Mathematical Structure of RelB Dynamics in the NF-κB Non-Canonical Pathway
Abstract
1. Introduction
2. Materials and Methods
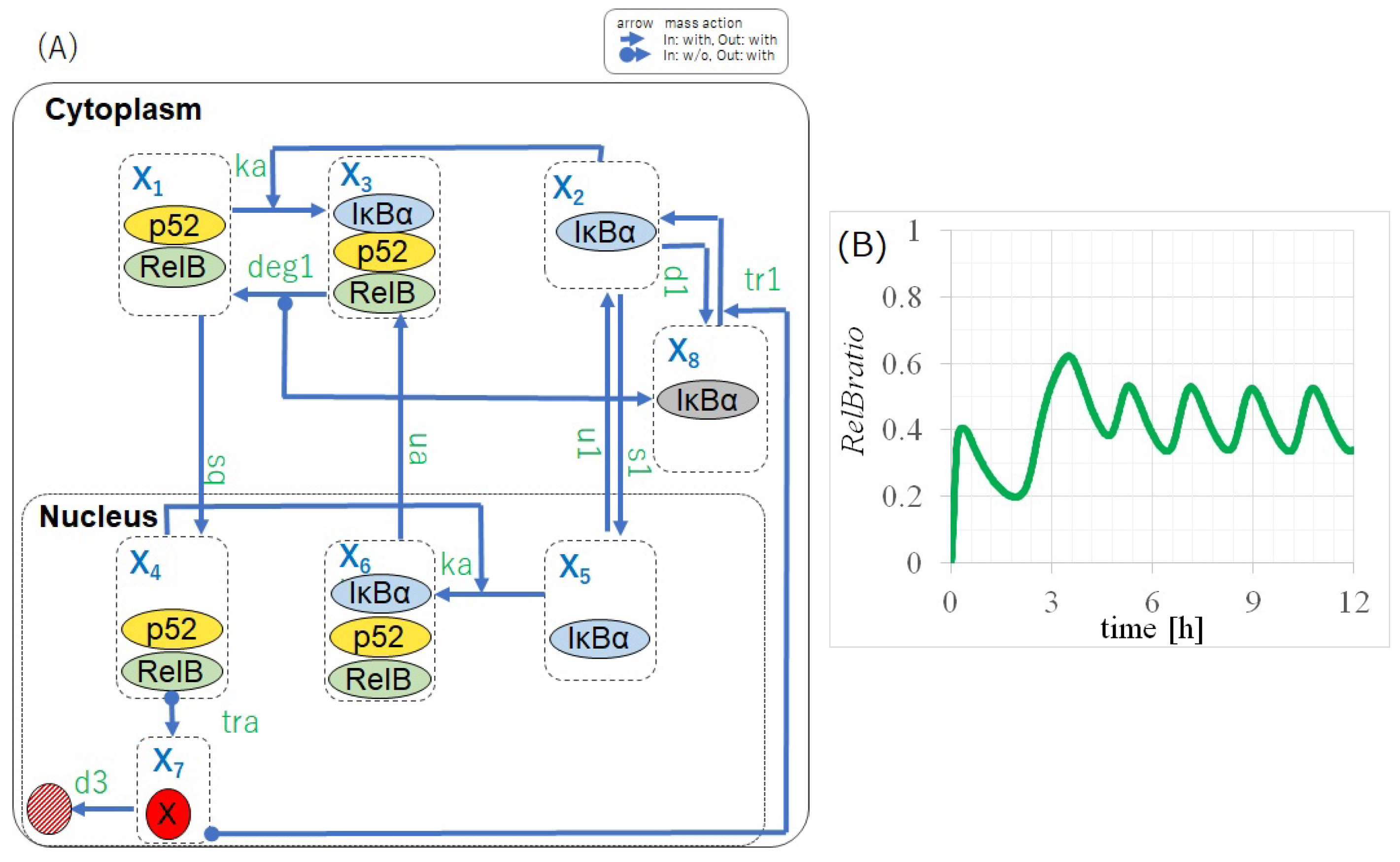
2.1. Negative Feedback from IB Is Essential for the Self-Sustained Oscillation of RelB
- (a) Certain elements activated via nuclear p50/RelB switched the activation state of IB (i.e., whether IB can bind to RelB).
