3.2.2. Methodologies for Building Ontologies
The analysis of the methodologies for building ontologies revealed that none of these approaches are fully mature [
54,
55,
56].
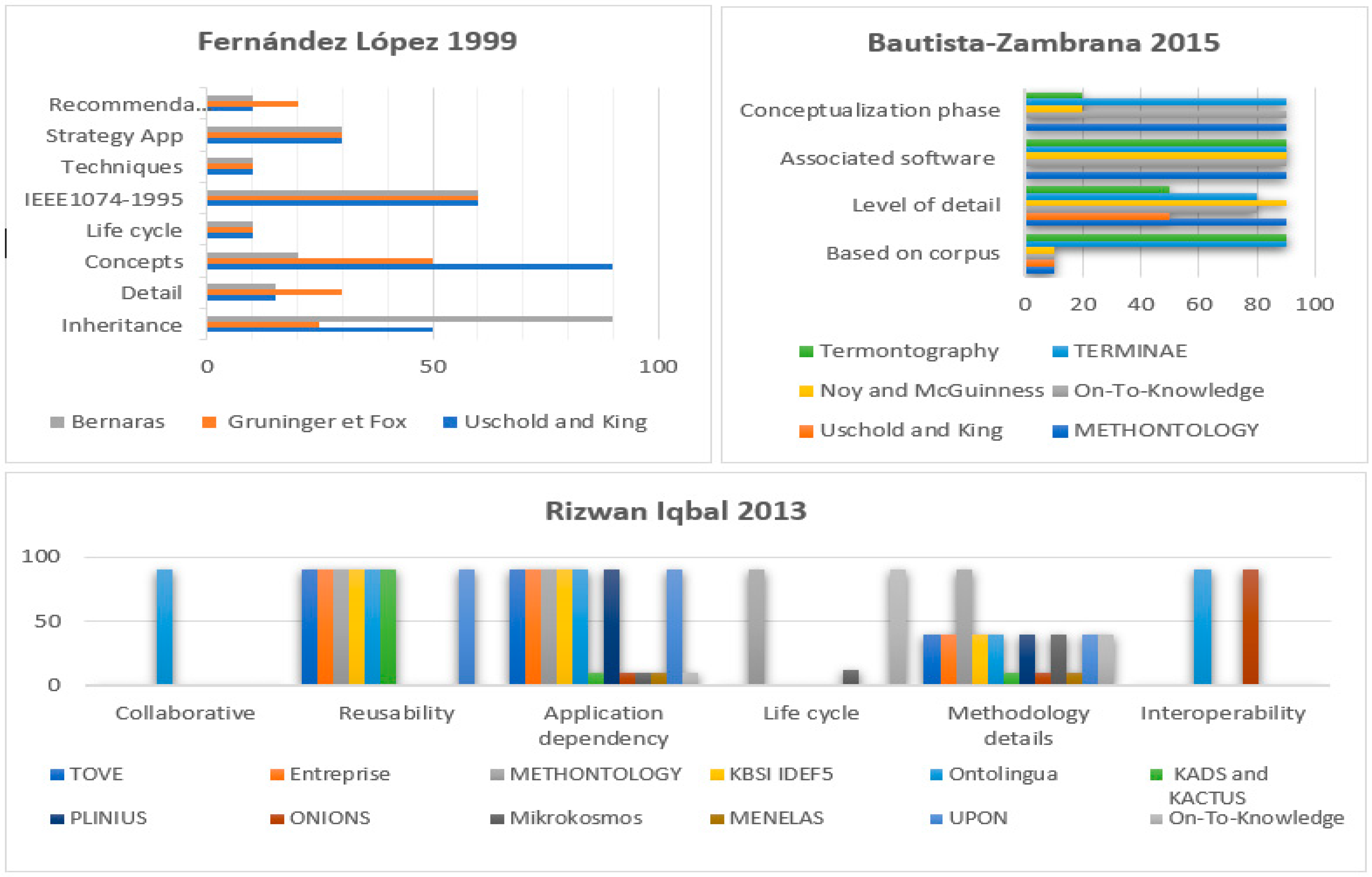
The histograms chart in
Figure 3 is a summary of three surveys that address existing methodologies. Lopez [
54] compared five methodologies: SENSUS, Bernaras, Uschold and King, METHONOLOGY, and Gruninger. Lopez established a set of criteria for analyzing each of these methodologies. The author believed that inheritance from knowledge engineering and specifying details of the methodology, whether in terms of the activities or proposed techniques, in addition to the strategy for building and identifying concepts are main points to be considered by the researchers while developing methodologies. Moreover, there are equally important characteristics for analysis such as the recommended life cycle, the differences between the methodology and the work proposed by the IEEE standard 1074–1995, and collaborative and distributive construction. For additional information, please refer to [
54]. Analogously, Rizwan [
56] conducted a broader critical study of twelve common methodologies based on six basic measures: (1) collaboration; (2) degree of reusability; (3) application dependency; (4) life cycle; (5) methodology details; and (6) interoperability. Zambrana et al. [
55] had a different point of view since his comparison focused on conceptualizations, development, and validation. Zambrana raised five questions to assess the six target methodologies. (1) “Are the ontology elements as concepts, relations, properties, etc. based on corpus work? (2) Who are the intended users of the methodology? (3) Does the methodology explicitly state which methods and techniques we should use to perform the different activities? (4) Does the methodology propose to perform a conceptualization activity? (5) Is there a program associated to the methodology that facilitates the different steps to be taken?”. Although some methods outperform others in some features, in general, it seems none of them comply perfectly with all the requirements. To solve this problem, the researchers sought to find standardized methodologies adaptable to different types of ontologies and in different application domains. Lopez states that one of the first attempts to unify two methodologies was described in [
57] but “the new synthesized methodology was not an actual methodology, it was a conception of a potential methodology”. Later, Sánchez [
58] combined two of the well-referenced methodologies METHONTOLOGY [
59] and Cyc 101 [
60] to obtain one of the most concrete methodologies for building medical ontologies (
Figure 4).
