BTK, NUTM2A, and PRPF19 Are Novel KMT2A Partner Genes in Childhood Acute Leukemia
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patient 1
2.2. Patient 2
2.3. Patient 3
3. Results
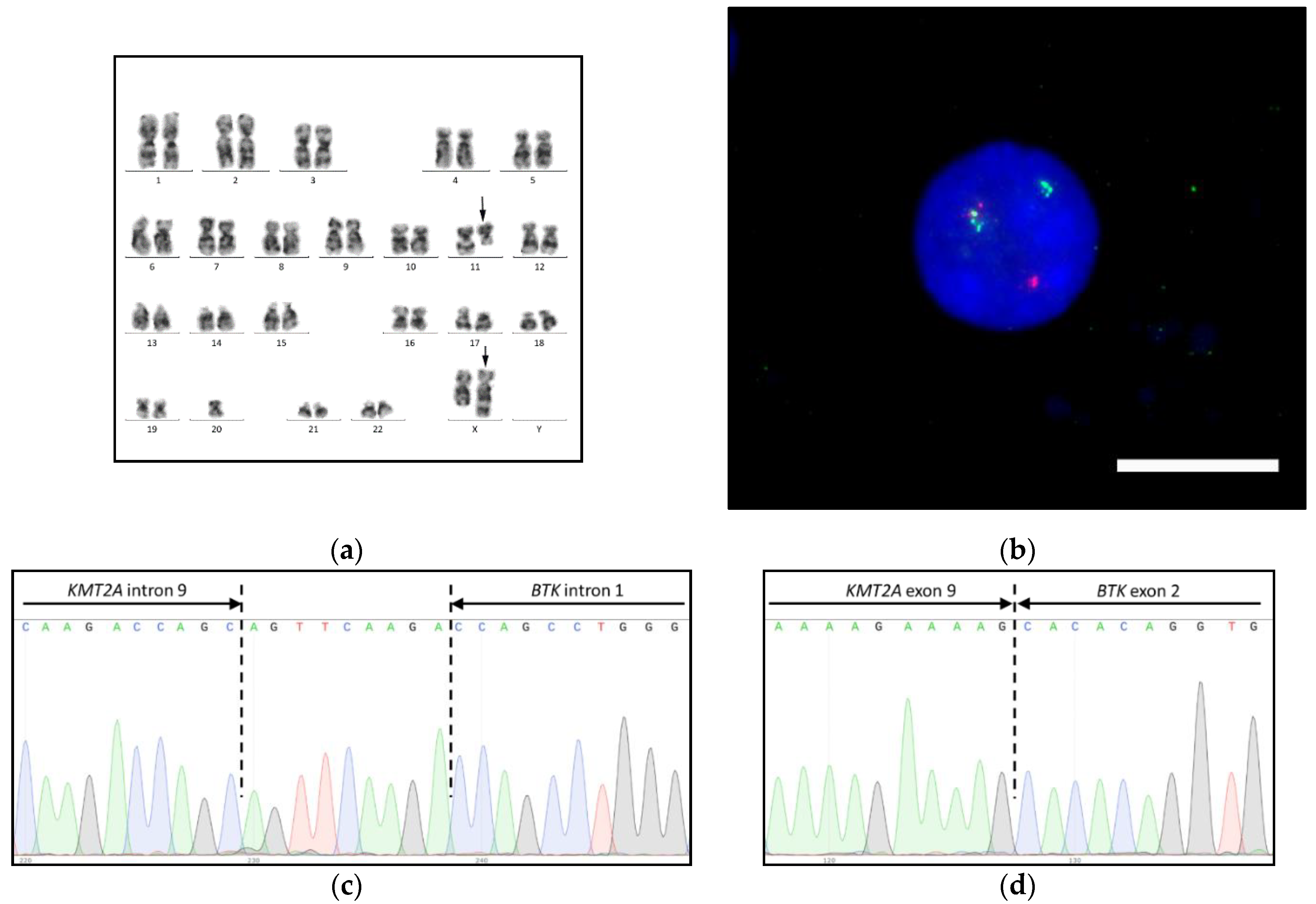
3.1. KMT2A-BTK Fusion Gene in AML with ins(X;11)(q26;q13q25)
3.2. KMT2A-NUTM2A Fusion Gene in T-ALL with t(10;11)(q22;q23.3)
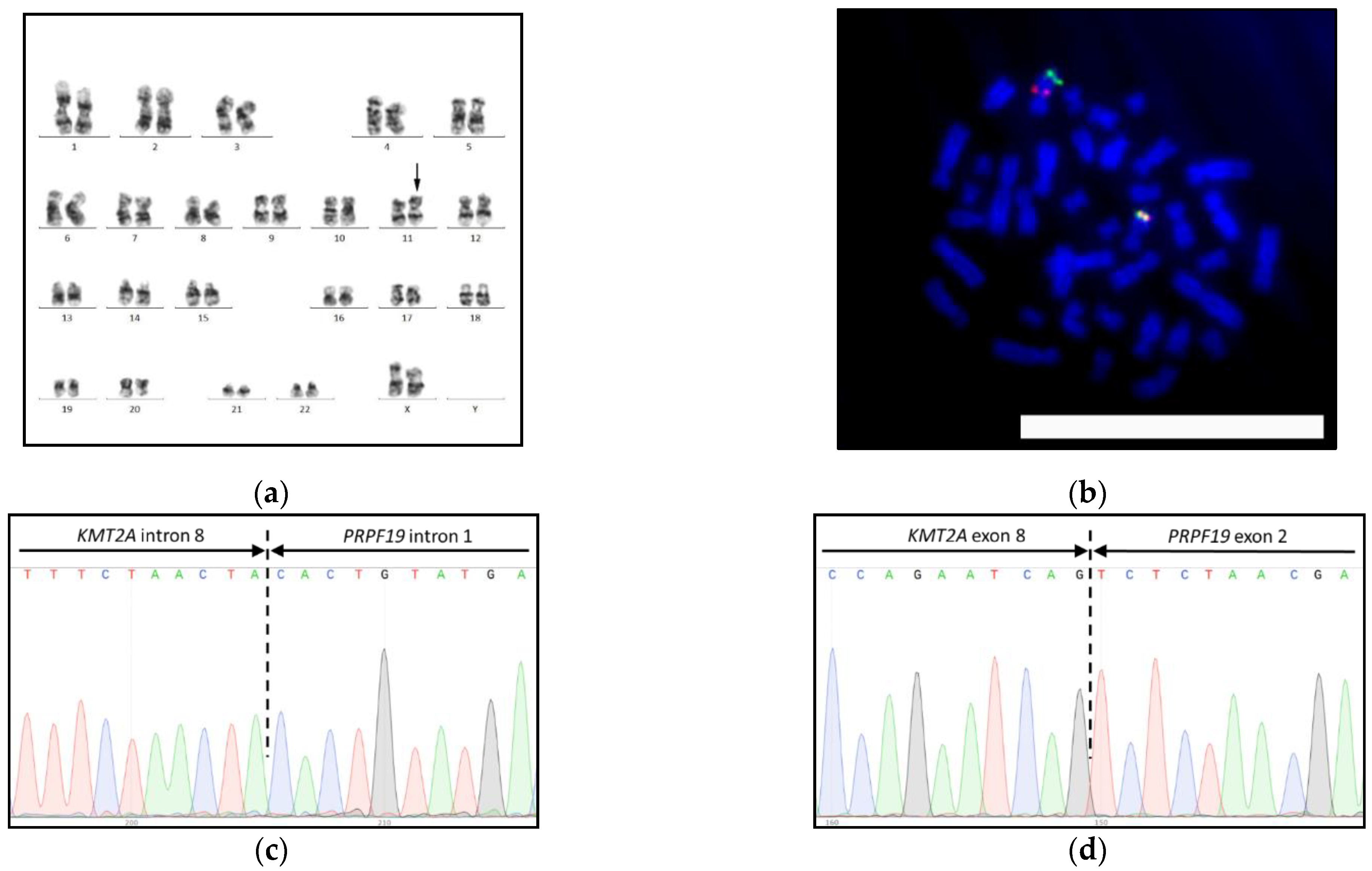
3.3. KMT2A-PRPF19 Fusion Gene in AML with inv(11)(q12.2q23.3)
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Swerdlow, S.H.; Campo, E.; Harris, N.L.; Jaffe, E.S.; Pileri, S.A.; Stein, H.; Thiele, J. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, 4th ed.; IARC: Lyon, France, 2017; pp. 153–155. [Google Scholar]
- Balgobind, B.V.; Raimondi, S.C.; Harbott, J.; Zimmermann, M.; Alonzo, T.A.; Auvrignon, A.; Beverloo, H.B.; Chang, M.; Creutzig, U.; Dworzak, M.N.; et al. Novel prognostic subgroups in childhood 11q23/MLL-rearranged acute myeloid leukemia: Results of an international retrospective study. Blood 2009, 114, 2489–2496. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chessells, J.M.; Harrison, C.; Kempski, H.; Webb, D.K.H.; Wheatley, K.; Hann, I.M.; Stevens, R.F.; Harrison, G.; Gibson, B.E. Clinical features, cytogenetics and outcome in acute lymphoblastic and myeloid leukaemia of infancy: Report from the MRC Childhood Leukaemia working party. Leukemia 2002, 16, 776–784. [Google Scholar] [CrossRef] [Green Version]
- Creutzig, U.; Zimmermann, M.; Reinhardt, D.; Rasche, M.; Von Neuhoff, C.; Alpermann, T.; Dworzak, M.; Perglerová, K.; Zemanova, Z.; Tchinda, J.; et al. Changes in cytogenetics and molecular genetics in acute myeloid leukemia from childhood to adult age groups. Cancer 2016, 122, 3821–3830. [Google Scholar] [CrossRef] [PubMed]
- Mullighan, C.G. Molecular genetics of B-precursor acute lymphoblastic leukemia. J. Clin. Investig. 2012, 122, 3407–3415. [Google Scholar] [CrossRef] [Green Version]
- Harrison, C.J.; Hills, R.; Moorman, A.; Grimwade, D.J.; Hann, I.; Webb, D.K.H.; Wheatley, K.; De Graaf, S.S.N.; Berg, E.V.D.; Burnett, A.K.; et al. Cytogenetics of Childhood Acute Myeloid Leukemia: United Kingdom Medical Research Council Treatment Trials AML 10 and 12. J. Clin. Oncol. 2010, 28, 2674–2681. [Google Scholar] [CrossRef] [PubMed]
