Reduced Antibiotic Resistance in the Rhizosphere of Lupinus albus in Mercury-Contaminated Soil Mediated by the Addition of PGPB
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area
2.2. PGPB Isolation, Selection and Characterization
2.3. Biological Assays
2.4. DNA Isolation
2.5. Data Analysis and Bioinformatics
2.6. Cenoantibiogram: AR Profile of the Microbial Community
2.7. Statistical Analysis
3. Results
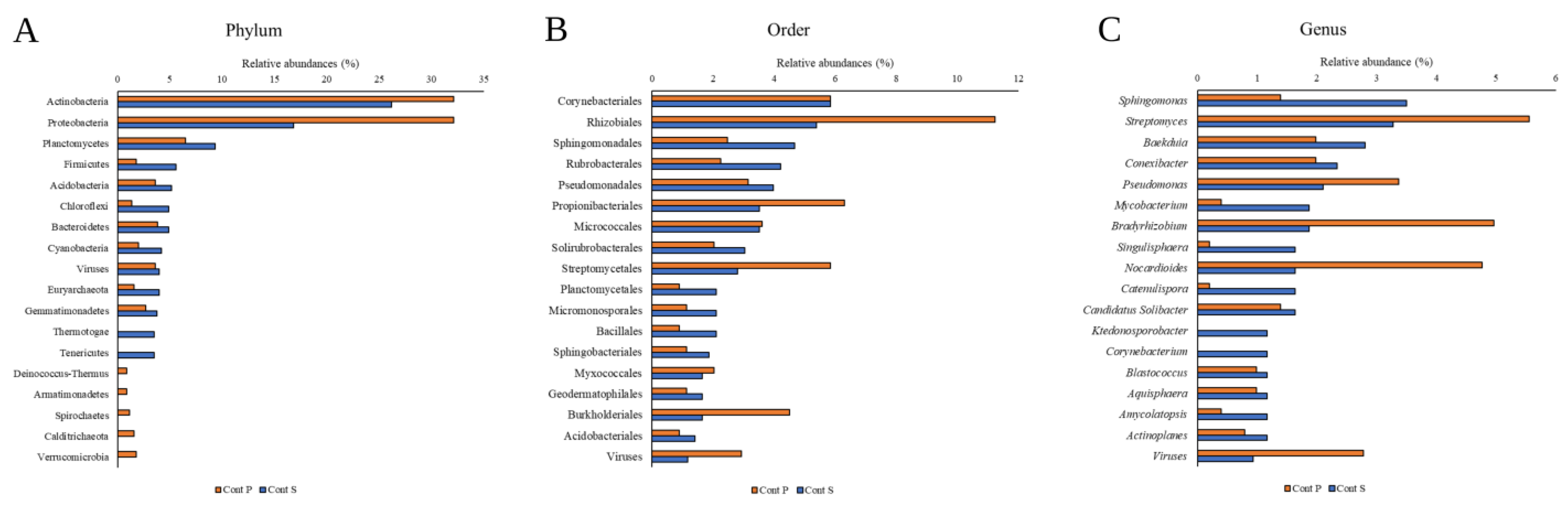
3.1. Metagenomic Analysis
3.2. Antibiogram
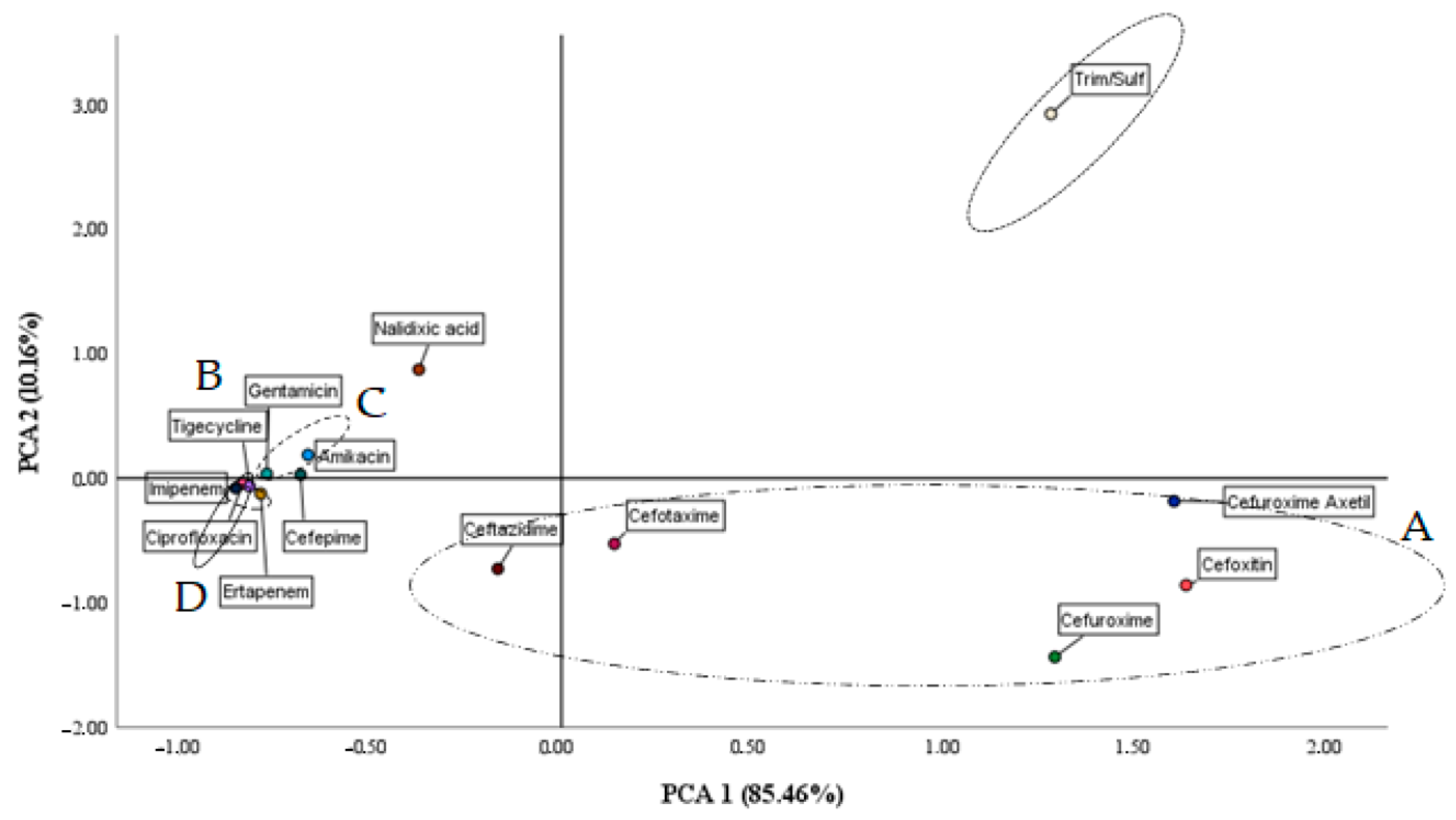
3.3. Cenoantibiogram
4. Discussion
- i.
