Datasets of Simulated Exhaled Aerosol Images from Normal and Diseased Lungs with Multi-Level Similarities for Neural Network Training/Testing and Continuous Learning
Abstract
:1. Summary
2. Data Description
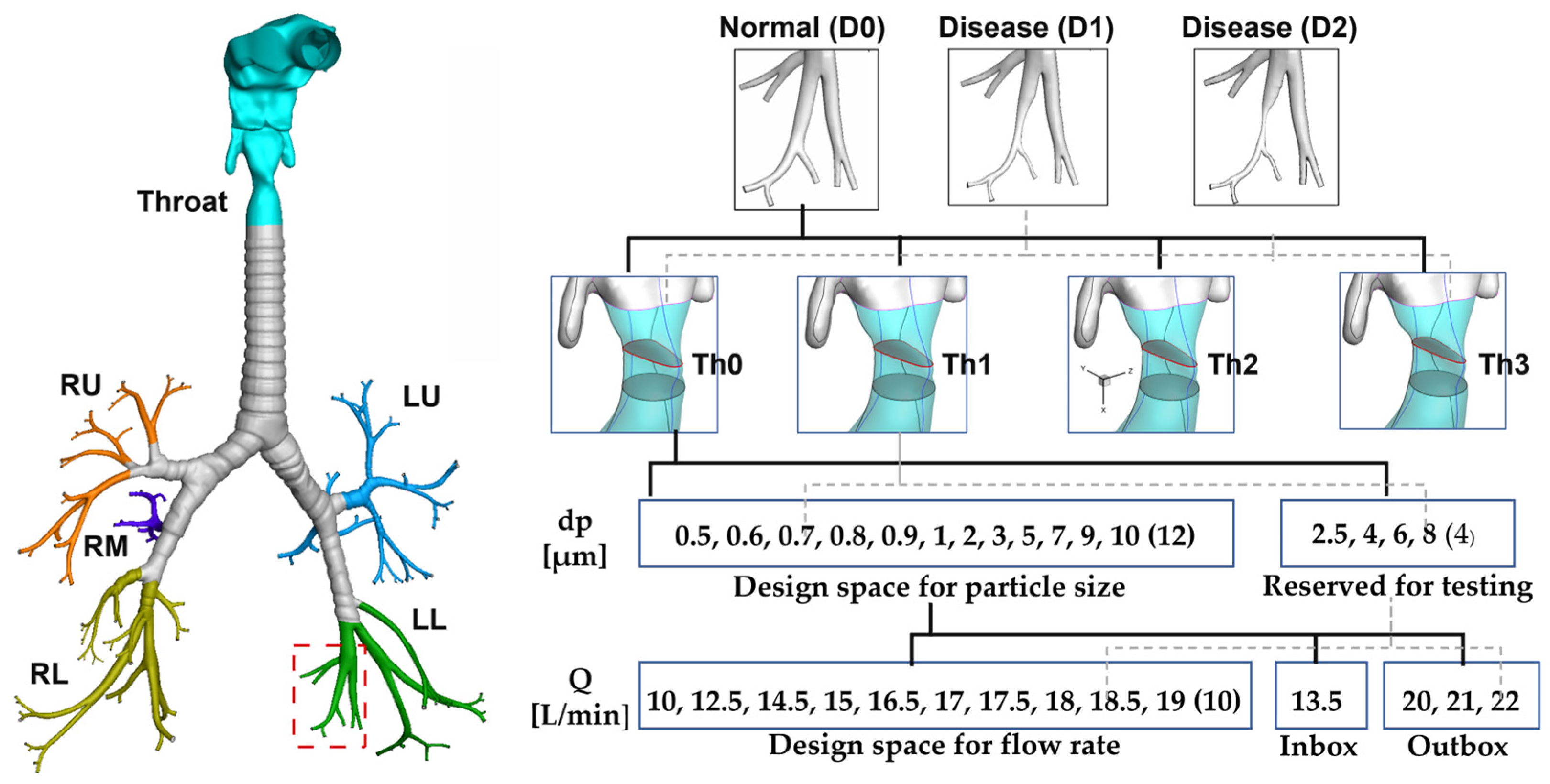
2.1. Data Achtiecture
2.2. Disease-Induced Disturbances to Particle Distributions
2.3. Results for Multi-Round Training and Multi-Level Testing
2.4. Learned Features from CNN Models
3. Methods
3.1. Normal and Diseased Lungs
3.2. Computational Mesh and Grid Independent Study
3.3. Boundary Conditions
3.4. Numerical Methods
4. User Notes
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hu, J.; Zhang, C.; Zhou, K.; Gao, S. Chest X-Ray diagnostic quality assessment: How much Is pixel-wise supervision needed? IEEE Trans. Med. Imaging 2022, 41, 1711–1723. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, R.M.; Atallah, R.P.; Struble, R.D.; Badgett, R.G. Lung cancer screening with low-dose CT: A meta-analysis. J. Gen. Intern. Med. 2020, 35, 3015–3025. [Google Scholar] [CrossRef] [PubMed]
- Tanino, A.; Kawamura, T.; Hamaguchi, M.; Tanino, R.; Nakao, M.; Hotta, T.; Tsubata, Y.; Hamaguchi, S.; Isobe, T. A novel model-based questionnaire based on low-dose CT screening data for chronic obstructive pulmonary disease diagnosis in Shimane, Japan. Int. J. Chron. Obstruct. Pulmon. Dis. 2021, 16, 1823–1833. [Google Scholar] [CrossRef] [PubMed]
- Duffy, S.W.; Field, J.K. Mortality reduction with low-dose CT screening for lung cancer. N. Engl. J. Med. 2020, 382, 572–573. [Google Scholar] [CrossRef] [PubMed]
- Das, N.; Topalovic, M.; Janssens, W. Artificial intelligence in diagnosis of obstructive lung disease: Current status and future potential. Curr. Opin. Pulm. Med. 2018, 24, 117–123. [Google Scholar] [CrossRef]
