Artificial Intelligence Analysis of Celiac Disease Using an Autoimmune Discovery Transcriptomic Panel Highlighted Pathogenic Genes including BTLA
Abstract
1. Introduction
2. Materials and Methods
2.1. Celiac Disease GSE164883 Dataset
2.2. GEOR Analysis
2.3. Transcriptome Panels
2.4. Gene Set Enrichment Analysis (GSEA)
2.5. Statistical Analyses
2.6. Immunohistochemical Analysis of BTLA in an Independent Series
3. Results
- A conventional analysis using GEO2R highlighted the genes differentially expressed between celiac disease and control.
- Gene set enrichment analysis (GSEA) identified the gene sets (pathways) that were associated with celiac disease, including the autoimmune discovery panel.
- Several Machine learning and artificial neural network analyses predicted celiac disease using the autoimmune discovery panel with high accuracy.
- Celiac disease was characterized by high expression of BTLA both at the gene expression level, and at protein level by immunohistochemistry in a validation series.
3.1. Gene Expression Analysis Using the GEO2R Software
3.2. Gene Set Enrichment Analysis (GSEA)
3.3. Artificial Intelligence Analysis
3.4. Differential Gene Expression of BTLA between Celiac Disease and Control Samples
3.5. Validation of BTLA by Immunohistochemistry in an Independent Series
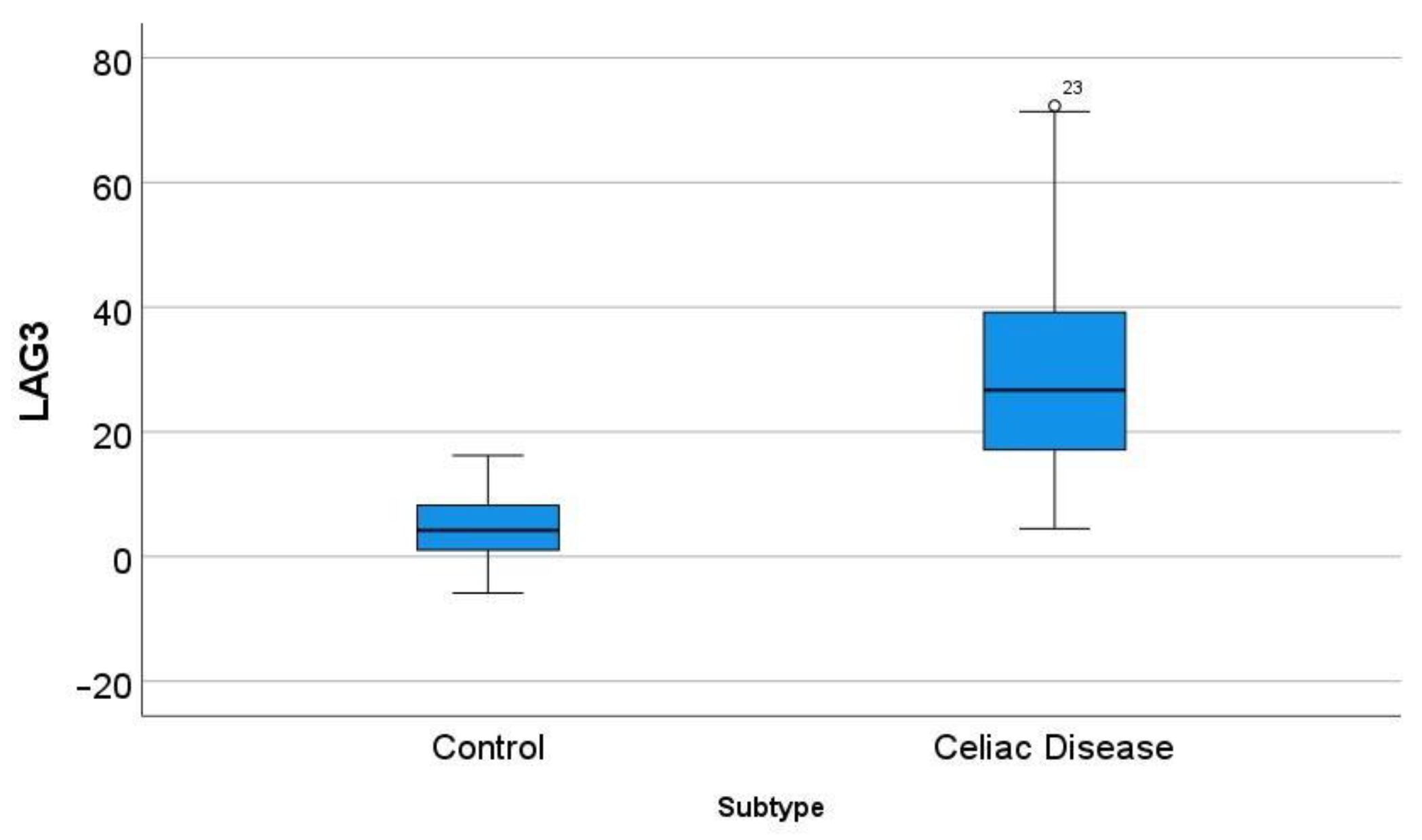
3.6. Differential Gene Expression of LAG3 between Celiac Disease and Control Samples
4. Discussion
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Age | Sex | Biopsy Location | Diagnosis | Marsh-Oberhuber Classification |
|---|---|---|---|---|
| 70 | Male | Duodenum | Celiac Disease | 3a |
| 62 | Male | Pylorus/duodenum | Celiac Disease/Chronic gastritis | 2 |
| 62 | Male | Duodenum | Celiac Disease | 2 |
| 78 | Female | Duodenum | Celiac Disease | 3b |
| 59 | Male | Duodenum | Celiac Disease | 3a |
| 44 | Female | Duodenum | Celiac Disease | 2 |
| 17 | Female | Duodenum | Celiac Disease | 3b |
| 56 | Female | Duodenum | Celiac Disease | 3a |
| 54 | Female | Duodenum | Celiac Disease | 2 |
| 58 | Female | Duodenum | Celiac Disease | 3b |
| 61 | Female | Duodenum | Celiac Disease | 3c |
| 45 | Male | Duodenum | Celiac Disease | 3a |
| 70 | Female | Duodenum | Celiac Disease | 2 |
| 40 | Female | Duodenum | Celiac Disease | 3a |
| 61 | Female | Duodenum | Celiac Disease | 3c |
| 44 | Female | Duodenum | Celiac Disease | 3a |
References
- Schuppan, D.; Dieterich, W. Epidemiology, Pathogenesis, and Clinical Manifestations of Celiac Disease in Adults. Available online: https://www.uptodate.com/contents/epidemiology-pathogenesis-and-clinical-manifestations-of-celiac-disease-in-adults?source=history_widget (accessed on 13 July 2022).
