GSN-HVNET: A Lightweight, Multi-Task Deep Learning Framework for Nuclei Segmentation and Classification
Abstract
1. Introduction
- We propose a novel, lightweight, multi-task deep learning framework containing a unified model for segmentation and classification of nuclei instances simultaneously with superior efficiency and accuracy.
- We propose the newly designed RGS and DGS to improve accuracy and compress the training model.
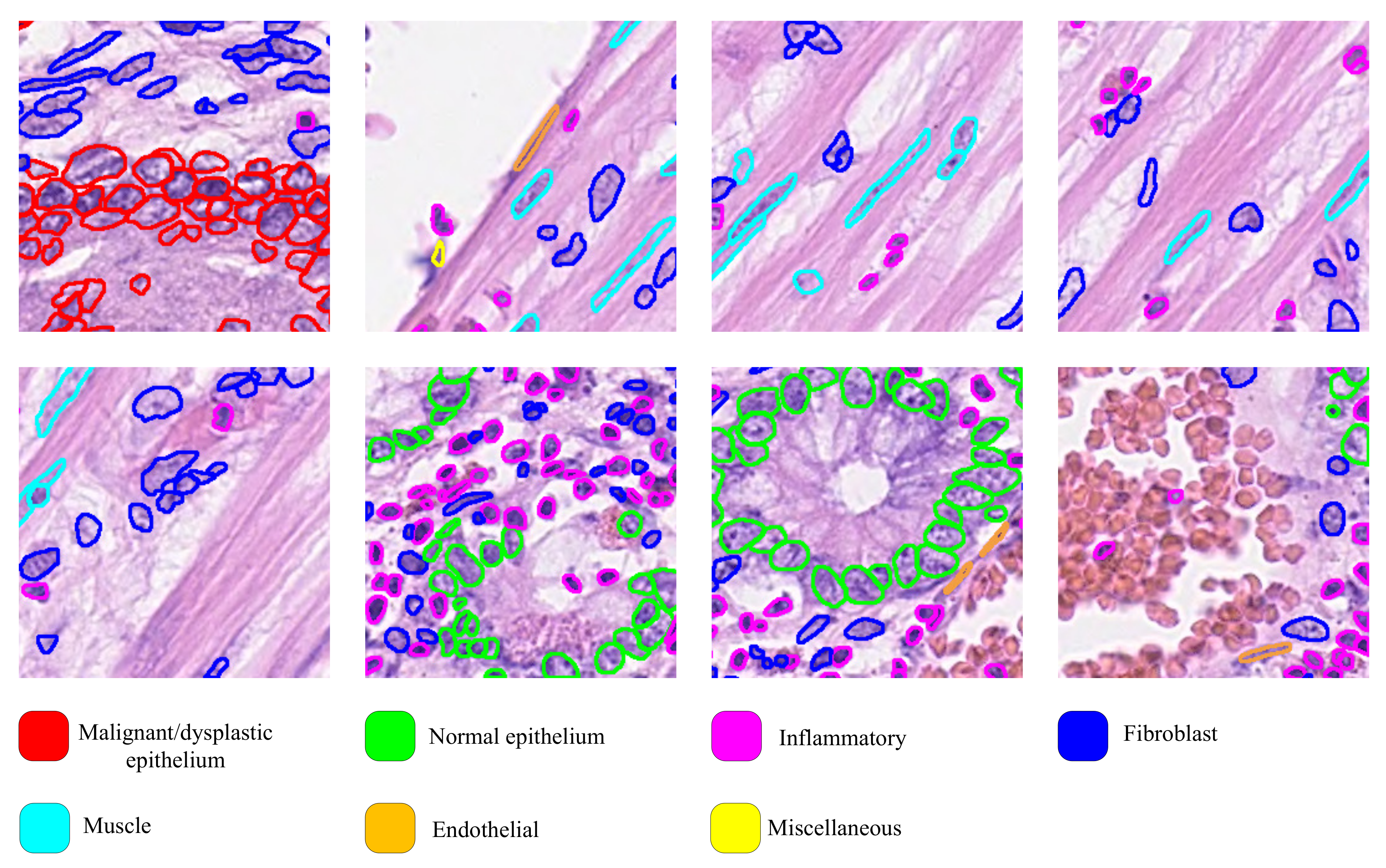
- We redefine the classification principle of the CoNSeP dataset so that the auxiliary diagnostic results have practical significance in pathological diagnosis.