- Balgobind, B.V.; Zwaan, C.M.; Pieters, R.; Heuvel-Eibrink, M.M.V.D. The heterogeneity of pediatric MLL-rearranged acute myeloid leukemia. Leukemia 2011, 25, 1239–1248. [Google Scholar] [CrossRef] [Green Version]
- Meyer, C.; Burmeister, T.; Gröger, D.; Tsaur, G.; Fechina, L.; Renneville, A.; Sutton, R.; Venn, N.C.; Emerenciano, M.; Pombo-De-Oliveira, M.S.; et al. The MLL recombinome of acute leukemias in 2017. Leukemia 2018, 32, 273–284. [Google Scholar] [CrossRef] [PubMed]
- Moorman, A.V.; Ensor, H.M.; Richards, S.M.; Chilton, L.; Schwab, C.; Kinsey, S.E.; Vora, A.; Mitchell, C.D.; Harrison, C. Prognostic effect of chromosomal abnormalities in childhood B-cell precursor acute lymphoblastic leukaemia: Results from the UK Medical Research Council ALL97/99 randomised trial. Lancet Oncol. 2010, 11, 429–438. [Google Scholar] [CrossRef]
- Emerenciano, M.; Meyer, C.; Mansur, M.B.; Marschalek, R.; Pombo-De-Oliveira, M.S. The Brazilian Collaborative Study Group of Infant Acute Leukaemia The distribution ofMLLbreakpoints correlates with outcome in infant acute leukaemia. Br. J. Haematol. 2013, 161, 224–236. [Google Scholar] [CrossRef] [PubMed]
- Zerkalenkova, E.; Lebedeva, S.; Kazakova, A.; Baryshev, P.; Meyer, C.; Marschalek, R.; Novichkova, G.; Maschan, M.; Maschan, A.; Olshanskaya, Y. A case of pediatric acute myeloid leukemia with t(11;16)(q23;q24) leading to a novel KMT2A-USP10 fusion gene. Genes, Chromosom. Cancer 2018, 57, 522–524. [Google Scholar] [CrossRef] [PubMed]
- Marschalek, R. Systematic Classification of Mixed-Lineage Leukemia Fusion Partners Predicts Additional Cancer Pathways. Ann. Lab. Med. 2016, 36, 85–100. [Google Scholar] [CrossRef] [Green Version]
- Winters, A.C.; Bernt, K.M. MLL-Rearranged Leukemias—An Update on Science and Clinical Approaches. Front. Pediatr. 2017, 5, 4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meyer, C.; Lopes, B.A.; Caye-Eude, A.; Cavé, H.; Arfeuille, C.; Cuccuini, W.; Sutton, R.; Venn, N.C.; Oh, S.H.; Tsaur, G.; et al. Human MLL/KMT2A gene exhibits a second breakpoint cluster region for recurrent MLL–USP2 fusions. Leuk. 2019, 33, 2306–2340. [Google Scholar] [CrossRef] [PubMed]
