Abstract
This study aimed to explore the molecular epidemiology of Staphylococcus aureus isolated from patients on mechanical ventilation and the participation of virulence factors in the development of ventilator-associated pneumonia (VAP). A prospective cohort study was conducted on patients under mechanical ventilation, with periodic visits for the collection of tracheal aspirates and clinical data. The S. aureus isolates were analyzed regarding resistance profile, virulence, expression of protein A and alpha-toxin using Western blot, clonal profile using PFGE, sequence type using MLST, and characterization and quantification of phenol-soluble modulins. Among the 270 patients in the study, 51 S. aureus strains were isolated from 47 patients. The incidence density of S. aureus and MRSA VAP was 2.35/1000 and 1.96/1000 ventilator days, respectively; of these, 45% (n = 5) were resistant to oxacillin, with 100% (n = 5) harboring SCCmec types II and IV. The most frequent among the tested virulence factors were icaA, hla, and hld. The clonal profile showed a predominance of sequence types originating from the community. Risk factors for VAP were the presence of solid tumors and the sea gene. In conclusion, patient-related risk factors, together with microbiological factors, are involved in the development of S. aureus VAP, which is caused by the patient’s own strains.
1. Introduction
Ventilator-associated pneumonia (VAP) is a type of nosocomial pneumonia that affects 9 to 27% of patients subjected to mechanical ventilation [1,2]. Mortality attributed to VAP varies according to the risk group studied, with surgical and long-stay patients being more likely to die from the disease [3]. Thus, the epidemiological and molecular study of factors related to the development of VAP is of great importance since it can help identify risk groups and develop new interventions.
The most common etiological agents of VAP are aerobic Gram-positive bacteria such as Staphylococcus aureus and Gram-negative bacteria such as Pseudomonas aeruginosa, Acinetobacter baumannii, and Klebsiella pneumoniae. S. aureus is the main causative agent of nosocomial pneumonia and the second most common agent associated with all nosocomial infections [4,5].
Pneumonias caused by methicillin-resistant S. aureus (MRSA) are generally difficult to treat and are associated with high rates of treatment failure [6]. MRSA strains carry the mecA gene, which confers resistance to beta-lactam antibiotics and is located on a mobile genetic element (staphylococcal cassette chromosome mec—SCCmec) of variable size and genetic composition. SCCmec typing is widely applied in epidemiological studies and can be useful for clinical purposes. There are currently 14 SCCmec types (I to XIV) [7]. Regarding their epidemiological distribution, types I to III are mostly found in hospitals, and the other types are predominantly community-associated. Haque et al. [6] found that SCCmec type II in conjunction with host factors is associated with mortality, high vancomycin minimum inhibitory concentrations (MIC), and clinical treatment failure in patients with nosocomial pneumonia caused by MRSA. This finding highlights the importance of epidemiological–molecular research on these microorganisms associated with VAP.
Staphylococcus aureus has a great ability to invade host tissues due to the production of different virulence factors. Several studies have addressed the role that each virulence factor plays in the severity of pneumonia and/or lung injury, which range from the investigation of bacterial cytotoxicity [6,8] to exploring the role of each virulence gene. Among the virulence factors that play an important role in pneumonia, alpha-hemolysin or alpha-toxin (Hla) is one of the most prominent cytotoxins that target erythrocytes, epithelial and endothelial cells, T cells, neutrophils, monocytes, and macrophages. Alpha-toxin is an amphipathic, water-soluble, pore-forming molecule that binds primarily to lipids present on the host cell membrane [9,10]. The main function of alpha-toxin is the causing osmotic swelling, rupture, lysis, and subsequently, cell death. It is highly inflammatory, causing pulmonary congestion [11].
Other factors, such as staphylococcal enterotoxins (SEs) and toxic shock syndrome toxin 1 (TSST-1), are superantigens involved in diverse human infections. To explore their participation in pneumonia, experimental studies using rabbits have shown that intrapulmonary instillation of SEB and SEC induces hemorrhage in the pulmonary tissue, causing symptoms of respiratory distress and lethal toxic shock syndrome. The administration of strains carrying the genes for the same enterotoxins resulted in pulmonary pathology and lethality similar to the group that received the purified toxins [12,13].
The participation of Panton–Valentine leukocidin (PVL) in pneumonia has been extensively reported in the literature, especially in pneumonia caused by community-acquired S. aureus strains [14,15]. PVL is a pore-forming toxin that targets neutrophils and mitochondria. It induces cell lysis, thereby releasing proinflammatory agents into the extracellular environment that cause tissue necrosis. When present in the lung, PVL is associated with necrotizing pneumonia [16]. Some strains that carry the PVL gene also harbor genes encoding exfoliative toxins (eta, etb, or etd) [17]. The role of exfoliative toxins in skin infections is well understood, but little is known about their participation in pneumonia; hence, research on S. aureus-associated pneumonia is needed.
Another important virulence factor in the pathogenesis of S. aureus pneumonia is biofilm formation, particularly in the area of the endotracheal tube. The icaADBC-mediated polysaccharide production is an important mechanism for biofilm formation and contributes to the early growth of bacteria [18]. The bap gene of S. aureus encodes a surface protein Bap (biofilm-associated protein) containing 2276 amino acids. Bap was identified as the main determinant of successful surface adhesion and intercellular adhesion during biofilm formation [18].
In addition to bacterial factors, the involvement of host-related factors in the development of VAP has been widely studied. The use of antimicrobials and antacids, opting for tracheostomy, supine position, parenteral nutrition, AIDS, male gender, lung disease, coma, and trauma are risk factors for the development of VAP [19].
In view of the above considerations, our study aimed to explore the molecular epidemiology of S. aureus isolated from patients on mechanical ventilation and to investigate the participation of virulence and patient-related factors in the development of VAP.
