (wzi-137) | ybt9/ ICEKp3 | IncFIA, IncFIB, IncFII,IncR,Col440,ColRNAI | IncF [K2:A13:B-] |
| RM2.31.T4 | 101 | KL17 (wzi-137) | ybt9/ ICEKp3 | IncFIA, IncFIB, IncFII,IncR,Col440,ColRNAI | IncF [K2:A13:B-] | |
| P6 | RM2.71.T0 | 101 | KL17 (wzi-137) | ybt9/ ICEKp3 | IncFIA, IncFIB, IncFII,IncR,Col440,ColRNAI | IncF [K2:A13:B-] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Errico, G.; Del Grosso, M.; Pagnotta, M.; Marra, M.; Carollo, M.; Cerquetti, M.; Fogato, E.; Cesana, E.; Gentiloni Silverj, F.; Zabzuni, D.; et al. Whole-Genome Sequencing and Molecular Analysis of Ceftazidime–Avibactam-Resistant KPC-Producing Klebsiella pneumoniae from Intestinal Colonization in Elderly Patients. Antibiotics 2023, 12, 1282. https://doi.org/10.3390/antibiotics12081282
Errico G, Del Grosso M, Pagnotta M, Marra M, Carollo M, Cerquetti M, Fogato E, Cesana E, Gentiloni Silverj F, Zabzuni D, et al. Whole-Genome Sequencing and Molecular Analysis of Ceftazidime–Avibactam-Resistant KPC-Producing Klebsiella pneumoniae from Intestinal Colonization in Elderly Patients. Antibiotics. 2023; 12(8):1282. https://doi.org/10.3390/antibiotics12081282
Chicago/Turabian StyleErrico, Giulia, Maria Del Grosso, Michela Pagnotta, Manuela Marra, Maria Carollo, Marina Cerquetti, Elena Fogato, Elisabetta Cesana, Flaminia Gentiloni Silverj, Dorjan Zabzuni, and et al. 2023. "Whole-Genome Sequencing and Molecular Analysis of Ceftazidime–Avibactam-Resistant KPC-Producing Klebsiella pneumoniae from Intestinal Colonization in Elderly Patients" Antibiotics 12, no. 8: 1282. https://doi.org/10.3390/antibiotics12081282