- Meyer, C.; Schneider, B.; Reichel, M.; Angermueller, S.; Strehl, S.; Schnittger, S.; Schoch, C.; Jansen, M.W.J.C.; Van Dongen, J.; Pieters, R.; et al. Diagnostic tool for the identification of MLL rearrangements including unknown partner genes. Proc. Natl. Acad. Sci. USA 2005, 102, 449–454. [Google Scholar] [CrossRef] [Green Version]
- Lomov, N.; Zerkalenkova, E.; Lebedeva, S.; Viushkov, V.; Rubtsov, M.A. Cytogenetic and molecular genetic methods for chromosomal translocations detection with reference to the KMT2A/MLL gene. Crit. Rev. Clin. Lab. Sci. 2021, 58, 180–206. [Google Scholar] [CrossRef] [PubMed]
- Afrin, S.; Zhang, C.R.C.; Meyer, C.; Stinson, C.L.; Pham, T.; Bruxner, T.J.C.; Venn, N.C.; Trahair, T.N.; Sutton, R.; Marschalek, R.; et al. Targeted Next-Generation Sequencing for Detecting MLL Gene Fusions in Leukemia. Mol. Cancer Res. 2018, 16, 279–285. [Google Scholar] [CrossRef] [Green Version]
- Nijs, J.D.; Gonggrijp, H.; Augustinus, E.; Leeksma, C. Hot bands: A simple G-banding method for leukemic metaphases. Cancer Genet. Cytogenet. 1985, 15, 373–374. [Google Scholar] [CrossRef]
- ISCN. An International System for Human Cytogenomic Nomenclature; Karger Publishers: Basel, Switzerland, 2020; Volume 160, pp. 7–8. [Google Scholar]
- Jansen, M.W.J.C.; Van Der Velden, V.H.J.; Van Dongen, J.J.M. Efficient and easy detection of MLL-AF4, MLL-AF9 and MLL-ENL fusion gene transcripts by multiplex real-time quantitative RT-PCR in TaqMan and LightCycler. Leukemia 2005, 19, 2016–2018. [Google Scholar] [CrossRef] [Green Version]
- Burmeister, T.; Meyer, C.; Gröger, D.; Hofmann, J.; Marschalek, R. Evidence-based RT-PCR methods for the detection of the 8 most common MLL aberrations in acute leukemias. Leuk. Res. 2015, 39, 242–247. [Google Scholar] [CrossRef]
- Prasad, R.; Gu, Y.; Alder, H.; Nakamura, T.; Canaani, O.; Saito, H.; Huebner, K.; Gale, R.P.; Nowell, P.C.; Kuriyama, K. Cloning of the ALL-1 fusion partner, the AF-6 gene, involved in acute myeloid leukemias with the t(6;11) chromosome translocation. Cancer Res. 1993, 53, 5624–5628. [Google Scholar] [PubMed]
- Zerkalenkova, E.; Lebedeva, S.; Kazakova, A.; Tsaur, G.; Starichkova, Y.; Timofeeva, N.; Soldatkina, O.; Aprelova, E.; Popov, A.; Ponomareva, N.; et al. Acute myeloid leukemia with t(10;11)(p11-12;q23.3): Results of Russian Pediatric AML registration study. Int. J. Lab. Hematol. 2018, 41, 287–292. [Google Scholar] [CrossRef]