- ii.
- iii.
- iv.
- v.
- vi.
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Liu, Y.-R.; Delgado-Baquerizo, M.; Bi, L.; Zhu, J.; He, J.-Z. Consistent Responses of Soil Microbial Taxonomic and Functional Attributes to Mercury Pollution across China. Microbiome 2018, 6, 183. [Google Scholar] [CrossRef]
- Hsu-Kim, H.; Eckley, C.S.; Achá, D.; Feng, X.; Gilmour, C.C.; Jonsson, S.; Mitchell, C.P. Challenges and Opportunities for Managing Aquatic Mercury Pollution in Altered Landscapes. Ambio 2018, 47, 141–169. [Google Scholar] [CrossRef]
- Bjørklund, G.; Tinkov, A.A.; Dadar, M.; Rahman, M.M.; Chirumbolo, S.; Skalny, A.V.; Skalnaya, M.G.; Haley, B.E.; Ajsuvakova, O.P.; Aaseth, J. Insights into the Potential Role of Mercury in Alzheimer’s Disease. J. Mol. Neurosci. 2019, 67, 511–533. [Google Scholar] [CrossRef]
- US Environmental Protection Agency. 2010 Biennial National Listing of Fish Advisories; US Environmental Protection Agency: Atlanta, GA, USA, 2011.
- Zhao, M.M.; Kou, J.; Chen, Y.; Xue, L.; Fan, T.T.; Wang, S. Bioremediation of Wastewater Containing Mercury Using Three Newly Isolated Bacterial Strains. J. Clean. Prod. 2021, 299, 126869. [Google Scholar] [CrossRef]
- González, D.; Robas, M.; Probanza, A.; Jiménez, P.A. Selection of Mercury-Resistant PGPR Strains Using the BMRSI for Bioremediation Purposes. Int. J. Environ. Res. Public Health 2021, 18, 9867. [Google Scholar] [CrossRef]
- Chellaiah, E.R. Cadmium (Heavy Metals) Bioremediation by Pseudomonas Aeruginosa: A Minireview. Appl. Water Sci. 2018, 8, 154. [Google Scholar] [CrossRef]
- Kara, A.K.; Fakıoğlu, Ö.; Kotan, R.; Atamanalp, M.; Alak, G. The Investigation of Bioremediation Potential of Bacillus Subtilis and B. Thuringiensis Isolates under Controlled Conditions in Freshwater. Arch. Microbiol. 2021, 203, 2075–2085. [Google Scholar] [CrossRef]
- Robas, M.; Jiménez, P.A.; González, D.; Probanza, A. Bio-Mercury Remediation Suitability Index: A Novel Proposal That Compiles the PGPR Features of Bacterial Strains and Its Potential Use in Phytoremediation. Int. J. Environ. Res. Public Health 2021, 18, 4213. [Google Scholar] [CrossRef]
- Robas, M.; Probanza, A.; González, D.; Jiménez, P.A. Mercury and Antibiotic Resistance Co-Selection in Bacillus Sp. Isolates from the Almadén Mining District. Int. J. Environ. Res. Public Health 2021, 18, 8304. [Google Scholar] [CrossRef]
- Liao, H.; Li, X.; Yang, Q.; Bai, Y.; Cui, P.; Wen, C.; Liu, C.; Chen, Z.; Tang, J.; Che, J. Herbicide Selection Promotes Antibiotic Resistance in Soil Microbiomes. Mol. Biol. Evol. 2021, 38, 2337–2350. [Google Scholar] [CrossRef]
- Yan, C.; Wang, F.; Liu, H.; Liu, H.; Pu, S.; Lin, F.; Geng, H.; Ma, S.; Zhang, Y.; Tian, Z. Deciphering the Toxic Effects of Metals in Gold Mining Area: Microbial Community Tolerance Mechanism and Change of Antibiotic Resistance Genes. Environ. Res. 2020, 189, 109869. [Google Scholar] [CrossRef]
- Mazhar, S.H.; Li, X.; Rashid, A.; Su, J.; Xu, J.; Brejnrod, A.D.; Su, J.-Q.; Wu, Y.; Zhu, Y.-G.; Zhou, S.G. Co-Selection of Antibiotic Resistance Genes, and Mobile Genetic Elements in the Presence of Heavy Metals in Poultry Farm Environments. Sci. Total Environ. 2021, 755, 142702. [Google Scholar] [CrossRef]
- Kumar, M.; Sarma, D.K.; Shubham, S.; Kumawat, M.; Verma, V.; Nina, P.B.; Jp, D.; Kumar, S.; Singh, B.; Tiwari, R.R. Futuristic Non-Antibiotic Therapies to Combat Antibiotic Resistance: A Review. Front. Microbiol. 2021, 12, 16. [Google Scholar] [CrossRef]
- Vikesland, P.; Garner, E.; Gupta, S.; Kang, S.; Maile-Moskowitz, A.; Zhu, N. Differential Drivers of Antimicrobial Resistance across the World. Acc. Chem. Res. 2019, 52, 916–924. [Google Scholar] [CrossRef]