2. Results
2.1. Microbial Isolation and Identification
Among 270 patients seen over the study period (November 2011 to August 2013), 51 S. aureus strains were isolated from 47 patients. Eleven strains were recovered from patients with VAP, and more than one isolate was identified in two patients, totaling thirteen S. aureus isolates, including 55% methicillin-susceptible S. aureus (MSSA) and 45% MRSA. The total incidence density of VAP was 12.9/1000 ventilator days, with 4.31/1000 ventilator days being caused by S. aureus (2.35/1000 ventilator days for MSSA and 1.96/1000 ventilator days for MRSA), 1.17/1000 ventilator days by Acinetobacter baumannii, 1.17/1000 ventilator days by Enterobacter spp., 0.78/1000 ventilator days by Klebsiella pneumoniae, 0.78/1000 ventilator days by Proteus mirabilis, and 1.56/1000 ventilator days by other agents. Identification of the etiological agent was not possible in some cases of VAP, which corresponded to an incidence density of 2.74/1000 ventilator days.
2.2. Detection of mecA Gene, Cassette Chromosome Typing and Antimicrobial Resistance
The mecA gene was detected in 45% (23/51) S. aureus isolates, 21% (5/23) isolated from patients with VAP. The most common cassette type among all isolates was SCCmec type II (n = 18; 78%), followed by type IV (n = 4; 17%) and type I (n = 1; 4%). Among the S. aureus isolates associated with VAP, two carried SCCmec type II and three carried SCCmec type IV.
Determination of MIC in the S. aureus isolates using the E-test revealed that none of the isolates had vancomycin MIC higher than 1.5 µg/mL. All isolates were susceptible to quinupristin-dalfopristin, tigecycline, linezolid, and daptomycin by the E-test. The disk diffusion method showed that 81% of the isolates were resistant to penicillin, 58% to erythromycin, 54% to clindamycin, 39% to oxacillin, and 39% to cefoxitin; 13% were positive in the D test. Among the mecA gene-positive isolates, 78% (18/23) were also resistant to erythromycin and clindamycin, showing a multidrug resistance profile. None of the MRSA strains were positive in the D test. All isolates were susceptible to gentamicin. Table 1 shows the MIC50 and MIC90 of the drugs. The data sheet with all the data for all 51 S. aureus strains isolated in the study can be seen in the Supplementary Materials Table S1.

Table 1.
Determination of minimum inhibitory concentration in S. aureus using the E-test.
2.3. Virulence Factors of S. aureus Isolates
Among the virulence factors studied, genes associated with biofilm formation (icaA) and hemolysins hla and hld were the most frequent, followed by the icaD and hlb genes (Table 2). There was no statistically significant association between the presence of the mecA gene and virulence factors. A slightly higher frequency of the sea, sec, icaC, and hlb genes was observed in mecA gene-positive isolates compared to negative isolates, but the association was not statistically significant.

Table 2.
Frequency of virulence genes and association with the presence of the mecA gene.
2.4. Clonal Characterization of S. aureus Isolates
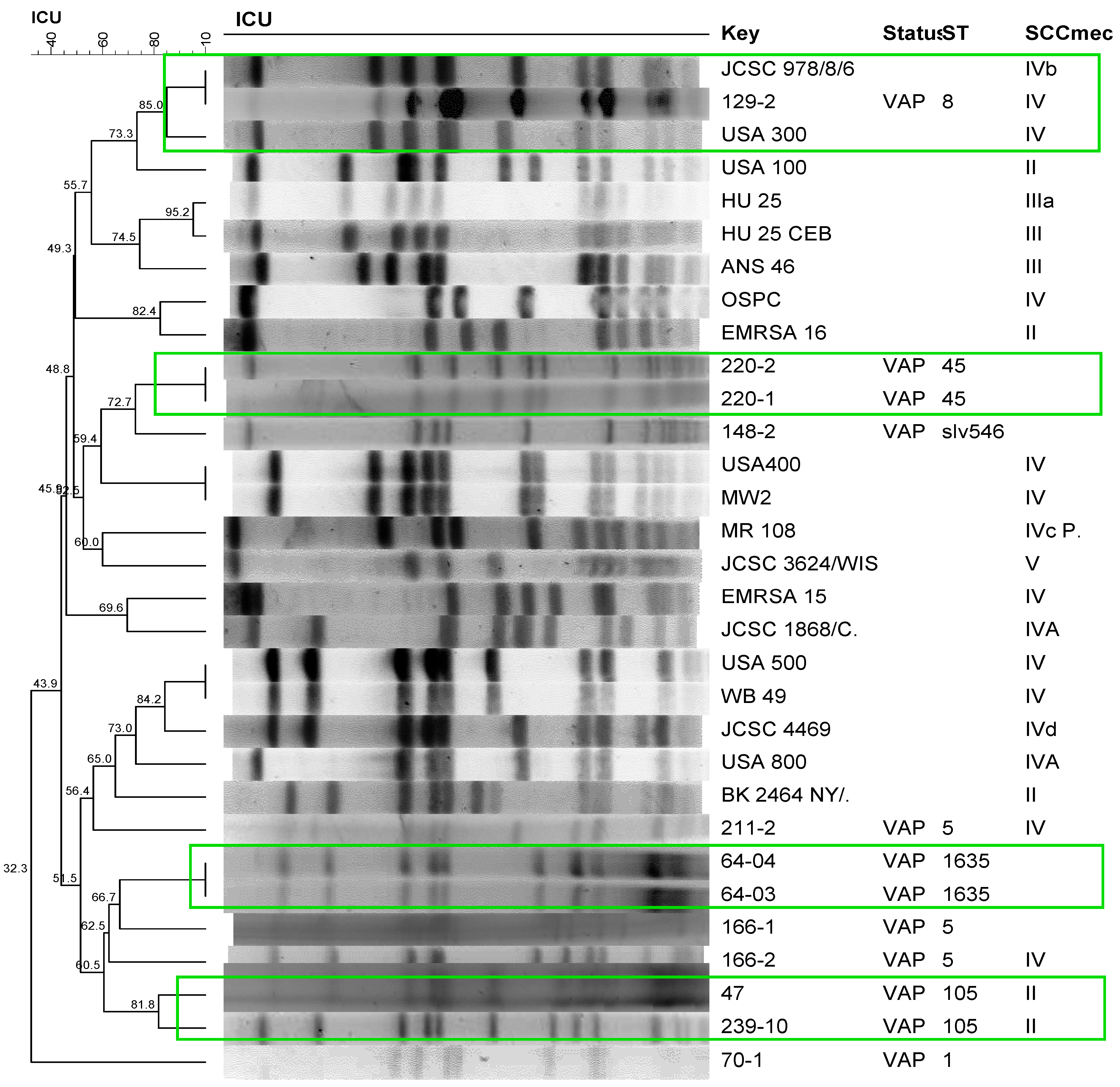
Clonal profile analysis of S. aureus isolated from patients with VAP using PFGE revealed a polyclonal profile (Figure 1). The most frequent STs were ST105, ST5, and ST398. The only strains that formed clusters were those isolated from the same patient during a period prior to the development of pneumonia, with the exception of one patient (166), who exhibited a different profile in the second sampling in which the isolate carried SCCmec IV but belonged to the same ST5 as the first isolate (Figure 1). Comparison of these isolates with international reference strains revealed the clustering of one S. aureus strain isolated in this study (129-2) with the international clone USA 300, with 85% similarity. Two isolates (47 and 239-10) formed a cluster with 81.8% similarity, which belonged to the same ST105 and harbored SCCmec II. The molecular characteristics, detection, and quantification of phenol-soluble modulins, alpha-toxin, and protein A of the isolates are shown in Table 3 and Table 4.