- Suzukawa, K.; Shimizu, S.; Nemoto, N.; Takei, N.; Taki, T.; Nagasawa, T. Identification of a Chromosomal Breakpoint and Detection of a Novel Form of an MLL-AF17 Fusion Transcript in Acute Monocytic Leukemia with t(11;17)(q23;q21). Int. J. Hematol. 2005, 82, 38–41. [Google Scholar] [CrossRef]
- Mitani, K.; Kanda, Y.; Ogawa, S.; Tanaka, T.; Inazawa, J.; Yazaki, Y.; Hirai, H. Cloning of several species of MLL/MEN chimeric cDNAs in myeloid leukemia with t(11;19)(q23;p13.1) translocation. Blood 1995, 85, 2017–2024. [Google Scholar] [CrossRef]
- Honigberg, L.A.; Smith, A.M.; Sirisawad, M.; Verner, E.; Loury, D.; Chang, B.; Li, S.; Pan, Z.; Thamm, D.; Miller, R.A.; et al. The Bruton tyrosine kinase inhibitor PCI-32765 blocks B-cell activation and is efficacious in models of autoimmune disease and B-cell malignancy. Proc. Natl. Acad. Sci. USA 2010, 107, 13075–13080. [Google Scholar] [CrossRef] [Green Version]
- Shinners, N.P.; Carlesso, G.; Castro, I.; Hoek, K.L.; Corn, R.A.; Woodland, R.L.; Scott, M.L.; Wang, D.; Khan, W.N. Bruton’s Tyrosine Kinase Mediates NF-κB Activation and B Cell Survival by B Cell-Activating Factor Receptor of the TNF-R Family. J. Immunol. 2007, 179, 3872–3880. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Conley, M.E.; Dobbs, K.; Farmer, D.M.; Kilic, S.; Paris, K.; Grigoriadou, S.; Coustan-Smith, E.; Howard, V.; Campana, D. Primary B Cell Immunodeficiencies: Comparisons and Contrasts. Annu. Rev. Immunol. 2009, 27, 199–227. [Google Scholar] [CrossRef] [PubMed]
- Tai, Y.-T.; Chang, B.Y.; Kong, S.-Y.; Fulciniti, M.; Yang, G.; Calle, Y.; Hu, Y.; Lin, J.; Zhao, J.-J.; Cagnetta, A.; et al. Bruton tyrosine kinase inhibition is a novel therapeutic strategy targeting tumor in the bone marrow microenvironment in multiple myeloma. Blood 2012, 120, 1877–1887. [Google Scholar] [CrossRef]
- Wang, M.L.; Rule, S.; Martin, P.; Goy, A.; Auer, R.; Kahl, B.S.; Jurczak, W.; Advani, R.; Romaguera, J.E.; Williams, M.E.; et al. Targeting BTK with Ibrutinib in Relapsed or Refractory Mantle-Cell Lymphoma. N. Engl. J. Med. 2013, 369, 507–516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woyach, J.A.; Bojnik, E.; Ruppert, A.S.; Stefanovski, M.R.; Goettl, V.M.; Smucker, K.A.; Smith, L.L.; Dubovsky, J.A.; Towns, W.H.; MacMurray, J.; et al. Bruton’s tyrosine kinase (BTK) function is important to the development and expansion of chronic lymphocytic leukemia (CLL). Blood 2014, 123, 1207–1213. [Google Scholar] [CrossRef] [PubMed]
