1. Introduction
In the past decades, antibiotic misuse and overuse in humans and livestock has led to the accelerated emergence of antibiotic-resistant organisms and the evolution of antibiotic-resistant strains, which have been serious risks to human security and animal health, as well as to the environment [
1]. Consequently, there is a common tendency to reduce or avoid using antibiotics in animal husbandry. Hence, it is crucial to find an ideal substitute for antibiotics. Nowadays, the use of prebiotics as a second-generation substitute for antibiotics has been the hotspot in the research on animal nutrition. Probiotics are “living microorganisms with low or no pathogenicity, which exerts beneficial effects on the host health when ingested in sufficient amounts [
2,
3]”. As one of the most important bacterial groups in the food industry, the application of lactic acid bacteria (
LAB) is considered to have beneficial effects on the health of animals and humans [
4], such as improved digestion [
5], strengthened mucosal immune barrier [
6], treatment of diarrhea [
7], anti-diabetic effects [
8], improved immunity [
9], maintenance of intestinal balance [
10], management of inflammatory bowel diseases [
10], and relieved constipation [
11]. However, the mode of action of probiotics on the host is still not fully understood. Therefore, studying the in vitro characteristics and evaluating their probiotic properties are essential for the optimization and use of probiotics.
LAB possess inhibitory characteristics against enteropathogens and regulate the intestinal microbial balance, thanks to the production of lactic acid and metabolites, such as antimicrobial compounds, antioxidants, and organic acids [
12]. These are considered important criteria in selecting potential probiotic strains because they are considered key ecological factors in determining the dominant bacteria in the intestinal ecosystem [
13]. However, in recent years, a growing body of research has revealed that functions of probiotics are highly strain-specific and their biological effects should be individually evaluated [
9]. Therefore, it is necessary to isolate and screen novel probiotics strain with potential health benefits. Dairy production systems are still important sources of probiotic strains [
14]. The cattle-yak has obvious heterosis, which is a hybrid offspring of yak (
Bos grunniens) and in cattle (
Bos taurus) [
15]. Cattle-yak milk, as a plateau specialty milk, has high quality, rich nutrition, high vitamin contents, and a fragrant smell. It is not only an important food and a dairy-processing raw material for plateau herdsmen, but is also a high-quality source of nutrients, such as fat, protein, lactose, and inorganic salt. It is expected to isolate probiotic strains with natural and excellent characteristics. However, there are few studies on probiotics. For the selection of a potential beneficial strain, candidate probiotic strains should be characterized by safety, function, and prebiotics, including antimicrobial susceptibility, adhesion to the intestinal mucosa and gastrointestinal transit, tolerance to the GI environment, acid potential, and antagonism against pathogens [
13]. In vitro methods are widely used to screen novel beneficial strains because examining in vivo efficacy is expensive and time-consuming. Moreover, it is necessary to analyze and evaluate the related characteristics of their probiotic function at the gene level to explore more potential biological functions and information [
16].
Therefore, the aim of this study was to identify the beneficial strains from the milk of cattle-yak, and to evaluate the probiotic characteristics of potential strains through a series of in vitro tests. Additionally, the whole genome sequence was also analyzed to provide insight into probiotic-associated genes for a better understanding of their potential biological functions and information.
4. Discussion
Improving human and animal health with probiotics has been the subject of intense scholarly debate. Numerous studies have shown that probiotics may adhere and survive in the gastrointestinal tract of humans and animals, and contribute to maintaining a microecological balance, promoting digestive and metabolic processes, as well as modulating the immune response, thereby enhancing host immunity, and improving human and animal health [
9,
13,
26]. However, because the effectiveness of probiotics is species or strain dependent, they should meet a series of characteristics, such as safety (antimicrobial susceptibility and hemolytic activity), functional (acid and bile salt tolerance and intestinal mucosa adhesion), as well as beneficial (antagonistic activity, growth, and acid-producing ability) characteristics [
12,
13,
17]. Hence, in this study, we focused on beneficial strains with potential probiotic properties isolated from cattle-yak milk using a series of in vitro tests, and the whole-genome sequencing was performed to reveal their potential biological functions and information.