- Kanjanasurat, I.; Tenghongsakul, K.; Purahong, B.; Lasakul, A. CNN-RNN network integration for the diagnosis of COVID-19 using Chest X-ray and CT images. Sensors 2023, 23, 1356. [Google Scholar] [CrossRef]
- Heuvelmans, M.A.; van Ooijen, P.M.A.; Ather, S.; Silva, C.F.; Han, D.; Heussel, C.P.; Hickes, W.; Kauczor, H.U.; Novotny, P.; Peschl, H.; et al. Lung cancer prediction by Deep Learning to identify benign lung nodules. Lung Cancer 2021, 154, 1–4. [Google Scholar] [CrossRef]
- Bharati, S.; Podder, P.; Mondal, M.R.H. Hybrid deep learning for detecting lung diseases from X-ray images. Inform. Med. Unlocked 2020, 20, 100391. [Google Scholar] [CrossRef]
- Xi, J.; Si, X.A.; Kim, J.; Mckee, E.; Lin, E.-B. Exhaled aerosol pattern discloses lung structural abnormality: A sensitivity study using computational modeling and fractal analysis. PLoS ONE 2014, 9, e104682. [Google Scholar] [CrossRef] [Green Version]
- Cavaleiro Rufo, J.; Paciência, I.; Mendes, F.C.; Farraia, M.; Rodolfo, A.; Silva, D.; de Oliveira Fernandes, E.; Delgado, L.; Moreira, A. Exhaled breath condensate volatilome allows sensitive diagnosis of persistent asthma. Allergy 2019, 74, 527–534. [Google Scholar] [CrossRef]
- Talaat, M.; Xi, J.; Tan, K.; Si, X.A.; Xi, J. Convolutional neural network classification of exhaled aerosol images for diagnosis of obstructive respiratory diseases. J. Nanotheranostics 2023, 4, 228–247. [Google Scholar] [CrossRef]
- Talaat, M.; Si, X.A.; Xi, J. CFD Lung Diagnosis|Kaggle. Available online: https://www.kaggle.com/datasets/mohamedtalaat92/cfd-lung-diagnosis. (accessed on 21 July 2023).