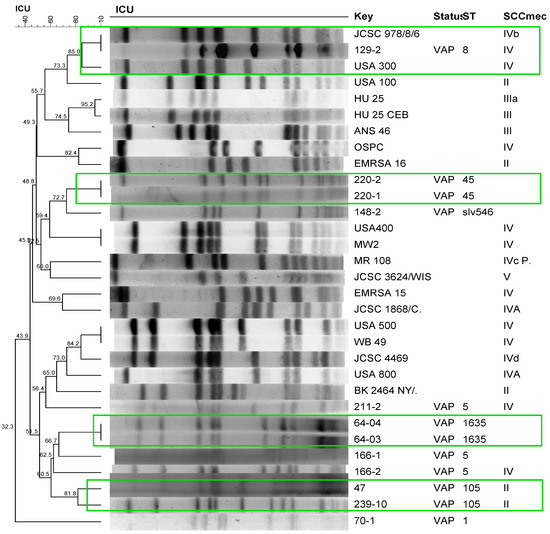

Figure 1.
Dendrogram of the PFGE of S. aureus strains isolated from patients with ventilator-associated pneumonia (VAP) generated by Dice analysis/UPGMA (BioNumerics, Applied Maths) and their molecular characterization by SCCmec typing and MLST. Green frame: Isolates showing >80% similarity after digestion with SmaI. International clones used as controls: JCSC 978/8/6, USA300, USA100, HU25, HU25CEB, ANS46, OSPC, EMRSA16, USA400, MW2, MR108, JCSC 3624/WIS, EMRSA15, JCSC1868/C, USA500, WB49, JCSC4469, USA800, BK2464NY/.

Table 3.
Characteristics of S. aureus strains isolated from patients with ventilator-associated pneumonia.

Table 4.
Detection and quantification of phenol-soluble modulins (PSM) in S. aureus strains isolated from patients with ventilator-associated pneumonia.
2.5. Risk Factors
The univariate Poisson regression model for analysis of the association of host and microbiological factors with the development of S. aureus VAP revealed a positive association with the sea and icaC genes and a negative association with central nervous system disease. However, in multivariate analysis, the only independent factors that were positively associated with S. aureus VAP were the presence of the sea gene and the presence of a solid tumor (Table 5).

Table 5.
Poisson regression model for analysis of the association of host and microbiological factors with the development of ventilator-associated pneumonia caused by S. aureus.
Regarding the death of patients with VAP caused by S. aureus as an outcome, the univariate Poisson model revealed a positive association between liver disease and the presence of the sea gene, while the presence of a central venous catheter was negatively associated. However, in multivariate analysis, the presence of the sea and hlb genes were the only independent factors that were positively associated with death (Table 6).

Table 6.
Poisson regression model for analysis of the association of host and microbiological factors with the death of patients due to ventilator-associated pneumonia caused by S. aureus.
3. Discussion
Nosocomial pneumonias, together with surgical site infections, are the main types of nosocomial infection, and S. aureus is the leading associated etiological agent, according to a survey conducted in the United States [4]. The present study showed an ID of S. aureus VAP of 2.35/1000 ventilator days, a value higher than that reported in the study by Lee et al. [20] in which the ID of VAP was 1.4/1000 ventilator days and S. aureus was the most frequently isolated microorganism.
In our study, the presence of the mecA gene, oxacillin resistance, or SCCmec type was not associated with the development of VAP or death. Some studies have shown an association between infection with MRSA methicillin-resistant strains and a longer ICU stay and higher mortality rates compared to patients who develop VAP caused by MSSA [21,22]. However, other studies found that methicillin resistance alone was not associated with recurrence, severity, or mortality in S. aureus VAP [23,24]. Thus, methicillin resistance does not seem to interfere with the mortality rate in VAP, a fact that raises speculation about whether other factors present in nosocomial strains may contribute to mortality. Interestingly, the frequency of the icaC, sec, sea, and hlb genes was slightly higher in mecA gene-positive isolates; however, there was no statistically significant association between the simultaneous presence of the mecA gene and virulence genes, with MSSA and MRSA isolates showing the same level of virulence.