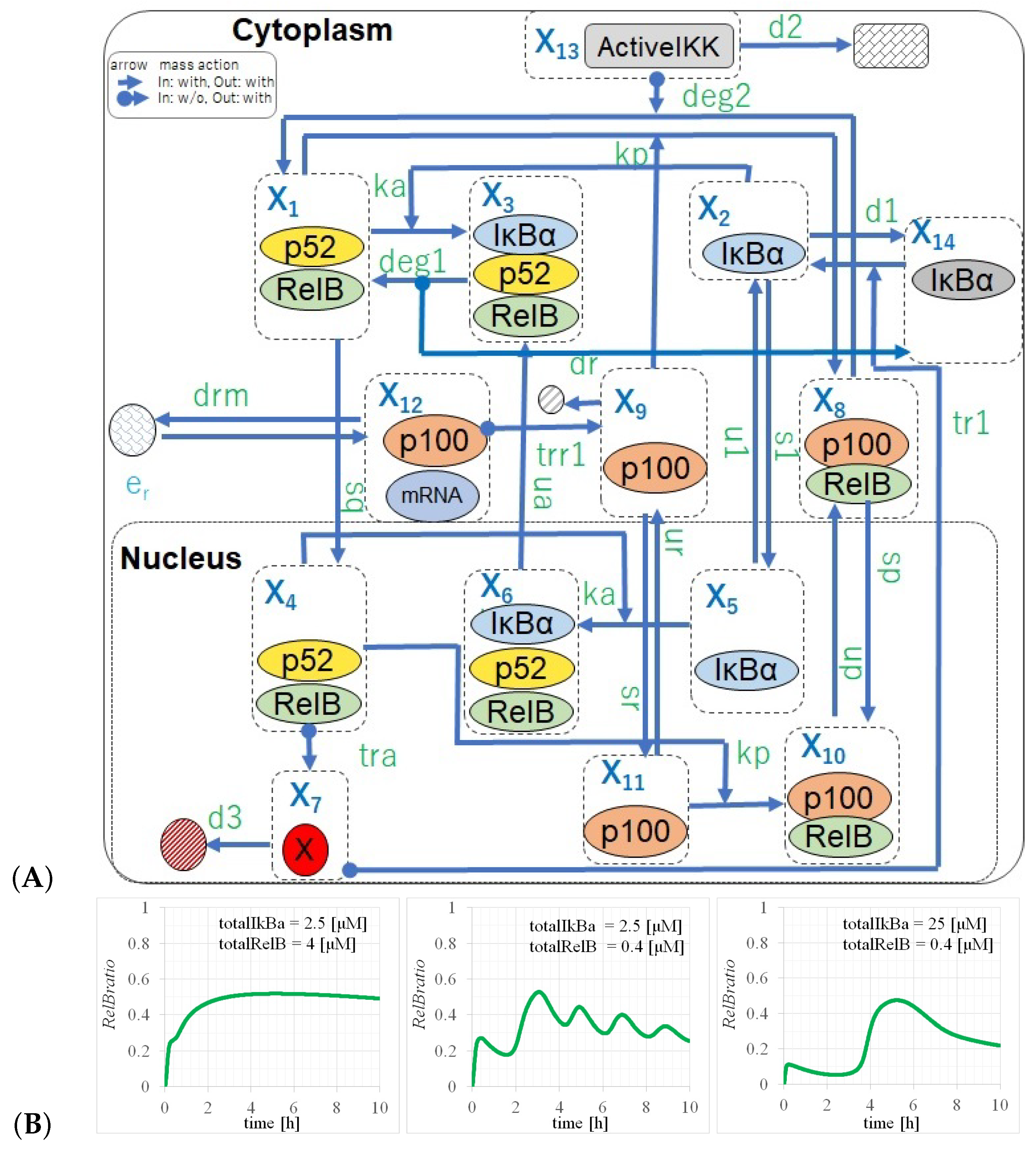
2.2. Full Non-Canonical Pathway Model
3. Results
3.1. Parameter Calibrations
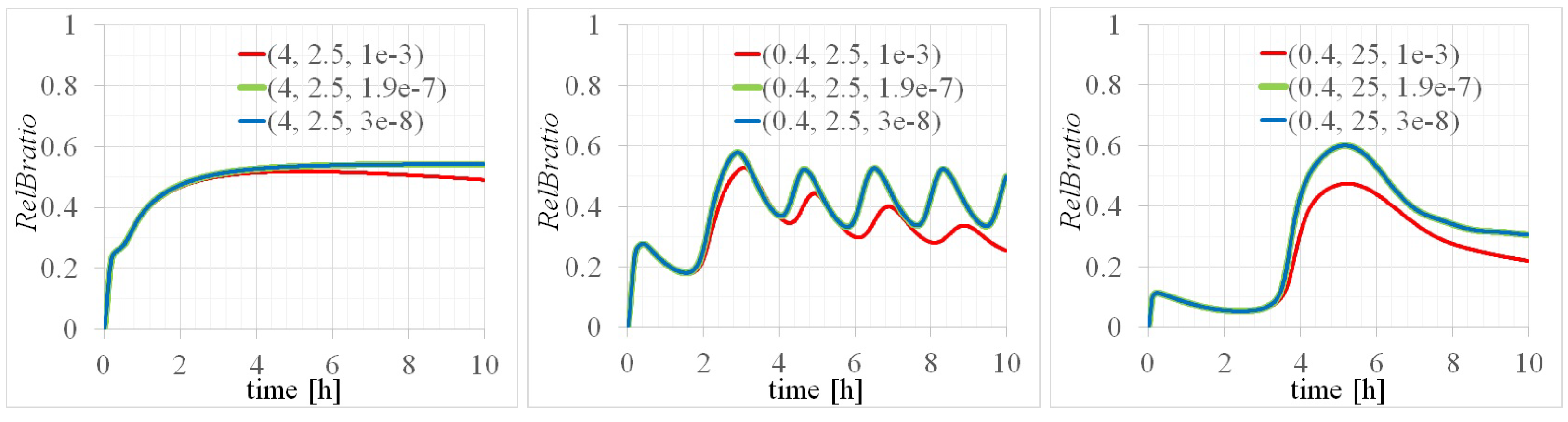
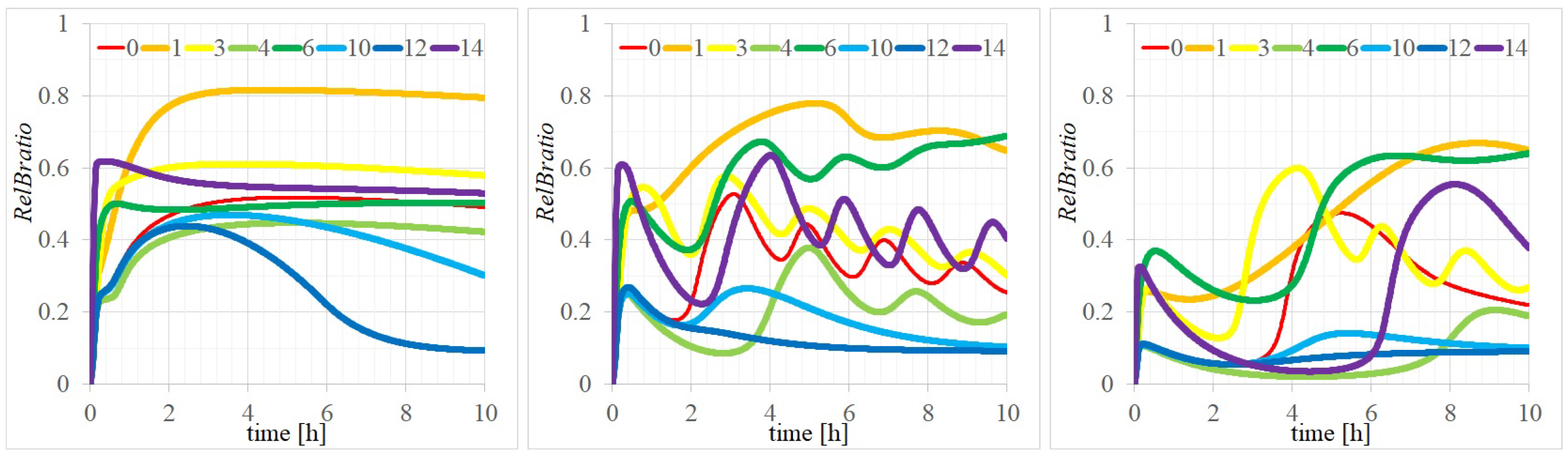
3.2. Three Types of RelB Dynamics Are Dependent on the Concentrations of Total and
4. Discussion
4.1. Dependency of RelB Dynamics on Concentrations of and