- Our experiments on the CoNSeP, Kumar, and CPM-17 datasets confirm the improvements to existing works [13,14]. Compared with the state-of-the-art HoVer-Net [13], the number of parameters is reduced by 64%. In addition, we try different batch sizes in our experiments and prove that batch size is no longer a strict limitation on the proposed network; even when a small batch is presented, the proposed network can maintain a high performance.
2. Related Work
2.1. Nuclei Segmentation
2.2. Nuclei Classification
3. Proposed Method
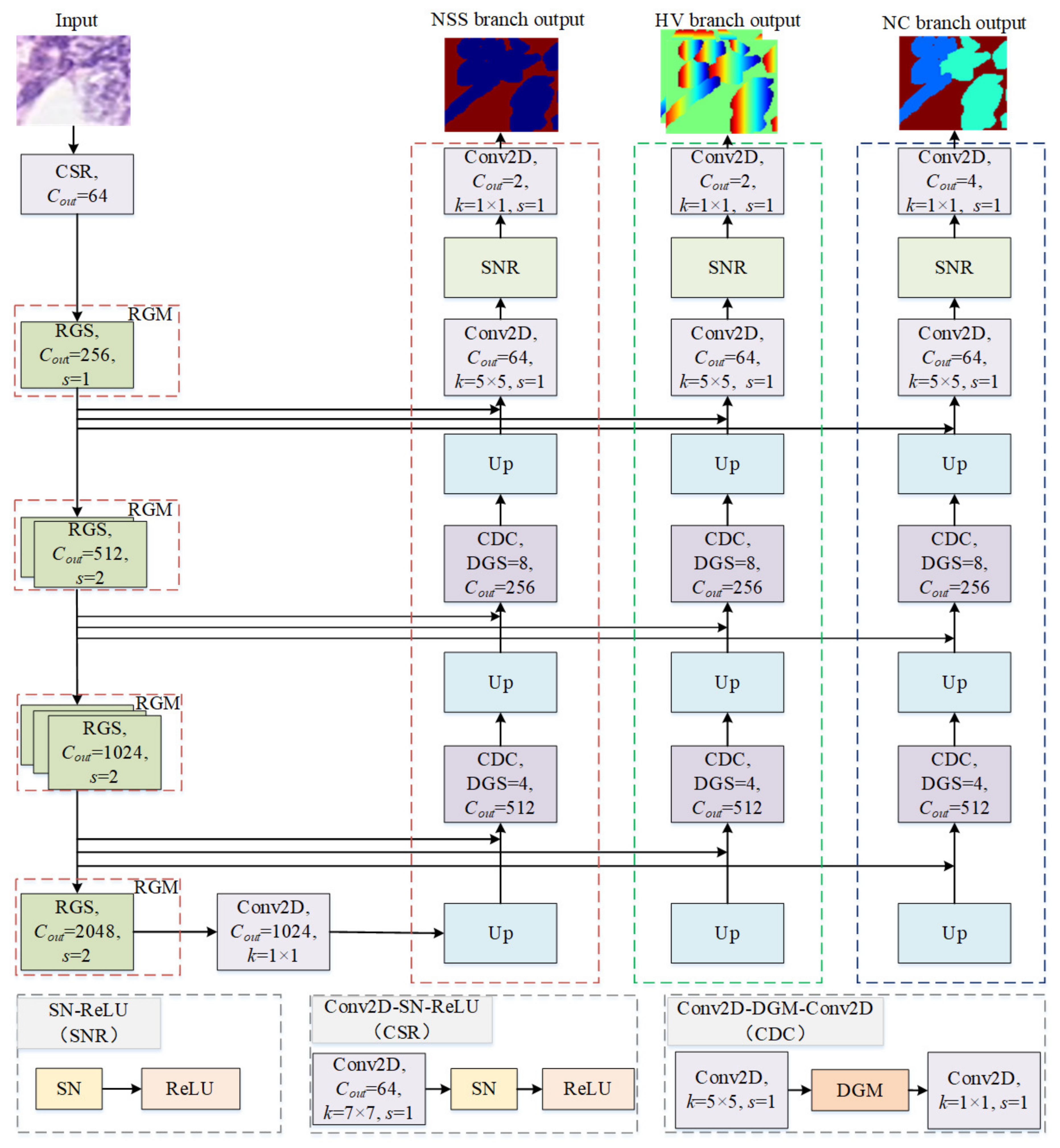
3.1. Network Architecture
3.1.1. Encoder
3.1.2. Ghost Block with Switchable Normalization
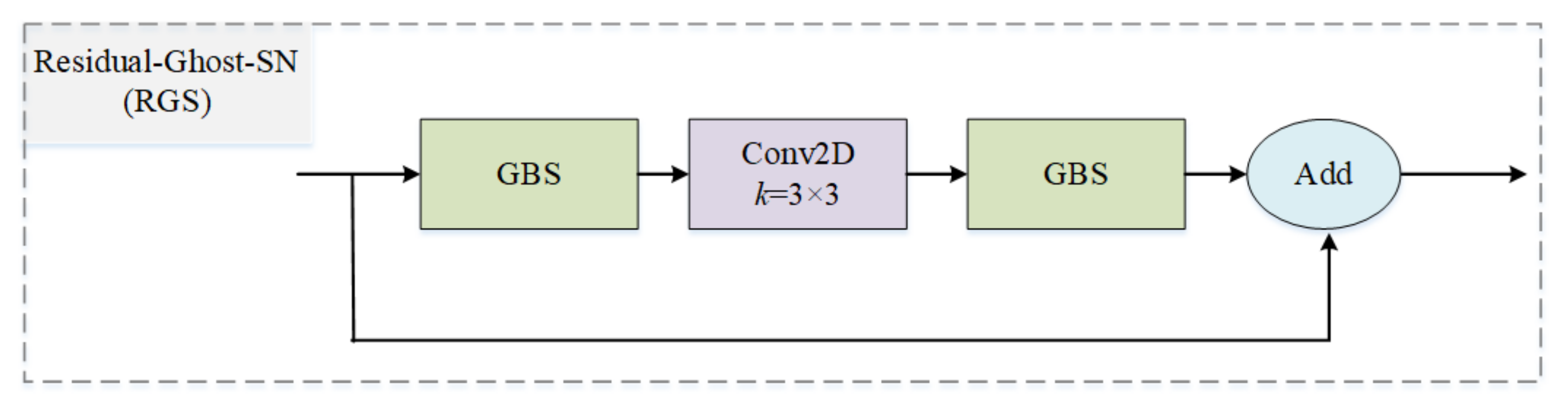
3.1.3. Residual Ghost Block with Switchable Normalization
3.1.4. Decoder
3.1.5. Dense Ghost Module with Switchable Normalization
3.1.6. Joint Loss Function of GSN-HVNET
3.2. Post-Processing
4. Experiment
4.1. Datasets and Implementation
4.2. Evaluation Metrics
4.2.1. Nuclei Instance Segmentation Evaluation
4.2.2. Nuclei Classification Evaluation
4.3. Experimental Results
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pantanowitz, L. Digital images and the future of digital pathology. J. Pathol. Inform. 2010, 1, 15. [Google Scholar] [CrossRef]
- Alsubaie, N.; Sirinukunwattana, K.; Raza, S.E.A.; Snead, D.; Rajpoot, N. A bottom-up approach for tumour differentiation in whole slide images of lung adenocarcinoma. In Medical Imaging 2018: Digital Pathology; Tomaszewski, J.E., Gurcan, M.N., Eds.; International Society for Optics and Photonics, SPIE: Houston, TX, USA, 2018; Volume 10581, p. 105810E. [Google Scholar] [CrossRef]