The wide variability in the clonal profiles among strains isolated from patients with VAP and the formation of clusters only for isolates from the same individual suggest that these isolates are endogenous and that their development occurs from strains that already colonize the patient. Only two isolates (47 and 239) that belonged to the same clone formed a cluster. Previous studies have shown that the pattern of microorganism colonization varies. Gram-positive bacteria colonize the trachea within the first 24 h of mechanical ventilation and are later replaced with Gram-negative bacteria and yeasts [25,26].
Sequence types characteristic of community settings were the types most frequently isolated from patients who developed VAP (ST45, ST5, ST8, ST1635, ST398, and SLV 546), with the exception of nosocomial clone ST105 carrying SCCmec II. One strain was grouped with USA300, which belonged to the same ST8 carrying SCCmec IV, and was negative for the PVL (lukSF-PV) genes. Classic studies of infection with USA300 mainly report necrotizing pneumonia [27] in community-dwelling healthy individuals without traditional factors for MRSA contraction [28]. Despite reports of pneumonia in community-dwelling patients, Pasquale et al. [29] recently described the involvement of USA300 in nosocomial pneumonia. In these cases, participation of this strain is of great clinical importance since, in addition to the difficulty in selecting antimicrobials for treatment, the strain can carry a wide range of virulence factors that can also influence the outcome [3,9].
The involvement of STs characteristic of community settings in nosocomial infections has been reported, especially an association of ST8 with outbreaks of difficult-to-treat infections [30]. This observation highlights the need for controlling the transmission of these community-associated strains among patients admitted to the same ICU.
Studies have discussed the participation of virulence factors in the development of VAP in an attempt to identify a marker that permits predicting the development and severity of these infections. The participation of a single virulence factor such as alpha-hemolysin or protein A has been reported to be important, if not crucial, for the development of pneumonia [9,31].
A previous study also found that patients with cancer, especially lung cancer, are at increased risk of developing VAP when compared to other patients [32]. The presence of sea in the lungs can elicit an inflammatory response through the production of IFN-γ by CD8 T cells, which causes lesions characteristic of acute respiratory stress syndrome [33]. This finding suggests that strains producing this enterotoxin pose a risk for developing pneumonia associated with severe symptoms.
The only independent factors associated with death were the presence of the sea and hlb genes. Among the four S. aureus strains isolated from patients with VAP who died from this condition, three carried the hlb gene and the other the sea gene. The presence of enterotoxin A has been linked to aggressive infection [33]. Similarly, the presence of hlb in lung tissue can increase neutrophil infiltration by modulating host factors, which contributes to the development of pneumonia [34]. Both death-related factors are known to activate an intense immune response that contributes to breathing problems and can quickly lead to the patient’s death.
The association between the presence of virulence genes and the development of VAP has been little explored, and studies are limited to the analysis of factors such as strain cytotoxicity [8] or few virulence factors in animal experiments [6,31], which offers an excellent perspective of the virulence factors studied. However, our study provides a practical view of virulence factors and new perspectives on the association of microbiological factors with VAP. Within this context, we found that the etiology of VAP is multifactorial, including microbiological factors and risk factors of the host itself.
The main strength of this study is the fact that the patients were followed up since intubation, which allows microbiological study and the analysis of clinical data throughout the period of development of VAP. The results suggest that, in patients admitted to the ICU, VAP can be caused by S. aureus strains that already colonize these patients. This suggestion is supported by the STs identified, which are characteristic of strains found in the community that carry important virulence factors involved in the development of VAP and subsequent death of the patient. The identification of patients with VAP previously colonized with S. aureus suggests that some type of screening may be useful in predicting the occurrence of infection. However, further studies are still needed to evaluate the best screening approach and its cost-effectiveness. In view of the high mortality associated with VAP, the results of this study also highlight the need for future studies that assess whether S. aureus decolonization of known carriers will affect the incidence of VAP.
4. Materials and Methods
4.1. Study Place and Sample
The study was conducted in two Specialized Intensive Care Units (ICU) of the University Hospital of the Botucatu Medical School (HC-FMB), with 16 and 9 beds, respectively, which admit adult clinical and surgical patients. Patients meeting the following criteria were included: patients undergoing mechanical ventilation started within 48 h of admission to the ICU or during their stay without a previous diagnosis of pneumonia whose families agreed to their participation in the study by signing the free informed consent form.
This is a prospective cohort study whose primary outcomes are VAP caused by S. aureus and death due to VAP. The subjects were followed up for a maximum period of eight weeks of ICU stay, twice weekly (Mondays and Thursdays), at an interval of 3.5 days. Clinical data and tracheal aspirates were collected during the evaluations. A doctor of the Committee for the Control of Healthcare-Related Infection (CCIRAS) made the diagnosis of VAP. The criteria recommended by the National Healthcare Safety Network (NHSN) of the Centers for Disease Control and Prevention (CDC) of the United States were used [35].
4.2. Collection of Clinical Data
The following data were obtained during the visit for patient inclusion: gender, age, underlying diseases and comorbidities, length of hospital stay prior to the current hospitalization, procedures, and presence of invasive devices (central venous catheters, indwelling urinary catheters, nasogastric tubes/nasoenteral tubes, and drains). Data comprising the two weeks preceding mechanical ventilation were also collected: past and current use of antimicrobials, history of infectious complications, and results of microbiological tests.
Data on length of stay, procedures, invasive devices, use of antimicrobials, infectious complications, and results of microbiological tests were also collected during the follow-up assessments.