4.2. Quantitative Consideration of the Formation of
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
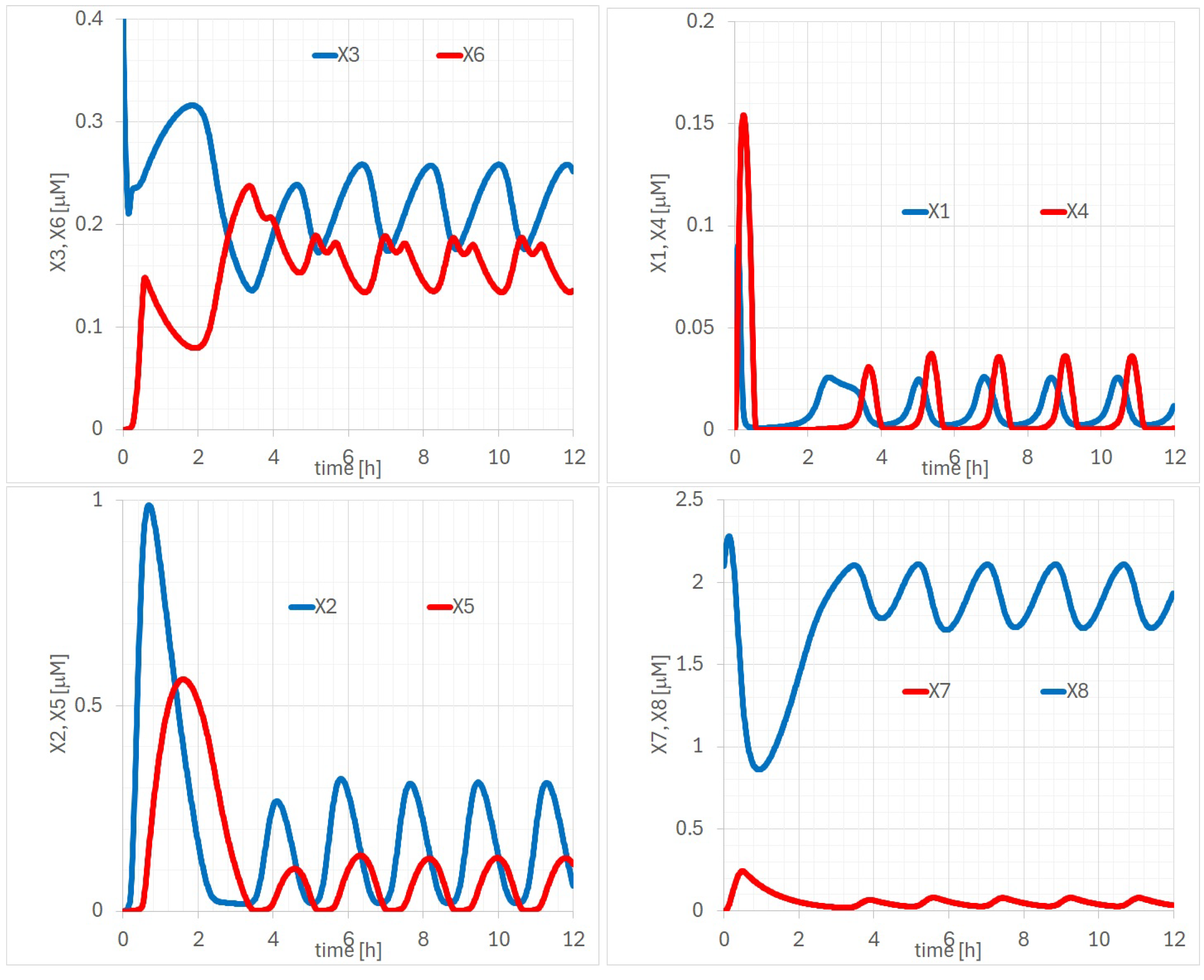
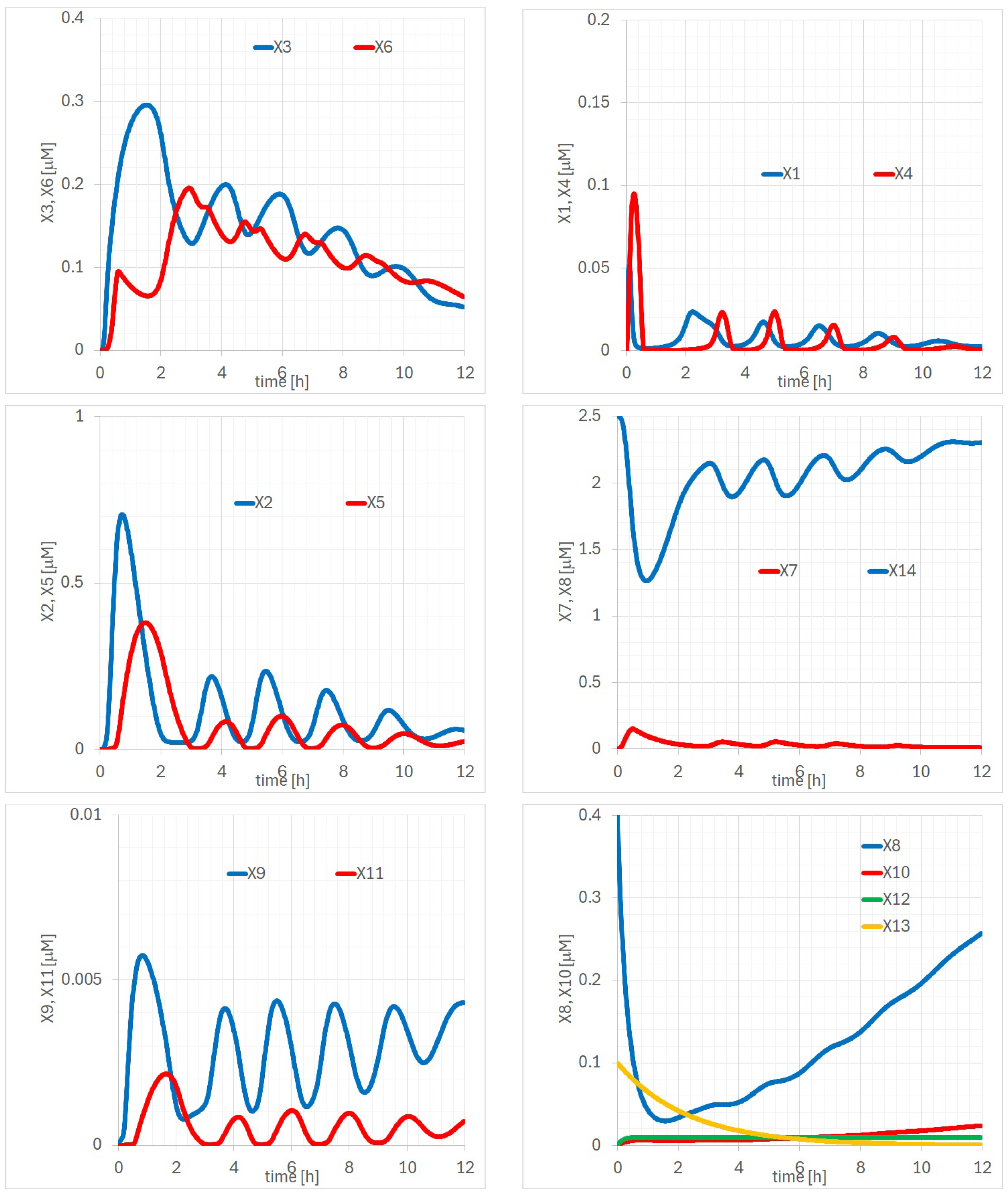
Appendix A. Detailed Analysis of Time Evolutions of with the 14 Parameters

Appendix B. Bifurcation Analysis
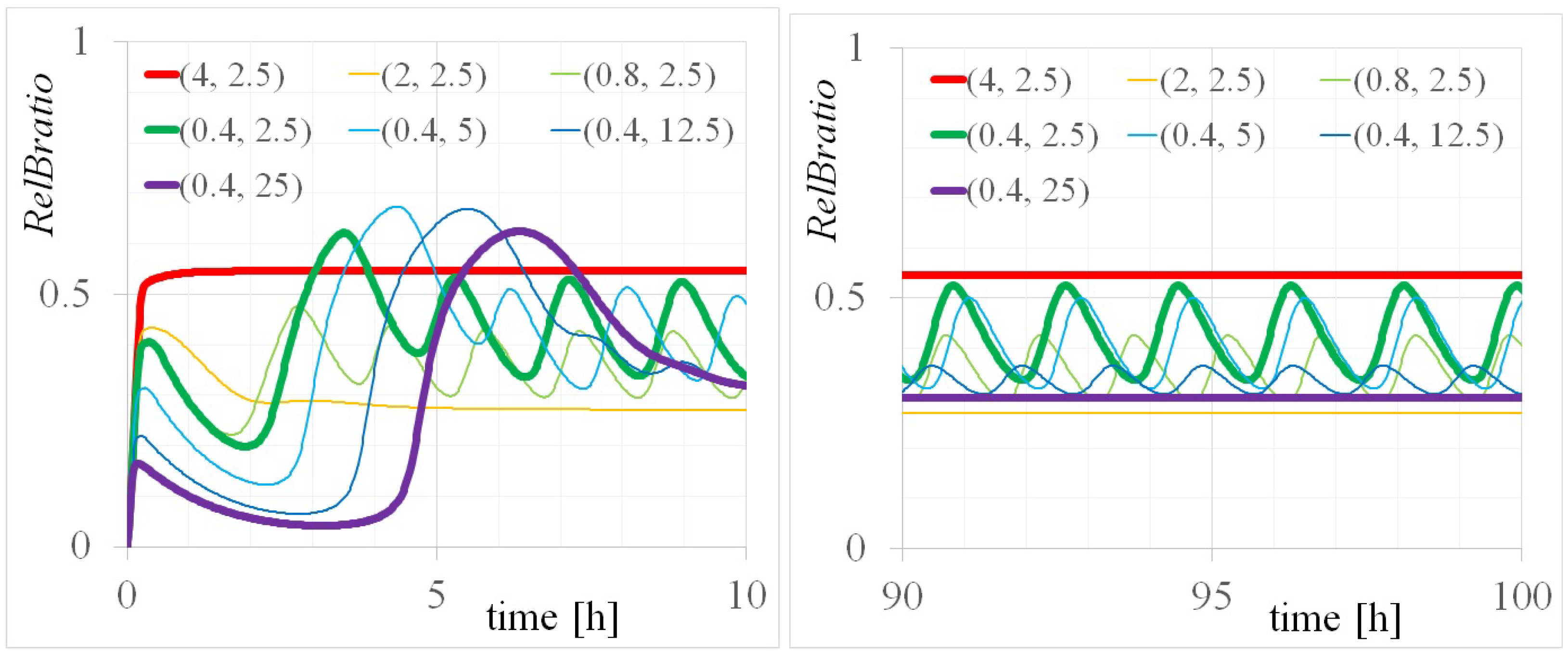
Appendix B.1. Algorithm for Finding the Steady State
Appendix B.2. Linearization and Eigenvalue Problems
- If all the eigenvalues of the Jacobian have negative real parts, then the equilibrium point () is asymptotically stable.
- If any eigenvalue of the Jacobian has a positive real part, then becomes unstable.
- When complex eigenvalues appear in the Jacobian, X approaches while oscillating.
