The Glyoxylate Cycle Is Involved in White-Opaque Switching in Candida albicans
Abstract
:1. Introduction
2. Materials and Methods
2.1. Strains and Growth Conditions
2.2. Genetic Procedures
2.3. Protein Extraction and Proteomics Analysis
2.4. In Vivo Procedures
3. Results
3.1. Analysis of Proteome in WOR1 Overexpressing Cells
3.2. WOR1 Is Involved in the Use of Non-Fermentable Two Carbon Sources
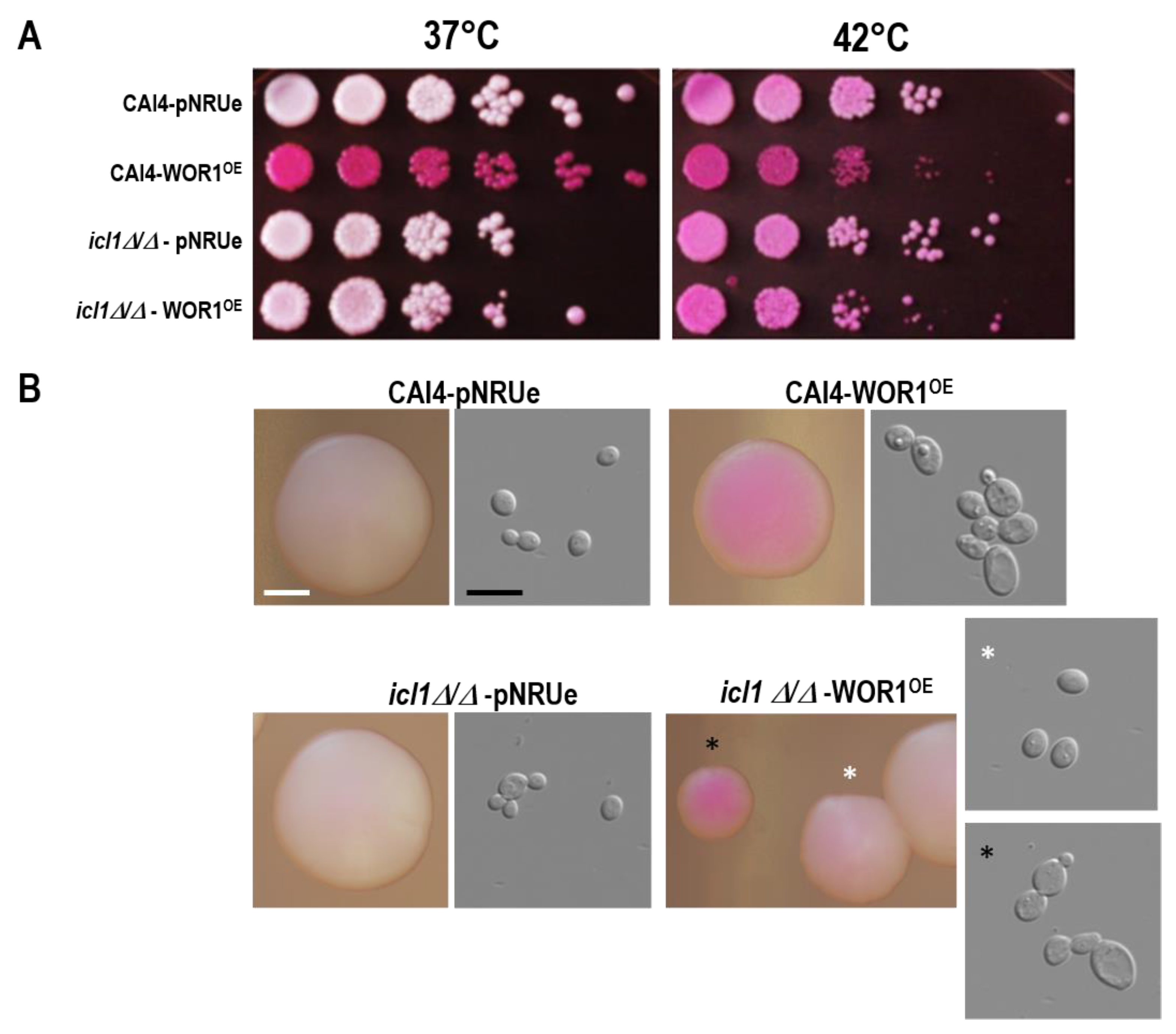
3.3. icl1Δ/Δ mutants Fail to Produce Phloxine B+ Cells in WOR1 Overexpressing Cells
3.4. icl1Δ/Δ Mutants Are Defective in Gastrointestical Colonization
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gudlaugsson, O.; Gillespie, S.; Lee, K.; Berg, J.V.; Hu, J.; Messer, S.; Herwaldt, L.; Pfaller, M.; Diekema, D. Attributable Mortality of Nosocomial Candidemia, Revisited. Clin. Infect. Dis. 2003, 37, 1172–1177. [Google Scholar] [CrossRef]
- Pappas, P.G.; Lionakis, M.S.; Arendrup, M.C.; Ostrosky-Zeichner, L.; Kullberg, B.J. Invasive candidiasis. Nat. Rev. Dis. Primers 2018, 4, 18026. [Google Scholar] [CrossRef]
- Brown, G.D.; Denning, D.W.; Gow, N.A.R.; Levitz, S.M.; Netea, M.G.; White, T.C. Hidden killers: Human fungal infections. Sci. Transl. Med. 2012, 4, 165rv113. [Google Scholar] [CrossRef] [Green Version]
- Mayer, F.L.; Wilson, D.; Hube, B. Candida albicans pathogenicity mechanisms. Virulence 2013, 4, 119–128. [Google Scholar] [CrossRef] [Green Version]
- Koh, A.Y. Murine models of Candida gastrointestinal colonization and dissemination. Eukaryot. Cell 2013, 12, 1416–1422. [Google Scholar] [CrossRef] [Green Version]
- Magill, S.S.; Swoboda, S.M.; Johnson, E.A.; Merz, W.G.; Pelz, R.K.; Lipsett, P.A.; Hendrix, C.W. The association between anatomic site of Candida colonization, invasive candidiasis, and mortality in critically ill surgical patients. Diagn. Microbiol. Infect. Dis. 2006, 55, 293–301. [Google Scholar] [CrossRef]
- Miranda, L.N.; van der Heijden, I.M.; Costa, S.F.; Sousa, A.P.; Sienra, R.A.; Gobara, S.; Santos, C.R.; Lobo, R.D.; Pessoa, V.P., Jr.; Levin, A.S. Candida colonisation as a source for candidaemia. J. Hosp. Infect. 2009, 72, 9–16. [Google Scholar] [CrossRef]
- Voss, A.; Hollis, R.J.; Pfaller, M.A.; Wenzel, R.P.; Doebbeling, B.N. Investigation of the sequence of colonization and candidemia in nonneutropenic patients. J. Clin. Microbiol. 1994, 32, 975–980. [Google Scholar] [CrossRef] [Green Version]
- Marco, F.; Lockhart, S.R.; Pfaller, M.A.; Pujol, C.; Rangel-Frausto, M.S.; Wiblin, T.; Blumberg, H.M.; Edwards, J.E.; Jarvis, W.; Saiman, L.; et al. Elucidating the origins of nosocomial infections with Candida albicans by DNA fingerprinting with the complex probe Ca3. J. Clin. Microbiol. 1999, 37, 2817–2828. [Google Scholar] [CrossRef] [Green Version]