4.3. Collection, Isolation, and Microbial Identification
Tracheal aspirate was collected by a physical therapist, doctor, or nurse and immediately sent to the Laboratory of Bacteriology. The clinical material was diluted 1:1 in 1% N-acetyl-L-cysteine (mucolytic), homogenized, and kept for 15 min at room temperature. A 0.1-mL aliquot of this solution was diluted in 9.9 mL sterile saline and seeded with a 0.01-mL calibrated loop onto sheep blood and Baird–Parker agar plates. After incubation for 24 h at 37 °C, the number of colony-forming units (CFU) was determined by multiplying the number of colonies by the corresponding dilution [36]. The cultures were classified as positive when the count was 106 CFU or higher. Microorganisms that grew in the culture medium were subjected to Gram staining for assessment of purity, observation of morphology, and specific staining. After confirmation of these features (Gram-positive cocci in clusters), the catalase and coagulase tube tests were performed according to Koneman et al. [37] for phenotypic identification of S. aureus. Catalase- and coagulase-positive isolates were subjected to DNA extraction (see item 4.6) for genotypic identification of S. aureus with PCR amplification of the Sa442 DNA fragment, which is specific for S. aureus, following the protocol described by Martineau et al. [38]. After the species was confirmed, the isolates were stored in nutrient broth with glycerol at −70 °C.
4.4. Antimicrobial Susceptibility Testing
The antimicrobial susceptibility/resistance profile of all S. aureus strains isolated in this study was evaluated using the Kirby–Bauer disk diffusion method using disks impregnated with gentamicin (10 µg), linezolid (30 µg), penicillin G (10 IU), erythromycin (15 µg), and clindamycin (2 µg), as well as the D test for identifying inducible clindamycin resistance, according to the criteria of the Clinical and Laboratory Standards Institute [39]. Methicillin susceptibility was determined using cefoxitin (30 µg) and oxacillin disks (1 µg).
4.5. Determination of Minimum Inhibitory Concentration (MIC)
The in vitro MIC of all S. aureus isolates against oxacillin, vancomycin, linezolid, daptomycin, quinupristin/dalfopristin, and tigecycline was determined with the E-test. The MIC results are expressed as the proportion of isolates susceptible to each drug according to the CLSI definition [39]. Isolates with intermediate values were classified as resistant.
4.6. DNA Extraction from S. aureus
Nucleic acid was extracted from all S. aureus strains isolated in the study. The isolates were cultured on blood agar, inoculated individually into BHI broth, and incubated for 24 h at 37 °C. Extraction was performed with the Illustra kit (GE Healthcare) according to the manufacturer’s instructions, and the extracted DNA was stored at −20 °C.
4.7. Detection of the mecA Gene
The parameters and primers described by Murakami et al. [40] were used for the detection of the mecA gene in the S. aureus isolates. International reference strains were included in the assays as positive (S. aureus ATCC 33591) and negative control (S. aureus ATCC 25923).
4.8. Classification of SCCmec in S. aureus
All MRSA isolates were subjected to SCCmec typing by multiplex PCR using the primers and parameters described by Milheiriço et al. [36]. The following strains were used as a control for SCCmec typing: COL for SCCmec type I; N315 for SCCmec type IA; PER34 for SCCmec type II; AN546 for SCCmec type III; HU25 for SCCmec type IIIA; and MW2 for SCCmec type IV.
4.9. Detection of Genes Encoding Virulence Factors
The genes for the following virulence factors were investigated in all S. aureus isolates with PCR using the primers and parameters described in Table 7: PVL (lukS-lukF-PV); toxic shock syndrome toxin (tst); staphylococcal enterotoxins A to C (sea, seb, and sec); biofilm (icaA, icaD, and bap); exfoliative toxins A and B (eta and etb); and hemolysins α, β, and δ (hla, hlb, and hld). International reference strains were included in all reactions as positive and negative control (Table 7).

Table 7.
Detection of genes encoding virulence factors in S. aureus.
4.10. Visualization of Amplified Products
The efficiency of the amplifications was monitored with electrophoresis on 2% agarose gel prepared in 1X TBE buffer and stained with SYBR Safe. The size of the amplified products was compared to 100-bp molecular markers, and the gel was photographed under UV transillumination.
4.11. Characterization and Quantification of Phenol-Soluble Modulins (PSM) with High-Performance Liquid Chromatography (HPLC)
For the detection and quantification of PSM, 10 µL of an overnight culture was incubated in 1 mL tryptic soy broth for 16 to 18 h, centrifuged at 14,000 rpm for 15 min at 4 °C, and 300 µL of the supernatant was reserved for reversed-phase HPLC-mass spectrometry as described previously [48]. PSMs were quantified by integrating the ion chromatograms extracted by the mass-to-charge ratios of doubly and triply charged ions of each PSM. The unit of PSM measurement is “amount arbitrary unit (A.U.)”.
4.12. Western Blot for the Quantification of spa and hla Expression
For analysis of protein A and alpha-hemolysin expression, the isolates were cultured overnight and 10 µL of this culture was added to 1 mL TSB and incubated for 8 h at 37 °C. The cultures were centrifuged, and a 300-µL aliquot of the supernatant was used for Western blot analysis. Supernatants were submitted at 95 °C for 5 min and loaded in the 12% SDS page gel. The applied voltage was 150 v, 400 mA for 1 h. The proteins were transferred to a nitrocellulose membrane, washed, and blocked with Rabbit antibody for alpha-toxin. The membranes were audiographed in the Typhon TRIO plus Variable Mode Imager®. GraphPad Prism® using a t-test analyzed the data.
4.13. Pulsed-Field Gel Electrophoresis (PFGE)
PFGE typing consisted of digestion with SmaI of chromosomal DNA from S. aureus isolates associated with VAP. The plug preparation protocol and parameters followed McDougal et al. [49]. The restriction fragments resulting from SmaI digestion were subjected to electrophoresis on 1% agarose gel in a CHEF-DR III system (Bio-Rad Laboratories, Hercules, CA, USA), with pulses alternating from 5 to 60 s at 6 V/cm and 13 °C for 23 h.