- Rada, M.; Barlev, N.; Macip, S. BTK: A two-faced effector in cancer and tumour suppression. Cell Death Dis. 2018, 9, 1064. [Google Scholar] [CrossRef]
- Rushworth, S.; Murray, M.Y.; Zaitseva, L.; Bowles, K.; MacEwan, D.J. Identification of Bruton’s tyrosine kinase as a therapeutic target in acute myeloid leukemia. Blood 2014, 123, 1229–1238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmidt, U.; Boucheron, N.; Unger, B.; Ellmeier, W. The Role of Tec Family Kinases in Myeloid Cells. Int. Arch. Allergy Immunol. 2004, 134, 65–78. [Google Scholar] [CrossRef]
- Tomasson, M.H.; Xiang, Z.; Walgren, R.; Zhao, Y.; Kasai, Y.; Miner, T.; Ries, R.; Lubman, O.; Fremont, D.H.; McLellan, M.D.; et al. Somatic mutations and germline sequence variants in the expressed tyrosine kinase genes of patients with de novo acute myeloid leukemia. Blood 2008, 111, 4797–4808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zaitseva, L.; Murray, M.Y.; Shafat, M.S.; Lawes, M.J.; MacEwan, D.; Bowles, K.; Rushworth, S.A. Ibrutinib inhibits SDF1/CXCR4 mediated migration in AML. Oncotarget 2014, 5, 9930–9938. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nimmagadda, S.C.; Frey, S.; Edelmann, B.; Hellmich, C.; Zaitseva, L.; König, G.M.; Kostenis, E.; Bowles, K.M.; Fischer, T. Bruton’s tyrosine kinase and RAC1 promote cell survival in MLL-rearranged acute myeloid leukemia. Leuk. 2017, 32, 846–849. [Google Scholar] [CrossRef] [Green Version]
- Pillinger, G.; Abdul-Aziz, A.; Zaitseva, L.; Lawes, M.; MacEwan, D.; Bowles, K.M.; Rushworth, S.A. Targeting BTK for the treatment of FLT3-ITD mutated acute myeloid leukemia. Sci. Rep. 2015, 5, 12949. [Google Scholar] [CrossRef] [Green Version]
- Huang, S.; Pan, J.; Jin, J.; Li, C.; Li, X.; Huang, J.; Huang, X.; Yan, X.; Li, F.; Yu, M.; et al. Abivertinib, a novel BTK inhibitor: Anti-Leukemia effects and synergistic efficacy with homoharringtonine in acute myeloid leukemia. Cancer Lett. 2019, 461, 132–143. [Google Scholar] [CrossRef]