- Wang, F.; Fu, Y.-H.; Sheng, H.-J.; Topp, E.; Jiang, X.; Zhu, Y.-G.; Tiedje, J.M. Antibiotic Resistance in the Soil Ecosystem: A One Health Perspective. Curr. Opin. Environ. Sci. Health 2021, 20, 100230. [Google Scholar] [CrossRef]
- Mojsoska, B.; Ghoul, M.; Perron, G.G.; Jenssen, H.; Alatraktchi, F.A. Changes in Toxin Production of Environmental Pseudomonas Aeruginosa Isolates Exposed to Sub-Inhibitory Concentrations of Three Common Antibiotics. PLoS ONE 2021, 16, e0248014. [Google Scholar] [CrossRef]
- Liu, Z.; Wang, W.; Zhu, Y.; Gong, Q.; Yu, W.; Lu, X. Antibiotics at Subinhibitory Concentrations Improve the Quorum Sensing Behavior of Chromobacterium Violaceum. FEMS Microbiol. Lett. 2013, 341, 37–44. [Google Scholar] [CrossRef]
- Chow, L.K.M.; Ghaly, T.M.; Gillings, M.R. A Survey of Sub-Inhibitory Concentrations of Antibiotics in the Environment. J. Environ. Sci. 2021, 99, 21–27. [Google Scholar] [CrossRef]
- Li, Y.; Xia, L.; Chen, J.; Lian, Y.; Dandekar, A.A.; Xu, F.; Wang, M. Resistance Elicited by Sub-Lethal Concentrations of Ampicillin Is Partially Mediated by Quorum Sensing in Pseudomonas Aeruginosa. Environ. Int. 2021, 156, 106619. [Google Scholar] [CrossRef]
- Bombaywala, S.; Mandpe, A.; Paliya, S.; Kumar, S. Antibiotic Resistance in the Environment: A Critical Insight on Its Occurrence, Fate, and Eco-Toxicity. Environ. Sci. Pollut. Res. 2021, 28, 24889–24916. [Google Scholar] [CrossRef]
- Woodford, N.; Turton, J.F.; Livermore, D.M. Multiresistant Gram-Negative Bacteria: The Role of High-Risk Clones in the Dissemination of Antibiotic Resistance. FEMS Microbiol. Rev. 2011, 35, 736–755. [Google Scholar] [CrossRef]
- Czekalski, N.; Berthold, T.; Caucci, S.; Egli, A.; Bürgmann, H. Increased Levels of Multiresistant Bacteria and Resistance Genes after Wastewater Treatment and Their Dissemination into Lake Geneva, Switzerland. Front. Microbiol. 2012, 3, 106. [Google Scholar] [CrossRef]
- Galler, H.; Luxner, J.; Petternel, C.; Reinthaler, F.F.; Habib, J.; Haas, D.; Kittinger, C.; Pless, P.; Feierl, G.; Zarfel, G. Multiresistant Bacteria Isolated from Intestinal Faeces of Farm Animals in Austria. Antibiotics 2021, 10, 466. [Google Scholar] [CrossRef]
- Sole, A. Multiresistant Organisms: Bacteria and Beyond. Curr. Opin. Organ Transplant. 2022, 27, 184–190. [Google Scholar] [CrossRef]
- World Health Organization. Global Action Plan on Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2015; ISBN 978-92-4-150976-3. [Google Scholar]
- Zhao, X.; Yu, Z.; Ding, T. Quorum-Sensing Regulation of Antimicrobial Resistance in Bacteria. Microorganisms 2020, 8, 425. [Google Scholar] [CrossRef]
- Stubbendieck, R.M.; Straight, P.D. Multifaceted Interfaces of Bacterial Competition. J. Bacteriol. 2016, 198, 2145–2155. [Google Scholar] [CrossRef]
- Nadell, C.D.; Bassler, B.L. A Fitness Trade-off between Local Competition and Dispersal in Vibrio Cholerae Biofilms. Proc. Natl. Acad. Sci. USA 2011, 108, 14181–14185. [Google Scholar] [CrossRef]
- Khare, A.; Tavazoie, S. Multifactorial Competition and Resistance in a Two-Species Bacterial System. PLoS Genet. 2015, 11, e1005715. [Google Scholar] [CrossRef]
- Li, Y.; Tian, X. Quorum Sensing and Bacterial Social Interactions in Biofilms: Bacterial Cooperation and Competition. In Stress and Environmental Regulation of Gene Expression and Adaptation in Bacteria; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2016; pp. 1195–1205. [Google Scholar]
- Komijani, M.; Shamabadi, N.S.; Shahin, K.; Eghbalpour, F.; Tahsili, M.R.; Bahram, M. Heavy Metal Pollution Promotes Antibiotic Resistance Potential in the Aquatic Environment. Environ. Pollut. 2021, 274, 116569. [Google Scholar] [CrossRef]