- Al-Toma, A.; Goerres, M.S.; Meijer, J.W.; Peña, A.S.; Crusius, J.B.; Mulder, C.J. Human Leukocyte Antigen–DQ2 Homozygosity and the Development of Refractory Celiac Disease and Enteropathy-Associated T-Cell Lymphoma. Clin. Gastroenterol. Hepatol. 2006, 4, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Pietzak, M.M.; Schofield, T.C.; McGinniss, M.J.; Nakamura, R.M. Stratifying Risk for Celiac Disease in a Large At-Risk United States Population by Using HLA Alleles. Clin. Gastroenterol. Hepatol. 2009, 7, 966–971. [Google Scholar] [CrossRef] [PubMed]
- Liu, E.; Lee, H.-S.; Aronsson, C.A.; Hagopian, W.A.; Koletzko, S.; Rewers, M.J.; Eisenbarth, G.S.; Bingley, P.J.; Bonifacio, E.; Simell, V.; et al. Risk of Pediatric Celiac Disease According to HLA Haplotype and Country. N. Engl. J. Med. 2014, 371, 42–49. [Google Scholar] [CrossRef] [PubMed]
- Houlston, R.S.; Ford, D. Genetics of coeliac disease. QJM Int. J. Med. 1996, 89, 737–744. [Google Scholar] [CrossRef]
- Houlston, R.S.; Tomlinson, I.P.; Ford, D.; Seal, S.; Marossy, A.M.; Ferguson, A.; Holmes, G.K.; Hosie, K.B.; Howdle, P.D.; Jewell, D.P.; et al. Linkage analysis of candidate regions for coeliac disease genes. Hum. Mol. Genet. 1997, 6, 1335–1339. [Google Scholar] [CrossRef][Green Version]
- Greco, L.; Corazza, G.; Babron, M.-C.; Clot, F.; Fulchignoni-Lataud, M.-C.; Percopo, S.; Zavattari, P.; Bouguerra, F.; Dib, C.; Tosi, R.; et al. Genome Search in Celiac Disease. Am. J. Hum. Genet. 1998, 62, 669–675. [Google Scholar] [CrossRef]
- Romanos, J.; van Diemen, C.C.; Nolte, I.M.; Trynka, G.; Zhernakova, A.; Fu, J.; Bardella, M.T.; Barisani, D.; McManus, R.; van Heel, D.; et al. Analysis of HLA and Non-HLA Alleles Can Identify Individuals at High Risk for Celiac Disease. Gastroenterology 2009, 137, 834–840.e3. [Google Scholar] [CrossRef]
- Trynka, G.; Zhernakova, A.; Romanos, J.; Franke, L.; Hunt, K.A.; Turner, G.; Bruinenberg, M.; Heap, G.A.; Platteel, M.; Ryan, A.W.; et al. Coeliac disease-associated risk variants in TNFAIP3 and REL implicate altered NF-kappa B signalling. Gut 2009, 58, 1078–1083. [Google Scholar] [CrossRef]
- Leonard, M.M.; Sapone, A.; Catassi, C.; Fasano, A. Celiac Disease and Nonceliac Gluten Sensitivity: A Review. JAMA 2017, 318, 647–656. [Google Scholar] [CrossRef]
- Caio, G.; Volta, U.; Sapone, A.; Leffler, D.A.; de Giorgio, R.; Catassi, C.; Fasano, A. Celiac disease: A comprehensive current review. BMC Med. 2019, 17, 142. [Google Scholar] [CrossRef]
- Oxentenko, A.S.; Rubio-Tapia, A. Celiac Disease. Mayo Clin. Proc. 2019, 94, 2556–2571. [Google Scholar] [CrossRef] [PubMed]
- Green, P.H.; Lebwohl, B.; Greywoode, R. Celiac disease. J. Allergy Clin. Immunol. 2015, 135, 1099–1106. [Google Scholar] [CrossRef]
- Lebwohl, B.; Sanders, D.S.; Green, P.H.R. Coeliac disease. Lancet 2018, 391, 70–81. [Google Scholar] [CrossRef]
- Naluai, A.T.; Ascher, H.; Nilsson, S.; Wahlström, J. Searching for genes influencing a complex disease: The case of coeliac disease. Eur. J. Hum. Genet. 2008, 16, 542–553. [Google Scholar] [CrossRef] [PubMed]
- Kahaly, G.J.; Frommer, L.; Schuppan, D. Celiac disease and endocrine autoimmunity—The genetic link. Autoimmun. Rev. 2018, 17, 1169–1175. [Google Scholar] [CrossRef]
- Kahaly, G.J.; Frommer, L.; Schuppan, D. Celiac Disease and Glandular Autoimmunity. Nutrients 2018, 10, 814. [Google Scholar] [CrossRef] [PubMed]
- Shewry, P.R.; Halford, N.G.; Belton, P.S.; Tatham, A.S. The structure and properties of gluten: An elastic protein from wheat grain. Philos. Trans. R. Soc. London Ser. B Biol. Sci. 2002, 357, 133–142. [Google Scholar] [CrossRef]
- Shan, L.; Molberg, Ø.; Parrot, I.; Hausch, F.; Filiz, F.; Gray, G.M.; Sollid, L.M.; Khosla, C. Structural Basis for Gluten Intolerance in Celiac Sprue. Science 2002, 297, 2275–2279. [Google Scholar] [CrossRef]
- Sollid, L.M. Coeliac disease: Dissecting a complex inflammatory disorder. Nat. Rev. Immunol. 2002, 2, 647–655. [Google Scholar] [CrossRef]
- Caminero, A.; Galipeau, H.J.; McCarville, J.L.; Johnston, C.W.; Bernier, S.P.; Russell, A.K.; Jury, J.; Herran, A.R.; Casqueiro, J.; Tye-Din, J.A.; et al. Duodenal Bacteria from Patients with Celiac Disease and Healthy Subjects Distinctly Affect Gluten Breakdown and Immunogenicity. Gastroenterology 2016, 151, 670–683. [Google Scholar] [CrossRef]
- van Heel, D.A.; Franke, L.; Hunt, K.A.; Gwilliam, R.; Zhernakova, A.; Inouye, M.; Wapenaar, M.C.; Barnardo, M.C.; Bethel, G.; Holmes, G.K.; et al. A genome-wide association study for celiac disease identifies risk variants in the region harboring IL2 and IL21. Nat. Genet. 2007, 39, 827–829. [Google Scholar] [CrossRef] [PubMed]