- Cortes, J.E.; Jonas, B.; Graef, T.; Luan, Y.; Stein, A.S. Clinical Experience With Ibrutinib Alone or in Combination With Either Cytarabine or Azacitidine in Patients With Acute Myeloid Leukemia. Clin. Lymphoma Myeloma Leuk. 2019, 19, 509–515.e1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, C.-H.; Mariño-Enriquez, A.; Ou, W.; Zhu, M.; Ali, R.; Chiang, S.; Amant, F.; Gilks, C.B.; Van De Rijn, M.; Oliva, E.; et al. The Clinicopathologic Features of YWHAE-FAM22 Endometrial Stromal Sarcomas. Am. J. Surg. Pathol. 2012, 36, 641–653. [Google Scholar] [CrossRef] [Green Version]
- Sugita, S.; Arai, Y.; Aoyama, T.; Asanuma, H.; Mukai, W.; Hama, N.; Emori, M.; Shibata, T.; Hasegawa, T. NUTM2A-CIC fusion small round cell sarcoma: A genetically distinct variant of CIC-rearranged sarcoma. Hum. Pathol. 2017, 65, 225–230. [Google Scholar] [CrossRef] [PubMed]
- Meyer, C.; Burmeister, T.; Strehl, S.; Schneider, B.; Hubert, D.; Zach, O.; Haas, O.; Klingebiel, T.; Dingermann, T.; Marschalek, R. Spliced MLL fusions: A novel mechanism to generate functional chimeric MLL-MLLT1 transcripts in t(11;19)(q23;p13.3) leukemia. Leukemia 2007, 21, 588–590. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.; Yan, C.; Hang, J.; Finci, L.I.; Lei, J.; Shi, Y. An Atomic Structure of the Human Spliceosome. Cell 2017, 169, 918–929.e14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bertram, K.; Agafonov, D.E.; Liu, W.-T.; Dybkov, O.; Will, C.L.; Hartmuth, K.; Urlaub, H.; Kastner, B.; Stark, H.; Lührmann, R. Cryo-EM structure of a human spliceosome activated for step 2 of splicing. Nat. Cell Biol. 2017, 542, 318–323. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Yan, C.; Zhan, X.; Li, L.; Lei, J.; Shi, Y. Structure of the human activated spliceosome in three conformational states. Cell Res. 2018, 28, 307–322. [Google Scholar] [CrossRef]
- Zhan, X.; Yan, C.; Zhang, X.; Lei, J.; Shi, Y. Structure of a human catalytic step I spliceosome. Science 2018, 359, 537–545. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fica, S.M.; Oubridge, C.; Wilkinson, M.E.; Newman, A.J.; Nagai, K. A human postcatalytic spliceosome structure reveals essential roles of metazoan factors for exon ligation. Science 2019, 363, 710–714. [Google Scholar] [CrossRef]