- Miao, J.; Yin, Z.; Yang, Y.; Liang, Y.; Xu, X.; Shi, H. Abundance and Dynamic Distribution of Antibiotic Resistance Genes in the Environment Surrounding a Veterinary Antibiotic Manufacturing Site. Antibiotics 2021, 10, 1361. [Google Scholar] [CrossRef]
- Wei, Z.; Feng, K.; Wang, Z.; Zhang, Y.; Yang, M.; Zhu, Y.-G.; Virta, M.P.; Deng, Y. High-Throughput Single-Cell Technology Reveals the Contribution of Horizontal Gene Transfer to Typical Antibiotic Resistance Gene Dissemination in Wastewater Treatment Plants. Environ. Sci. Technol. 2021, 55, 11824–11834. [Google Scholar] [CrossRef]
- Zhou, G.; Qiu, X.; Wu, X.; Lu, S. Horizontal Gene Transfer Is a Key Determinant of Antibiotic Resistance Genes Profiles during Chicken Manure Composting with the Addition of Biochar and Zeolite. J. Hazard. Mater. 2021, 408, 124883. [Google Scholar] [CrossRef]
- Kittredge, H.A.; Dougherty, K.M.; Evans, S.E. Dead but Not Forgotten: How Extracellular DNA, Moisture, and Space Modulate the Horizontal Transfer of Extracellular Antibiotic Resistance Genes in Soil. Appl. Environ. Microbiol. 2022, 88, e02280-21. [Google Scholar] [CrossRef]
- Li, W.; Zhang, W.-G.; Zhang, M.-S.; Lei, Z.-F.; Li, P.-F.; Ma, Y.; Gao, Y. Environmentally Relevant Concentrations of Mercury Facilitate the Horizontal Transfer of Plasmid-Mediated Antibiotic Resistance Genes. Sci. Total Environ. 2022, 852, 158272. [Google Scholar] [CrossRef]
- Mora, M.R.; Gómez, P.A.J.; Valbuena, C.; Probanza, A. Effect of the Type of Vitis Vinifera Cultivation in the Cenophenoresistome and Metabolic Profiling (CLPP) of Edaphic Bacterial Communities. J. Agric. Sci. Technol. A 2017, 7, 522–536. [Google Scholar]
- Millán, R.; Carpena, R.; Schmid, T.; Sierra, M.; Moreno, E.; Peñalosa, J.; Gamarra, R.; Esteban, E. Rehabilitación de Suelos Contaminados Con Mercurio: Estrategias Aplicables En El Área de Almadén. Rev. Ecosistemas 2007, 16, 6. [Google Scholar]
- Ehmann, A. The Van Urk-Salkowski Reagent—A Sensitive and Specific Chromogenic Reagent for Silica Gel Thin-Layer Chromatographic Detection and Identification of Indole Derivatives. J. Chromatogr. A 1977, 132, 267–276. [Google Scholar] [CrossRef]
- Glick, B.R. The Enhancement of Plant Growth by Free-Living Bacteria. Can. J. Microbiol. 1995, 41, 109–117. [Google Scholar] [CrossRef]
- Alexander, D.B.; Zuberer, D.A. Use of Chrome Azurol S Reagents to Evaluate Siderophore Production by Rhizosphere Bacteria. Biol. Fertil. Soils 1991, 12, 39–45. [Google Scholar] [CrossRef]
- De Freitas, J.; Banerjee, M.; Germida, J. Phosphate-Solubilizing Rhizobacteria Enhance the Growth and Yield but Not Phosphorus Uptake of Canola (Brassica Napus L.). Biol. Fertil. Soils 1997, 24, 358–364. [Google Scholar] [CrossRef]
- Lemos, M.L.; Toranzo, A.E.; Barja, J.L. Antibiotic Activity of Epiphytic Bacteria Isolated from Intertidal Seaweeds. Microb. Ecol. 1985, 11, 149–163. [Google Scholar] [CrossRef] [PubMed]
- Abdel Latef, A.A.H.; Abu Alhmad, M.F.; Abdelfattah, K.E. The Possible Roles of Priming with ZnO Nanoparticles in Mitigation of Salinity Stress in Lupine (Lupinus Termis) Plants. J. Plant Growth Regul. 2017, 36, 60–70. [Google Scholar] [CrossRef]
- Velasco, A.G.-V.; Probanza, A.; Mañero, F.G.; Ramos, B.; García, J.L. Functional Diversity of Rhizosphere Microorganisms from Different Genotypes of Arabidopsis Thaliana. Community Ecol. 2009, 10, 111–119. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High throughput Sequence Data. 2010. 2017. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 26 January 2023).