Step 1: Determine the Domain and Scope of the Ontology
The scope refers to the domain of interest which is to be described in this ontology. This step must draw the boundary or limitations that constrain the initial purpose of the conceptualization domain. Formally, ontologies developers should make good use scenarios or try to ask straightforward questions. Researchers have proposed a set of questions for mapping objectives with an ontology to determine its domain, scope, contribution, and structure.
- ▪
What is the domain that the ontology will cover?
The chronic obstructive pulmonary disease is the domain of this ontology.
- ▪
What is the purpose of this ontology?
This ontology is designed for preventive management of COPD patients. The main purpose is to facilitate the systematic extraction of information from detailed observations. Our ontology is dedicated to support a personalized system for COPD patient. This ontology provides real-time monitoring and recommendations to help patients cease contact with risk factors and prevent progressive respiratory impairment and allows physicians to be kept informed of the patient’s condition.
- ▪
Who will use the ontology?
Potential users of this ontology are physicians and patients.
- ▪
What types of questions should the information in the ontology provide answers for?
The COPDology must provide answers to questions such as:
What data should be collected to supervise the patient?
How often should the patient take a measurement?
Should the acquired data be transmitted to the healthcare site?
How should the data be analyzed?
Should an alarm be triggered according to the evaluation results?
Which actions should be performed if an alarm is triggered?
Step 2: Ontology Reuse
There is almost always the possibility an ontology has been modeled before from a third party that provides a useful starting point to be fully or partially reused. Reusing existing ontologies is necessary to save time and effort, to interact with the tools that use other ontologies or to exploit ontologies that have been validated through use in applications. For example, we can reuse ontology libraries (DAML and Ontolingua) or high-level ontologies such as general or domain-specific ontologies. Indeed, the ontological templates targeting remote monitoring of lung diseases is not well planned. Lasierra et al. [
24] proposed an approach to provide clinical management at a personal level in home-based telemonitoring scenarios by developing an ontology-driven solution that enables a wide range of services such as core health indicators, real-time alerts, and medication reminders. Paganelli et al. [
26] described an ontology-based context model and a related context management middleware providing a reusable and extensible application framework for monitoring and assisting patients at home. Mcheick et al. [
61] proposed a context-aware system to derive relevant attributes and early detection of COPD exacerbations but their use of ontology was only to realize a general architecture of application.
Although there is poor ontological coverage of pulmonary diseases, there are many global references of terminologies for standardizing the storage, retrieval, and exchange of electronic health data that can be considered as a fundamental point of building our ontology, especially concerning the medical glossary. Adhering to shared knowledge principles in such kind of projects requires reusing the standard clinical and medical abbreviations and terminology. In this work, we reuse a wide range of terms provided in Systematized Nomenclature of Medicine Clinical Terms (SNOMED CT), and the Global Medical Device Nomenclature (GMDN). The terms that we used are explained in patient ontology paragraph (page 12).
Step 3: Development of a Conceptual Model
Enumerate key terms in the ontology: Enumerating important terms, such as needed nouns and verbs, is a crucial step to make statements or to explain the context. The nouns are divided into concepts, attributes or instances. Concepts are considered nouns standing on their own; attributes can describe the type of things, and instances are nouns of specific things. Then, verbs describe relations between nouns. Medical ontologies often use coding terminology standards to label values of clinic data items such as symptoms, diseases, drugs, and laboratory measurements. There are several coding systems that overlap highly but with varying degree of generality and specificity in coding terms such as Systematized Nomenclature of Medicine-Clinical Terms (SNOMED-CT), Logical Observation Identifiers, Names and Codes (LOINC), The International Statistical Classification of Diseases and Related Health Problems (ICD), Common Classification of Medical Procedures (CCAM) and the Global Medical Device Nomenclature (GMDN).
Classes and class hierarchy definition: This step aims to classify the proposed concepts in a hierarchy as a form of a taxonomic architecture. This phase of ontology development starts vertically by defining classes which are selected to build COPDology. When the ontology has many elements, we must taxonomize the concepts. To achieve such a taxonomy, we can use one of the categorization methods. As we mentioned earlier, there are three different methods; we used a top-down approach to develop the class hierarchy, through representing the core concepts (main classes) and subclasses as classes in the COPDology. Classes (concepts) have a direct relation with patient needs to detect abnormal status and dangerous activity.
Class properties definition: Properties are used to describe the attributes or the relationships of the classes. There are four types of properties: (1) intrinsic, (2) extrinsic, (3) parts and (4) relationships to other individuals. Defining properties of classes is a requirement to realize the true value of ontology. Classes and their sub-classes do not provide sufficient information or rather do not have the ability to properly represent the relationship among the different elements. Practically, there are two types of properties: object properties and datatype properties. Object properties play important roles in connecting classes where the starting point class is called domain and the endpoint is called range. On the other hand, datatype properties only connect the concept to a specific value, for example, String, Integer, Boolean, etc.
Define the facet of slots: Slimani [
62] defined the slot as a word that should be assigned to a class, for example, a name, a price, etc. The slot may have different kinds of facets that outline the value type, permitted values, cardinalities, and other features, which may be added as needed. In our COPDology, most of the slot values are String, Float, Integer, and Boolean.
Create instances: An instance is an individual of a class; defining an instance requires primarily choosing a class, creating an individual instance, and then filling in the slot values. Below is a small set of classes and their possible instances (Patient: John; Disease: COPD; Location: home). These Individuals are also interpreted as instances of classes. The information depicted for the individuals has been taken from medical guidelines and research papers of COPD domain.