- Lu, C.; Romo-Bucheli, D.; Wang, X.; Janowczyk, A.; Ganesan, S.; Gilmore, H.; Rimm, D.; Madabhushi, A. Nuclear shape and orientation features from H&E images predict survival in early-stage estrogen receptor-positive breast cancers. Lab. Investig. 2018, 98, 1438–1448. [Google Scholar] [CrossRef]
- Javed, S.; Fraz, M.M.; Epstein, D.; Snead, D.; Rajpoot, N.M. Cellular community detection for tissue phenotyping in histology images. In Computational Pathology and Ophthalmic Medical Image Analysis; Springer: Berlin/Heidelberg, Germany, 2018; pp. 120–129. [Google Scholar] [CrossRef]
- Sirinukunwattana, K.; Snead, D.; Epstein, D.; Aftab, Z.; Mujeeb, I.; Tsang, Y.W.; Cree, I.; Rajpoot, N. Novel digital signatures of tissue phenotypes for predicting distant metastasis in colorectal cancer. Sci. Rep. 2018, 8, 13692. [Google Scholar] [CrossRef] [PubMed]
- Corredor, G.; Wang, X.; Zhou, Y.; Lu, C.; Fu, P.; Syrigos, K.; Rimm, D.L.; Yang, M.; Romero, E.; Schalper, K.A.; et al. Spatial architecture and arrangement of tumor-infiltrating lymphocytes for predicting likelihood of recurrence in early-stage non–small cell lung cancer. Clin. Cancer Res. 2019, 25, 1526–1534. [Google Scholar] [CrossRef]
- Sharma, H.; Zerbe, N.; Heim, D.; Wienert, S.; Behrens, H.M.; Hellwich, O.; Hufnagl, P. A multi-resolution approach for combining visual information using nuclei segmentation and classification in histopathological images. In Proceedings of the VISAPP, Berlin, Germany, 11–14 March 2015; Volume 3, pp. 37–46. [Google Scholar]