- Prieto, D.; Correia, I.; Pla, J.; Román, E. Adaptation of Candida albicans to commensalism in the gut. Future Microbiol. 2016, 11, 567–583. [Google Scholar] [CrossRef]
- Witchley, J.N.; Penumetcha, P.; Abon, N.V.; Woolford, C.A.; Mitchell, A.P.; Noble, S.M. Candida albicans Morphogenesis Programs Control the Balance between Gut Commensalism and Invasive Infection. Cell Host Microbe 2019, 25, 432–443.e6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neville, B.A.; d’Enfert, C.; Bougnoux, M.E. Candida albicans commensalism in the gastrointestinal tract. FEMS Yeast Res. 2015, 15, fov081. [Google Scholar] [CrossRef] [Green Version]
- Noble, S.M. Candida albicans specializations for iron homeostasis: From commensalism to virulence. Curr. Opin. Microbiol. 2013, 16, 708–715. [Google Scholar] [CrossRef] [Green Version]
- Perez, J.C.; Kumamoto, C.A.; Johnson, A.D. Candida albicans commensalism and pathogenicity are intertwined traits directed by a tightly knit transcriptional regulatory circuit. PLoS Biol. 2013, 11, e1001510. [Google Scholar] [CrossRef] [Green Version]
- Pierce, J.V.; Kumamoto, C.A. Variation in Candida albicans EFG1 expression enables host-dependent changes in colonizing fungal populations. MBio 2012, 3, e00117-12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vautier, S.; Drummond, R.A.; Chen, K.; Murray, G.I.; Kadosh, D.; Brown, A.J.; Gow, N.A.; MacCallum, D.M.; Kolls, J.K.; Brown, G.D. Candida albicans colonization and dissemination from the murine gastrointestinal tract: The influence of morphology and Th17 immunity. Cell. Microbiol. 2015, 17, 445–450. [Google Scholar] [CrossRef] [Green Version]
- Prieto, A.D.; Román, E.; Correia, I.; Pla, J. The HOG pathway is critical for the colonization of the mouse gastrointestinal tract by Candida albicans. PLoS ONE 2014, 9, e87128. [Google Scholar]
- Ramírez-Zavala, B.; Mottola, A.; Haubenreißer, J.; Schneider, S.; Allert, S.; Brunke, S.; Ohlsen, K.; Hube, B.; Morschhäuser, J. The Snf1-activating kinase Sak1 is a key regulator of metabolic adaptation and in vivo fitness of Candida albicans. Mol. Microbiol. 2017, 104, 989–1007. [Google Scholar] [CrossRef] [Green Version]
- Román, E.; Huertas, B.; Prieto, D.; Diez-Orejas, R.; Pla, J. TUP1-mediated filamentation in Candida albicans leads to inability to colonize the mouse gut. Future Microbiol. 2018, 13, 857–867. [Google Scholar] [CrossRef] [Green Version]
- Soll, D.R. The role of phenotypic switching in the basic biology and pathogenesis of Candida albicans. J. Oral Microbiol. 2014, 6. [Google Scholar] [CrossRef]