- Trynka, G.; Hunt, K.A.; Bockett, N.A.; Romanos, J.; Mistry, V.; Szperl, A.; Bakker, S.F.; Bardella, M.T.; Bhaw-Rosun, L.; Castillejo, G.; et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat. Genet. 2011, 43, 1193–1201. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Achury, J.; Zhernakova, A.; Pulit, S.; Trynka, G.; Hunt, K.A.; Romanos, J.; Raychaudhuri, S.; Van Heel, D.A.; Wijmenga, C.; De Bakker, P.I. Fine mapping in the MHC region accounts for 18% additional genetic risk for celiac disease. Nat. Genet. 2015, 47, 577–578. [Google Scholar] [CrossRef] [PubMed]
- Hunt, K.A.; Zhernakova, A.; Turner, G.; Heap, G.A.; Franke, L.; Bruinenberg, M.; Romanos, J.; Dinesen, L.C.; Ryan, A.W.; Panesar, D.; et al. Newly identified genetic risk variants for celiac disease related to the immune response. Nat. Genet. 2008, 40, 395–402. [Google Scholar] [CrossRef]
- Dubois, P.C.; Trynka, G.; Franke, L.; Hunt, K.A.; Romanos, J.; Curtotti, A.; Zhernakova, A.; Heap, G.A.; Adány, R.; Aromaa, A.; et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 2010, 42, 295–302. [Google Scholar] [CrossRef]
- Mansour, H.; Banaganapalli, B.; Nasser, K.K.; Al-Aama, J.Y.; Shaik, N.A.; Saadah, O.I.; Elango, R. Genome-Wide Association Study-Guided Exome Rare Variant Burden Analysis Identifies IL1R1 and CD3E as Potential Autoimmunity Risk Genes for Celiac Disease. Front. Pediatr. 2022, 10, 837957. [Google Scholar] [CrossRef]
- Wijmenga, C.; Zhernakova, A. The importance of cohort studies in the post-GWAS era. Nat. Genet. 2018, 50, 322–328. [Google Scholar] [CrossRef]
- Lindfors, K.; Ciacci, C.; Kurppa, K.; Lundin, K.E.A.; Makharia, G.K.; Mearin, M.L.; Murray, J.A.; Verdu, E.F.; Kaukinen, K. Coeliac disease. Nat. Rev. Dis. Prim. 2019, 5, 3. [Google Scholar] [CrossRef]
- Dieterich, W.; Ehnis, T.; Bauer, M.; Donner, P.; Volta, U.; Riecken, E.O.; Schuppan, D. Identification of tissue transglutaminase as the autoantigen of celiac disease. Nat. Med. 1997, 3, 797–801. [Google Scholar] [CrossRef]
- Di Niro, R.; Snir, O.; Kaukinen, K.; Yaari, G.; Lundin, K.E.; Gupta, N.T.; Kleinstein, S.H.; Cols, M.; Cerutti, A.; Mäki, M.; et al. Responsive population dynamics and wide seeding into the duodenal lamina propria of transglutaminase-2-specific plasma cells in celiac disease. Mucosal Immunol. 2016, 9, 254–264. [Google Scholar] [CrossRef]
- Iversen, R.; Snir, O.; Stensland, M.; Kroll, J.E.; Steinsbø, Ø.; Korponay-Szabó, I.R.; Lundin, K.E.A.; de Souza, G.A.; Sollid, L.M. Strong Clonal Relatedness between Serum and Gut IgA despite Different Plasma Cell Origins. Cell Rep. 2017, 20, 2357–2367. [Google Scholar] [CrossRef] [PubMed]
- Rauhavirta, T.; Hietikko, M.; Salmi, T.; Lindfors, K. Transglutaminase 2 and Transglutaminase 2 Autoantibodies in Celiac Disease: A Review. Clin. Rev. Allergy Immunol. 2019, 57, 23–38. [Google Scholar] [CrossRef] [PubMed]
- Malamut, G.; El Machhour, R.; Montcuquet, N.; Martin-Lannerée, S.; Dusanter-Fourt, I.; Verkarre, V.; Mention, J.-J.; Rahmi, G.; Kiyono, H.; Butz, E.A.; et al. IL-15 triggers an antiapoptotic pathway in human intraepithelial lymphocytes that is a potential new target in celiac disease–associated inflammation and lymphomagenesis. J. Clin. Investig. 2010, 120, 2131–2143. [Google Scholar] [CrossRef]
- Salvati, V.M.; MacDonald, T.T.; Bajaj-Elliott, M.; Borrelli, M.; Staiano, A.; Auricchio, S.; Troncone, R.; Monteleone, G. Interleukin 18 and associated markers of T helper cell type 1 activity in coeliac disease. Gut 2002, 50, 186–190. [Google Scholar] [CrossRef]
- Kutlu, T.; Brousse, N.; Rambaud, C.; Le Deist, F.; Schmitz, J.; Cerf-Bensussan, N. Numbers of T cell receptor (TCR) alpha beta+ but not of TcR gamma delta+ intraepithelial lymphocytes correlate with the grade of villous atrophy in coeliac patients on a long term normal diet. Gut 1993, 34, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Maiuri, L.; Ciacci, C.; Raia, V.; Vacca, L.; Ricciardelli, I.; Raimondi, F.; Auricchio, S.; Quaratino, S.; Londei, M. FAS engagement drives apoptosis of enterocytes of coeliac patients. Gut 2001, 48, 418–424. [Google Scholar] [CrossRef] [PubMed]
- Oberhuber, G.; Vogelsang, H.; Stolte, M.; Muthenthaler, S.; Kummer, J.A.; Radaszkiewicz, T. Evidence that intestinal intraepithelial lymphocytes are activated cytotoxic T cells in celiac disease but not in giardiasis. Am. J. Pathol. 1996, 148, 1351–1357. [Google Scholar]