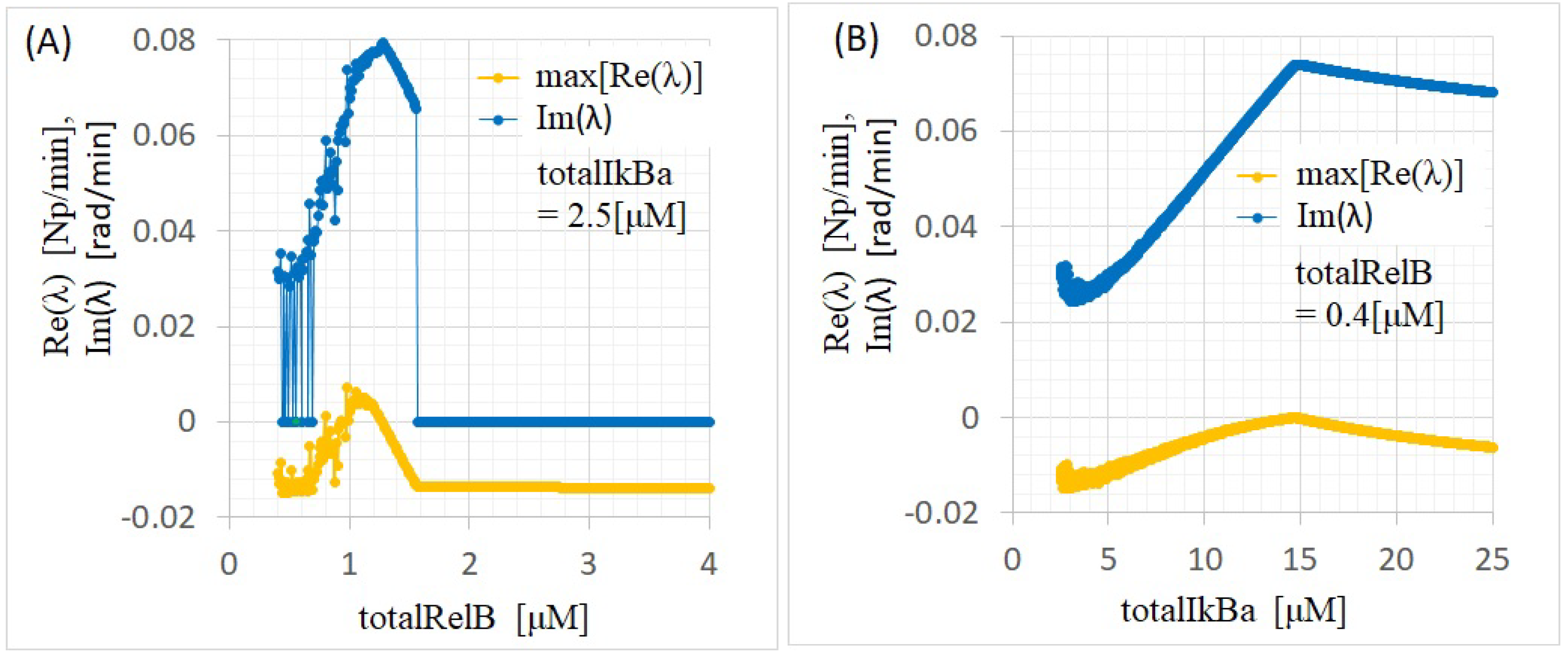
| totalRelB [M] | totalIkBa [M] | [rad/min] | Equivalent Period: 2 [h] |
|---|---|---|---|
| 0.8 | 2.5 | 0.0590 | 1.77 |
| 0.4 | 2.5 | 0.0316 | 3.32 |
| 0.4 | 5.0 | 0.0286 | 3.66 |
| 0.4 | 12.5 | 0.0636 | 1.65 |
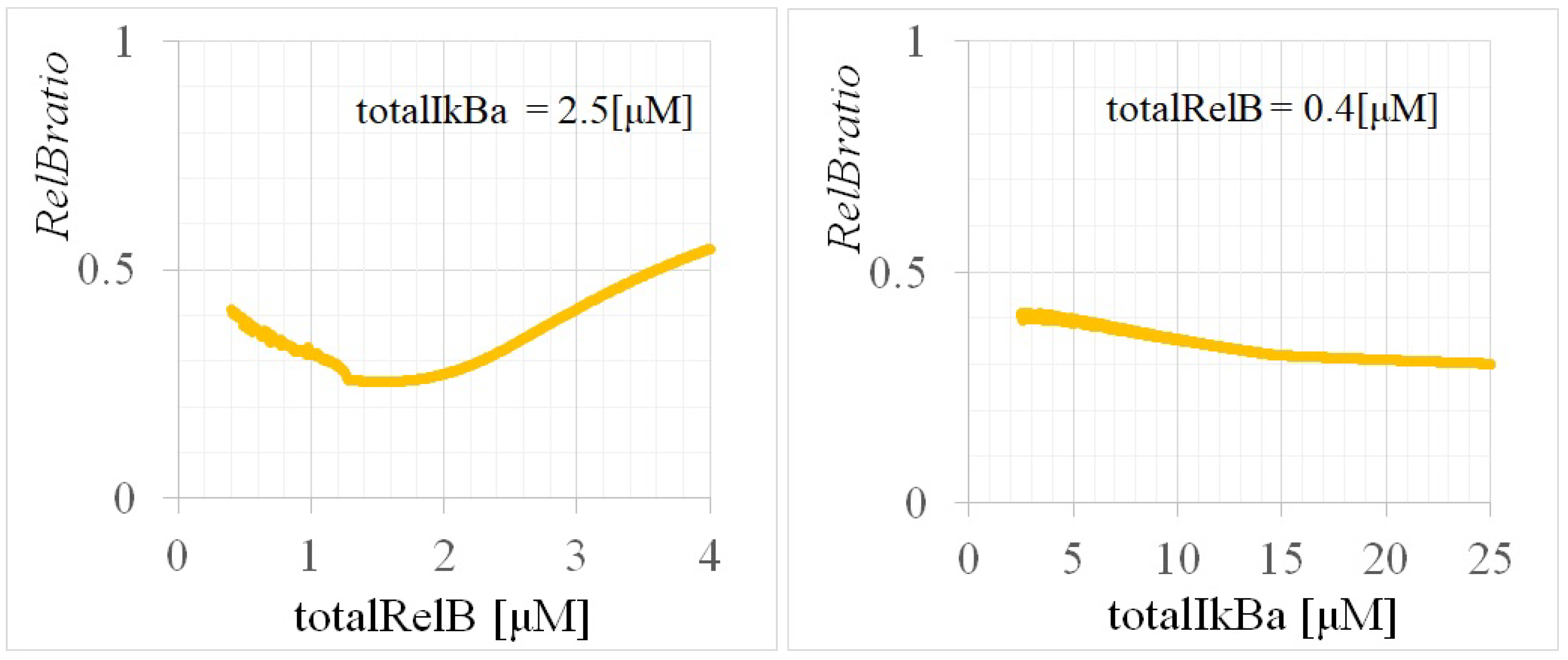
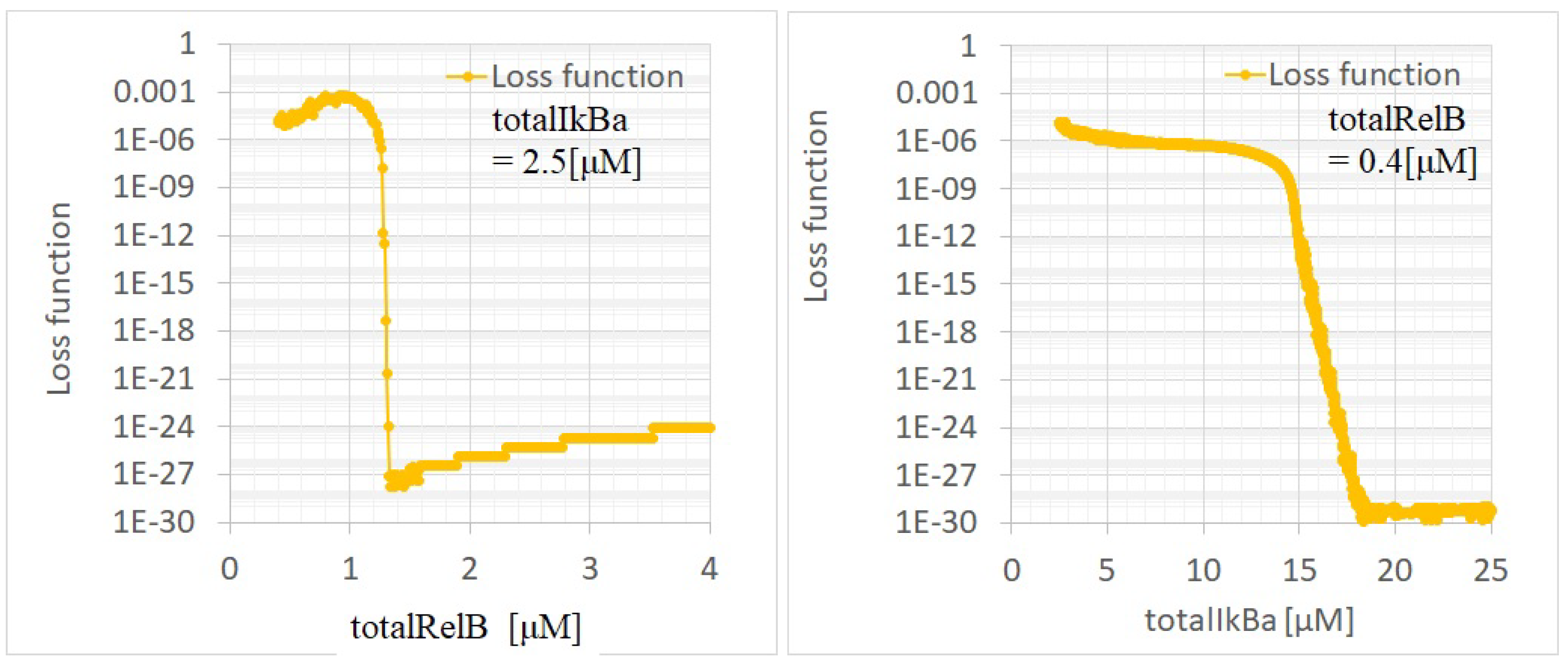
Appendix C. Experimental Methods and Data Details
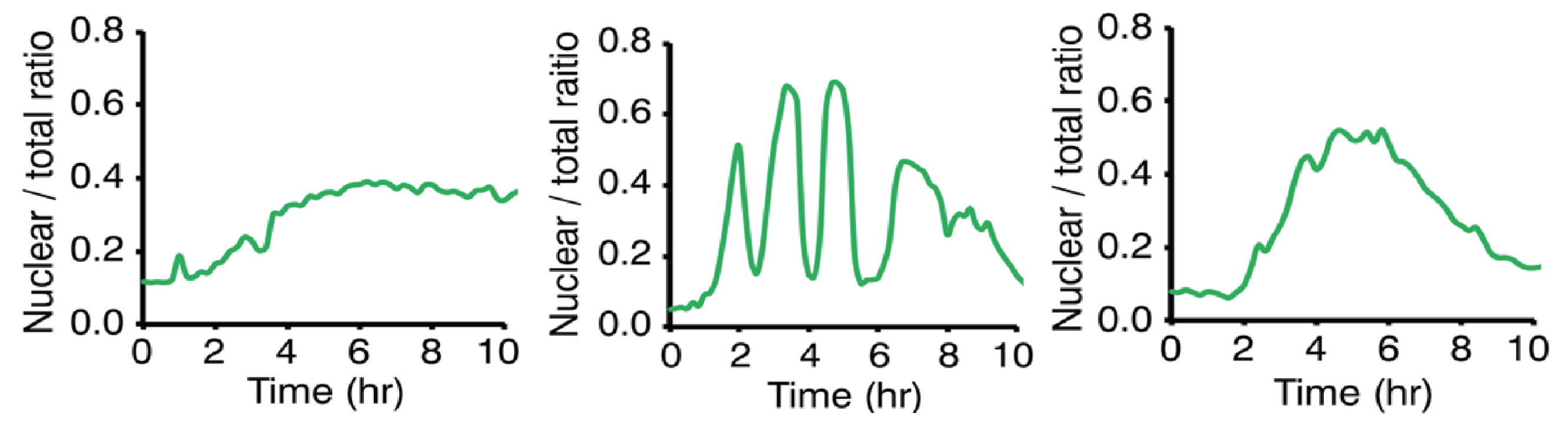
- Cell preparation: MEF (mouse embryonic fibroblast) cells stably expressing RelB-Venus were used. These cells were modified with a retroviral vector in which the nuclear localization signal (NLS) from the nuclear fluorescent protein EYFP-Nuc was fused to the 3’ end of mCherry [20].
- Antibody treatment: Cells were treated with 0.3 g/mL anti-LTR antibody to induce RelB nuclear translocation. This treatment activates the non-canonical NF-B pathway [20].
- Time-lapse imaging: Cells were observed using time-lapse imaging, and the fluorescence intensity in the nucleus and the entire cell was measured. These data show the nuclear fluorescence intensity of RelB, particularly the time evolution following stimulation with 0.3 g/mL anti-LTR antibody. The experiment observed patterns of RelB nuclear translocation and oscillation after stimulation [20].