Development of COPD Ontology domain
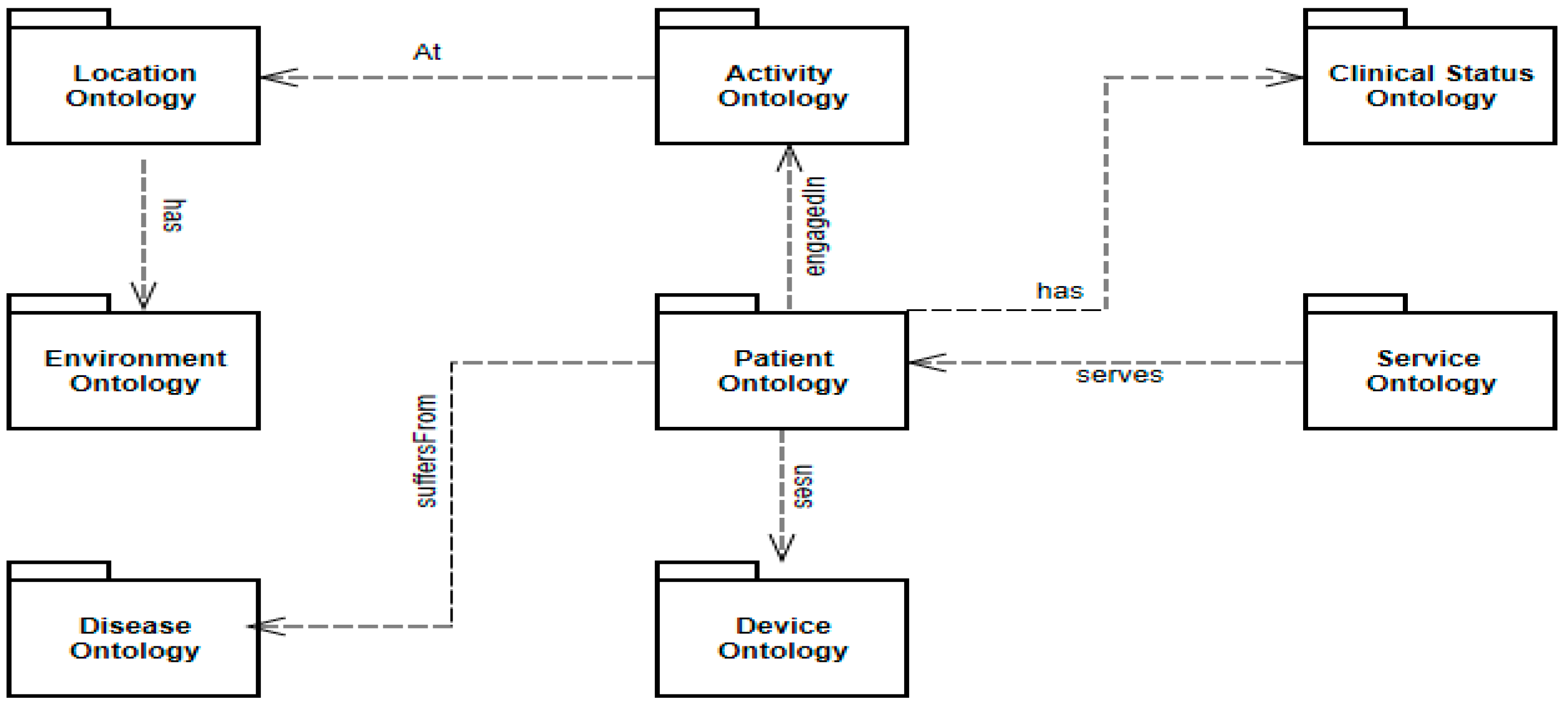
Chronic Obstructive Pulmonary Disease ontology (COPDology) is a model of specific medical domain collected from many research papers and relevant guidelines as well as information obtained from pneumologists that were interviewed and asked about care plans of COPD. This ontology contains concepts related to the disease, environment, equipment, patient data (personal information, symptoms, risk factors and clinical tests results) and treatment. The ontology was implemented with Protégé in OWL format. COPDology consists of 680 classes, 276 object properties, 310 datatype properties, 5000 instances and a set of inference rules that guide the diagnosis and risk assessment process. The knowledge base or COPDology that we provide in this work consists of a set of interrelated ontologies describing the physical and abstract objects in the domain scope. The ontologies we created to support the necessary health surveillance of COPD patient contain primarily patient, clinical status, devices, activities, environment, services, location, and disease.
Figure 5 depicts these different ontologies including their distribution and their general relationships.