- Hüe, S.; Mention, J.-J.; Monteiro, R.C.; Zhang, S.; Cellier, C.; Schmitz, J.; Verkarre, V.; Fodil, N.; Bahram, S.; Cerf-Bensussan, N.; et al. A Direct Role for NKG2D/MICA Interaction in Villous Atrophy during Celiac Disease. Immunity 2004, 21, 367–377. [Google Scholar] [CrossRef]
- Bhagat, G.; Naiyer, A.J.; Shah, J.G.; Harper, J.; Jabri, B.; Wang, T.C.; Green, P.H.; Manavalan, J.S. Small intestinal CD8+TCRgammadelta+NKG2A+ intraepithelial lymphocytes have attributes of regulatory cells in patients with celiac disease. J. Clin. Investig. 2008, 118, 281–293. [Google Scholar] [CrossRef]
- Abadie, V.; Discepolo, V.; Jabri, B. Intraepithelial lymphocytes in celiac disease immunopathology. Semin. Immunopathol. 2012, 34, 551–566. [Google Scholar] [CrossRef]
- Perez, F.; Ruera, C.N.; Miculan, E.; Carasi, P.; Chirdo, F.G. Programmed Cell Death in the Small Intestine: Implications for the Pathogenesis of Celiac Disease. Int. J. Mol. Sci. 2021, 22, 7426. [Google Scholar] [CrossRef] [PubMed]
- Setty, M.; Discepolo, V.; Abadie, V.; Kamhawi, S.; Mayassi, T.; Kent, A.; Ciszewski, C.; Maglio, M.; Kistner, E.; Bhagat, G.; et al. Distinct and Synergistic Contributions of Epithelial Stress and Adaptive Immunity to Functions of Intraepithelial Killer Cells and Active Celiac Disease. Gastroenterology 2015, 149, 681–691.e10. [Google Scholar] [CrossRef] [PubMed]
- Krzystek-Korpacka, M.; Kempiński, R.; Bromke, M.A.; Neubauer, K. Oxidative Stress Markers in Inflammatory Bowel Diseases: Systematic Review. Diagnostics 2020, 10, 601. [Google Scholar] [CrossRef] [PubMed]
- Wacklin, P.; Laurikka, P.; Lindfors, K.; Collin, P.; Salmi, T.; Lähdeaho, M.-L.; Saavalainen, P.; Mäki, M.; Mättö, J.; Kurppa, K.; et al. Altered Duodenal Microbiota Composition in Celiac Disease Patients Suffering from Persistent Symptoms on a Long-Term Gluten-Free Diet. Am. J. Gastroenterol. 2014, 109, 1933–1941. [Google Scholar] [CrossRef]
- Sánchez, E.; Donat, E.; Ribes-Koninckx, C.; Fernández-Murga, M.L.; Sanz, Y. Duodenal-Mucosal Bacteria Associated with Celiac Disease in Children. Appl. Environ. Microbiol. 2013, 79, 5472–5479. [Google Scholar] [CrossRef]
- D’Argenio, V.; Casaburi, G.; Precone, V.; Pagliuca, C.; Colicchio, R.; Sarnataro, D.; Discepolo, V.; Kim, S.M.; Russo, I.; Del Vecchio Blanco, G.; et al. Metagenomics Reveals Dysbiosis and a Potentially Pathogenic N. flavescens Strain in Duodenum of Adult Celiac Patients. Am. J. Gastroenterol. 2016, 111, 879–890. [Google Scholar] [CrossRef]
- Girbovan, A.; Sur, G.; Samasca, G.; Lupan, I. Dysbiosis a risk factor for celiac disease. Med. Microbiol. Immunol. 2017, 206, 83–91. [Google Scholar] [CrossRef]
- Chibbar, R.; Dieleman, L.A. The Gut Microbiota in Celiac Disease and Probiotics. Nutrients 2019, 11, 2375. [Google Scholar] [CrossRef]
- Nomura, K.; Ishikawa, D.; Okahara, K.; Ito, S.; Haga, K.; Takahashi, M.; Arakawa, A.; Shibuya, T.; Osada, T.; Kuwahara-Arai, K.; et al. Bacteroidetes Species Are Correlated with Disease Activity in Ulcerative Colitis. J. Clin. Med. 2021, 10, 1749. [Google Scholar] [CrossRef]
- Austin, A.S.; Logan, R.F.; Thomason, K.; Holmes, G.K. Cigarette smoking and adult coeliac disease. Scand. J. Gastroenterol. 2002, 37, 978–982. [Google Scholar] [CrossRef]
- Rubio-Tapia, A.; Ludvigsson, J.F.; Brantner, T.L.; Murray, J.A.; Everhart, J.E. The Prevalence of Celiac Disease in the United States. Am. J. Gastroenterol. 2012, 107, 1538–1544. [Google Scholar] [CrossRef] [PubMed]
- Choung, R.S.; Larson, S.A.; Khaleghi, S.; Rubio-Tapia, A.; Ovsyannikova, I.G.; King, K.S.; Larson, J.J.; Lahr, B.D.; Poland, G.A.; Camilleri, M.J.; et al. Prevalence and Morbidity of Undiagnosed Celiac Disease from a Community-Based Study. Gastroenterology 2017, 152, 830–839.e5. [Google Scholar] [CrossRef] [PubMed]
- Schuppan, D.; Hahn, E.G. Celiac disease and its link to type 1 diabetes mellitus. J. Pediatr. Endocrinol. Metab. 2001, 14 (Suppl. S1), 597–605. [Google Scholar] [CrossRef] [PubMed]
- Acerini, C.L.; Ahmed, M.L.; Ross, K.M.; Sullivan, P.B.; Bird, G.; Dunger, D.B. Coeliac disease in children and adolescents with IDDM: Clinical characteristics and response to gluten-free diet. Diabet. Med. 1998, 15, 38–44. [Google Scholar] [CrossRef]
- Cronin, C.C.; Feighery, A.; Ferriss, J.B.; Liddy, C.; Shanahan, F.; Feighery, C. High prevalence of celiac disease among patients with insulin-dependent (type I) diabetes mellitus. Am. J. Gastroenterol. 1997, 92, 2210–2212. [Google Scholar]