The gels were stained with Gel Red (Biotium, San Francisco, CA, USA) for 1 h, washed in Milli Q water for another hour, and photographed under UV transillumination. The profiles were analyzed with the BioNumerics software (Applied Maths, Sint-Martens-Latem, Belgium) using the Dice similarity coefficient and the UPGMA method (1.2 tolerance and 1% optimization) for cluster analysis of isolates. Clusters were defined when similarity was ≥80%. International clones were kindly provided by Antônio Carlos Campos Pignatari, Special Laboratory of Clinical Microbiology, Department of Infectology, Federal University of São Paulo-Escola Paulista de Medicina and Agnes Marie Sá Figueiredo, Paulo de Góes Institute of Microbiology, Federal University of Rio de Janeiro, Brazil.
4.14. Multilocus Sequence Typing (MLST)
The constitutive genes of S. aureus associated with VAP were amplified and sequenced to determine allele composition and sequence type (ST) using the parameters proposed by Enright et al. [50]. Seven housekeeping genes were used (arcC, aroE, yqiL, gmk, tpi, glp, and pta). Sequencing was performed using Macrogen (Rockville, MD, USA), and sequences were analyzed using SeqBuider® and the official website mlst.org.
4.15. Statistical Analysis
Poisson regression of host and microbiological factors was performed to evaluate their relationship with the development of VAP and death, using backward selection (the first model included all variables, and the multivariate model included variables with p > 0.1, removing the variable with the highest p-value at each step and including only those with p < 0.1, i.e., significant, and marginally significant variables). The values obtained for the production of PSM, alpha-toxin, and protein A (spa) were dichotomized by the median for inclusion in the model (‘yes’ for a value equal to or greater than the median and ‘no’ for a lower value). Data were stored in Epi Info® 3.5.2 (Centers for Disease Control and Prevention) and Excel and were analyzed using Epi Info and SPSS®. The incidence density of S. aureus and MRSA VAP was calculated using the following equation:
where ID = incidence density, NVAP = number of patients with VAP, 1000 = standard coefficient, ΣnVis = sum of the number of visits of patients (728), and 3.5 = interval between visits in days.
5. Conclusions
Our results allow us to suggest that in ICU patients, VAP is mainly caused by S. aureus strains that already colonize these patients. This suggestion is supported by the STs identified, which are characteristic of strains found in the community that produce important virulence factors involved in the development of VAP and subsequent death.
Supplementary Materials
The following supporting information can be downloaded at https://www.mdpi.com/article/10.3390/antibiotics12081336/s1, Table S1. The data sheet with all the data for all 51 S. aureus strains isolated in the study.
Author Contributions
Conceptualization, M.F.B., M.O., M.d.L.R.d.S.d.C. and C.M.C.B.F.; investigation L.M.A., M.F.B., R.d.S.C., M.T.S., C.F.R. and H.-S.J.; software, C.M.C.B.F.; formal analysis C.M.C.B.F.; writing—original draft preparation, M.F.B.; writing—review and editing, L.M.A. and M.d.L.R.d.S.d.C.; supervision, C.M.C.B.F., M.O. and M.d.L.R.d.S.d.C.; project administration, M.F.B. All authors have read and agreed to the published version of the manuscript.
Funding
This study was funded by the National Council for Scientific and Technological Development (CNPq—Grant 303603/2020-8) to M.d.L.R.d.S.d.C., Coordination for Improvement of Higher Education Personnel (CAPES—Grant 99999.007555/2013-00) to M.F.B. and the Intramural Research Program of the National Institute of Allergy and Infectious Diseases (NIAID), NIH to M.O. (project number ZIA-AI000904).
Institutional Review Board Statement
The present study was approved by the Research Ethics Committee of the Botucatu Medical School, São Paulo State, Brazil (Approval number 290/11).
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
The data presented in this study are original and have not been published in scientific journals. The only document that contains these data is the doctoral thesis of Mariana Fávero Bonesso, openly available in [Institutional Repository of UNESP] at [http://hdl.handle.net/11449/139372] (accessed on 1 June 2023).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Papazian, L.; Klompas, M. Ventilator-associated pneumonia in adults: A narrative review. Intensive Care Med. 2020, 46, 888–906. [Google Scholar] [CrossRef]
- Isac, C.; Samson, H.R. Prevention of VAP: Endless evolving evidences–systematic literature review. Nurs. Forum 2021, 56, 905–915. [Google Scholar] [CrossRef]
- Melsen, W.G.; Rovers, M.M. Attributable mortality of ventilator-associated pneumonia: A meta-analysis of individual patient data from randomised prevention studies. Lancet Infect. Dis. 2013, 13, 665–671. [Google Scholar] [CrossRef]
- Magill, S.S.; Edwards, J.R. Emerging Infections Program Healthcare-Associated Infections and Antimicrobial Use Prevalence Survey Team. Multistate point-prevalence survey of health care-associated infections. N. Engl. J. Med. 2014, 370, 1198–1208. [Google Scholar] [CrossRef]
- Zaragoza, R.; Vidal-Cortés, P. Update of the treatment of nosocomial pneumonia in the ICU. Crit. Care 2020, 24, 383. [Google Scholar] [CrossRef]
- Haque, N.Z.; Arshad, S. Analysis of pathogen and host factors related to clinical outcomes in patients with hospital-acquired pneumonia due to methicillin-resistant Staphylococcus aureus. J. Clin. Microbiol. 2012, 50, 1640–1644. [Google Scholar] [CrossRef]
- IWG-SCC. International Working Group on the Staphylococcal Cassette Chromosome Elements. Web Page on the Internet. Available online: https://www.sccmec.org/index.php/en/sccmmcc-structure/sccmmcc-types-from1-toxiv-smn (accessed on 28 April 2023).