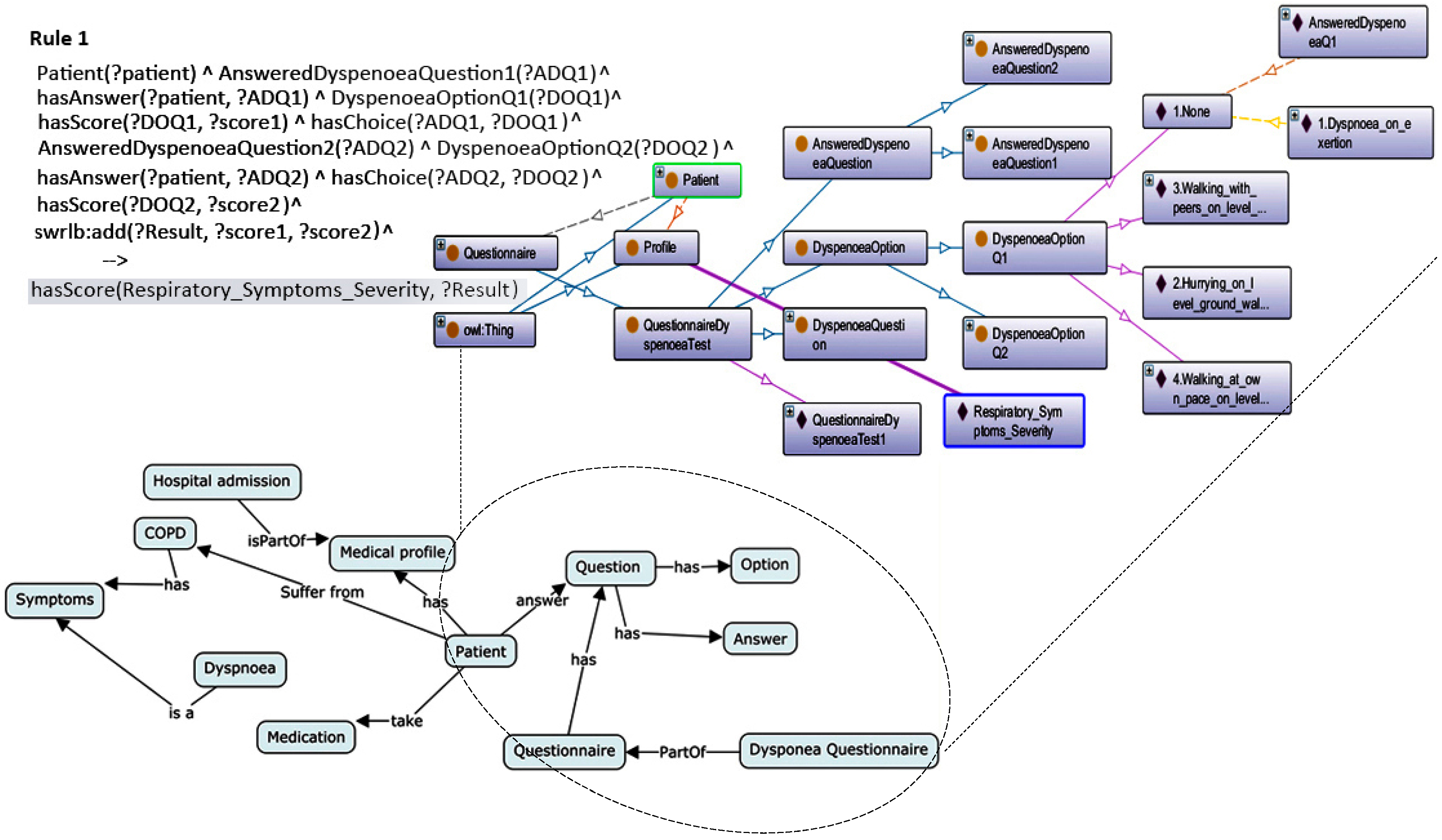
Patient ontology
The patient’s ontology consists of three main branches: physical factors, psychological factors and the personal information of the patient. Physical factors refer to vital signs that have a direct relation with COPD. Recognizing these elements was not an easy task; we needed hours of research and meetings with lung specialists. The results of this effort are 15 key elements, namely temperature, heart rate, FEV1, PH Level, Paco2, BUN, sodium level, hematocrit, diastolic pressure, systolic pressure, oxygen saturation, respiration rate, body height, body weight, and glucose. For psychological factors, recent studies [
1,
10] have confirmed that there is a proportional relationship between the deterioration of mental and physical conditions. The most prominent psychological states are depression, stress, and anxiety. As for the profile, it is limited to some personal information such as name, age, occupation, gender, race, nationality, telephone, address, and habits. In
Table 3, we present some of the used classes with their corresponding codes in SNOMED CT.
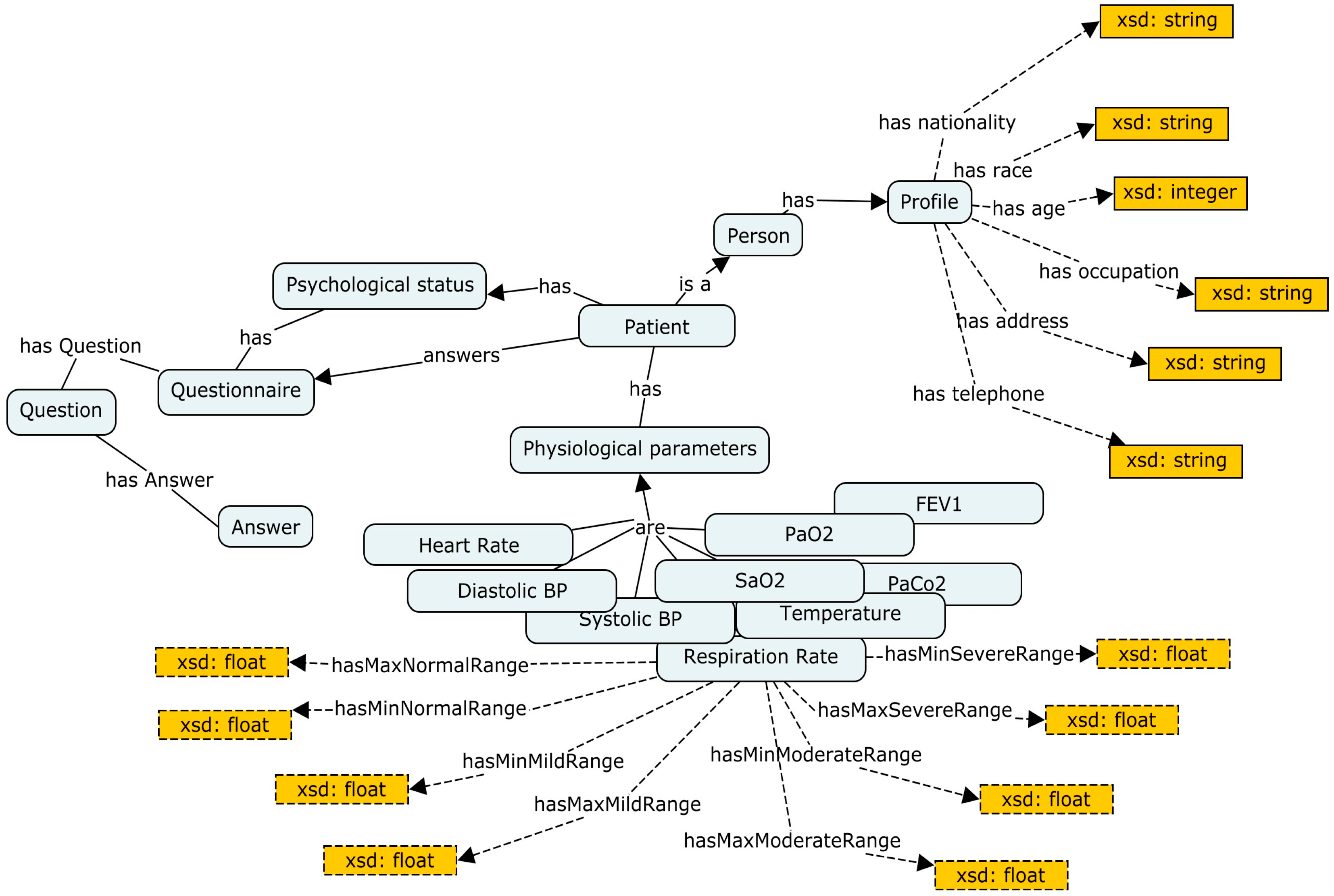
Due to some technical limitations, we only browse a general representation of the main components, as shown in
Figure 6.