- Talal, A.H.; Murray, J.A.; Goeken, J.A.; Sivitz, W.I. Celiac disease in an adult population with insulin-dependent diabetes mellitus: Use of endomysial antibody testing. Am. J. Gastroenterol. 1997, 92, 1280–1284. [Google Scholar]
- Counsell, C.E.; Taha, A.; Ruddell, W.S. Coeliac disease and autoimmune thyroid disease. Gut 1994, 35, 844–846. [Google Scholar] [CrossRef]
- Badenhoop, K.; Dieterich, W.; Segni, M.; Hofmann, S.; Hüfner, M.; Usadel, K.H.; Hahn, E.G.; Schuppan, D. HLA DQ2 and/or DQ8 Is Associated With Celiac Disease–Specific Autoantibodies to Tissue Transglutaminase in Families With Thyroid Autoimmunity. Am. J. Gastroenterol. 2001, 96, 1648–1649. [Google Scholar] [CrossRef]
- Ciacci, C.; Cavallaro, R.; Iovino, P.; Sabbatini, F.; Palumbo, A.; Amoruso, D.; Tortora, R.; Mazzacca, G. Allergy prevalence in adult celiac disease. J. Allergy Clin. Immunol. 2004, 113, 1199–1203. [Google Scholar] [CrossRef]
- Zauli, D.; Grassi, A.; Granito, A.; Foderaro, S.; De Franceschi, L.; Ballardini, G.; Bianchi, F.; Volta, U. Prevalence of silent coeliac disease in atopics. Dig. Liver Dis. 2000, 32, 775–779. [Google Scholar] [CrossRef]
- Volta, U.; De Giorgio, R.; Granito, A.; Stanghellini, V.; Barbara, G.; Avoni, P.; Liguori, R.; Petrolini, N.; Fiorini, E.; Montagna, P.; et al. Anti-ganglioside antibodies in coeliac disease with neurological disorders. Dig. Liver Dis. 2006, 38, 183–187. [Google Scholar] [CrossRef] [PubMed]
- Cervio, E.; Volta, U.; Verri, M.; Boschi, F.; Pastoris, O.; Granito, A.; Barbara, G.; Parisi, C.; Felicani, C.; Tonini, M.; et al. Sera of Patients With Celiac Disease and Neurologic Disorders Evoke a Mitochondrial-Dependent Apoptosis In Vitro. Gastroenterology 2007, 133, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Wierdsma, N.J.; Nijeboer, P.; de van der Schueren, M.A.; Berkenpas, M.; van Bodegraven, A.A.; Mulder, C.J. Refractory celiac disease and EATL patients show severe malnutrition and malabsorption at diagnosis. Clin. Nutr. 2016, 35, 685–691. [Google Scholar] [CrossRef] [PubMed]
- García-Hoz, C.; Crespo, L.; Lopez, N.; De Andrés, A.; Ríos León, R.; Santón, A.; Garriga, M.; Butz, E.; León, F.; Roy Ariño, G. The Intracellular Intensity of CD3 on Aberrant Intraepithelial Lymphocytes Is a Prognostic Factor of the Progression to Overt Lymphoma in Refractory Celiac Disease Type II (Pre-Enteropathy-Associated T Cell Lymphoma). Dig. Dis. 2020, 38, 490–499. [Google Scholar] [CrossRef]
- Soderquist, C.R.; Lewis, S.K.; Gru, A.A.; Vlad, G.; Williams, E.S.; Hsiao, S.; Mansukhani, M.M.; Park, D.C.; Bacchi, C.E.; Alobeid, B.; et al. Immunophenotypic Spectrum and Genomic Landscape of Refractory Celiac Disease Type II. Am. J. Surg. Pathol. 2021, 45, 905–916. [Google Scholar] [CrossRef]
- Chibbar, R.; Nostedt, J.; Mihalicz, D.; Deschenes, J.; McLean, R.; Dieleman, L.A. Refractory Celiac Disease Type II: A Case Report and Literature Review. Front. Med. 2020, 7, 564875. [Google Scholar] [CrossRef]
- Liu, H.; Brais, R.; Lavergne-Slove, A.; Jeng, Q.; Payne, K.; Ye, H.; Liu, Z.; Carreras, J.; Huang, Y.; Bacon, C.M.; et al. Continual monitoring of intraepithelial lymphocyte immunophenotype and clonality is more important than snapshot analysis in the surveillance of refractory coeliac disease. Gut 2010, 59, 452–460. [Google Scholar] [CrossRef]
- Mulder, C.J.; Wahab, P.J.; Moshaver, B.; Meijer, J.W. Refractory coeliac disease: A window between coeliac disease and enteropathy associated T cell lymphoma. Scand. J. Gastroenterol. Suppl. 2000, 232, 32–37. [Google Scholar]
- Wolf, J.; Willscher, E.; Loeffler-Wirth, H.; Schmidt, M.; Flemming, G.; Zurek, M.; Uhlig, H.H.; Händel, N.; Binder, H. Deciphering the Transcriptomic Heterogeneity of Duodenal Coeliac Disease Biopsies. Int. J. Mol. Sci. 2021, 22, 2551. [Google Scholar] [CrossRef]
- Carreras, J.; Nakamura, N.; Hamoudi, R. Artificial Intelligence Analysis of Gene Expression Predicted the Overall Survival of Mantle Cell Lymphoma and a Large Pan-Cancer Series. Healthcare 2022, 10, 155. [Google Scholar] [CrossRef]
- Carreras, J.; Hamoudi, R.; Nakamura, N. Artificial Intelligence Analysis of Gene Expression Data Predicted the Prognosis of Patients with Diffuse Large B-Cell Lymphoma. Tokai J. Exp. Clin. Med. 2020, 45, 37–48. [Google Scholar] [PubMed]