- González, D.; Blanco, C.; Probanza, A.; Jiménez, P.A.; Robas, M. Evaluation of the PGPR Capacity of Four Bacterial Strains and Their Mixtures, Tested on Lupinus Albus Var. Dorado Seedlings, for the Bioremediation of Mercury-Polluted Soils. Processes 2021, 9, 1293. [Google Scholar] [CrossRef]
- Kaur, I.; Gaur, V.K.; Regar, R.K.; Roy, A.; Srivastava, P.K.; Gaur, R.; Manickam, N.; Barik, S.K. Plants Exert Beneficial Influence on Soil Microbiome in a HCH Contaminated Soil Revealing Advantage of Microbe-Assisted Plant-Based HCH Remediation of a Dumpsite. Chemosphere 2021, 280, 130690. [Google Scholar] [CrossRef] [PubMed]
- Manoj, S.R.; Karthik, C.; Kadirvelu, K.; Arulselvi, P.I.; Shanmugasundaram, T.; Bruno, B.; Rajkumar, M. Understanding the Molecular Mechanisms for the Enhanced Phytoremediation of Heavy Metals through Plant Growth Promoting Rhizobacteria: A Review. J. Environ. Manag. 2020, 254, 109779. [Google Scholar] [CrossRef]
- He, S.; Li, Y.; Guo, H.; Lu, L.; Yang, C. Combined Effect of Ryegrass and Hyphomicrobium Sp. GHH on the Remediation of EE2-Cd Co-Contaminated Soil. J. Soils Sediments 2020, 20, 425–434. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, D.; Pan, X.; Lee, D.-J.; Al-Misned, F.A.; Mortuza, M.G.; Gadd, G.M. Aerobic and Anaerobic Biosynthesis of Nano-Selenium for Remediation of Mercury Contaminated Soil. Chemosphere 2017, 170, 266–273. [Google Scholar] [CrossRef]
- Priyadarshanee, M.; Das, S. Bioremediation Potential of Biofilm Forming Multi-Metal Resistant Marine Bacterium Pseudomonas Chengduensis PPSS-4 Isolated from Contaminated Site of Paradip Port, Odisha. J. Earth Syst. Sci. 2021, 130, 125. [Google Scholar] [CrossRef]
- Kumari, W.M.N.H.; Thiruchittampalam, S.; Weerasinghe, M.S.S.; Chandrasekharan, N.V.; Wijayarathna, C.D. Characterization of a Bacillus Megaterium Strain with Metal Bioremediation Potential and in Silico Discovery of Novel Cadmium Binding Motifs in the Regulator, CadC. Appl. Microbiol. Biotechnol. 2021, 105, 2573–2586. [Google Scholar] [CrossRef]
- Faits, T.; Odom-Mabey, A.R.; Castro-Nallar, E.; Crandall, K.A.; Johnson, W.E. Metagenomic Profiling Pipelines Improve Taxonomic Classification for 16S Amplicon Sequencing Data. bioRxiv 2022. [Google Scholar]
- Liu, Y.-X.; Qin, Y.; Chen, T.; Lu, M.; Qian, X.; Guo, X.; Bai, Y. A Practical Guide to Amplicon and Metagenomic Analysis of Microbiome Data. Protein Cell 2021, 12, 315–330. [Google Scholar] [CrossRef]
- González, D.; Robas, M.; Fernández, V.; Bárcena, M.; Probanza, A.; Jiménez, P.A. Comparative Metagenomic Study of Rhizospheric and Bulk Mercury-Contaminated Soils in the Mining District of Almadén. Front. Microbiol. 2022, 13, 797444. [Google Scholar] [CrossRef] [PubMed]
- Mora, M.; Probanza, A.; Gómez, P. Metagenomic Study of the Taxonomic Profile of Rhizobacterial Communities in Soils Contaminated with Mercury. Acta Sci. Microbiol. 2020, 3, 1–13. [Google Scholar] [CrossRef]
- López-Casaperalta, P.; Febres-Molina, C.; Aguilar-Pineda, J.A.; Bernabe-Ortiz, J.C.; Fernandez-F, F. Peruvian Native Bacterial Strains as Potential Bioremediation Agents in Hg-Polluted Soils by Artisanal Mining Activities in Southern Peru. Sustainability 2022, 14, 10272. [Google Scholar] [CrossRef]
- Imron, M.F.; Kurniawan, S.B.; Abdullah, S.R.S. Resistance of Bacteria Isolated from Leachate to Heavy Metals and the Removal of Hg by Pseudomonas Aeruginosa Strain FZ-2 at Different Salinity Levels in a Batch Biosorption System. Sustain. Environ. Res. 2021, 31, 14. [Google Scholar] [CrossRef]
- Mora, M.R.; Pastrana, V.M.F.; Reguero, D.G.; Oliva, L.L.G.; Lobo, A.P.; Gómez, P.A.J. Oxidative Stress Protection and Growth Promotion Activity of Pseudomonas Mercuritolerans Sp. Nov., in Forage Plants under Mercury Abiotic Stress Conditions. Front. Microbiol. 2022, 13, 1032901. [Google Scholar] [CrossRef]
- González-Reguero, D.; Robas-Mora, M.; Probanza, A.; Jiménez, P.A. Evaluation of the Oxidative Stress Alleviation in Lupinus Albus Var. Orden Dorado by the Inoculation of Four Plant Growth-Promoting Bacteria and Their Mixtures in Mercury-Polluted Soils. Front. Microbiol. 2022, 13, 907557. [Google Scholar] [CrossRef]