Table 4 lists some of the object and data properties that describe the patient’s ontology. As we can see, each of these properties has its own characteristics that specify its domain and range.
Environment Ontology
Environmental factors have enormous potential to affect our body. COPD is one of the most sensitive diseases to the surrounding environment. The environmental factors that negatively affect COPD patients include ambient air, weather, and pollution. Ambient air is a gas mixture composed of N
2 and O
2, with extremely small quantities of CO
2, argon and some inert gases [
63], such as neon, hydrogen, methane, xenon, krypton, and helium. In general, the density of these gases can be changed by changing either the pressure or the temperature [
64], which may pose significant risks to the respiratory system of patients. Furthermore, according to some statistical surveys [
65,
66], lung disease symptoms are widely affected by weather conditions such as extreme temperatures, humidity, pressure, and precipitation. For more comprehensiveness and precision in observation, we have added climate and type of weather.
Figure 7 is a simple review of the most prominent elements in this ontology.
Devices ontology
This ontology includes computing hardware devices such as Personal Digital Assistants (PDAs) and sensors. Basically, this ontology covers the mobile device used to collect and send data as well as all fixed and portable biomedical equipment used by the patients to monitor their vital signs in addition to environmental sensors to detect any change in the environment.
Figure 8 shows the types of devices found in this ontology. Biomedical parameters are sensed by body thermometer, pulse oximeter, blood pressure monitor, weighing scale, body composition analyzer, peak flow, basic ECG, a respiration rate monitor, and accelerometer. The environmental information can be obtained by the thermometer, hygrometer, air quality sensors, barometer, and GPS.
Activity Ontology
Identifying the current activity of the patient adds more accuracy to the medical applications. In this context, it is important to know what physical activity a person is doing. It would also be useful to identify possible movements and places to be visited as well as the means used during such activities.
Figure 9 provides a part of concepts and relations used to realize activity’s ontology.
Location Ontology
Location is considered the backbones of all these sub-ontologies. Location awareness described in
Figure 10 serves to determine the physical parameters to be measured, where relevant contextual information varies between indoor and outdoor space.
Disease ontology
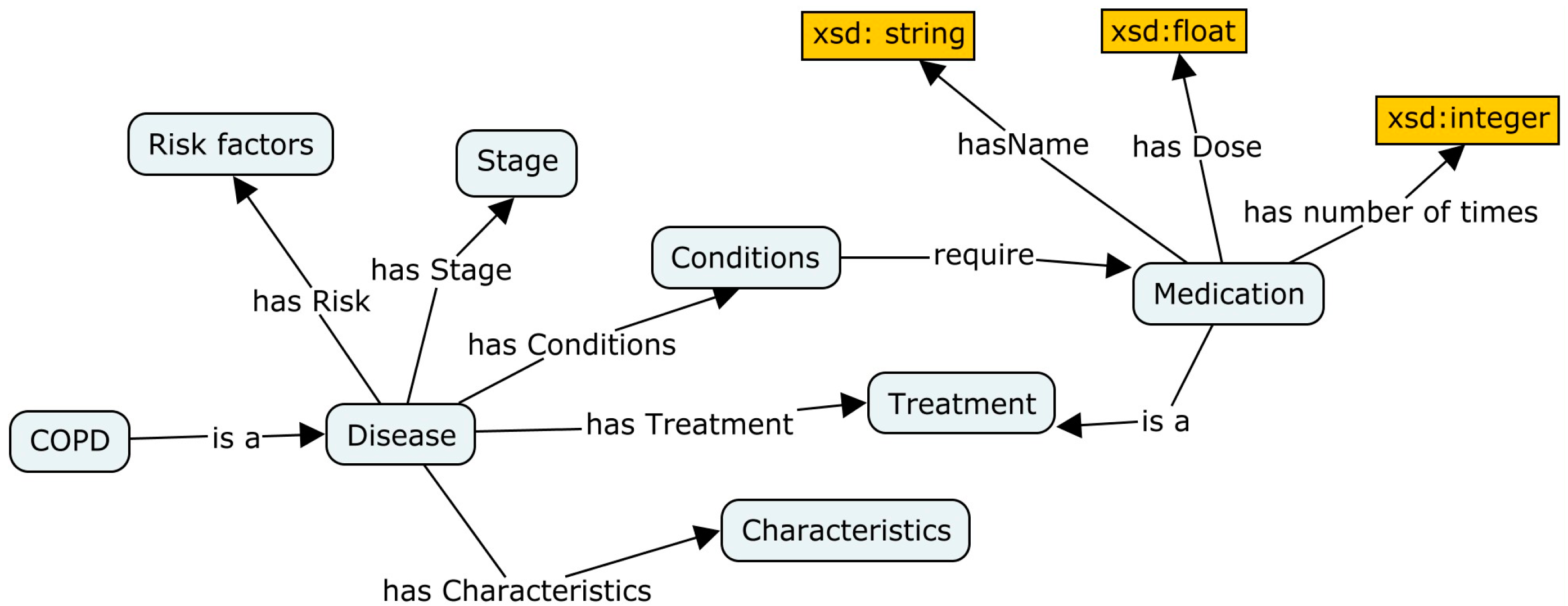
To present personalized care suited to patient status, we need to understand the nature of the disease. This sub-ontology primarily aims to provide efficient administration of treatment. A disease ontology comprises type of illness, stage, treatment, risk factors, conditions and physical characteristics of the disease (see
Figure 11).