- Carreras, J.; Kikuti, Y.Y.; Miyaoka, M.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; Nakamura, N.; Hamoudi, R. A Combination of Multilayer Perceptron, Radial Basis Function Artificial Neural Networks and Machine Learning Image Segmentation for the Dimension Reduction and the Prognosis Assessment of Diffuse Large B-Cell Lymphoma. AI 2021, 2, 106–134. [Google Scholar] [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Miyaoka, M.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; Shiraiwa, S.; Hamoudi, R.; et al. A Single Gene Expression Set Derived from Artificial Intelligence Predicted the Prognosis of Several Lymphoma Subtypes; and High Immunohistochemical Expression of TNFAIP8 Associated with Poor Prognosis in Diffuse Large B-Cell Lympho-ma. AI 2020, 1, 342–360. [Google Scholar] [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Roncador, G.; Miyaoka, M.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; Shiraiwa, S.; et al. High Expression of Caspase-8 Associated with Improved Survival in Diffuse Large B-Cell Lymphoma: Machine Learning and Artificial Neural Networks Analyses. BioMedInformatics 2021, 1, 18–46. [Google Scholar] [CrossRef]
- Carreras, J.; Hiraiwa, S.; Kikuti, Y.Y.; Miyaoka, M.; Tomita, S.; Ikoma, H.; Ito, A.; Kondo, Y.; Roncador, G.; Garcia, J.F.; et al. Artificial Neural Networks Predicted the Overall Survival and Molecular Subtypes of Diffuse Large B-Cell Lymphoma Using a Pancancer Immune-Oncology Panel. Cancers 2021, 13, 6384. [Google Scholar] [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Miyaoka, M.; Roncador, G.; Garcia, J.F.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; et al. Integrative Statistics, Machine Learning and Artificial Intelligence Neural Network Analysis Correlated CSF1R with the Prognosis of Diffuse Large B-Cell Lymphoma. Hemato 2021, 2, 182–206. [Google Scholar] [CrossRef]
- Carreras, J.; Hamoudi, R. Artificial Neural Network Analysis of Gene Expression Data Predicted Non-Hodgkin Lymphoma Subtypes with High Accuracy. Mach. Learn. Knowl. Extr. 2021, 3, 720–739. [Google Scholar] [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Miyaoka, M.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; Nakamura, N.; Hamoudi, R. Artificial Intelligence Analysis of the Gene Expression of Follicular Lymphoma Predicted the Overall Survival and Correlated with the Immune Microenvironment Response Signatures. Mach. Learn. Knowl. Extr. 2020, 2, 647–671. [Google Scholar] [CrossRef]
- Carreras, J.; Kikuti, Y.Y.; Miyaoka, M.; Hiraiwa, S.; Tomita, S.; Ikoma, H.; Kondo, Y.; Ito, A.; Hamoudi, R.; Nakamura, N. The Use of the Random Number Generator and Artificial Intelligence Analysis for Dimensionality Reduction of Follicular Lymphoma Transcriptomic Data. BioMedInformatics 2022, 2, 268–280. [Google Scholar] [CrossRef]
- Carreras, J. Artificial Intelligence Analysis of Ulcerative Colitis Using an Autoimmune Discovery Transcriptomic Panel. Healthcare 2022, 10, 1476. [Google Scholar] [CrossRef]
- Uhde, M.; Yu, X.; Bunin, A.; Brauner, C.; Lewis, S.K.; Lebwohl, B.; Krishnareddy, S.; Alaedini, A.; Reizis, B.; Ghosh, S.; et al. Phenotypic shift of small intestinal intra-epithelial type 1 innate lymphoid cells in celiac disease is associated with enhanced cytotoxic potential. Clin. Exp. Immunol. 2020, 200, 163–175. [Google Scholar] [CrossRef] [PubMed]
- van Wanrooij, R.L.; Müller, D.M.; Neefjes-Borst, E.A.; Meijer, J.; Koudstaal, L.G.; Heideman, D.A.; Bontkes, H.J.; von Blomberg, B.M.; Bouma, G.; Mulder, C.J. Optimal Strategies to Identify Aberrant Intra-Epithelial Lymphocytes in Refractory Coeliac Disease. J. Clin. Immunol. 2014, 34, 828–835. [Google Scholar] [CrossRef] [PubMed]
- Verbeek, W.H.; Goerres, M.S.; von Blomberg, B.M.; Oudejans, J.J.; Scholten, P.E.; Hadithi, M.; Al-Toma, A.; Schreurs, M.W.; Mulder, C.J. Flow cytometric determination of aberrant intra-epithelial lymphocytes predicts T-cell lymphoma development more accurately than T-cell clonality analysis in Refractory Celiac Disease. Clin. Immunol. 2008, 126, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Müller, L.; Szaflarska-Popławska, A. Immunologiczne mechanizmy w chorobie trzewnej [Immunological mechanisms of celiac disease]. Przegl. Lek. 2005, 62, 123–127. (In Polish) [Google Scholar] [PubMed]
- Mazzarella, G. Effector and suppressor T cells in celiac disease. World J. Gastroenterol. 2015, 21, 7349–7356. [Google Scholar] [CrossRef]
- IBM Cloud Education. Machine Learning. 15 July 2020. IBM Cloud Learn Hub. IBM Corporation. Available online: https://www.ibm.com/cloud/learn/machine-learning (accessed on 13 July 2022).