- Bhatia, M.; Khan, M.L.; Qureshi, A. Microbial Remediation of Mercury-Contaminated Soils. Microbes Microb. Biotechnol. Green Remediat. 2022, 633–649. [Google Scholar] [CrossRef]
- Garbeva, P.v.; Van Veen, J.A.; Van Elsas, J.D. Microbial Diversity in Soil: Selection of Microbial Populations by Plant and Soil Type and Implications for Disease Suprressiveness. Annu. Rev. Phytopathol. 2004, 42, 243. [Google Scholar] [CrossRef]
- Hawkes, C.V.; DeAngelis, K.M.; Firestone, M.K. CHAPTER 1—Root Interactions with Soil Microbial Communities and Processes. In The Rhizosphere; Cardon, Z.G., Whitbeck, J.L., Eds.; Academic Press: Burlington, VT, USA, 2007; pp. 1–29. ISBN 978-0-12-088775-0. [Google Scholar]
- Ambrosini, A.; de Souza, R.; Passaglia, L.M. Ecological Role of Bacterial Inoculants and Their Potential Impact on Soil Microbial Diversity. Plant Soil 2016, 400, 193–207. [Google Scholar] [CrossRef]
- Chaudhary, P.; Khati, P.; Chaudhary, A.; Maithani, D.; Kumar, G.; Sharma, A. Cultivable and Metagenomic Approach to Study the Combined Impact of Nanogypsum and Pseudomonas Taiwanensis on Maize Plant Health and Its Rhizospheric Microbiome. PLoS ONE 2021, 16, e0250574. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Su, J.-Q.; Yang, X.-R.; Zhu, Y.-G. Distinct Rhizosphere Effect on Active and Total Bacterial Communities in Paddy Soils. Sci. Total Environ. 2019, 649, 422–430. [Google Scholar] [CrossRef] [PubMed]
- Wu, A.-L.; Jiao, X.-Y.; Wang, J.-S.; Dong, E.-W.; Guo, J.; Wang, L.-G.; Sun, A.-Q.; Hu, H.-W. Sorghum Rhizosphere Effects Reduced Soil Bacterial Diversity by Recruiting Specific Bacterial Species under Low Nitrogen Stress. Sci. Total Environ. 2021, 770, 144742. [Google Scholar] [CrossRef]
- Zhang, P.; Li, B.; Wu, J.; Hu, S. Invasive Plants Differentially Affect Soil Biota through Litter and Rhizosphere Pathways: A Meta-analysis. Ecol. Lett. 2019, 22, 200–210. [Google Scholar] [CrossRef]
- Enebe, M.C.; Babalola, O.O. Effects of Inorganic and Organic Treatments on the Microbial Community of Maize Rhizosphere by a Shotgun Metagenomics Approach. Ann. Microbiol. 2020, 70, 49. [Google Scholar] [CrossRef]
- Eze, M.O.; Thiel, V.; Hose, G.C.; George, S.C.; Daniel, R. Metagenomic Insight into the Plant Growth-Promoting Potential of a Diesel-Degrading Bacterial Consortium for Enhanced Rhizoremediation Application. bioRxiv 2021. [Google Scholar] [CrossRef]
- Tara, N.; Saharan, B.S. Plant Growth Promoting Traits Shown by Bacteria Brevibacterium Frigrotolerans SMA23 Isolated from Aloe Vera Rhizosphere. Agric. Sci. Dig.-Res. J. 2017, 37, 226–231. [Google Scholar]
- Raufa, B.; Mazhar, R.; Javed, A.; Tehmeena, M.; Shehzad, M.; Tariq, S.; Munis, F.; Chaudhary, H. Biocontrol Potential of Bacillus Gibsonii and Brevibacterium Frigoritolerans in Suppression of Fusarium Stalk Rot of Maize: A Sustainable Approach. Asian J. Agric. Biol. 2019, 7, 320–333. [Google Scholar]
- Sultan, I.; Ali, A.; Gogry, F.A.; Rather, I.A.; Sabir, J.S.; Haq, Q.M. Bacterial Isolates Harboring Antibiotics and Heavy-Metal Resistance Genes Co-Existing with Mobile Genetic Elements in Natural Aquatic Water Bodies. Saudi J. Biol. Sci. 2020, 27, 2660–2668. [Google Scholar] [CrossRef]
- Jin, M.; Zhao, Q.; Zhou, Z.; Zhu, L.; Zhang, Z.; Jiang, L. Draft Genome Sequence of a Potential Organic Phosphorus-Degrading Bacterium Brevibacterium Frigoritolerans GD44, Isolated from Radioactive Soil in Xinjiang, China. Curr. Microbiol. 2020, 77, 2896–2903. [Google Scholar] [CrossRef] [PubMed]
- Hiebert, N.; Carrau, T.; Bartling, M.; Vilcinskas, A.; Lee, K.-Z. Identification of Entomopathogenic Bacteria Associated with the Invasive Pest Drosophila Suzukii in Infested Areas of Germany. J. Invertebr. Pathol. 2020, 173, 107389. [Google Scholar] [CrossRef] [PubMed]
- Radisic, V.; Marathe, N.P. Genomic Characterisation of Multidrug-Resistant Bacillus Toyonensis Strain 4HC1 Isolated from Marine Plastic in Norway. J. Glob. Antimicrob. Resist. 2021, 26, 249–251. [Google Scholar] [CrossRef] [PubMed]