- Wang, P.; Hu, X.; Li, Y.; Liu, Q.; Zhu, X. Automatic cell nuclei segmentation and classification of breast cancer histopathology images. Signal Process. 2016, 122, 1–13. [Google Scholar] [CrossRef]
- Girshick, R. Fast r-cnn. In Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 7–13 December 2015; pp. 1440–1448. [Google Scholar] [CrossRef]
- Kendall, A.; Gal, Y.; Cipolla, R. Multi-task learning using uncertainty to weigh losses for scene geometry and semantics. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 7482–7491. [Google Scholar] [CrossRef]
- Zhou, Y.; Chen, H.; Li, Y.; Liu, Q.; Xu, X.; Wang, S.; Yap, P.T.; Shen, D. Multi-task learning for segmentation and classification of tumors in 3D automated breast ultrasound images. Med. Image Anal. 2021, 70, 101918. [Google Scholar] [CrossRef] [PubMed]
- Shan, P.; Wang, Y.; Fu, C.; Song, W.; Chen, J. Automatic skin lesion segmentation based on FC-DPN. Comput. Biol. Med. 2020, 123, 103762. [Google Scholar] [CrossRef]
- Graham, S.; Vu, Q.D.; Raza, S.E.A.; Azam, A.; Tsang, Y.W.; Kwak, J.T.; Rajpoot, N. Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med. Image Anal. 2019, 58, 101563. [Google Scholar] [CrossRef]
- Raza, S.E.A.; Cheung, L.; Shaban, M.; Graham, S.; Epstein, D.; Pelengaris, S.; Khan, M.; Rajpoot, N.M. Micro-Net: A unified model for segmentation of various objects in microscopy images. Med. Image Anal. 2019, 52, 160–173. [Google Scholar] [CrossRef]
- Abdulla, W. Mask R-CNN for Object Detection and Instance Segmentation on Keras and TensorFlow. 2017. Available online: https://github.com/matterport/Mask_RCNN (accessed on 31 January 2023).