Clinical status ontology
The ontology of clinical status contains the medical history of patients including physical exam findings, diagnostic test results, family diseases and medications that a patient has taken in the past or is currently taking. This ontology improves the performance of healthcare systems where it provides the ability for treatment to be monitored and achieve high-quality care.
Figure 12 shows some fragments of this ontology.
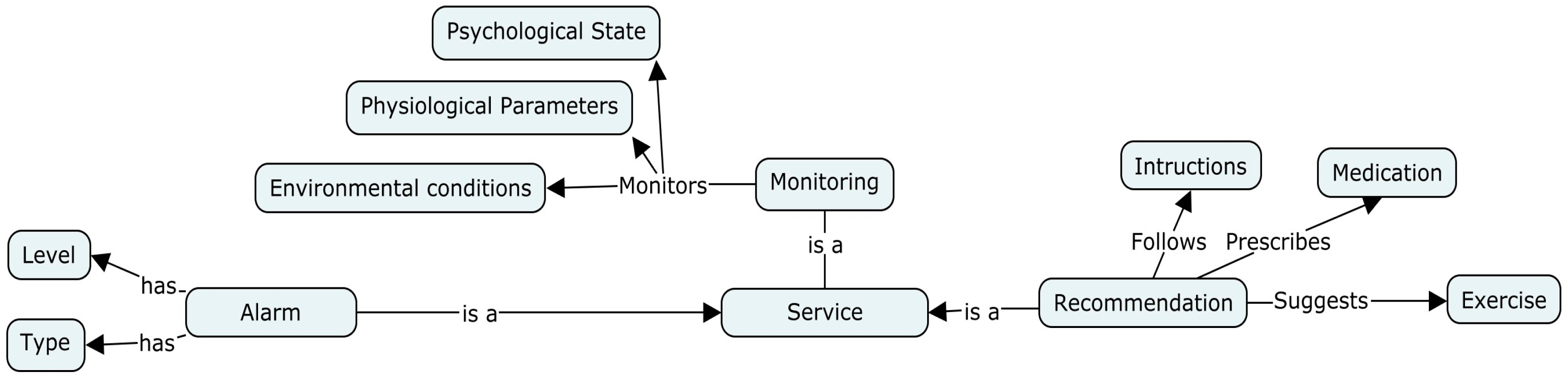
Service ontology
Essentially, this ontology-based model is designed to provide services and interact with patients to control precarious or suspicious situations. Hence, a service is considered a major component of the proposed ontology. These medical services are divided into three basic services: monitoring, triggering alarms and recommendation (
Figure 13).
Step 4: Implementation
The development of ontologies in the medical world is a complex task that requires considerable effort and collaboration between health care professionals and ontology engineers. Clinical decision support necessarily needs methods that verify the correct representation of activities in terms of effects and ontological responses. For knowledge representation, the tools and techniques are essential to support design work. The implementation and the validation of the logical and structural aspects of ontology can be automatically realized with specialized tools. Protégé is one of the best-known open source editors to develop ontologies [
67]. Protégé has been distributed originally for biomedical informatics research at the Stanford University School of Medicine. This tool is specifically dedicated to the OWL but it is a highly extensible editor, capable of handling a wide variety of formats [
67]. Our ontology was formalized using OWL DL because it is highly expressive and thus we can apply all standard automatic reasoning techniques. Protégé contains built-in reasoners such as FaCT++, Pellet, HermiT, ELK, jcel, Ontop, Mastro and RACER used for describing logic. Choosing a good reasoner is also an essential step towards delivering an effective ontological framework. Abburu [
68] conducted a comparison between some of the popular reasoners developed in the last few years. This survey describes these reasoners with their important features such as completeness, expressivity, native profile, incremental classification, rule support, platforms, justifications, ABOX reasoning, OWL API, protégé support, Jena support, etc. Abburu [
68] explains four types of reasoners that may support rules: RACER, Peller, Hermit, and ELK. Racer does not support Jena and is commercial. Hermit does not present explanations for the inconsistency that exists in the ontologies and cannot work with Jena API. ELK is a reasoner for OWL 2 EL ontologies, which is not the case of this project. Based on the previous assessment, we used Pellet in our ontology.
Step 5: Evaluation of COPDology
There are many ontology evaluation methods such as those proposed by Jonathan [
69], Pérez et al. [
70] and Lovrenčić et al. [
71]. These studies present robust approaches for ontology evaluation based on criteria and measures or metrics. The authors defined the evaluation criteria as general qualities for making a technical judgment of the content [
69]. These criteria include consistency, classification, completeness, conciseness, expandability, and sensitiveness. In contrast, the authors found that ontology evaluation measures are primarily oriented towards the structural aspects.
Ontology Evaluation Criteria
Criteria evaluation was performed on our ontology, with the following results:
Consistency: This refers to evaluating the logical consistency of an ontology by checking invariants [
70]. Running the automatic consistency reasoner check proves that COPDology is consistent and coherent; there is no inferred conflicting knowledge from other definitions, axioms, and formal definitions, and no contradictory knowledge can be inferred from all definitions and axioms.