Antimicrobial activity is one of the important criteria for selecting novel potential probiotic strains [
26]. Previous studies have shown that
LAB possesses extremely inhibitory characteristics against foodborne pathogenic bacteria [
20,
27,
28]. Recent studies also suggest that
LAB strains from milk had different degrees of inhibitory ability against both Gram-negative and Gram-positive pathogens [
29,
30], in agreement with our results. In this study, the strains C4 and C5 showed no zones of inhibition, while
LAB strains CY2 and CY12 showed better antimicrobial activity against
E. coli (29 mm; 28 mm),
S. aureus (24 mm; 27 mm),
S.agalactis (27 mm; 29 mm), and
S. typhii (23 mm; 25 mm) than ATCC 7469 and the other strains. The potential antimicrobial mechanisms may include forming the immune system, competing nutrients, as well as producing organic acids, bacteriocins, and antioxidants, etc. Moreover, antimicrobial activity related to the growth rate and acid-producing ability of
Lactobacillus are very crucial indicators to select potential beneficial microorganisms, which must have high acid-producing ability and growth velocity to compete for adhesion sites and nutrients within the GI ecosystem, further exerting their probiotic effects and inhibiting pathogenic bacteria [
31,
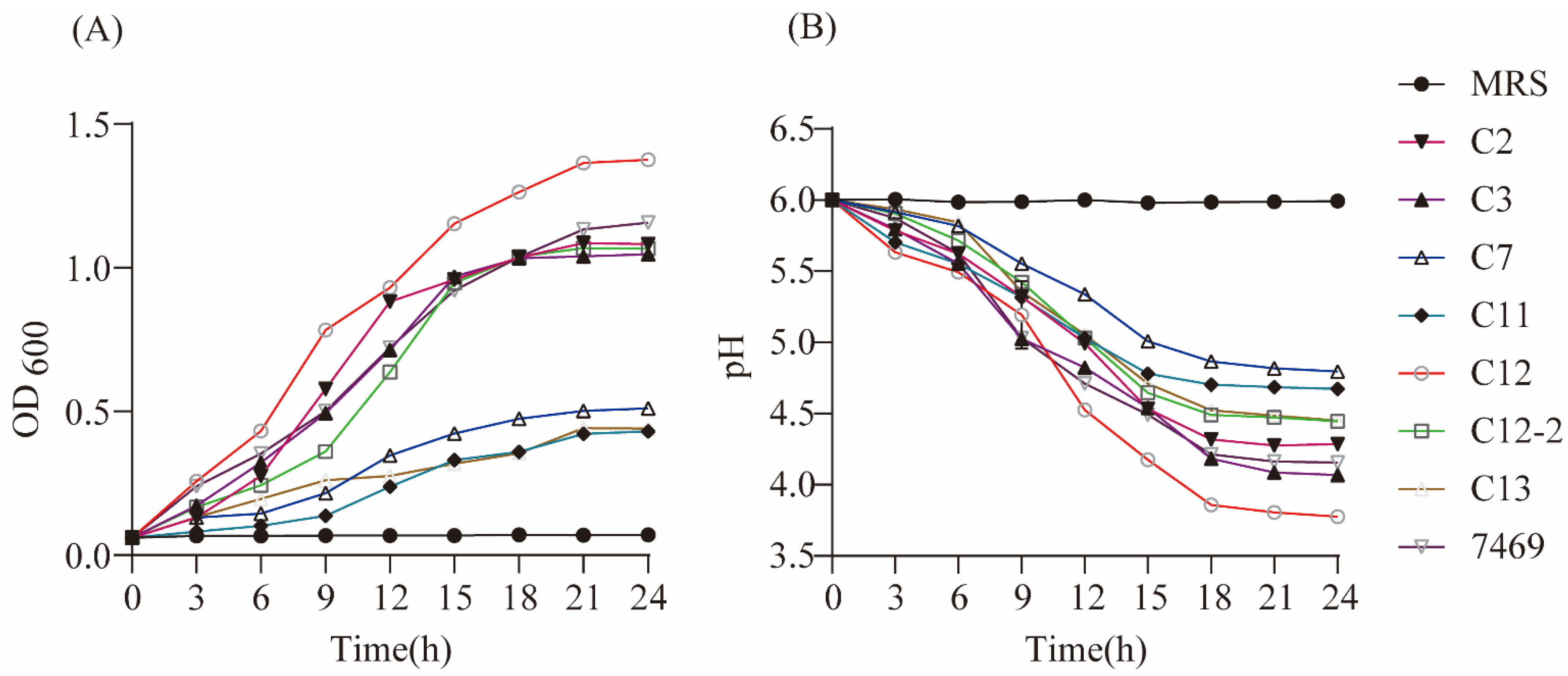
32]. Results of the present study suggest that the strain CY12 had a growth rate and acid-producing ability superior to that of ATCC 7469 and the other strains, consistent with our antimicrobial activity assay.
The presence of bile salts and high acidic conditions constitute the biggest barriers to the survival of
Lactobacillus in the gastrointestinal tract of the host [
12,
33]. Consequently, the capacity to tolerate both bile salts and acid conditions are two crucial characteristics considered for a candidate beneficial strain [
18,
34]. A previous study isolated
L. rhamnosus Z5 from human breast milk and reported that the strain had super resistance to bile salts and acid, but its growth was prominently inhibited at higher concentrations of bile salts [
35]. Another study also reported that the growth of
L. rhamnosus GG was significantly inhibited by bile salts at 0.15% [
36]. These findings were in agreement with our results: in the present study, we observed that the strains CY2, CY12, and ATCC 7469 had higher survivability at higher concentrations of bile salts and low pH, but were significantly inhibited at higher concentrations of bile salts, indicating that the strains CY2 and CY12 could survive in the GI tract. High osmotic pressure and surfactant activity of bile salts are potential mechanisms for disrupting phospholipids and proteins in the bacterial cell membrane, causing disruption of the bacterial membrane and causing DNA damage or death, further reducing the strains’ survival [
37,
38]. However, it has been reported that when
L. rhamnosus JL-1 was exposed to bile salts for 4 h, the strain had a survival rate as high as 90.01% [
9]; this may be due to the
LAB being isolated from the intestinal tract, and could be more adaptable to the GI ecosystem than other
Lactobacillus [
12].