Appendix C.1. Empirical Evidence for the Non-Canonical NF-κB Pathway
References
- Cheong, R.; Hoffmann, A.; Levchenko, A. Understanding NF-κB signaling via mathematical modeling. Mol. Syst. Biol. 2008, 4, 192. [Google Scholar] [CrossRef] [PubMed]
- Basak, S.; Behar, M.; Hoffmann, A. Lessons from mathematically modeling the NF-κB pathway. Immunol. Rev. 2012, 246, 221–238. [Google Scholar] [CrossRef] [PubMed]
- Shih, V.F.-S.; Tsui, R.; Caldwell, A.; Hoffmann, A. A single NFκB system for both canonical and non-canonical signaling. Cell Res. 2011, 21, 86–102. [Google Scholar] [CrossRef] [PubMed]
- Pomerantz, J.-L.; Baltimore, D. Two pathways to NF-κB. Mol. Cell 2002, 10, 693–695. [Google Scholar] [CrossRef]
- Oeckinghaus, A.; Ghosh, S. The NF-κB family of transcription factors and its regulation. Cold Spring Harb. Perspect. Biol. 2009, 1, a000034. [Google Scholar] [CrossRef] [PubMed]
- Hayden, M.S.; Ghosh, S. Shared principles in NF-κB signaling. Cell 2008, 132, 344–362. [Google Scholar] [CrossRef] [PubMed]
- Basak, S.; Shih, V.F.-S.; Hoffmann, A. Generation and activation of multiple dimeric transcription factors within the NF-κB signaling system. Mol. Cell. Biol. 2008, 28, 3139–3150. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, A.; Natoli, G.; Ghosh, S. Transcriptional regulation via the NF-κB signaling module. Oncogene 2006, 25, 6706–6716. [Google Scholar] [CrossRef]
- Mitchell, S.; Vargas, J.; Hoffmann, A. Signaling via the NFκB system. WIREs Syst. Biol. Med. 2016, 8, 227–241. [Google Scholar] [CrossRef] [PubMed]
- Iwai, K. Diverse ubiquitin signaling in NF-κB activation. Trends Cell Biol. 2012, 22, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Vallabhapurapu, S.; Matsuzawa, A.; Zhang, W.; Tseng, P.-H.; Keats, J.J.; Wang, H.; Vignali, D.A.; Bergsagel, P.L.; Karin, M. Nonredundant and complementary functions of TRAF2 and TRAF3 in a ubiquitination cascade that activates NIKdependent alternative NF-κB signaling. Nat. Immunol. 2008, 9, 1364. [Google Scholar] [CrossRef] [PubMed]
- Xiao, G.; Harhaj, E.W.; Sun, S.-C. NF-κB-inducing kinase regulates the processing of NF-κB2 p100. Mol. Cell 2001, 7, 401–409. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D.E.; Ihekwaba, A.E.C.; Elliott, M.; Johnson, J.R.; Gibney, C.A.; Foreman, B.E.; Nelson, G.; See, V.; Horton, C.A.; Spiller, D.G.; et al. Oscillations in NF-κB Signaling Control the Dynamics of Gene Expression. Science 2004, 306, 704–709. [Google Scholar] [CrossRef] [PubMed]
- Ashall, L.; Horton, C.A.; Nelson, D.E.; Paszek, P.; Harper, C.V.; Sillitoe, K.; Ryan, S.; Spiller, D.G.; Unitt, J.F.; Broomhead, D.S.; et al. Pulsatile Stimulation Determines Timing and Specificity of NF-κB–Dependent Transcription. Science 2009, 324, 242–247. [Google Scholar] [CrossRef] [PubMed]
- Hatanaka, N.; Seki, T.; Inoue, J.; Tero, A.; Suzuki, T. Critical roles of IκBα and RelA phosphorylation in transitional oscillation in NF-κB signaling module. J. Theor. Biol. 2019, 462, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, A.; Leung, T.H.; Baltimore, D. Genetic analysis of NF-κB/Rel transcription factors defines functional specificities. EMBO J. 2003, 22, 5530–5539. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, B.B. Signalling pathways of the TNF superfamily: A double-edged sword. Nat. Rev. Immunol. 2003, 9, 745–756. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.-C. Non-canonical NF-κB signaling pathway. Cell Res. 2011, 21, 71–85. [Google Scholar] [CrossRef] [PubMed]
- Seki, T.; Yamamoto, M.; Taguchi, Y.; Miyauchi, M.; Akiyama, N.; Yamaguchi, N.; Gohda, J.; Akiyama, T.; Inoue, J. Visualization of RelB expression and activation at the single-cell level during dendritic cell maturation in Relb-Venus knock-in mice. J. Biochem. 2015, 158, 485–495. [Google Scholar]
- Seki, T. Analysis of Spatiotemporal Control Mechanism of Transcription Factor NF-κB. Ph.D. Thesis, Tokyo University, Tokyo, Japan, 2015. [Google Scholar]
- Lee, D.; Jayaraman, A.; Kwon, J. Identification of cell-to-cell heterogeneity through systems engineering approaches. AIChE J. 2020, 66, e16925. [Google Scholar] [CrossRef]
- Lee, D.; Jayaraman, A.; Kwon, J. Development of a hybrid model for a partially known intracellular signaling pathway through correction term estimation and neural network modeling. PLOS Comp. Biol. 2020, 16, e1008472. [Google Scholar] [CrossRef] [PubMed]
- Shih, V.F.-S.; Davis-Turak, J.; Macal, M.; Huang, J.Q.; Ponomarenko, J. Control of RelB during dendritic cell activation integrates canonical and noncanonical NF-κB pathways. Nat. Immunol. 2012, 13, 1162–1172. [Google Scholar] [CrossRef] [PubMed]
- Banoth, B.; Chatterjee, B.; Vijayaragavan, B.; Prasad, M.V.R.; Roy, P.; Basak, S. Stimulus-selective crosstalk via the NF-κB signaling system reinforces innate immune response to alleviate gut infection. eLife 2015, 4, e05648. [Google Scholar] [CrossRef] [PubMed]
- Dobrzanski, P.; Ryseck, R.; Bravo, R. Differential interactions of Rel -NF-κB complexes with IκBα determine pools of constitutive and inducible NF-κB activity. EMBO J. 1994, 13, 4608–4616. [Google Scholar] [CrossRef] [PubMed]