Classification: It is one of the most important reasoning services provided by all OWL reasoners. Ontology classification means computing all entailed class subsumptions between named classes [
72]. Unfortunately, when an ontology evolves or even slightly modified, the reasoners repeat the whole reasoning process. For large and complex ontologies, this might take a considerable amount of time [
73]. For some purposes, the reasoner should be executed often, and then time response becomes a critical issue. Wang et al. [
74] suggested reducing disjoint statements that may cause performance problems, as this may restrict the reasoner too much. In this project, our result was reasonable, where the total classification time in Pellet was 26,849 ms which is near real-time and therefore the reasoner could be working transparently without slowing down the response.
Completeness: An ontology is called complete if all the stated information is explicitly defined or can be inferred from other definitions and axioms [
71]. In terms of design features and providing sufficient information to answer the competency questions, COPDology is complete, but since some information related to lung diseases is difficult to obtain due to lack of studies, this application ontology is not complete, as it does not provide comprehensive medical service of all aspects of a patient’s life.
Conciseness: This attribute determines whether an ontology has redundant terms. In this work, we reduced the size of the representation as much as possible to avoid having any unnecessary concepts, whether explicit redundancies or implicit redundancies (inferred). Therefore, COPDology is concise.
Expandability: It is an indicator that ontology is smoothly expandable without significant modifications in the case of adding new knowledge to existing structures [
71]. Development of COPDology showed that hierarchy of core concepts does not have to be considerably altered. The division of the representation field in several parts promotes the expansion of the ontology. Practically, modification or creation of new classes and axioms does not influence other parts, which means that this ontology was built with expansion capabilities.
Sensitiveness: An ontology is considered sensitive if minimal changes in definition affect directly a set of coherent concepts and well-defined relations [
71]. As explained above, alteration of a set of concepts or adding new definitions does not influence other axioms and classes, therefore COPDology is not sensitive.
Ontology Evaluation Measures
This type of evaluation focuses on the complexity and formality of structure by respecting three basic levels: vocabulary level, taxonomy level, and nontaxonomic level [
75,
76,
77]. The purpose of the evaluation is to estimate the internal maturity level of the ontologies. In this context, Zhang [
78,
79] proposed a set of metrics to measure the ontology complexity on both class and ontology levels through combinations of dimensional characteristics.
Ontology-level evaluation
Srinivasulu et al. [
80] suggested four ontology-level metrics to describe the complexes of an ontology on holistic intention: size of vocabulary, edge node ratio, tree impurity and entropy of graph.
- 1.
Size of vocabulary (SOV): This metric includes the total number of created classes, instances and properties in the ontology; the SOV is defined as:
where
represents the number of named classes, while
and
are the number of properties and instances, respectively [
79].
- 2.
Edge node ratio (ENR): ENR represents the connectivity density which increases proportionally with the increment of the number of edges between nodes (classes and individuals). ENR is measured as follows:
where the number of edges |E| is divided by the number of nodes
.
- 3.
Tree impurity (TIP): This indicator is mainly used to discover how far an ontology inheritance hierarchy digresses from a tree; the TIP is measured as in Equation (3):
where
represent the suite of relations and concepts in the inheritance hierarchy, respectively.
- 4.
Entropy of ontology graph (EOG): This norm is an indicator of the graph complexity [
80]. It is calculated directly by the application of the logarithm function to a probability distribution over the ontology graph:
where p(i) represents the probability mass function for a concept to have i relations. Arithmetically, p(i) can be calculated for each vertex (concepts) in the ontology graph by dividing the degree of the vertex (i.e., properties) connected to that concept over the sum of all degrees of V vertices:
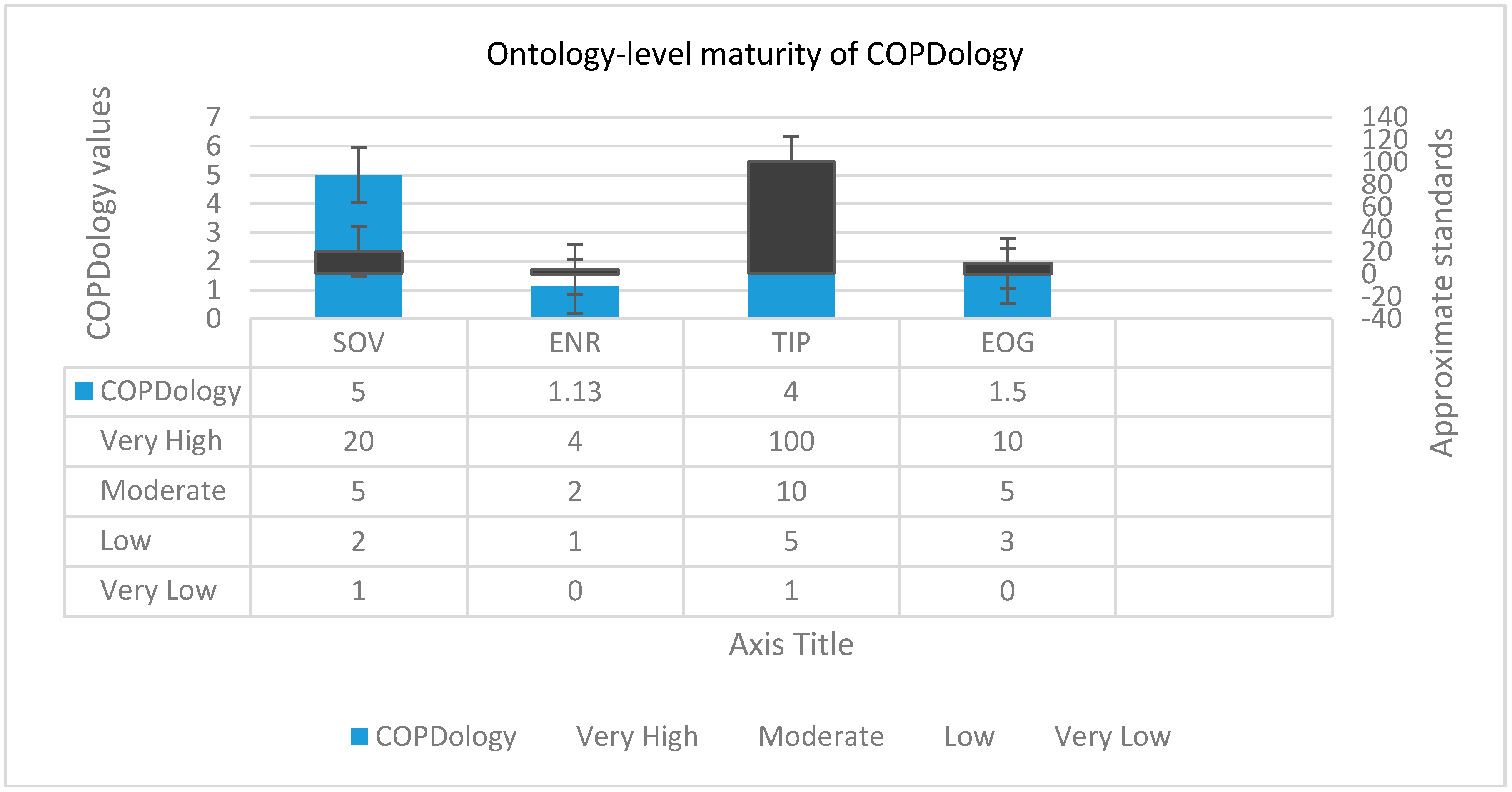
Typically, designers compare their ontology against a “gold standard” which is considered to serve as a reference. The metrics values presented in
Figure 14 belong to some well-constructed ontologies [
81,
82,