- Song, W.; Zheng, Y.; Fu, C.; Shan, P. A novel batch image encryption algorithm using parallel computing. Inf. Sci. 2020, 518, 211–224. [Google Scholar] [CrossRef]
- Song, W.; Fu, C.; Zheng, Y.; Cao, L.; Tie, M. A practical medical image cryptosystem with parallel acceleration. J. Ambient. Intell. Humaniz. Comput. 2022, 1–15. [Google Scholar]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef]
- Song, W.; Fu, C.; Zheng, Y.; Cao, L.; Tie, M.; Sham, C.W. Protection of image ROI using chaos-based encryption and DCNN-based object detection. Neural Comput. Appl. 2022, 34, 5743–5756. [Google Scholar]
- Song, W.; Fu, C.; Zheng, Y.; Tie, M.; Liu, J.; Chen, J. A parallel image encryption algorithm using intra bitplane scrambling. Math. Comput. Simul. 2023, 204, 71–88. [Google Scholar] [CrossRef]
- Vincent, L.; Soille, P. Watersheds in digital spaces: An efficient algorithm based on immersion simulations. IEEE Comput. Archit. Lett. 1991, 13, 583–598. [Google Scholar] [CrossRef]
- Yang, X.; Li, H.; Zhou, X. Nuclei segmentation using marker-controlled watershed, tracking using mean-shift, and Kalman filter in time-lapse microscopy. IEEE Trans. Circuits Syst. Regul. Pap. 2006, 53, 2405–2414. [Google Scholar] [CrossRef]
- Mohammed, J.G.; Boudier, T. Classified region growing for 3D segmentation of packed nuclei. In Proceedings of the 2014 IEEE 11th International Symposium on Biomedical Imaging (ISBI), Beijing, China, 29 April–2 May 2014; pp. 842–845. [Google Scholar]
- Lu, Z.; Carneiro, G.; Bradley, A.P. An improved joint optimization of multiple level set functions for the segmentation of overlapping cervical cells. IEEE Trans. Image Process. 2015, 24, 1261–1272. [Google Scholar] [CrossRef]
- Nguyen, K.; Sarkar, A.; Jain, A.K. Prostate cancer grading: Use of graph cut and spatial arrangement of nuclei. IEEE Trans. Med. Imaging 2014, 33, 2254–2270. [Google Scholar] [CrossRef]
- Mitra, S.; Dey, S.; Das, N.; Chakrabarty, S.; Nasipuri, M.; Naskar, M.K. Identification of Benign and Malignant Cells from cytological images using superpixel based segmentation approach. In Proceedings of the Annual Convention of the Computer Society of India, Kolkata, India, 19–21 January 2018; Springer: Berlin/Heidelberg, Germany, 2018; pp. 257–269. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; Springer: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar] [CrossRef]
- Graham, S.; Rajpoot, N.M. SAMS-NET: Stain-aware multi-scale network for instance-based nuclei segmentation in histology images. In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; pp. 590–594. [Google Scholar] [CrossRef]
- Naylor, P.; Laé, M.; Reyal, F.; Walter, T. Segmentation of nuclei in histopathology images by deep regression of the distance map. IEEE Trans. Med. Imaging 2018, 38, 448–459. [Google Scholar] [CrossRef]
- Zhou, Y.; Onder, O.F.; Dou, Q.; Tsougenis, E.; Chen, H.; Heng, P.A. Cia-net: Robust nuclei instance segmentation with contour-aware information aggregation. In Proceedings of the International Conference on Information Processing in Medical Imaging, Hong Kong, China, 2–7 June 2019; Springer: Berlin/Heidelberg, Germany, 2019; pp. 682–693. [Google Scholar] [CrossRef]
- Jones, T.R.; Carpenter, A.E.; Lamprecht, M.R.; Moffat, J.; Silver, S.J.; Grenier, J.K.; Castoreno, A.B.; Eggert, U.S.; Root, D.E.; Golland, P.; et al. Scoring diverse cellular morphologies in image-based screens with iterative feedback and machine learning. Proc. Natl. Acad. Sci. USA 2009, 106, 1826–1831. [Google Scholar] [CrossRef]