- Lu, J.; Xi, J.; Langenderfer, J.E. Sensitivity analysis and uncertainty quantification in pulmonary drug delivery of orally inhaled pharmaceuticals. J. Pharm. Sci. 2017, 106, 3303–3315. [Google Scholar] [CrossRef]
- Force, U.P.S.T. Screening for lung cancer: US preventive services task force recommendation statement. JAMA 2021, 325, 962–970. [Google Scholar]
- Henschke, C.; McCauley, D.I.; Yankelevitz, D.F. Early lung cancer action project: Overall design and findings from baseline screening. Lancet 1999, 354, 99–105. [Google Scholar] [CrossRef]
- Sone, S.; Li, F.; Yang, Z.G. Results of three-year mass screening programme for lung cancer using low-dose spiral computed tomography scanner. Br. J. Cancer 2001, 84, 25–32. [Google Scholar] [CrossRef]
- Swensen, S. Screening for cancer with computed tomography. BMJ 2003, 326, 894–895. [Google Scholar] [CrossRef]
- Gao, Y.; Cui, Y. Deep transfer learning for reducing health care disparities arising from biomedical data inequality. Nat. Commun. 2020, 11, 5131. [Google Scholar] [CrossRef]
- Ayana, G.; Dese, K.; Choe, S.W. Transfer learning in breast cancer diagnoses via ultrasound imaging. Cancers 2021, 13, 738. [Google Scholar] [CrossRef]
- Valverde, J.M.; Imani, V.; Abdollahzadeh, A.; De Feo, R.; Prakash, M.; Ciszek, R.; Tohka, J. Transfer learning in magnetic resonance brain imaging: A systematic review. J. Imaging 2021, 7, 66. [Google Scholar] [CrossRef]
- Loey, M.; Manogaran, G.; Taha, M.H.N.; Khalifa, N.E.M. Fighting against COVID-19: A novel deep learning model based on YOLO-v2 with ResNet-50 for medical face mask detection. Sustain. Cities Soc. 2021, 65, 102600. [Google Scholar] [CrossRef]
- Benali Amjoud, A.; Amrouch, M. Convolutional neural networks backbones for object detection. In Image and Signal Processing; Springer: Cham, Switzerland, 2020; Volume 12119. [Google Scholar]
- Tan, M.; Le, Q. EfficientNet: Rethinking model scaling for convolutional neural networks. In Proceedings of the 36th ICML, Long Beach, CA, USA, 9–15 June 2019; pp. 6105–6114. [Google Scholar]
- Tan, M.; Le, Q. Efficientnetv2: Smaller models and faster training. In Proceedings of the 38th ICML, Online, 18–24 July 2021; pp. 10096–10106. [Google Scholar]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv 2017, arXiv:1704.04861. [Google Scholar]
- Sandler, M.; Howard, A.; Zhu, M.; Zhmoginov, A.; Chen, L.-C. Mobilenetv2: Inverted residuals and linear bottlenecks. In Proceedings of the IEEE CVPR, Salt Lake City, UT, USA, 18–23 June 2018; pp. 4510–4520. [Google Scholar]
- Si, X.A.; Xi, J. Deciphering exhaled aerosol fingerprints for early diagnosis and personalized therapeutics of obstructive respiratory diseases in small airways. J. Nanotheranostics 2021, 2, 94–117. [Google Scholar] [CrossRef]
- Xi, J.; Zhao, W. Correlating exhaled aerosol images to small airway obstructive diseases: A study with dynamic mode decomposition and machine learning. PLoS ONE 2019, 14, e0211413. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolutional neural networks. Commun. ACM 2017, 60, 84–90. [Google Scholar] [CrossRef] [Green Version]
- Xi, J.; Zhao, W.; Yuan, J.E.; Cao, B.; Zhao, L. Multi-resolution classification of exhaled aerosol images to detect obstructive lung diseases in small airways. Comput. Biol. Med. 2017, 87, 57–69. [Google Scholar] [CrossRef]
- Xi, J.; Wang, Z.; Talaat, K.; Glide-Hurst, C.; Dong, H. Numerical study of dynamic glottis and tidal breathing on respiratory sounds in a human upper airway model. Sleep Breath. 2018, 22, 463–479. [Google Scholar] [CrossRef] [Green Version]
