Transcriptomic Profiles of Long Noncoding RNAs and Their Target Protein-Coding Genes Reveals Speciation Adaptation on the Qinghai-Xizang (Tibet) Plateau in Orinus
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Samples
2.2. Overall Transcriptome Mapping and Assembly
2.3. Pipeline for lncRNA Identification
2.4. Differential Expression Analysis
2.5. Prediction of Cis-Regulated Target Genes of DElncRNAs
2.6. Gene Ontology Enrichment Analysis
2.7. Transcription Factor Survey
3. Results
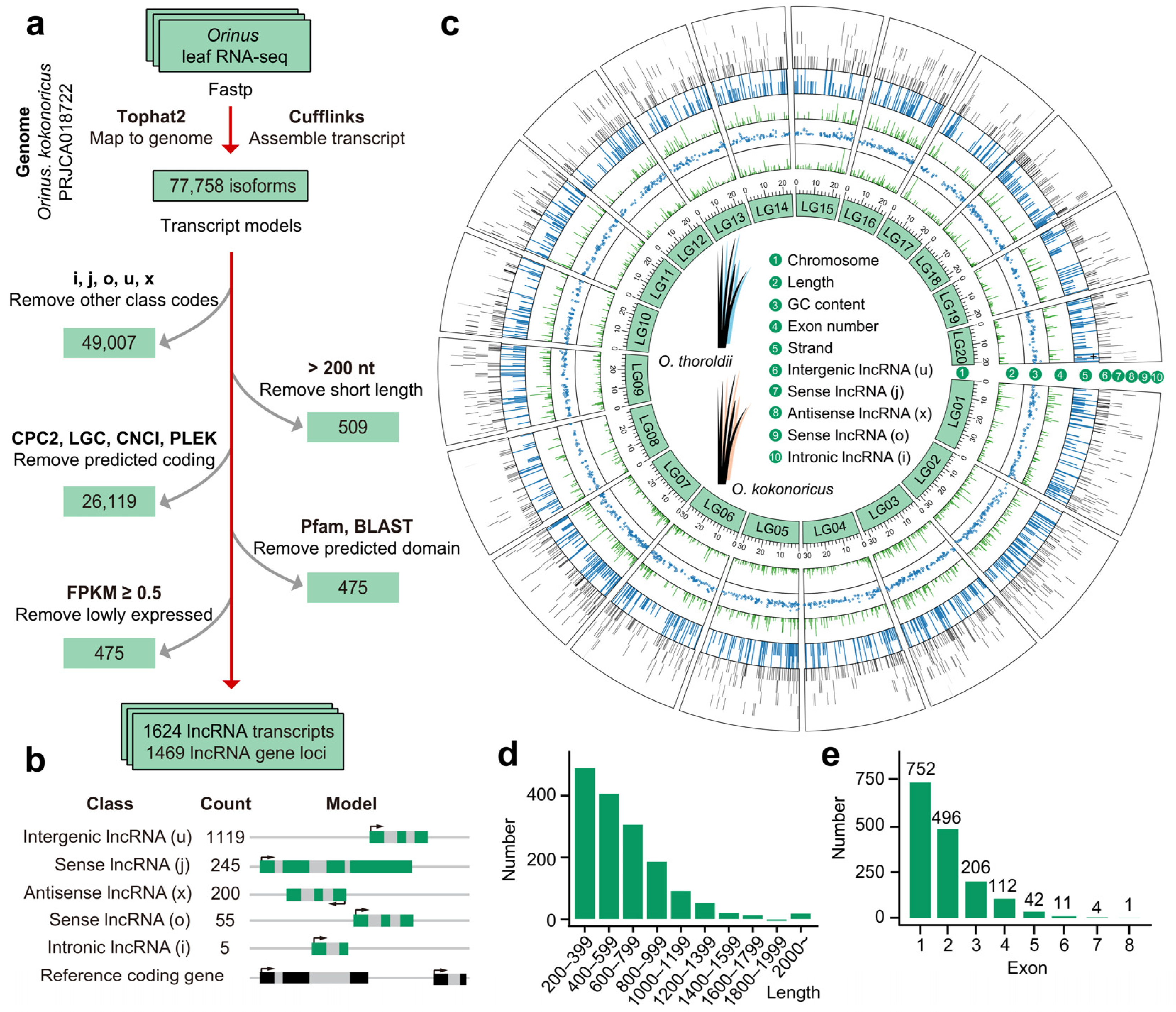
3.1. LncRNA Screening Lays the Foundation for the Further Understanding of the Molecular and Biological Functions of Non-Coding Genes in Orinus
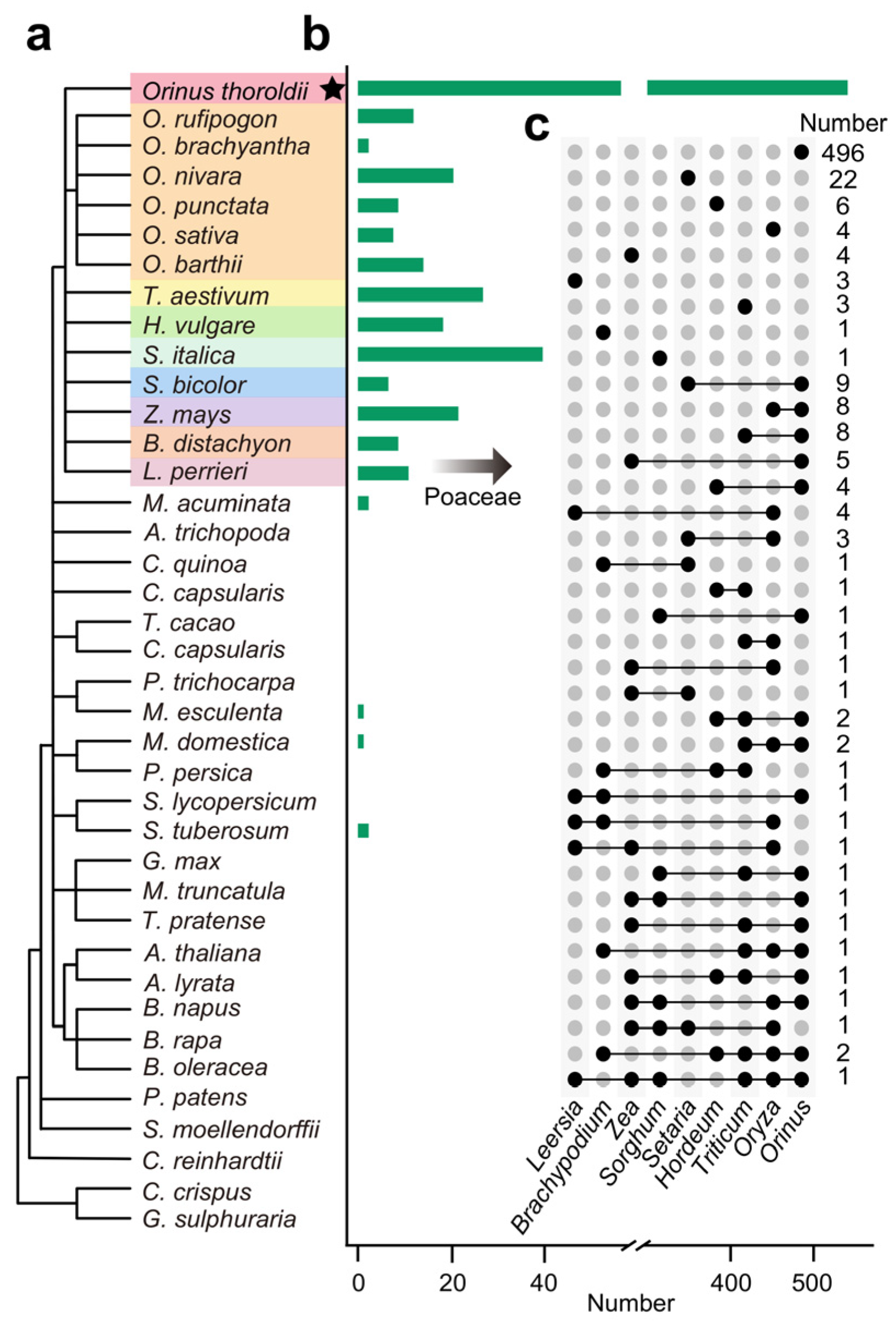
3.2. The Limited Sequence Conservation of lncRNAs in Orinus Implies That Species-Specific Noncoding RNAs Might Contribute to Speciation Adaptation Evolution
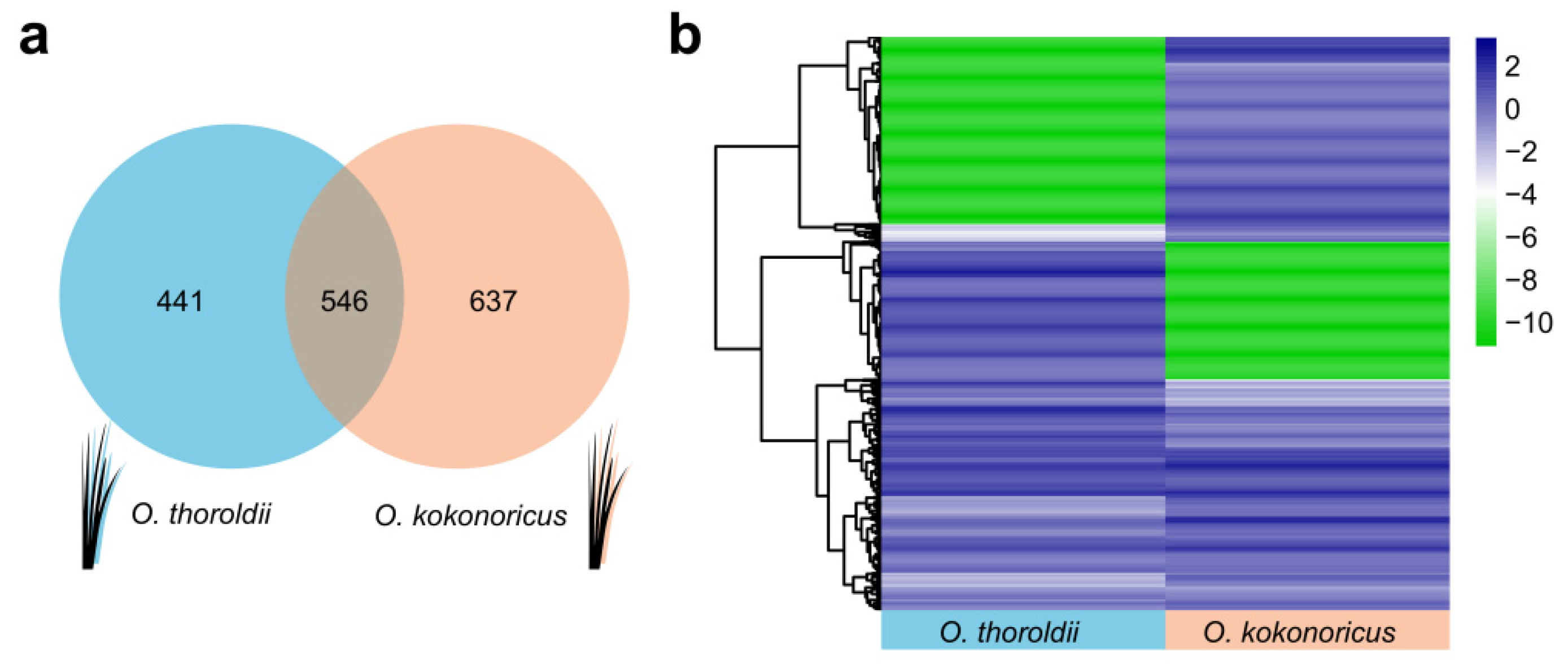
3.3. DElncRNAs from Two Species in Orinus Provide Evidence for the Analysis of Speciation within the Genus
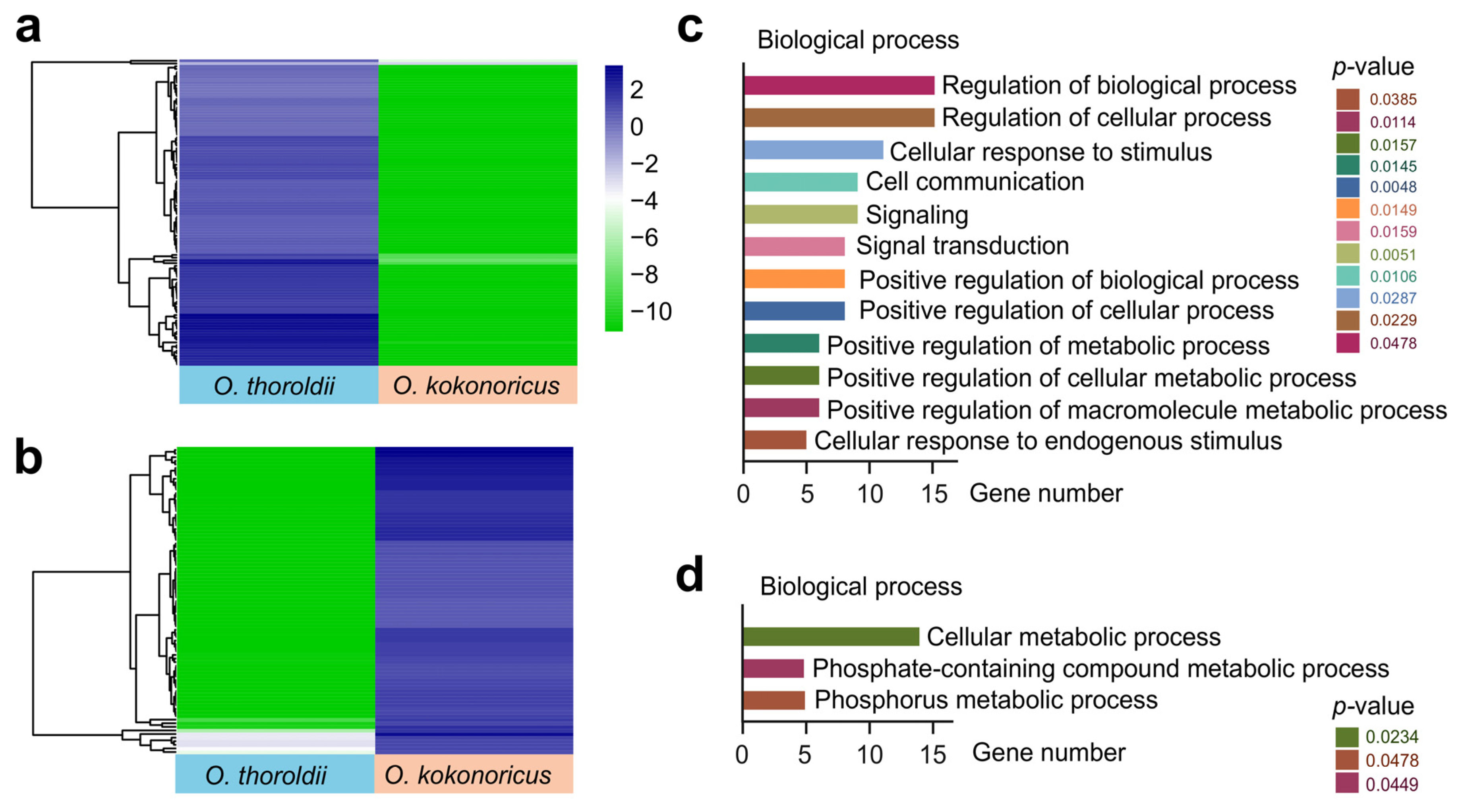
3.4. Through Potential Cis-Regulation Patterns, DElncRNAs, and Their Target PCGs in Orinus Participate in Different Biological Processes
3.5. Detailed Investigation of Potential LncRNA Target Genes Provides Further Understanding of Speciation Adaptation about O. thoroldii
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef] [PubMed]
- Ponjavic, J.; Ponting, C.P.; Lunter, G. Functionality or transcriptional noise? Evidence for selection within long noncoding RNAs. Genome Res. 2007, 17, 556–565. [Google Scholar] [CrossRef] [PubMed]
- Pauli, A.; Rinn, J.L.; Schier, A.F. Noncoding RNAs as regulators of embryogenesis. Nat. Rev. Genet. 2011, 12, 136–149. [Google Scholar] [CrossRef] [PubMed]