- Fiedler, G.; Schneider, C.; Igbinosa, E.O.; Kabisch, J.; Brinks, E.; Becker, B.; Stoll, D.A.; Cho, G.-S.; Huch, M.; Franz, C.M. Antibiotics Resistance and Toxin Profiles of Bacillus Cereus-Group Isolates from Fresh Vegetables from German Retail Markets. BMC Microbiol. 2019, 19, 250. [Google Scholar] [CrossRef]
- Tvrzova, L.; Schumann, P.; Spröer, C.; Sedláček, I.; Páčová, Z.; Šedo, O.; Zdráhal, Z.; Steffen, M.; Lang, E. Pseudomonas Moraviensis Sp. Nov. and Pseudomonas Vranovensis Sp. Nov., Soil Bacteria Isolated on Nitroaromatic Compounds, and Emended Description of Pseudomonas Asplenii. Int. J. Syst. Evol. Microbiol. 2006, 56, 2657–2663. [Google Scholar] [CrossRef]
- Rasmussen, L.D.; Sørensen, S.J. The Effect of Longterm Exposure to Mercury on the Bacterial Community in Marine Sediment. Curr. Microbiol. 1998, 36, 291–297. [Google Scholar] [CrossRef]
- Vega, N.M.; Gore, J. Collective Antibiotic Resistance: Mechanisms and Implications. Curr. Opin. Microbiol. 2014, 21, 28–34. [Google Scholar] [CrossRef]
- Song, T.; Zhu, C.; Xue, S.; Li, B.; Ye, J.; Geng, B.; Li, L.; Sardar, M.F.; Li, N.; Feng, S. Comparative Effects of Different Antibiotics on Antibiotic Resistance during Swine Manure Composting. Bioresour. Technol. 2020, 315, 123820. [Google Scholar] [CrossRef]
- Tetz, G.; Tetz, V. Overcoming Antibiotic Resistance with Novel Paradigms of Antibiotic Selection. Microorganisms 2022, 10, 2383. [Google Scholar] [CrossRef]
- Armalytė, J.; Skerniškytė, J.; Bakienė, E.; Krasauskas, R.; Šiugždinienė, R.; Kareivienė, V.; Kerzienė, S.; Klimienė, I.; Sužiedėlienė, E.; Ružauskas, M. Microbial Diversity and Antimicrobial Resistance Profile in Microbiota from Soils of Conventional and Organic Farming Systems. Front. Microbiol. 2019, 10, 892. [Google Scholar] [CrossRef]
- Duan, M.; Zhang, Y.; Zhou, B.; Wang, Q.; Gu, J.; Liu, G.; Qin, Z.; Li, Z. Changes in Antibiotic Resistance Genes and Mobile Genetic Elements during Cattle Manure Composting after Inoculation with Bacillus Subtilis. Bioresour. Technol. 2019, 292, 122011. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Ben, W.; Qiang, Z.; Zhang, J. Removal of Antibiotic Resistance Genes in Pig Manure Composting Influenced by Inoculation of Compound Microbial Agents. Bioresour. Technol. 2020, 317, 123966. [Google Scholar] [CrossRef] [PubMed]
- Shamsizadeh, Z.; Ehrampoush, M.H.; Nikaeen, M.; Mohammadi, F.; Mokhtari, M.; Gwenzi, W.; Khanahmad, H. Antibiotic Resistance and Class 1 Integron Genes Distribution in Irrigation Water-Soil-Crop Continuum as a Function of Irrigation Water Sources. Environ. Pollut. 2021, 289, 117930. [Google Scholar] [CrossRef] [PubMed]
- Kaviani Rad, A.; Balasundram, S.K.; Azizi, S.; Afsharyzad, Y.; Zarei, M.; Etesami, H.; Shamshiri, R.R. An Overview of Antibiotic Resistance and Abiotic Stresses Affecting Antimicrobial Resistance in Agricultural Soils. Int. J. Environ. Res. Public Health 2022, 19, 4666. [Google Scholar] [CrossRef]
- Li, Z.; Sun, A.; Liu, X.; Chen, Q.-L.; Bi, L.; Ren, P.-X.; Shen, J.-P.; Jin, S.; He, J.-Z.; Hu, H.-W.; et al. Climate Warming Increases the Proportions of Specific Antibiotic Resistance Genes in Natural Soil Ecosystems. J. Hazard. Mater. 2022, 430, 128442. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Wang, J.; Li, Z.; Guo, S.; Li, K.; Xu, P.; Ok, Y.S.; Jones, D.L.; Zou, J. Antibiotics and Antibiotic Resistance Genes in Agricultural Soils: A Systematic Analysis. Crit. Rev. Environ. Sci. Technol. 2022, 53, 847–864. [Google Scholar] [CrossRef]

| Strain | HgCl2 Tolerance (µg mL−1) | BMRSI | Strain Origin | 16S rRNA Identification |
|---|---|---|---|---|
| A1 | 140 | 6.54 | Avena sativa | Brevibacterium frigoritolerans |
| A2 | 140 | 7.30 | Bulk soil | Bacillus toyonensis |
| B1 | 140 | 7.20 | Bulk soil | Pseudomonas mercuritolerans |
| B2 | 140 | 6.92 | Avena sativa | Pseudomonas baetica |
| CS1 | CS2 | CS3 | CS4 | CS5 | CS6 | |
|---|---|---|---|---|---|---|
| Strains | A1+B1 | A1+A2 | A1+B2 | B1+A2 | B1+B2 | A2+B2 |
| Soil | D | H’ |
|---|---|---|
| Cont S | 0.985 | 4.464 |
| Cont P | 0.982 | 3.998 |
| A1 | A2 | B1 | B2 | |
|---|---|---|---|---|
| 16S rRNA identification | Brevibacterium frigoritolerans | Bacillus toyonensis | Pseudomonas mercuritolerans | Pseudomonas baetica |
