Diagnostic Allele-Specific PCR for the Identification of Candida auris Clades
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sequence Analysis for Species- and Clade-Specific Allele Selection
2.2. Strains and Media
2.3. DNA Extraction
2.4. PCR and Sequencing
2.5. Allele-Specific Primer Design
2.6. AS-PCR
3. Results
3.1. C. auris Identification
3.2. C. auris Clade Identification
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Name | NCBI Accession Nr. | Strain | Size (bp) |
|---|---|---|---|
| Candida auris | NR_154998.1 | CBS 10913 | 342 |
| Candida haemulonii | JX459664.1 | CBS 6590 | 479 |
| Candida haemulonii var. vulnera | JX459687.1 | CNMCL7256 | 322 |
| Candida pseudohaemulonii | EU881972.1 | C4368 | 327 |
| Candida duobushaemulonii | JX459667.1 | CBS 7799 | 487 |
| Candida albicans | NR_125332.1 | CBS 562 | 462 |
| Candida tropicalis | NR_111250.1 | CBS 94 | 454 |
| Candida lusitaniae | NR_130677.1 | CBS 6936 | 307 |
| Candida glabrata | NR_130691.1 | NRRL Y-65 | 655 |
| Candida dubliniensis | NR_119386.1 | CBS 7987 | 460 |
| Candida parapsilosis | NR_130673.1 | ATCC 22019 | 461 |
| Candida orthopsilosis | NR_130661.1 | ATCC 96139 | 436 |
| Candida metapsilosis | NR_165186.1 | CBS 10907 | 472 |
| Candida sake | NR_151807.1 | CBS 159 | 432 |
| Candida inconspicua | NR_111116.1 | CBS 180 | 379 |
| Candida castellii | NR_154958.1 | CBS 4332 | 628 |
| Candida bracarensis | NR_136973.1 | CBS 10154 | 704 |
| Candida pararugosa | MF797780.1 | S200 | 324 |
| Candida guilliermondii (Meyerozyma guillermondii) | NR_111247.1 | CBS 2030 | 532 |
| Candida krusei (Pichia kudriavzevii) | NR_131315.1 | ATCC 6258 | 435 |