- Schaukowitch, K.; Kim, T.K. Emerging epigenetic mechanisms of long noncoding RNAs. Neuroscience 2014, 264, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Long, H.; Zhou, C.; Zheng, S.; Wu, H.; Guo, T.; Wu, Q.; Zhong, T.; Wang, T. Long noncoding RNA Braveheart promotes cardiogenic differentiation of mesenchymal stem cells in vitro. Stem Cell Res. Ther. 2017, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Peng, W.X.; Koirala, P.; Mo, Y.Y. LncRNA-mediated regulation of cell signaling in cancer. Oncogene 2017, 36, 5661–5667. [Google Scholar] [CrossRef] [PubMed]
- Ewart, K.M.; Johnson, R.N.; Ogden, R.; Joseph, L.; Frankham, G.J.; Lo, N. Museum specimens provide reliable SNP data for population genomic analysis of a widely distributed but threatened cockatoo species. Mol. Ecol. Resour. 2019, 19, 1578–1592. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Li, J.; Lian, B.; Gu, H.; Li, Y.; Qi, Y. Global identification of Arabidopsis lncRNAs reveals the regulation of MAF4 by a natural antisense RNA. Nat. Commun. 2018, 9, 5056. [Google Scholar] [CrossRef]
- Ulitsky, I.; Bartel, D.P. lincRNAs: Genomics, evolution, and mechanisms. Cell 2013, 154, 26–46. [Google Scholar] [CrossRef]
- Sarropoulos, I.; Marin, R.; Cardoso-Moreira, M.; Kaessmann, H. Developmental dynamics of lncRNAs across mammalian organs and species. Nature 2019, 571, 510–514. [Google Scholar] [CrossRef]
- Wilusz, J.E.; Sunwoo, H.; Spector, D.L. Long noncoding RNAs: Functional surprises from the RNA world. Genes Dev. 2009, 23, 1494–1504. [Google Scholar] [CrossRef]
- Marquardt, S.; Raitskin, O.; Wu, Z.; Liu, F.; Sun, Q.; Dean, C. Functional consequences of splicing of the antisense transcript COOLAIR on FLC transcription. Mol. Cell 2014, 54, 156–165. [Google Scholar] [CrossRef]
- Hu, X.; Kong, X.; Wang, C.; Ma, L.; Zhao, J.; Wei, J.; Zhang, X.; Loake, G.J.; Zhang, T.; Huang, J.; et al. Proteasome-mediated degradation of FRIGIDA modulates flowering time in Arabidopsis during vernalization. Plant Cell 2014, 26, 4763–4781. [Google Scholar] [CrossRef]