- Yuan, Y.; Failmezger, H.; Rueda, O.M.; Ali, H.R.; Gräf, S.; Chin, S.F.; Schwarz, R.F.; Curtis, C.; Dunning, M.J.; Bardwell, H.; et al. Quantitative image analysis of cellular heterogeneity in breast tumors complements genomic profiling. Sci. Transl. Med. 2012, 4, 157ra143. [Google Scholar] [CrossRef] [PubMed]
- Sirinukunwattana, K.; Raza, S.E.A.; Tsang, Y.W.; Snead, D.R.; Cree, I.A.; Rajpoot, N.M. Locality sensitive deep learning for detection and classification of nuclei in routine colon cancer histology images. IEEE Trans. Med. Imaging 2016, 35, 1196–1206. [Google Scholar] [CrossRef]
- Luo, P.; Ren, J.; Peng, Z.; Zhang, R.; Li, J. Differentiable Learning-to-Normalize via Switchable Normalization. In Proceedings of the International Conference on Learning Representation (ICLR), Vancouver, BC, Canada, 30 April–3 May 2019. [Google Scholar]
- Han, K.; Wang, Y.; Tian, Q.; Guo, J.; Xu, C.; Xu, C. Ghostnet: More features from cheap operations. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Seattle, WA, USA, 13–19 June 2020; pp. 1580–1589. [Google Scholar] [CrossRef]
- Huang, X.; Belongie, S. Arbitrary style transfer in real-time with adaptive instance normalization. In Proceedings of the IEEE International Conference on Computer Vision, Venice, Italy, 22–29 October 2017; pp. 1501–1510. [Google Scholar] [CrossRef]
- Ba, J.L.; Kiros, J.R.; Hinton, G.E. Layer normalization. arXiv 2016, arXiv:1607.06450. [Google Scholar] [CrossRef]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the International Conference on Machine Learning, Lille, France, 7–9 July 2015; pp. 448–456. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar] [CrossRef]
- Kumar, N.; Verma, R.; Sharma, S.; Bhargava, S.; Vahadane, A.; Sethi, A. A dataset and a technique for generalized nuclear segmentation for computational pathology. IEEE Trans. Med. Imaging 2017, 36, 1550–1560. [Google Scholar] [CrossRef]
- Vu, Q.D.; Graham, S.; Kurc, T.; To, M.N.N.; Shaban, M.; Qaiser, T.; Koohbanani, N.A.; Khurram, S.A.; Kalpathy-Cramer, J.; Zhao, T.; et al. Methods for segmentation and classification of digital microscopy tissue images. Front. Bioeng. Biotechnol. 2019, 7, 53. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Delving deep into rectifiers: Surpassing human-level performance on imagenet classification. In Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 7–13 December 2015; pp. 1026–1034. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar] [CrossRef]
- Kirillov, A.; He, K.; Girshick, R.; Rother, C.; Dollár, P. Panoptic segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 15–20 June 2019; pp. 9404–9413. [Google Scholar] [CrossRef]
- Chen, H.; Qi, X.; Yu, L.; Heng, P.A. DCAN: Deep contour-aware networks for accurate gland segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 2487–2496. [Google Scholar] [CrossRef]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar] [CrossRef]





| Notation | Definition |
|---|---|
| The value of a pixel before normalization. | |
| , | The value of a pixel after normalization. |
| , | Scale and shift parameter |
| , | A set of pixels, and the number of pixels in . |
| L, | L denotes the loss function and represents its parameters. |
| I | The input image. |
| The HV distance of nuclei pixels to their mass centers. | |
| The regression output of HV branch. | |
| The pixel-wise and softmax predictions of NSS branch. | |
| The pixel-wise and softmax predictions of NC branch. | |
| E | The energy landspace. |
| The whole measurement for nuclei type classification and nuclei instance segmentation. | |
| False-positive, false-negative. | |
| True-positive, true-negative. |
| CoNSeP | CPM-17 | Kumar | |