- Tuch, B.B.; Mitrovich, Q.M.; Homann, O.R.; Hernday, A.D.; Monighetti, C.K.; De La Vega, F.M.; Johnson, A.D. The transcriptomes of two heritable cell types illuminate the circuit governing their differentiation. PLoS Genet. 2010, 6, e1001070. [Google Scholar] [CrossRef] [Green Version]
- Lan, C.Y.; Newport, G.; Murillo, L.A.; Jones, T.; Scherer, S.; Davis, R.W.; Agabian, N. Metabolic specialization associated with phenotypic switching in Candida albicans. Proc. Natl. Acad. Sci. USA 2002, 99, 14907–14912. [Google Scholar] [CrossRef] [Green Version]
- Tsong, A.E.; Miller, M.G.; Raisner, R.M.; Johnson, A.D. Evolution of a combinatorial transcriptional circuit: A case study in yeasts. Cell 2003, 115, 389–399. [Google Scholar] [CrossRef] [Green Version]
- Kvaal, C.; Lachke, S.A.; Srikantha, T.; Daniels, K.; McCoy, J.; Soll, D.R. Misexpression of the opaque-phase-specific gene PEP1 (SAP1) in the white phase of Candida albicans confers increased virulence in a mouse model of cutaneous infection. Infect. Immun. 1999, 67, 6652–6662. [Google Scholar] [CrossRef] [Green Version]
- Kvaal, C.A.; Srikantha, T.; Soll, D.R. Misexpression of the white-phase-specific gene WH11 in the opaque phase of Candida albicans affects switching and virulence. Infect. Immun. 1997, 65, 4468–4475. [Google Scholar] [CrossRef] [Green Version]
- Srikantha, T.; Borneman, A.R.; Daniels, K.J.; Pujol, C.; Wu, W.; Seringhaus, M.R.; Gerstein, M.; Yi, S.; Snyder, M.; Soll, D.R. TOS9 regulates white-opaque switching in Candida albicans. Eukaryot. Cell 2006, 5, 1674–1687. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pande, K.; Chen, C.; Noble, S.M. Passage through the mammalian gut triggers a phenotypic switch that promotes Candida albicans commensalism. Nat. Genet. 2013, 45, 1088–1091. [Google Scholar] [CrossRef] [Green Version]
- Huang, G.; Wang, H.; Chou, S.; Nie, X.; Chen, J.; Liu, H. Bistable expression of WOR1, a master regulator of white-opaque switching in Candida albicans. Proc. Natl. Acad. Sci. USA 2006, 103, 12813–12818. [Google Scholar] [CrossRef] [Green Version]
- Zordan, R.E.; Galgoczy, D.J.; Johnson, A.D. Epigenetic properties of white-opaque switching in Candida albicans are based on a self-sustaining transcriptional feedback loop. Proc. Natl. Acad. Sci. USA 2006, 103, 12807–12812. [Google Scholar] [CrossRef] [Green Version]
- Zordan, R.E.; Miller, M.G.; Galgoczy, D.J.; Tuch, B.B.; Johnson, A.D. Interlocking transcriptional feedback loops control white-opaque switching in Candida albicans. PLoS Biol. 2007, 5, e256. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hernday, A.D.; Lohse, M.B.; Fordyce, P.M.; Nobile, C.J.; DeRisi, J.L.; Johnson, A.D. Structure of the transcriptional network controlling white-opaque switching in Candida albicans. Mol. Microbiol. 2013, 90, 22–35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lohse, M.B.; Ene, I.V.; Craik, V.B.; Hernday, A.D.; Mancera, E.; Morschhauser, J.; Bennett, R.J.; Johnson, A.D. Systematic Genetic Screen for Transcriptional Regulators of the Candida albicans White-Opaque Switch. Genetics 2016, 203, 1679–1692. [Google Scholar] [CrossRef] [Green Version]