- Helliwell, C.A.; Robertson, M.; Finnegan, E.J.; Buzas, D.M.; Dennis, E.S. Vernalization-repression of Arabidopsis FLC requires promoter sequences but not antisense transcripts. PLoS ONE 2011, 6, e21513. [Google Scholar] [CrossRef]
- Qin, T.; Zhao, H.; Cui, P.; Albesher, N.; Xiong, L. A Nucleus-Localized Long Noncoding RNA Enhances Drought and Salt Stress Tolerance. Plant Physiol. 2017, 175, 1321–1336. [Google Scholar] [CrossRef] [PubMed]
- Wunderlich, M.; Gross-Hardt, R.; Schöffl, F. Heat shock factor HSFB2a involved in gametophyte development of Arabidopsis thaliana and its expression is controlled by a heat-inducible long noncoding antisense RNA. Plant Mol. Biol. 2014, 85, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Dong, Q.; Deng, M.; Lin, D.; Xiao, J.; Cheng, P.; Xing, L.; Niu, Y.; Gao, C.; Zhang, W.; et al. The vernalization-induced long noncoding RNA VAS functions with the transcription factor TaRF2b to promote TaVRN1 expression for flowering in hexaploid wheat. Mol. Plant 2021, 14, 1525–1538. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Xin, Z.; Huang, Y.; Yu, J. Climate suitability assessment on the Qinghai-Tibet Plateau. Sci. Total Environ. 2022, 816, 151653. [Google Scholar] [CrossRef]
- Wang, J.; Wang, Y.; Qu, M.; Feng, Y.; Wu, B.; Lu, Q.; He, N.; Li, J. Testing the Functional and Phylogenetic Assembly of Plant Communities in Gobi Deserts of Northern Qinghai-Tibet Plateau. Front. Plant Sci. 2022, 13, 952074. [Google Scholar] [CrossRef]
- Su, X.; Liu, Y.P.; Chen, Z.; Chen, K.L. Evaluation of candidate barcoding markers in Orinus (Poaceae). Genet. Mol. Res. 2016, 15, 27714. [Google Scholar] [CrossRef] [PubMed]
- Su, X.; Wu, G.; Li, L.; Liu, J. Species delimitation in plants using the Qinghai-Tibet Plateau endemic Orinus (Poaceae: Tridentinae) as an example. Ann. Bot. 2015, 116, 35–48. [Google Scholar] [CrossRef]