|---|---|---|---|
| Total numbers of nuclei | 24,319 | 7570 | 21,623 |
| Labeled nuclei | 24,319 | 0 | 0 |
| Number of images | 41 | 32 | 30 |
| Origin | UHCW | TCGA | TCGA |
| Magnification | 40× | 40× & 20 × | 40× |
| Size of images | 1000 × 1000 | 500 × 500 to 600 × 600 | 1000 × 1000 |
| Seg/Class | Seg&Class | Seg | Seg |
| Number of cancer types | 1 | 4 | 8 |
| Method | Seg/Class | Parameters |
|---|---|---|
| HoVer-Net [13] | Seg | 42.94M |
| HoVer-Net [13] | Seg&Class | 52.20M |
| Micro-Net [14] | Seg&Class | 183.67M |
| DIST [30] | Seg&Class | 8.81M |
| DCAN [47] | Seg | 39.54M |
| SegNet [48] | Seg | 28.07M |
| FCN8 [49] | Seg | 128.05M |
| U-Net [28] | Seg&Class | 35.23M |
| Mask-RCNN [15] | Seg&Class | 44.17M |
| Our proposed | Seg | 15.03M |
| Our proposed | Seg&Class | 32.52M |
| Batch Size | Our Proposed | HoVer-Net | Micro-Net | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Dice | Dice | Dice | |||||||
| CoNSeP | Kumar | CPM-17 | CoNSeP | Kumar | CPM-17 | CoNSeP | Kumar | CPM-17 | |
| 1 | 0.821 | 0.851 | 0.865 | 0.816 | 0.794 | 0.843 | 0.752 | 0.759 | 0.828 |
| 2 | 0.830 | 0.844 | 0.870 | 0.806 | 0.804 | 0.875 | 0.764 | 0.785 | 0.857 |
| 3 | 0.839 | 0.842 | 0.870 | 0.835 | 0.819 | 0.879 | 0.758 | 0.794 | 0.859 |
| Method | CoNSeP | Kumar | CPM-17 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Dice | AJI | PQ | Dice | AJI | PQ | Dice | AJI | PQ | |
| HoVer-Net [13] | 0.838 | 0.525 | 0.494 | 0.826 | 0.618 | 0.597 | 0.869 | 0.705 | 0.697 |
| SegNet [48] | 0.796 | 0.194 | 0.270 | 0.811 | 0.377 | 0.407 | 0.857 | 0.491 | 0.531 |
| FCN8 [49] | 0.756 | 0.123 | 0.163 | 0.797 | 0.281 | 0.312 | 0.840 | 0.397 | 0.435 |
| U-Net [28] | 0.724 | 0.482 | 0.328 | 0.758 | 0.556 | 0.478 | 0.813 | 0.643 | 0.578 |
| DIST [30] | 0.798 | 0.495 | 0.386 | 0.789 | 0.559 | 0.443 | 0.826 | 0.616 | 0.504 |
| DCAN [47] | 0.733 | 0.289 | 0.256 | 0.792 | 0.525 | 0.492 | 0.828 | 0.561 | 0.545 |
| Micro-Net [14] | 0.784 | 0.518 | 0.421 | 0.797 | 0.560 | 0.519 | 0.857 | 0.668 | 0.661 |
| Mask-RCNN [15] | 0.740 | 0.474 | 0.460 | 0.760 | 0.546 | 0.509 | 0.850 | 0.684 | 0.674 |
| CIA-Net [31] | - | - | - | 0.818 | 0.620 | 0.577 | - | - | - |
| Our proposed | 0.861 | 0.602 | 0.566 | 0.879 | 0.635 | 0.644 | 0.899 | 0.701 | 0.683 |
| Method | ||||
|---|---|---|---|---|
| HoVer-Net [13] | 0.784 | 0.488 | 0.525 | 0.517 |
| Micro-Net [14] | 0.812 | 0.487 | 0.549 | 0.546 |
| DIST [30] | 0.782 | 0.489 | 0.569 | 0.526 |
| Mask-RCNN [15] | 0.701 | 0.413 | 0.568 | 0.514 |
| Our proposed | 0.820 | 0.514 | 0.572 | 0.519 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, T.; Fu, C.; Tian, Y.; Song, W.; Sham, C.-W. GSN-HVNET: A Lightweight, Multi-Task Deep Learning Framework for Nuclei Segmentation and Classification. Bioengineering 2023, 10, 393. https://doi.org/10.3390/bioengineering10030393
Zhao T, Fu C, Tian Y, Song W, Sham C-W. GSN-HVNET: A Lightweight, Multi-Task Deep Learning Framework for Nuclei Segmentation and Classification. Bioengineering. 2023; 10(3):393. https://doi.org/10.3390/bioengineering10030393
Chicago/Turabian StyleZhao, Tengfei, Chong Fu, Yunjia Tian, Wei Song, and Chiu-Wing Sham. 2023. "GSN-HVNET: A Lightweight, Multi-Task Deep Learning Framework for Nuclei Segmentation and Classification" Bioengineering 10, no. 3: 393. https://doi.org/10.3390/bioengineering10030393
APA StyleZhao, T., Fu, C., Tian, Y., Song, W., & Sham, C.-W. (2023). GSN-HVNET: A Lightweight, Multi-Task Deep Learning Framework for Nuclei Segmentation and Classification. Bioengineering, 10(3), 393. https://doi.org/10.3390/bioengineering10030393