- Stevenson, J.S.; Liu, H. Regulation of white and opaque cell-type formation in Candida albicans by Rtt109 and Hst3. Mol. Microbiol. 2011, 81, 1078–1091. [Google Scholar] [CrossRef] [Green Version]
- Srikantha, T.; Tsai, L.; Daniels, K.; Klar, A.J.; Soll, D.R. The histone deacetylase genes HDA1 and RPD3 play distinct roles in regulation of high-frequency phenotypic switching in Candida albicans. J. Bacteriol. 2001, 183, 4614–4625. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hnisz, D.; Schwarzmuller, T.; Kuchler, K. Transcriptional loops meet chromatin: A dual-layer network controls white-opaque switching in Candida albicans. Mol. Microbiol. 2009, 74, 1–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stoldt, V.R.; Sonneborn, A.; Leuker, C.E.; Ernst, J.F. Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J. 1997, 16, 1982–1991. [Google Scholar] [CrossRef] [Green Version]
- Sonneborn, A.; Tebarth, B.; Ernst, J.F. Control of white-opaque phenotypic switching in Candida albicans by the Efg1p morphogenetic regulator. Infect. Immun. 1999, 67, 4655–4660. [Google Scholar] [CrossRef] [Green Version]
- Prieto, D.; Román, E.; Alonso-Monge, R.; Pla, J. Overexpression of the Transcriptional Regulator WOR1 Increases Susceptibility to Bile Salts and Adhesion to the Mouse Gut Mucosa in Candida albicans. Front. Cell. Infect. Microbiol. 2017, 7, 389. [Google Scholar] [CrossRef]
- Fonzi, W.A.; Irwin, M.Y. Isogenic strain construction and gene mapping in Candida albicans. Genetics 1993, 134, 717–728. [Google Scholar] [CrossRef]
- Román, E.; Correia, I.; Salazin, A.; Fradin, C.; Jouault, T.; Poulain, D.; Liu, F.T.; Pla, J. The Cek1-mediated MAP kinase pathway regulates exposure of α−(1,2) and β−(1,2)-mannosides in the cell wall of Candida albicans modulating immune recognition. Virulence 2016, 7, 558–577. [Google Scholar] [CrossRef] [Green Version]
- Lorenz, M.C.; Fink, G.R. The glyoxylate cycle is required for fungal virulence. Nature 2001, 412, 83–86. [Google Scholar] [CrossRef]
- Ramirez, M.A.; Lorenz, M.C. Mutations in alternative carbon utilization pathways in Candida albicans attenuate virulence and confer pleiotropic phenotypes. Eukaryot. Cell 2007, 6, 280–290. [Google Scholar] [CrossRef] [Green Version]