- IBM. IBM SPSS Neural Networks 26; IBM: Armonk, NY, USA, 2019. [Google Scholar]
- IBM. IBM SPSS Neural Networks; New Tools for Building Predictive Models; YTD03119-GBEN-01; IBM: Somers, NY, USA, 2012. [Google Scholar]
- Schumann, M.; Siegmund, B.; Schulzke, J.D.; Fromm, M. Celiac Disease: Role of the Epithelial Barrier. Cell. Mol. Gastroenterol. Hepatol. 2017, 3, 150–162. [Google Scholar] [CrossRef]
- Shalimar, D.M.; Das, P.; Sreenivas, V.; Gupta, S.D.; Panda, S.K.; Makharia, G.K. Mechanism of Villous Atrophy in Celiac Disease: Role of Apoptosis and Epithelial Regeneration. Arch. Pathol. Lab. Med. 2013, 137, 1262–1269. [Google Scholar] [CrossRef]
- Györy, I.; Fejér, G.; Ghosh, N.; Seto, E.; Wright, K.L. Identification of a Functionally Impaired Positive Regulatory Domain I Binding Factor 1 Transcription Repressor in Myeloma Cell Lines. J. Immunol. 2003, 170, 3125–3133. [Google Scholar] [CrossRef]
- Høydahl, L.S.; Richter, L.; Frick, R.; Snir, O.; Gunnarsen, K.S.; Landsverk, O.J.B.; Iversen, R.; Jeliazkov, J.; Gray, J.J.; Bergseng, E.; et al. Plasma Cells Are the Most Abundant Gluten Peptide MHC-expressing Cells in Inflamed Intestinal Tissues From Patients with Celiac Disease. Gastroenterology 2019, 156, 1428–1439.e10. [Google Scholar] [CrossRef]
- Pohjanen, V.M.; Kokkonen, T.S.; Arvonen, M.; Augustin, M.A.; Patankar, M.; Turunen, S.; Vähäsalo, P.; Karttunen, T.J. Decreased Expression of Protease Inhibitor 9, a Granzyme B Inhibitor, in Celiac Disease: A Potential Mechanism in Enterocyte Destruction and Villous Atrophy. Int. J. Immunopathol. Pharmacol. 2013, 26, 897–905. [Google Scholar] [CrossRef]
- Huard, B.; Tournier, M.; Hercend, T.; Triebel, F.; Faure, F. Lymphocyte-activation gene 3/major histocompatibility complex class II interaction modulates the antigenic response of CD4+ T lymphocytes. Eur. J. Immunol. 1994, 24, 3216–3221. [Google Scholar] [CrossRef] [PubMed]
- Gagliani, N.; Magnani, C.F.; Huber, S.; Gianolini, M.E.; Pala, M.; Licona-Limon, P.; Guo, B.; Herbert, D.R.; Bulfone, A.; Trentini, F.; et al. Coexpression of CD49b and LAG-3 identifies human and mouse T regulatory type 1 cells. Nat. Med. 2013, 19, 739–746. [Google Scholar] [CrossRef] [PubMed]
- Bauché, D.; Joyce-Shaikh, B.; Jain, R.; Grein, J.; Ku, K.S.; Blumenschein, W.M.; Ganal-Vonarburg, S.C.; Wilson, D.C.; McClanahan, T.K.; Malefyt, R.W.; et al. LAG3+ Regulatory T Cells Restrain Interleukin-23-Producing CX3CR1+ Gut-Resident Macrophages during Group 3 Innate Lymphoid Cell-Driven Colitis. Immunity 2018, 49, 342–352.e5. [Google Scholar] [CrossRef] [PubMed]
- Gianfrani, C.; Levings, M.K.; Sartirana, C.; Mazzarella, G.; Barba, G.; Zanzi, D.; Camarca, A.; Iaquinto, G.; Giardullo, N.; Auricchio, S.; et al. Gliadin-specific type 1 regulatory T cells from the intestinal mucosa of treated celiac patients inhibit pathogenic T cells. J. Immunol. 2006, 177, 4178–4186. [Google Scholar] [CrossRef] [PubMed]
- van Leeuwen, M.A.; Costes, L.M.M.; van Berkel, L.A.; Simons-Oosterhuis, Y.; du Pré, M.F.; Kozijn, A.E.; Raatgeep, H.C.; Linden-bergh-Kortleve, D.J.; van Rooijen, N.; Koning, F.; et al. Macrophage-mediated gliadin degradation and concomitant IL-27 production drive IL-10- and IFN-γ-secreting Tr1-like-cell differentiation in a murine model for gluten tolerance. Mucosal Immunol. 2017, 10, 635–649. [Google Scholar] [CrossRef]
- Villarino, A.V.; Sciumè, G.; Davis, F.P.; Iwata, S.; Zitti, B.; Robinson, G.W.; Hennighausen, L.; Kanno, Y.; O’Shea, J.J. Subset- and tissue-defined STAT5 thresholds control homeostasis and function of innate lymphoid cells. J. Exp. Med. 2017, 214, 2999–3014. [Google Scholar] [CrossRef]
- Gilbert, S.; Zhang, R.; Denson, L.; Moriggl, R.; Steinbrecher, K.; Shroyer, N.; Lin, J.; Han, X. Enterocyte STAT5 promotes mucosal wound healing via suppression of myosin light chain kinase-mediated loss of barrier function and inflammation. EMBO Mol. Med. 2012, 4, 109–124. [Google Scholar] [CrossRef]
- Rubin, C.E.; Brandborg, L.L.; Phelps, P.C.; Taylor, H.C., Jr. Studies of celiac disease I. The apparent identical and specific nature of the duodenal and proximal jejunal lesion in celiac disease and idiopathic sprue. Gastroenterology 1960, 38, 28–49. [Google Scholar] [CrossRef]
- Hujoel, I.A.; Murray, J.A. Refractory Celiac Disease. Curr. Gastroenterol. Rep. 2020, 22, 18. [Google Scholar] [CrossRef]
- Oberhuber, G.; Granditsch, G.; Vogelsang, H. The histopathology of coeliac disease: Time for a standardized report scheme for pathologists. Eur. J. Gastroenterol. Hepatol. 1999, 11, 1185–1194. [Google Scholar] [CrossRef]
- Kelly, C.P.; Lamont, J.T.; Grover, S. Diagnosis of Celiac Disease in Adults. UpToDate. 7 April 2022. Available online: https://www.uptodate.com/contents/diagnosis-of-celiac-disease-in-adults?search=celiac%20disease&source=search_result&selectedTitle=1~150&usage_type=default&display_rank=1#H3181992152 (accessed on 13 July 2022).