- Almaden, V.; Tsui, R.; Liu, Y.C.; Birnbaum, H.; Shokhirev, M.N.; Ngo, K.A.; Davis-Turak, J.C.; Otero, D.; Basak, S.; Rickert, R.C.; et al. A Pathway Switch Directs BAFF Signaling to Distinct NFκB Transcription Factors in Maturing and Proliferating B Cells. Cell Rep. 2014, 9, 2098–2111. [Google Scholar] [CrossRef] [PubMed]
- Lewis, J.; Lakshmivarahan, S.; Dhall, S. Dynamic Data Assimilation: A Least Squares Approach; Cambridge University Press: Cambridge, UK, 2006. [Google Scholar]
- Meza, J. Steepest Descent; Technical Report in Lawrence Berkeley National Lab; Lawrence Berkeley National Laboratory: Berkeley, CA, USA, 2010. [Google Scholar]
- Vanderplaats, G.N. Numerical Optimization Techniques for Engineering Design: With Applications, 3rd ed.; Vanderplaats Research and Development, Inc.: Colorado Springs, CO, USA, 2001. [Google Scholar]
| Value | Molecule p100KO-M | NC-FM |
|---|---|---|
| p52/RelB | p52/RelB | |
| IB | IB | |
| IB/p52/RelB | IB/p52/RelB | |
| p52/RelBi | p52/RelBi | |
| IB | IB | |
| IB/p52/RelBi | IB/p52/RelBi | |
| unknown | unknown | |
| inactive IB | p100/RelB | |
| - | p100 | |
| - | p100/RelBi | |
| - | p100i | |
| - | p100 mRNA | |
| - | active IKK | |
| - | inactive IB |
| Pathway | Description | |
|---|---|---|
| (i) | p50/RelB → p50/RelBi | Cytoplasmic p50/RelB moves into the nucleus |
| (ii) | IB IB | IB shuttles between the nucleus and cytoplasm |
| (iii) | p50/RelB/IB → p50/RelB/IB | Nuclear p50/RelB exports to the cytoplasm by binding to IB |
| (iv) | p50/RelB/IB → p50/RelB | IB of p50/RelB/IB degrades in the cytoplasm |
| Symbol | Value | Unit | Description |
|---|---|---|---|
| 0.2 * | 1/min | p52/RelB nuclear import | |
| 30 * | 1/M min | p52/RelB association | |
| 0.00678 ‡ | 1/min | IB degradation | |
| 0.018 ‡ | 1/min | IB nuclear import | |
| 0.2448 ‡ | 1/M min | IB activation | |
| 0.012 ‡ | 1/min | IB nuclear export | |
| 0.1 * | M/min | p52/RelB-dependent (unknown factor) transcription | |
| 0.0168 ‡ | 1/min | IB mRNA degradation | |
| 0.012 * | 1/min | IB/p52/RelB nuclear export | |
| 0.1 * | 1/min | IB of IB/NF-B dissociation | |
| 50 * | 1/M min | p52/RelB replacement with p100/RelB | |
| 0.001 * | 1/min | p100/RelB nuclear import | |
| 0.01 * | 1/min | p100/RelB nuclear export | |
| 0.01 * | 1/min | p100 nuclear import | |
| 0.01 * | 1/min | p100 nuclear export | |
| 0.1 * | 1/min | p100 protein translation | |
| 0.1 * | 1/min | p100 degradation | |
| 0.1 * | 1/min | p100 mRNA degradation | |
| 0.001 * | M/min | p100 mRNA transcription | |
| 0.5 * | 1/M min | p100 processing | |
| 0.0072 ‡ | 1/min | IKK degradation | |
| 0.4 * | M | Total amount of RelB in a cell in the oscillation event | |
| 2.5 * | M | Total amount of IB in a cell in the oscillation event | |
| 0.1 * | M | Total amount of activeIKK in a cell |
| Symbol | PAV | EMV | Unit | Description |
|---|---|---|---|---|
| 0.2 * | 5.4 † | 1/min | p52/RelB nuclear import | |
| 30 * | 51 † | 1/M min | p52/RelB association | |
| 0.00678 ‡ | as left | 1/min | IB degradation | |
| 0.018 ‡ | as left | 1/min | IB nuclear import | |
| 0.2448 ‡ | as left | 1/M min | IB activation | |
| 0.012 ‡ | as left | 1/min | IB nuclear export | |
| 0.1 * | 0.01 * | M/min | p52/RelB-dependent (unknown factor) transcription | |
| 0.0168 ‡ | as left | 1/min | IB mRNA degradation | |
| 0.012 * | as left | 1/min | IB/p52/RelB nuclear export | |
| 0.1 * | 0.032 † | 1/min | IB of IB/NF-B dissociation | |
| 50 * | 0.13 † | 1/M min | p52/RelB replacement with p100/RelB | |
| 0.001 * | 0.028 † | 1/min | p100/RelB nuclear import | |
| 0.01 * | 0.42 † | 1/min | p100/RelB nuclear export | |
| 0.01 * | 0.045 † | 1/min | p100 nuclear import | |
| 0.01 * | 0.012 † | 1/min | p100 nuclear export | |
| 0.1 * | 0.5 # | 1/min | p100 protein translation | |
| 0.1 * | 0.4 # | 1/min | p100 degradation | |
| 0.1 * | 1.6 × # | 1/min | p100 mRNA degradation | |
| 0.001 * | 1.9 × # | M/min | p100 mRNA transcription | |
| 0.5 * | 4.2 # | 1/M min | p100 processing | |
| 0.0072 ‡ | as left | 1/min | IKK degradation | |
| 0.4 * | 0.08 $ | M | Total amount of RelB in a cell in the oscillation event | |
| 2.5 * | 0.6 * | M | Total amount of IB in a cell in the oscillation event | |
| 0.1 * | as left | M | Total amount of activeIKK in a cell |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Umegaki, T.; Hatanaka, N.; Suzuki, T. Mathematical Structure of RelB Dynamics in the NF-κB Non-Canonical Pathway. Math. Comput. Appl. 2024, 29, 62. https://doi.org/10.3390/mca29040062
Umegaki T, Hatanaka N, Suzuki T. Mathematical Structure of RelB Dynamics in the NF-κB Non-Canonical Pathway. Mathematical and Computational Applications. 2024; 29(4):62. https://doi.org/10.3390/mca29040062
Chicago/Turabian StyleUmegaki, Toshihito, Naoya Hatanaka, and Takashi Suzuki. 2024. "Mathematical Structure of RelB Dynamics in the NF-κB Non-Canonical Pathway" Mathematical and Computational Applications 29, no. 4: 62. https://doi.org/10.3390/mca29040062
APA StyleUmegaki, T., Hatanaka, N., & Suzuki, T. (2024). Mathematical Structure of RelB Dynamics in the NF-κB Non-Canonical Pathway. Mathematical and Computational Applications, 29(4), 62. https://doi.org/10.3390/mca29040062