- Qu, K.; Liu, A.; Yin, M.; Mu, W.; Wu, S.; Hu, H.; Chen, J.; Su, X.; Dou, Q.; Ren, G. A genome assembly for Orinus kokonorica provides insights into the origin.; adaptive evolution and further diversification of two closely related grass genera. Commun. Biol. 2023, 6, 1223. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; García, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef] [PubMed]
- Ben Amor, B.; Wirth, S.; Merchan, F.; Laporte, P.; d’Aubenton-Carafa, Y.; Hirsch, J.; Maizel, A.; Mallory, A.; Lucas, A.; Deragon, J.M.; et al. Novel long non-protein coding RNAs involved in Arabidopsis differentiation and stress responses. Genome Res. 2009, 19, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Fedak, H.; Palusinska, M.; Krzyczmonik, K.; Brzezniak, L.; Yatusevich, R.; Pietras, Z.; Kaczanowski, S.; Swiezewski, S. Control of seed dormancy in Arabidopsis by a cis-acting noncoding antisense transcript. Proc. Natl. Acad. Sci. USA 2016, 113, E7846–E7855. [Google Scholar] [CrossRef] [PubMed]
- Su, X.; Liu, T.; Liu, Y.P.; Harris, A.J.; Chen, J.Y. Adaptive radiation in Orinus, an endemic alpine grass of the Qinghai-Tibet Plateau.; based on comparative transcriptomic analysis. J. Plant Physiol. 2022, 277, 153786. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.J.; Yang, D.C.; Kong, L.; Hou, M.; Meng, Y.Q.; Wei, L.; Gao, G. CPC2: A fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res. 2017, 45, W12–W16. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.K.; Wang, H. Computational Analysis Predicts Hundreds of Coding lncRNAs in Zebrafish. Biology 2021, 10, 371. [Google Scholar] [CrossRef]
- Zhao, J.; Song, X.; Wang, K. lncScore: Alignment-free identification of long noncoding RNA from assembled novel transcripts. Sci. Rep. 2016, 6, 34838. [Google Scholar] [CrossRef]
- Li, A.; Zhang, J.; Zhou, Z. PLEK: A tool for predicting long non-coding RNAs and messenger RNAs based on an improved k-mer scheme. BMC Bioinform. 2014, 15, 311. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Gan, L.; Sun, P.; Wang, J.; Li, D.; Cao, Y.; Fu, Y.; Li, P.; Bai, X.; Li, K.; et al. The long non-coding RNA LNC_000397 negatively regulates PRRSV replication through induction of interferon-stimulated genes. Virol. J. 2022, 19, 40. [Google Scholar] [CrossRef] [PubMed]