- Shapiro, R.S.; Chavez, A.; Porter, C.B.M.; Hamblin, M.; Kaas, C.S.; DiCarlo, J.E.; Zeng, G.; Xu, X.; Revtovich, A.V.; Kirienko, N.V.; et al. A CRISPR-Cas9-based gene drive platform for genetic interaction analysis in Candida albicans. Nature Microbiol. 2018, 3, 73–82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Min, K.; Ichikawa, Y.; Woolford, C.A.; Mitchell, A.P. Candida albicans Gene Deletion with a Transient CRISPR-Cas9 System. mSphere 2016, 1, e00130-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Román, E.; Coman, I.; Prieto, D.; Alonso-Monge, R.; Pla, J. Implementation of a CRISPR-Based System for Gene Regulation in Candida albicans. mSphere 2019, 4, e00001-19. [Google Scholar] [CrossRef] [Green Version]
- Vyas, V.K.; Barrasa, M.I.; Fink, G.R. A CRISPR system permits genetic engineering of essential genes and gene families. Sci. Adv. 2015, 1, e1500248. [Google Scholar] [CrossRef] [Green Version]
- Martin, H.; Rodriguez-Pachon, J.M.; Ruiz, C.; Nombela, C.; Molina, M. Regulatory mechanisms for modulation of signaling through the cell integrity Slt2-mediated pathway in Saccharomyces cerevisiae. J. Biol. Chem. 2000, 275, 1511–1519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rico-San Román, L.; Horcajo, P.; Regidor-Cerrillo, J.; Fernández-Escobar, M.; Collantes-Fernández, E.; Gutiérrez-Blázquez, D.; Hernáez-Sánchez, M.L.; Saeij, J.P.J.; Ortega-Mora, L.M. Comparative tachyzoite proteome analyses among six Neospora caninum isolates with different virulence. Int. J. Parasitol. 2020, 50, 377–388. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.N.; Morschhauser, J. Tetracycline-inducible gene expression and gene deletion in Candida albicans. Eukaryot. Cell 2005, 4, 1328–1342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ene, I.V.; Lohse, M.B.; Vladu, A.V.; Morschhauser, J.; Johnson, A.D.; Bennett, R.J. Phenotypic Profiling Reveals that Candida albicans Opaque Cells Represent a Metabolically Specialized Cell State Compared to Default White Cells. MBio 2016, 7. [Google Scholar] [CrossRef] [Green Version]
- Piekarska, K.; Hardy, G.; Mol, E.; van den Burg, J.; Strijbis, K.; van Roermund, C.; van den Berg, M.; Distel, B. The activity of the glyoxylate cycle in peroxisomes of Candida albicans depends on a functional beta-oxidation pathway: Evidence for reduced metabolite transport across the peroxisomal membrane. Microbiology 2008, 154, 3061–3072. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gunsalus, K.T.; Tornberg-Belanger, S.N.; Matthan, N.R.; Lichtenstein, A.H.; Kumamoto, C.A. Manipulation of Host Diet To Reduce Gastrointestinal Colonization by the Opportunistic Pathogen Candida albicans. mSphere 2016, 1, e00020-15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prieto, D.; Pla, J. Distinct stages during colonization of the mouse gastrointestinal tract by Candida albicans. Front. Microbiol. 2015, 6, 792. [Google Scholar] [CrossRef] [PubMed]