| Factors | Pathophysiology | References |
|---|---|---|
| Dietary gluten | ① Gluten of wheat, rye, and barley. Gliadins and glutenins are rich in proline, which makes them resistant to proteolysis by gastric and pancreatic enzymes. Various long gliadin peptides activate the immune system (“33mer”). Undigested peptides may also affect intestinal microbiota. | [18,19,20,21] |
| Genetics | ① Genetic predisposition: HLA-DQ2 and HLA-DQ8 contribute to 20%–40% of the genetic risk. They are class II MHC expressed by antigen-presenting cells (APCs). | [22,23,24] |
| ② Forty-two non-HLA regions have been associated with celiac disease. It is estimated that they account for 15% of the genetic risk: IL18R1, IL18RAP, STAT4, CD28, CTLA4, ICOS, CCR4, CCR1, CCR2, CCR3, CD3E, IL1R1, IL12A, IL2, IL21, TNFAIP3, ELMO1, PRKCQ, SOCS1, ICOSLG, and IRAK1. These genes belong to cytokine-cytokine receptor activation, JAK-STAT pathway, T-cell receptor signaling, intestinal immune network for IgA production, NF-KB signaling, and cell adhesion molecules. Of note, many of these genes belong to the immune checkpoint and immune-oncology pathway. | [22,23,25,26,27,28] | |
| Immune | ① Generation of gluten-specific T-cell responses: presence of gluten-specific CD4-positive T lymphocytes, antibodies against gliadin and de enzyme TG2, and pro-inflammatory cytokines. | [29,30] |
| ② Generation of autoantibodies: activation and differentiation into plasma cells of gluten-specific and TG2-specific B lymphocytes, generation of autoantibodies that are both circulating and deposited in the mucosa. These autoantibodies are responsible for the increased permeability of the epithelial barrier. | [31,32,33] | |
| ③ Cytokines in the intestinal mucosal immune system: IFN gamma and IL-21 are produced by gluten-specific CD4-positive T lymphocytes. Secretion of IL-15, IL-18, and inhibition of FOXP3-positive regulatory T lymphocytes (Tregs). | [34,35] | |
| ④ Intraepithelial lymphocytes (IELs): increased in celiac disease and their amount correlates with mucosal atrophy. IELs display cytotoxic transformation and induce apoptosis of intestinal epithelial cells through FAS-L, perforin, granzyme B, and NKG2D. NKG2D interacts with MICA on epithelial cells. | [36,37,38,39,40,41,42] | |
| ⑤ Innate immune activation: dysregulation of the production of IL-15 and activation of the innate immune response, including the induction of epithelial stress. | [43,44] | |
| Environmental | ① Microorganisms: intestinal dysbiosis (unbalanced intestinal microbiota) and increased prevalence of specific microbial virulence genes isolated from celiac disease patients. | [45,46,47,48,49,50] |
| ② Others, such as smoking | [51] |
| Model | Overall Accuracy (%) | No. Genes (Fields) Used | Most Relevant Genes |
|---|---|---|---|
| C5 | 100 | 1 | IFNG |
| Logistic regression | 100 | 737 | (Refer to Table 3) |
| Discriminant | 100 | 737 | - |
| LSVM 1 | 100 | 737 | CASP1, IL18, ARPC2, CASP3, KLF4, GBP1, SULT1A1, RNASET2, MIF, and PIGR |
| SVM | 100 | 737 | - |
| XGBoost linear | 100 | 737 | - |
| XGBoost tree | 100 | 737 | - |
| CHAID | 100 | 2 | BATF, GBP1 |
| C&R tree | 100 | 6 | IFNG |
| Random forest 1 | 100 | 737 | CXCL10, PRDM1, GZMB, STAT2, IL12RB1, LAG3, PTPN22, TMEM50B, IFI35, PRDX5, GALC, C1QBP, RIPK2, IFNG, CSF2, STAT5A, TNPO3, IQCB1, and DEXI |
| Neural network 1 | 100 | 737 | CXCL2, IL7R, PLCH2, CCL23, MBD2, CSF3R, MUC1, GPR183, CD226, and PNMT |
| KNN algorithm | 96 | 737 | - |
| Quest | 96 | 6 | STAT1 |
| Random trees 1 | 86 | 737 | BTLA, CARD14, CASP10, CCL13, CCL5, CCR7, CXCL10, CXCL9, CXCR6, ELMO1, and EXTL |
| Bayesian network | 58 | 737 | - |
| Equation for Predicting Celiac Disease |
|---|
| −0.1765 × AAMP + −0.008 × ABHD6 + −0.1178 × ACKR2 + −1.725 × ACOXL + 0.6231 × ACSL6 + 0.0009441 × ADA + 1.16 × ADAM30 + 0.04882 × ADCY3 + 1.108 × ADCY7 + 0.2923 × AFF3 + −0.5828 × AGAP2 + 0.6009 × AHI1 + 0.3013 × AHR + −0.002197 × AIRE + −0.7633 × ANKRD55 + 0.06059 × ANTXR2 + 0.2416 × APEH + 1.215 × APOBEC3G + −2.377 × ARG1 + −0.2806 × ARHGAP30 + 0.0796 × ARID5B + −0.0168 × ARPC2 + −0.009025 × ATF4 + −0.08039 × ATG16L1 + −0.156 × ATG5 + 0.09123 × ATM + 0.003949 × B2M + 0.02826 × B3GNT2 + −0.2021 × BABAM2 + 1.132 × BACH2 + −0.6567 × BAD + −0.2759 × BANK1 + −0.09905 × BATF + 0.617 × BATF3 + 0.1081 × BCL10 + −0.1113 × BCL3 + 0.2034 × BCL6 + 0.7125 × BID + 0.3596 × BLK + 0.1998 × BLNK + −0.4926 × BORCS5 + −3.589 × BSN + 1.291 × BTK + −1.079 × BTLA + −1.254 × BTNL2 + −0.1576 × C1orf53 + 0.004046 × C1QBP + −49.65 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carreras, J. Artificial Intelligence Analysis of Celiac Disease Using an Autoimmune Discovery Transcriptomic Panel Highlighted Pathogenic Genes including BTLA. Healthcare 2022, 10, 1550. https://doi.org/10.3390/healthcare10081550
Carreras J. Artificial Intelligence Analysis of Celiac Disease Using an Autoimmune Discovery Transcriptomic Panel Highlighted Pathogenic Genes including BTLA. Healthcare. 2022; 10(8):1550. https://doi.org/10.3390/healthcare10081550
Chicago/Turabian StyleCarreras, Joaquim. 2022. "Artificial Intelligence Analysis of Celiac Disease Using an Autoimmune Discovery Transcriptomic Panel Highlighted Pathogenic Genes including BTLA" Healthcare 10, no. 8: 1550. https://doi.org/10.3390/healthcare10081550
APA StyleCarreras, J. (2022). Artificial Intelligence Analysis of Celiac Disease Using an Autoimmune Discovery Transcriptomic Panel Highlighted Pathogenic Genes including BTLA. Healthcare, 10(8), 1550. https://doi.org/10.3390/healthcare10081550