- Zheng, K.; Wu, X.; Xue, X.; Li, W.; Wang, Z.; Chen, J.; Zhang, Y.; Qiao, F.; Zhao, H.; Zhang, F.; et al. Transcriptome Screening of Long Noncoding RNAs and Their Target Protein-Coding Genes Unmasks a Dynamic Portrait of Seed Coat Coloration Associated with Anthocyanins in Tibetan Hulless Barley. Int. J. Mol. Sci. 2023, 24, 10587. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef] [PubMed]
- Ke, J.; Du, X.; Cui, J.; Yu, L.; Li, H. LncRNA and mRNA expression associated with myasthenia gravis in patients with thymoma. Thorac. Cancer 2022, 13, 15–23. [Google Scholar] [CrossRef]
- Zheng, K.; Wang, Z.; Pang, L.; Song, Z.; Zhao, H.; Wang, Y.; Wang, B.; Han, S. Systematic Identification of Methyl Jasmonate-Responsive Long Noncoding RNAs and Their Nearby Coding Genes Unveils Their Potential Defence Roles in Tobacco BY-2 Cells. Int. J. Mol. Sci. 2022, 23, 15568. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Tian, F.; Yang, D.C.; Meng, Y.Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2017, 45, D1040–D1045. [Google Scholar] [CrossRef]
- Ding, J.; Lu, Q.; Ouyang, Y.; Mao, H.; Zhang, P.; Yao, J.; Xu, C.; Li, X.; Xiao, J.; Zhang, Q. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc. Natl. Acad. Sci. USA 2012, 109, 2654–2659. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Csorba, T.; Skourti-Stathaki, K.; Proudfoot, N.J.; Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 2013, 340, 619–621. [Google Scholar] [CrossRef]
- Zou, C.; Wang, Q.; Lu, C.; Yang, W.; Zhang, Y.; Cheng, H.; Feng, X.; Prosper, M.A.; Song, G. Transcriptome analysis reveals long noncoding RNAs involved in fiber development in cotton (Gossypium arboreum). Sci. China Life Sci. 2016, 59, 164–171. [Google Scholar] [CrossRef]