| Cefuroxime | 1 | 16 | 16 | 16 |
| Cefuroxim eAxetil | 1 | 8 | 16 | 16 |
| Cefoxitin | 8 | 8 | 16 | 16 |
| Cefotaxime | 1 | 8 | 8 | 8 |
| Ceftacidime | 0.5 | 8 | 4 | 4 |
| Cefepime | 1 | 1 | 2 | 2 |
| Ertapenem | 0.5 | 0.5 | 0.5 | 0.5 |
| Imipenem | 0.25 | 0.25 | 0.25 | 0.25 |
| Amikacin | 2 | 2 | 2 | 2 |
| Gentamicin | 1 | 1 | 1 | 1 |
| Nalidixic Acid | 4 | 4 | 2 | 2 |
| Ciprofloxacin | 0.5 | 0.5 | 0.25 | 0.5 |
| Tigecyclina | 0.5 | 0.5 | 0.5 | 0.5 |
| Trimethoprim/Sulfomethoxazole | 20 | 20 | 20 | 20 |
| Treatment | Cont S | Cont P | A1 | A2 | B1 | B2 | CS1 | CS2 | CS3 | CS4 | CS5 | CS6 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cefuroxime | 64 | 32 | 8 | 32 | 32 | 32 | 8 | 8 | 8 | 32 | 32 | 16 |
| Cefuroxime Axetil | 64 | 32 | 8 | 32 | 32 | 32 | 16 | 16 | 16 | 32 | 32 | 16 |
| Cefoxitin | 64 | 32 | 8 | 32 | 32 | 32 | 8 | 16 | 16 | 32 | 32 | 32 |
| Cefotaxime | 16 | 16 | 1 | 8 | 16 | 16 | 2 | 8 | 8 | 8 | 16 | 16 |
| Ceftacidime | 16 | 8 | 0.5 | 8 | 8 | 8 | 0.5 | 4 | 4 | 8 | 16 | 16 |
| Cefepime | 4 | 2 | 1 | 2 | 2 | 2 | 2 | 1 | 2 | 2 | 2 | 2 |
| Ertapenem | 4 | 4 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 |
| Imipenem | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 | 0.25 |
| Amicacin | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Gentamicin | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| Nalidixic Acid | 8 | 8 | 4 | 4 | 2 | 2 | 4 | 8 | 8 | 2 | 2 | 2 |
| Ciprofloxacin | 0.5 | 0.5 | 0.5 | 0.5 | 0.25 | 0.5 | 0.5 | 0.5 | 0.25 | 0.25 | 0.25 | 0.5 |
| Tigecyclina | 2 | 1 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 |
| Trimethoprime/Sulfomethoxazole | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 |
| Component | Total | % SD | % Accumulated |
|---|---|---|---|
| 1 | 10.26 | 85.46 | 85.46 |
| 2 | 1.22 | 10.16 | 95.64 |
| Treatment | Cont S | Cont P | A1 | A2 | B1 | B2 | CS1 | CS2 | CS3 | CS4 | CS5 | CS6 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cefuroxime | 64 | 32 | 8 a,b | 32 | 32 | 32 | 8 a,b | 8 a,b | 8 a | 32 | 32 | 16 a |
| Cefuroxime Axetil | 64 | 32 | 8 a,b | 32 | 32 | 32 | 16 a,b | 16 a,b | 16 a,b | 32 | 32 | 16 a,b |
| Cefoxitin | 64 | 32 | 8 a,b | 32 | 32 | 32 | 8 a,b | 16 a | 16 a | 32 | 32 | 32 |
| Cefotaxime | 16 | 16 | 1 a,b | 8 a,b | 16 | 16 | 2 a,b | 8 a,b | 8 a,b | 8 a,b | 16 | 16 |
| Ceftazidime | 16 | 8 | 1 a,b | 8 | 8 | 8 | 1 a,b | 4 a | 4 a | 8 | 16 | 16 |
| Cefepime | 4 | 2 | 1 a,b | 2 a | 2 a | 2 a | 2 a | 1 a,b | 2 a | 2 a | 2 a | 2 a |
| Ertapenem | 4 | 4 | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b |
| Nalidixic Acid | 8 | 8 | 4 | 4 | 2 a,b | 2 a,b | 4 | 8 | 8 | 2 a,b | 2 a,b | 2 a,b |
| Ciprofloxacin | 0.5 | 0.5 | 0.5 | 0.5 | 0.25 a,b | 0.5 | 0.5 | 0.5 | 0.25 a,b | 0.25 a,b | 0.25 a,b | 0.5 |
| Tigecycline | 2 | 1 | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b | 0.5 a,b |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
González-Reguero, D.; Robas-Mora, M.; Fernández-Pastrana, V.M.; Probanza-Lobo, A.; Jiménez-Gómez, P.A. Reduced Antibiotic Resistance in the Rhizosphere of Lupinus albus in Mercury-Contaminated Soil Mediated by the Addition of PGPB. Biology 2023, 12, 801. https://doi.org/10.3390/biology12060801
González-Reguero D, Robas-Mora M, Fernández-Pastrana VM, Probanza-Lobo A, Jiménez-Gómez PA. Reduced Antibiotic Resistance in the Rhizosphere of Lupinus albus in Mercury-Contaminated Soil Mediated by the Addition of PGPB. Biology. 2023; 12(6):801. https://doi.org/10.3390/biology12060801
Chicago/Turabian StyleGonzález-Reguero, Daniel, Marina Robas-Mora, Vanesa M. Fernández-Pastrana, Agustín Probanza-Lobo, and Pedro Antonio Jiménez-Gómez. 2023. "Reduced Antibiotic Resistance in the Rhizosphere of Lupinus albus in Mercury-Contaminated Soil Mediated by the Addition of PGPB" Biology 12, no. 6: 801. https://doi.org/10.3390/biology12060801
APA StyleGonzález-Reguero, D., Robas-Mora, M., Fernández-Pastrana, V. M., Probanza-Lobo, A., & Jiménez-Gómez, P. A. (2023). Reduced Antibiotic Resistance in the Rhizosphere of Lupinus albus in Mercury-Contaminated Soil Mediated by the Addition of PGPB. Biology, 12(6), 801. https://doi.org/10.3390/biology12060801