- Rose, H.R.; Holzman, R.S. Cytotoxic Virulence Predicts Mortality in Nosocomial Pneumonia Due to Methicillin-Resistant Staphylococcus aureus. J. Infect. Dis. 2015, 211, 1862–1874. [Google Scholar] [CrossRef]
- Lacoma, A.; Laabei, M. Genotypic and Phenotypic Characterization of Staphylococcus aureus Isolates from the Respiratory Tract in Mechanically-Ventilated Patients. Toxins 2021, 13, 122. [Google Scholar] [CrossRef]
- Berube, B.J.; Bubeck Wardenburg, J. Staphylococcus aureus α-toxin: Nearly a century of intrigue. Toxins 2013, 5, 1140–1166. [Google Scholar] [CrossRef]
- Spaulding, A.R.; Salgado-Pabón, W. Staphylococcal and streptococcal superantigen exotoxins. Clin. Microbiol. Rev. 2013, 26, 422–447. [Google Scholar] [CrossRef]
- Krakauer, T. Staphylococcal Superantigens: Pyrogenic Toxins Induce Toxic Shock. Toxins 2019, 11, 178. [Google Scholar] [CrossRef]
- Strandberg, K.L.; Rotschafer, J.H. Staphylococcal superantigens cause lethal pulmonary disease in rabbits. J. Infect. Dis. 2010, 202, 1690–1697. [Google Scholar] [CrossRef]
- Kreienbuehl, L.; Charbonney, E. 2011. Community-acquired necrotizing pneumonia due to methicillin-sensitive Staphylococcus aureus secreting Panton-Valentine leukocidin: A review of case reports. Ann. Intensive Care 2012, 1, 52. [Google Scholar] [CrossRef]
- Larsen, S.A.H.; Kyhl, K. Life-Threatening Necrotizing Pneumonia with Panton–Valentine Leukocidin-Producing, Methicillin-Sensitive Staphylococcus aureus in a Healthy Male Co-Infected with Influenza B. Infect. Dis. Rep. 2022, 14, 12–19. [Google Scholar] [CrossRef]
- Lina, G.; Piémont, Y. Involvement of Panton-Valentine Leukocidin–Producing Staphylococcus aureus in Primary Skin Infections and Pneumonia. Clin. Infect. Dis. 1999, 29, 1128–1132. [Google Scholar] [CrossRef]
- Liassine, N.; Auckenthaler, R. Community-acquired methicillin-resistant Staphylococcus aureus isolated in Switzerland contains the Panton-Valentine leukocidin or exfoliative toxin genes. J. Clin. Microbiol. 2004, 42, 825–828. [Google Scholar] [CrossRef]
- Peng, Q.; Tang, X.; Dong, W.; Sun, N.; Yuan, W. A Review of Biofilm Formation of Staphylococcus aureus and Its Regulation Mechanism. Antibiotics 2023, 12, 12. [Google Scholar] [CrossRef]
- Lanks, C.W.; Musani, A.I. Community-acquired Pneumonia and Hospitalacquired Pneumonia. Med. Clin. N. Am. 2019, 103, 487–501. [Google Scholar] [CrossRef]
- Lee, M.S.; Walker, V. The Epidemiology of Ventilator-Associated Pneumonia in a Network of Community Hospitals: A Prospective Multicenter Study. Infect. Control. Hosp. Epidemiol. Off. J. Soc. Hosp. Epidemiol. Am. 2013, 34, 657–662. [Google Scholar] [CrossRef][Green Version]
- Pickens, C.; Wunderink, R.G. Methicillin-Resistant Staphylococcus aureus Hospital-Acquired Pneumonia/Ventilator-Associated Pneumonia. Semin. Respir. Crit. Care Med. 2022, 43, 304–309. [Google Scholar] [CrossRef]
- Rasigade, J.P.; Laurent, F. Lethal necrotizing pneumonia caused by an ST398 Staphylococcus aureus strain. Emerg. Infect. Dis. Aug. 2010, 16, 1330. [Google Scholar] [CrossRef]
- Klein, E.Y.; Jiang, W. National Costs Associated With Methicillin-Susceptible and Methicillin-Resistant Staphylococcus aureus Hospitalizations in the United States, 2010–2014. Clin. Infect. Dis. 2019, 68, 22–28. [Google Scholar] [CrossRef]
- Zahar, J.R.; Clec’h, C. Is methicillin resistance associated with a worse prognosis in Staphylococcus aureus ventilator-associated pneumonia? Clin. Infect. Dis. 2005, 41, 1224–1231. [Google Scholar] [CrossRef]
- Modi, A.R. Hospital-acquired and ventilator-associated pneumonia: Diagnosis, management, and prevention. Clevel. Clin. J. Med. 2020, 87, 633–639. [Google Scholar] [CrossRef]
- Cardeñosa, C.J.; Solé-Violán, J. Role of different routes of tracheal colonization in the development of pneumonia in patients receiving mechanical ventilation. Chest 1999, 116, 462–470. [Google Scholar]
- Toro, C.M.; Janvier, J. Community-associated methicillin-resistant Staphylococcus aureus necrotizing pneumonia without evidence of antecedent viral upper respiratory infection. Can. J. Infect. Dis. Med. Microbiol. 2014, 25, e76–e82. [Google Scholar] [CrossRef]
- King, M.D.; Humphrey, B.J.; Wang, Y.F.; Kourbatova, E.V.; Ray, S.M.; Blumberg, H.M. Emergence of community-acquired methicillin-resistant Staphylococcus aureus USA 300 clone as the predominant cause of skin and soft tissue infections. Ann. Intern. Med. 2006, 144, 309–317. [Google Scholar] [CrossRef]
- Pasquale, T.R.; Jabrocki, B. Emergence of methicillin-resistant Staphylococcus aureus USA300 genotype as a major cause of late-onset nosocomial pneumonia in intensive care patients in the USA. Int. J. Infect. Dis. 2013, 17, e398–e403. [Google Scholar] [CrossRef]