- Talaat, M.; Si, X.A.; Dong, H.; Xi, J. Leveraging statistical shape modeling in computational respiratory dynamics: Nanomedicine delivery in remodeled airways. Comput. Methods Programs Biomed. 2021, 204, 106079. [Google Scholar] [CrossRef]
- Xi, J.; Talaat, M.; Si, X.A.; Chandra, S. The application of statistical shape modeling for lung morphology in aerosol inhalation dosimetry. J. Aerosol Sci. 2021, 151, 105623. [Google Scholar] [CrossRef]
- Xiao, Q.; Stewart, N.J.; Willmering, M.M.; Gunatilaka, C.C.; Thomen, R.P.; Schuh, A.; Krishnamoorthy, G.; Wang, H.; Amin, R.S.; Dumoulin, C.L.; et al. Human upper-airway respiratory airflow: In vivo comparison of computational fluid dynamics simulations and hyperpolarized 129Xe phase contrast MRI velocimetry. PLoS ONE 2021, 16, e0256460. [Google Scholar] [CrossRef]







| Training | Testing | |||
|---|---|---|---|---|
| Level 1 | Level 2 | Level 3 | ||
| Round 1 | 90% Base | 10% Base | Inbox | Outbox |
| Round 2: (plus 90% Base) | Th1, Th2, Th3 | 10% Base | Inbox | Outbox |
| Round 3 (plus 90% Base, and Th1, Th2, Th3) | 25% Outbox | 10% Base | Inbox | Outbox |
| 50% Outbox | 10% Base | Inbox | Outbox | |
| 75% Outbox | 10% Base | Inbox | Outbox | |
| 2—Classes | 3—Classes | ||||||
|---|---|---|---|---|---|---|---|
| Network | Accuracy (%) | Level 1 | Inbox | Outbox | Level 1 | Inbox | Outbox |
| AlexNet | Round 1 | 100 | 98.88 | 58.49 | 99.24 | 99.24 | 47.07 |
| Round 2 | 98.6 | 98.69 | 68.06 | 99.3 | 99.3 | 54.01 | |
| Round 3 | |||||||
| 25% Outbox | 98.73 | 98.13 | 97.38 | 98.09 | 98.09 | 93.06 | |
| 50% Outbox | 99.36 | 97.75 | 99.07 | 92.99 | 92.99 | 94.14 | |
| 75% Outbox | 99.36 | 98.5 | 99.69 | 100 | 100 | 98.46 | |
| ResNet–50 | Round 1 | 100 | 99.63 | 65.12 | 99.24 | 82.77 | 60.65 |
| Round 2 | 99.3 | 99.44 | 65.9 | 100 | 91.01 | 58.18 | |
| Round 3 | |||||||
| 25% Outbox | 99.36 | 99.63 | 97.84 | 97.45 | 84.83 | 95.06 | |
| 50% Outbox | 100 | 97.94 | 99.85 | 99.36 | 88.2 | 97.38 | |
| 75% Outbox | 99.36 | 98.69 | 98.77 | 98.73 | 84.46 | 98.92 | |
| MobileNet | Round 1 | 99.24 | 96.63 | 60.19 | 97.73 | 73.4 | 45.8 |
| Round 2 | 97.9 | 95.88 | 71.14 | 95.1 | 77.34 | 54.48 | |
| Round 3 | |||||||
| 25% Outbox | 98.1 | 97.57 | 94.14 | 94.27 | 78.28 | 87.35 | |
| 50% Outbox | 98.73 | 95.69 | 98.15 | 99.24 | 76.78 | 95.06 | |
| 75% Outbox | 100 | 95.51 | 98.3 | 96.18 | 75.09 | 96.91 | |
| EfficientNet | Round 1 | 96.97 | 91.2 | 61.27 | 90.15 | 70.22 | 41.98 |
| Round 2 | 97.9 | 93.07 | 61.88 | 93 | 68.73 | 56.33 | |
| Round 3 | |||||||
| 25% Outbox | 93.63 | 90.64 | 87.5 | 84.18 | 68.73 | 80.09 | |
| 50% Outbox | 96.18 | 91.76 | 95.68 | 89.81 | 68.35 | 90.28 | |
| 75% Outbox | 98.1 | 89.14 | 97.99 | 89.1 | 65.92 | 95.06 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Talaat, M.; Si, X.; Xi, J. Datasets of Simulated Exhaled Aerosol Images from Normal and Diseased Lungs with Multi-Level Similarities for Neural Network Training/Testing and Continuous Learning. Data 2023, 8, 126. https://doi.org/10.3390/data8080126
Talaat M, Si X, Xi J. Datasets of Simulated Exhaled Aerosol Images from Normal and Diseased Lungs with Multi-Level Similarities for Neural Network Training/Testing and Continuous Learning. Data. 2023; 8(8):126. https://doi.org/10.3390/data8080126
Chicago/Turabian StyleTalaat, Mohamed, Xiuhua Si, and Jinxiang Xi. 2023. "Datasets of Simulated Exhaled Aerosol Images from Normal and Diseased Lungs with Multi-Level Similarities for Neural Network Training/Testing and Continuous Learning" Data 8, no. 8: 126. https://doi.org/10.3390/data8080126