- Aksit, A.; van der Klei, I.J. Yeast peroxisomes: How are they formed and how do they grow? Int. J. Biochem. Cell Biol. 2018, 105, 24–34. [Google Scholar] [CrossRef] [PubMed]




| Strain Name | Common Name in Manuscript | Background Strain and Genotype | Source |
|---|---|---|---|
| CAI4 | ura3Δ::imm434/ura3Δ::imm434 | [39] | |
| CAI4-pNRUe | [CAI4] ADH1/adh1::tTATETPR-myc-URA3 | [40] | |
| CAI4-RFP | [CAI4] ADH1/adh1:: tTATETPR-dTOM2-URA3 | [38] | |
| CAI4-WOR1OE | [CAI4] ADH1/adh1:: tTATETPR-WOR1-myc-URA3 | [38] | |
| MLC9 | [CAI4] icl1::hisG/icl1::hisG | [41] | |
| MRC10 | icl1Δ/Δ | [CAI4] icl1::hisG/icl1::hisG RPS10/rps10::URA3 | [42] |
| MRC11 | icl1Δ/Δ-pICL1 | [icl1Δ/Δ] RPS10/rps10::ICL1-URA3 | [42] |
| SHV1 | icl1Δ/Δ-pICL1-GFP | [icl1Δ/Δ-pICL1] ADH1/adh1:: tTATETPR--GFP-myc-SAT1 | This study |
| SHV2 | icl1Δ/Δ-RFP | [icl1Δ/Δ] ADH1/adh1:: tTATETPR-dTOM2-SAT1 | This study |
| SHV3 | icl1Δ/Δ-pNRUe | [icl1Δ/Δ] ADH1/adh1::tTATETPR-myc-URA3 | This study |
| SHV4 | icl1Δ/Δ-WOR1OE | [icl1Δ/Δ] ADH1/adh1:: tTATETPR-WOR1-myc-URA3 | This study |
| SHV5 | icl1Δ/Δ-mtla1Δ | [icl1Δ/Δ] mtla1∆::SAT1 | This study |
| SHV6 | icl1Δ/Δ-pICL1-mtla1Δ | [icl1Δ/Δ-pICL1] mtla1∆::SAT1 | This study |
| Primer Name | Sequence (5′→3′) |
|---|---|
| SNR52/F | AAGAAAGAAAGAAAACCAGGAGTGAA |
| SNR52/R_MTL1 | CTCACGCTTCAATTGTAAGACAAATTAAAAATAGTTTACGCAAGTC |
| sgRNA/F_MTL1 | TCTTACAATTGAAGCGTGAGGTTTTAGAGCTAGAAATAGCAAGTTAAA |
| sgRNA/R | ACAAATATTTAAACTCGGGACCTGG |
| SNR52/N | GCGGCCGCAAGTGATTAGACT |
| sgRNA/N | GCAGCTCAGTGATTAAGAGTAAAGATGG |
| MATa1_del_F | TTTCTGCGTATTGTGATAAATAACTTTTTTTCCCTCTAAAAATATTGATTAGAGGCACAAAATAAAAATCACCTTCAACCCGTCAAAACTAGAGAATAATAAAGAAAACG |
| MATa1_del_R | CCCACCAAGACATGTTACGAATAGATCTATTAGTTAGCAATATTCTGTTTGATAATACATACCCAAACTCTTATTTGGGAGCAGGACCACCTTTGATTGTAAATAG |
| Comp_del_F | CACGTATAAAACTAGACCTCAAGTCTCG |
| oRS108 | ATGAACTCAGAAATAGAAAGTAGC |
| oRS109 | CTAGGTTGAATTTGAACTTGATTT |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vico, S.H.; Prieto, D.; Monge, R.A.; Román, E.; Pla, J. The Glyoxylate Cycle Is Involved in White-Opaque Switching in Candida albicans. J. Fungi 2021, 7, 502. https://doi.org/10.3390/jof7070502
Vico SH, Prieto D, Monge RA, Román E, Pla J. The Glyoxylate Cycle Is Involved in White-Opaque Switching in Candida albicans. Journal of Fungi. 2021; 7(7):502. https://doi.org/10.3390/jof7070502
Chicago/Turabian StyleVico, Susana Hidalgo, Daniel Prieto, Rebeca Alonso Monge, Elvira Román, and Jesús Pla. 2021. "The Glyoxylate Cycle Is Involved in White-Opaque Switching in Candida albicans" Journal of Fungi 7, no. 7: 502. https://doi.org/10.3390/jof7070502