- Schuenck, R.P.; Nouér, S.A. Polyclonal presence of non-multiresistant methicillin-resistant Staphylococcus aureus isolates carrying SCCmec IV in health care-associated infections in a hospital in Rio de Janeiro, Brazil. Diagn. Microbiol. Infect. Dis. 2009, 64, 434–441. [Google Scholar] [CrossRef]
- Wardenburg, J.B.; Patel, R.J. Surface proteins and exotoxins are required for the pathogenesis of Staphylococcus aureus pneumonia. Infect. Immun. 2007, 75, 1040–1044. [Google Scholar] [CrossRef]
- Park, S.A.; Cho, S.S. Factors influencing ventilator-associated pneumonia in cancer patients. Asian Pac. J. Cancer Prev. 2014, 15, 5787–5791. [Google Scholar] [CrossRef] [PubMed]
- Muralimohan, G.; Rossi, R.J. Inhalation of Staphylococcus aureus enterotoxin A induces IFN-gamma and CD8 T cell-dependent airway and interstitial lung pathology in mice. J. Immunol. 2008, 181, 3698–3705. [Google Scholar] [CrossRef]
- Hayashida, A.; Bartlett, A.H. Staphylococcus aureus Beta-Toxin Induces Lung Injury through Syndecan-1. Am. J. Pathol. 2009, 174, 509–518. [Google Scholar] [CrossRef]
- Horan, T.C.; Andrus, M. CDC/NHSN surveillance definition of health care–associated infection and criteria for specific types of infections in the acute care setting. Am. J. Infect. Control 2008, 36, 309–332. [Google Scholar] [CrossRef] [PubMed]
- Milheirico, C.; Oliveira, D.C. Update to the Multiplex PCR Strategy for Assignment of mec Element Types in Staphylococcus aureus. Antimicrob. Agents Chemother. 2007, 51, 4537. [Google Scholar] [CrossRef]
- Konemman, E.V.; Allen, S.D.; Sowell, V.R.; Sommer, H.M. Introdução à Microbiologia Médica. In Diagnóstico Microbiológico: Texto e Atlas Colorido, 5th ed.; Medsi: Rio de Janeiro, Brazil, 2001. [Google Scholar]
- Martineau, F.; Picard, F.J.; Roy, P.H.; Ouellette, M.; Bergeron, M.G. Species-Specific and Ubiquitous-DNA-Based Assays for Rapid Identification of Staphylococcus aureus. J. Clin. Microbiol. 1998, 36, 618–623. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Disk Susceptibility Tests, 31st ed.; CLSI Supplement M100; CLSI: Wayne, PA, USA, 2021. [Google Scholar]
- Murakami, K.; Minamide, K. Identification of methicillin-resistant strains of staphylococci by polymerase chain reaction. J. Clin. Microb. 1991, 29, 2240–2244. [Google Scholar] [CrossRef]
- Johnson, W.M.; Tyler, S.D. Detection of genes for enterotoxins, exfoliative toxins, and toxic shock syndrome toxin 1 in Staphylococcus aureus by the polymerase chain reaction. J. Clin. Microbiol. 1991, 29, 426–430. [Google Scholar] [CrossRef]
- Cunha, M.D.L.R.S.; Calsolari, R.O. Detection of enterotoxin and toxic shock syndrome toxin 1 genes in Staphylococcus, with emphasis on coagulase-negative staphylococci. Microbiol. Immunol. 2007, 51, 381–390. [Google Scholar] [CrossRef]
- Rohde, H.; Burandt, E.C. Polysaccharide intercellular adhesin or protein factors in biofilm accumulation of Staphylococcus epidermidis and Staphylococcus aureus isolated from prosthetic hip and knee joint infections. Biomaterials 2007, 28, 1711–1720. [Google Scholar] [CrossRef]
- Arciola, C.R.; Gamberini, S. A multiplex PCR method for the detection of all five individual genes of ica locus in Staphylococcus epidermidis. A survey on 400 clinical isolates from prosthesis-associated infections. J. Biomed. Mater. Res. A 2005, 75, 408–413. [Google Scholar] [CrossRef] [PubMed]
- Cucarella, C.; Tormo, M.A. Role of Biofilm–Associated Protein Bap in the Pathogenesis of Bovine Staphylococcus aureus. Infect. Immun. 2004, 72, 2177–2185. [Google Scholar] [CrossRef] [PubMed]
- Jarraud, S.; Mougel, C. Relationships between Staphylococcus aureus genetic background, virulence factors, agr groups (alleles), and human disease. Infect. Immun. 2002, 70, 631–641. [Google Scholar] [CrossRef] [PubMed]
- Marconi, C.; Cunha, M.L.R.S. Standardization of the PCR technique for the detection of delta toxin in Staphylococcus spp. J. Venom. Anim. Toxins Incl. Trop. Dis. 2005, 11, 117–128. [Google Scholar] [CrossRef]
- Joo, H.S.; Otto, M. The isolation and analysis of phenol-soluble modulins of Staphylococcus epidermidis. Methods Mol. Biol. 2014, 1106, 93–100. [Google Scholar] [PubMed]
- McDougal, L.K.; Steward, C.D. Pulsed-Field Gel Electrophoresis Typing of Oxacillin-Resistant Staphylococcus aureus Isolates from the United States: Establishing a National Database. J. Clin. Microbiol. 2003, 41, 5113–5120. [Google Scholar] [CrossRef]
- Enright, M.C.; Day, N.P.J. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J. Clin. Microbiol. 2000, 38, 1008–1015. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).