83]. The SOV of COPDology exceeds 5000 of components constituted from huge sets of concepts, parameters, patient medical records, etc. Therefore, it would be very useful for semantic developers, specifically those interested in the biomedical domain, to reuse this ontology rather than try to build a new ontology for COPD from scratch. On the other hand, the ontologies with large vocabularies would require a considerable amount of time and effort to build and maintain [
84]. The edge node ratio (ENR) value is somehow higher than normal, which means that our ontology is complex and needs further modularization to minimize the effort required for understanding and management. The TIP is a rational indicator of how well an ontology is organized through inheritance relationships. A TIP = 0 indicates the inheritance hierarchy graph is structured as a tree. The higher is the TIP, the greater does the ontology inheritance hierarchy drift away from the rooted tree, thus the greater is its complexity. The total value of our COPDology TIP reaches 4, which means that this inheritance hierarchy deviated relatively from the traditional shape of the tree. The last metric in this level is the entropy of ontology graph (EOG), where 0 corresponds to the least value of EOG when classes have the same distribution of relations, which can only be obtained if all nodes of the ontology sub-graphs have equal number of edges. The practical interpretation of small EOG is indicative of less complex ontology in terms of relation distribution [
85]. The EOG of COPDology is almost 1.5, thus it has relatively good structure.
Class level evaluation
Zakaria [
76] combined eight metric functions to measure complexness at class-level. These metrics are the number of classes, number of inheritances, number of properties, number of root classes, average population, class richness, relationship richness, and inheritance richness.
- 1.
Number of classes (NOC): The NOC metric is simply a count of the defined classes in the ontology [
85].
- 2.
Number of instances (NOI): The NAO criterion is a census of the instances created in the ontology.
- 3.
Number of properties (NOP): As its name implies, NOP is the number of properties found in an ontology [
84].
- 4.
Number of root classes (NORC): This metric corresponds to the number of non-rooted classes or the concepts that do not have super-classes in their upper layer. Let us consider C the classes in ontology:
- 5.
Average Population (AP): This variable measures the mean distribution of instances across all classes. Theoretically, AP is defined as follow:
According to the rules set [
80], this metric has been proposed as an indication of whether there is sufficient information in the ontology.
- 6.
Class Richness (CR): This value is the ratio between the number of non-empty classes that have instances
and the total number of classes. CR percentage give us an idea of how many instances are related to classes defined in the graph.
- 7.
Relationship Richness (RR): This metric represents the number of relationships divided by the sum of the number of subclasses and the number of relationships [
80]:
where |P| is considered the overall count of relationships and |SC| is the tally of subclasses or the number of inheritance relationships.
- 8.
Inheritance richness (IR): The IR describes the distribution of knowledge overall levels of the ontology’s inheritance tree. The inheritance richness of the schema (IRs) is known as the average number of subclasses per class. Formally, this value is calculated from the equation:
Table 5 summarizes the class-level evaluation of our COPDology. NOC and NOI were quite high at 180 and 4000 respectively. The number of properties in the NOP indicates a strong reasoning system [
85]. As mentioned above, the NORC is the number of root classes in the COPDology. The higher is the NORC value, more diverse is the ontology [
84]. COPDology has a high NORC value, existing of 12 root classes is a proof that this ontology has a large structure. The high AP value (AP = 6.5) is a good indication that COPDology has sufficient information to query data from the built framework. Since AP and CR are correlated, it is obvious that our ontology achieved only 0.80 for the CR metric. Therefore, this indicates that the majority of the ontology classes have instances. Usually, an ontology that contains many descriptive relationships or non-typical relationships such as class-subclass is richer than taxonomies that have a category–subcategory hierarchy. In this work, the COPDology is very rich in COPD content where its RR arrived at the threshold of 0.4. The inheritance richness has been proposed to distinguish a horizontal ontology from a vertical ontology. COPDology has high IR which might reflect vertical nature and a very detailed type of knowledge.
In this section, we have present twelve metrics to examine the maturity of COPDology. Clearly, such interpretation proves that our ontology is mature and valid to be reused and extended.