- Zhang, W.; Han, Z.; Guo, Q.; Liu, Y.; Zheng, Y.; Wu, F.; Jin, W. Identification of maize long noncoding RNAs responsive to drought stress. PLoS ONE 2014, 9, e98958. [Google Scholar]
- Liu, N.; Xu, Y.; Li, Q.; Cao, Y.P.; Yang, D.; Liu, S.; Wang, X.; Mi, Y.; Liu, Y.; Ding, C.; et al. A lncRNA fine-tunes salicylic acid biosynthesis to balance plant immunity and growth. Cell Host Microbe 2022, 30, 1124–1138.e8. [Google Scholar]
- Ulitsky, I.; Shkumatava, A.; Jan, C.H.; Sive, H.; Bartel, D.P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 2011, 147, 1537–1550. [Google Scholar] [CrossRef]
- Sarkar, N.K.; Kundnani, P.; Grover, A. Functional analysis of Hsp70 superfamily proteins of rice (Oryza sativa). Cell Stress Chaperones 2013, 18, 427–437. [Google Scholar] [CrossRef] [PubMed]
- Ross, C.J.; Ulitsky, I. Discovering functional motifs in long noncoding RNAs. Wiley Interdiscip. Rev. RNA 2022, 13, e1708. [Google Scholar] [CrossRef]
- Deng, P.; Liu, S.; Nie, X.; Weining, S.; Wu, L. Conservation analysis of long noncoding RNAs in plants. Sci. China Life Sci. 2018, 61, 190–198. [Google Scholar] [CrossRef] [PubMed]
- Brosnan, C.A.; Voinnet, O. The long and the short of noncoding RNAs. Curr. Opin. Cell Biol. 2009, 21, 416–425. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.R.; Mondal, T.; Mohammad, F.; Enroth, S.; Redrup, L.; Komorowski, J.; Nagano, T.; Mancini-Dinardo, D.; Kanduri, C. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell 2008, 32, 232–246. [Google Scholar] [CrossRef]
- Nagano, T.; Mitchell, J.A.; Sanz, L.A.; Pauler, F.M.; Ferguson-Smith, A.C.; Feil, R.; Fraser, P. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 2008, 322, 1717–1720. [Google Scholar] [CrossRef]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Hah, N.; Murakami, S.; Nagari, A.; Danko, C.G.; Kraus, W.L. Enhancer transcripts mark active estrogen receptor binding sites. Genome Res. 2013, 23, 1210–1223. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Froberg, J.E.; Lee, J.T. Long noncoding RNAs: Fresh perspectives into the RNA world. Trends Biochem. Sci. 2014, 39, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Liu, F.; Wang, Y.; Liu, X. A novel long noncoding RNA CIL1 enhances cold stress tolerance in Arabidopsis. Plant Sci. 2022, 323, 111370. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, K.; Takasaki, H.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. NAC transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta 2012, 1819, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Koyama, H.; Bhati, K.K.; Alok, A. The biotechnological importance of the plant-specific NAC transcription factor family in crop improvement. J. Plant Res. 2021, 134, 475–495. [Google Scholar] [CrossRef] [PubMed]
- Yan, P.; Du, Q.; Chen, H.; Guo, Z.; Wang, Z.; Tang, J.; Li, W.X. Biofortification of iron content by regulating a NAC transcription factor in maize. Science 2023, 382, 1159–1165. [Google Scholar] [CrossRef]
- Li, X.; Wang, Q.; Guo, C.; Sun, J.; Li, Z.; Wang, Y.; Yang, A.; Pu, W.; Guo, Y.; Gao, J.; et al. NtNAC053, A Novel NAC Transcription Factor, Confers Drought and Salt Tolerances in Tobacco. Front. Plant Sci. 2022, 13, 817106. [Google Scholar] [CrossRef]
- Hao, Y.; Zong, X.; Ren, P.; Qian, Y.; Fu, A. Basic Helix-Loop-Helix (bHLH) Transcription Factors Regulate a Wide Range of Functions in Arabidopsis. Int. J. Mol. Sci. 2021, 22, 7152. [Google Scholar] [CrossRef]
- Li, C.; Yan, C.; Sun, Q.; Wang, J.; Yuan, C.; Mou, Y.; Shan, S.; Zhao, X. The bHLH transcription factor AhbHLH112 improves the drought tolerance of peanut. BMC Plant Biol. 2021, 21, 540. [Google Scholar] [CrossRef]