- David, C.J.; Boyne, A.R.; Millhouse, S.R.; Manley, J. The RNA polymerase II C-terminal domain promotes splicing activation through recruitment of a U2AF65-Prp19 complex. Genes Dev. 2011, 25, 972–983. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuraoka, I.; Ito, S.; Wada, T.; Hayashida, M.; Lee, L.; Saijo, M.; Nakatsu, Y.; Matsumoto, M.; Matsunaga, T.; Handa, H.; et al. Isolation of XAB2 Complex Involved in Pre-mRNA Splicing, Transcription, and Transcription-coupled Repair. J. Biol. Chem. 2008, 283, 940–950. [Google Scholar] [CrossRef] [Green Version]
- Maréchal, A.; Li, J.-M.; Ji, X.Y.; Wu, C.-S.; Yazinski, S.A.; Nguyen, D.; Liu, S.; Jiménez, A.E.; Jin, J.; Zou, L. PRP19 Transforms into a Sensor of RPA-ssDNA after DNA Damage and Drives ATR Activation via a Ubiquitin-Mediated Circuitry. Mol. Cell 2014, 53, 235–246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beck, B.D.; Park, S.-J.; Lee, Y.-J.; Roman, Y.; Hromas, R.A.; Lee, S.-H. Human Pso4 Is a Metnase (SETMAR)-binding Partner That Regulates Metnase Function in DNA Repair. J. Biol. Chem. 2008, 283, 9023–9030. [Google Scholar] [CrossRef] [Green Version]

| Rearrangement | Fusion Gene | Fusion Transcript |
|---|---|---|
| ins(X;11)(q26;q13q25)/KMT2A-BTK | MN687943 | MN238630 (exon 9-exon 2) |
| t(10;11)(q22;q23.3)/KMT2A-NUTM2A | MT721855 (forward) MT721856 (reciprocal) | MT721854 |
| inv(11)(q12.2q23.3)/KMT2A-PRPF19 | MZ443566 | MZ443565 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zerkalenkova, E.; Lebedeva, S.; Borkovskaia, A.; Soldatkina, O.; Plekhanova, O.; Tsaur, G.; Maschan, M.; Maschan, A.; Novichkova, G.; Olshanskaya, Y. BTK, NUTM2A, and PRPF19 Are Novel KMT2A Partner Genes in Childhood Acute Leukemia. Biomedicines 2021, 9, 924. https://doi.org/10.3390/biomedicines9080924
Zerkalenkova E, Lebedeva S, Borkovskaia A, Soldatkina O, Plekhanova O, Tsaur G, Maschan M, Maschan A, Novichkova G, Olshanskaya Y. BTK, NUTM2A, and PRPF19 Are Novel KMT2A Partner Genes in Childhood Acute Leukemia. Biomedicines. 2021; 9(8):924. https://doi.org/10.3390/biomedicines9080924
Chicago/Turabian StyleZerkalenkova, Elena, Svetlana Lebedeva, Aleksandra Borkovskaia, Olga Soldatkina, Olga Plekhanova, Grigory Tsaur, Michael Maschan, Aleksey Maschan, Galina Novichkova, and Yulia Olshanskaya. 2021. "BTK, NUTM2A, and PRPF19 Are Novel KMT2A Partner Genes in Childhood Acute Leukemia" Biomedicines 9, no. 8: 924. https://doi.org/10.3390/biomedicines9080924