A crucial property of
Lactobacillus intended for the oral route of administration is the capability to survive the gastric and intestinal conditions, including the presence of lysozyme, acid pH values with pepsin, pancreatin, and bile salts [
39,
40]. From this study, The results showed that all the
Lactobacillus strains had good tolerance to pepsin and trypsin simulated gastrointestinal juice, and the strains CY2 and CY12 showed high tolerance (survival percentage >80%) to artificial gastric juice at pH 2.0 for 2 h. Meanwhile, they also had better tolerance abilities to simulated intestinal juice for 2, 4, 6, and 8 h. In contrast, the survival percentages of the other strains decreased significantly. The survival rates of CY2 and CY12 in the gastrointestinal juice conditions were higher than the survival percentage of previously reported LAB, such as
L. rhamnosus and
Lactobacillus casei [
35,
41,
42]. Another study reported that the
LAB showed strong tolerance to the simulated GI environment, with survival rates of >80% at pH 2.0 to 3.0, which is consistent with our results [
43].
It has been reported that candidate probiotics should not serve as antibiotic resistance genes for hosts [
31,
44]. Therefore, the assessment of antimicrobial sensitivity is considered essential for the safe use of potential probiotics. From this study, we evaluated the antimicrobial susceptibility of seven selected strains using the disk diffusion method, and found that all studied strains were susceptible to antibiotics such as penicillin, chloramphenicol, erythrocin, tetracycline, ciprofloxacin, amoxicillin, ampicillin, and cefazolin, supporting most of the
LAB strains exhibited sensitivity to tetracycline, penicillin, and chloramphenicol, indicating that lactic acid bacteria generally had low resistance to these antibiotics [
13,
31,
45]. However, we observed that all selected strains are resistant to oxacillin and vancomycin. It is well known that
Lactobacillus strains are usually resistance to β-lactam antibiotics such as oxacillin, ceftriaxone, and ampicillin, due to the presence of β-lactamase in lactic acid bacteria [
26,
28]. Likewise, resistance to glycopeptides (vancomycin) has been noted in
LAB and is associated with congenital resistance to membrane permeability (in most cases), possibly through a mechanism of resistant efflux [
29]. Similar results found from previous studies also showed resistance to kanamycin, vancomycin, and ampicillin [
44,
46,
47]. In addition, hemolytic activity is considered to be an important indicator of the safety of potential beneficial strains; microbial probiotic strains are not expected to dissolve red blood cells when ingested in humans or animals. Like most
LAB strains [
48], our results found that all strains showed non-hemolytic activity and, thus, are safe for use. However, it is necessary to further perform in vivo safety assays.
The adhesion to the mucosal surfaces and epithelial cells is considered a crucial characteristic for candidate probiotics selection and plays an important role in competitively excluding or inhibiting pathogens [
26,
49]. Most
Lactobacillus produce cell surface proteins and help bacteria bind to the gastrointestinal epithelium, thereby improving immunity and antagonism against pathogens [
50]. The previous study demonstrated that the
L. rhamnosus GG strain had a higher adherence capacity to Caco-2 cells [
13]; similar results have been observed in the present study, whereby the strain CY12 exhibited higher adhesion capacity than the other strains, with over 80% adhesion capacity. However, whether the selected strains could inhibit invasion and intracellular survival of bacteria needs to be further researched. In this study, the CY12 strain was selected according to its probiotic features and safety evaluation. Genome-wide sequencing was performed on the
L. rhamnosus CY12 strain to better understand its potential biological functions. According to the definition of bacterial genome size by Ochman and Davalos [
51], the genome size of
L. rhamnosus CY12 isolated in this study is 2,506,167 bp, which indicates a medium-sized genome; this type of bacteria usually has a strong metabolic ability and tolerance [
52] and can adapt to different ecological environments. Moreover, based on the GO, KEGG, and COG annotation results, we also found that the genes were outstanding in the immune system process, biological adhesion, metabolic pathways, and galactose metabolism; glycine, serine, and threonine metabolism; energy production and conversion; amino acid transport and metabolism; and carbohydrate transport and metabolism, indicating that the strains were involved in the immune system process, adhesion, and had a metabolic ability and tolerance [
16]. These results are consistent with our noted beneficial characteristics of the
L. rhamnosus CY12 strain. However, there were still some genes for which we were unable to obtain COG functional annotation, which requires us to further explore these unknown functional genes in future studies.