- Li, Z.; Liu, C.; Zhang, Y.; Wang, B.; Ran, Q.; Zhang, J. The bHLH family member ZmPTF1 regulates drought tolerance in maize by promoting root development and abscisic acid synthesis. J. Exp. Bot. 2019, 70, 5471–5486. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.S.; Liang, D.; Shuai, P.; Xia, X.L.; Yin, W.L. The salt- and drought-inducible poplar GRAS protein SCL7 confers salt and drought tolerance in Arabidopsis thaliana. J. Exp. Bot. 2010, 61, 4011–4019. [Google Scholar] [CrossRef] [PubMed]
- Habib, S.; Waseem, M.; Li, N.; Yang, L.; Li, Z. Overexpression of SlGRAS7 Affects Multiple Behaviors Leading to Confer Abiotic Stresses Tolerance and Impacts Gibberellin and Auxin Signaling in Tomato. Int. J. Genom. 2019, 2019, 4051981. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Min, Q.; Zheng, K.; Liu, T.; Wang, Z.; Xue, X.; Li, W.; Liu, Y.; Zhang, Y.; Qiao, F.; Chen, J.; et al. Transcriptomic Profiles of Long Noncoding RNAs and Their Target Protein-Coding Genes Reveals Speciation Adaptation on the Qinghai-Xizang (Tibet) Plateau in Orinus. Biology 2024, 13, 349. https://doi.org/10.3390/biology13050349
Min Q, Zheng K, Liu T, Wang Z, Xue X, Li W, Liu Y, Zhang Y, Qiao F, Chen J, et al. Transcriptomic Profiles of Long Noncoding RNAs and Their Target Protein-Coding Genes Reveals Speciation Adaptation on the Qinghai-Xizang (Tibet) Plateau in Orinus. Biology. 2024; 13(5):349. https://doi.org/10.3390/biology13050349
Chicago/Turabian StyleMin, Qinyue, Kaifeng Zheng, Tao Liu, Zitao Wang, Xiuhua Xue, Wanjie Li, Yuping Liu, Yanfen Zhang, Feng Qiao, Jinyuan Chen, and et al. 2024. "Transcriptomic Profiles of Long Noncoding RNAs and Their Target Protein-Coding Genes Reveals Speciation Adaptation on the Qinghai-Xizang (Tibet) Plateau in Orinus" Biology 13, no. 5: 349. https://doi.org/10.3390/biology13050349
APA StyleMin, Q., Zheng, K., Liu, T., Wang, Z., Xue, X., Li, W., Liu, Y., Zhang, Y., Qiao, F., Chen, J., Su, X., & Han, S. (2024). Transcriptomic Profiles of Long Noncoding RNAs and Their Target Protein-Coding Genes Reveals Speciation Adaptation on the Qinghai-Xizang (Tibet) Plateau in Orinus. Biology, 13(5), 349. https://doi.org/10.3390/biology13050349