| Candida famata (Debaryomyces hansenii) | NR_120016.1 | JCM 1990 | 580 |
| Candida kefyr (Kluyveromyces marxianus) | NR_111251.1 | CBS 712 | 628 |
| Candida norvegensis (Pichia norvegensis) | MH396411.1 | URM 7762 | 434 |
| Lodderomyces elongisporus | NR_111593.1 | ATCC 11503 | 496 |
| Yarrowia lipolytica | NR_111212.1 | CBS 6124 | 285 |
| Nakaseomyces delphensis | NR_167682.1 | CBS 2170 | 709 |
| Nakaseomyces bacillisporus | NR_138202.1 | NRRL Y-17846 | 662 |
| Kodameae ohmeri | NR_121464.1 | CBS 5367 | 348 |
| Diutina catenulata | NR_077200.1 | CBS 565 | 343 |
| Rhodotorula mucilaginosa (R. rubra) | NR_073296.1 | CBS 316 | 557 |
| Rhodotorula glutinis | MH665424.1 | R.g0726 | 582 |
| Saccharomyces cerevisiae | NR_111007.1 | CBS 1171 | 613 |
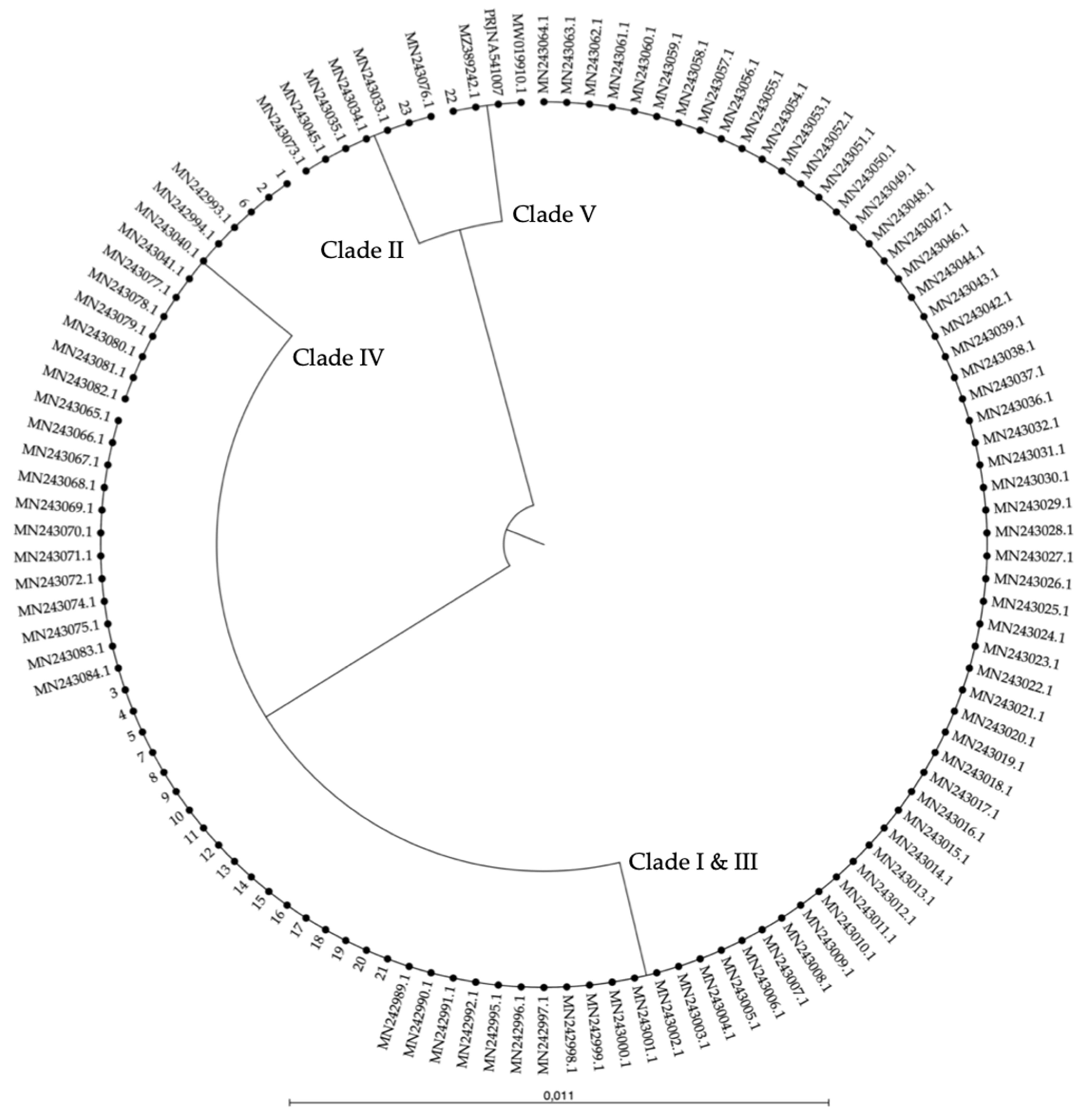
| Nr. | Strain Name | Origin | Clade | Typing Reference |
|---|---|---|---|---|
| 1 | B11244 | Venezuela | IV | [2,13,17] |
| 2 | B11245 | Venezuela | IV | |
| 3 | B11221 | South Africa | III | [2,3,13,17] |
| 4 | B11222 | South Africa | III | |
| 5 | CDC_390 | India | I | |
| 6 | - | Colombia | IV | |
| 7 | B11098 | Pakistan | I | |
| 8 | B11203 | India | I | |
| 9 | - | India | I | |
| 10 | - | India | I | |
| 11 | - | India | I | |
| 12 | - | India | I | |
| 13 | VPCI 714/P/14 | India | I | |
| 14 | B8441 | Pakistan | I | [2,3,13,17] |
| 15 | 317052804 | Oman | I | |
| 16 | VPCI 514/P/14 | India | I | |
| 17 | VPCI 1133/P/13 | India | I | |
| 18 | B11109 | India | I | |
| 19 | VPCI 1131/P/13 | India | I | |
| 20 | OS299 | Belgium | I | |
| 21 | MRU224 | South Africa | III | |
| 22 | IFRC2087/B18474 | Iran | V | [12,17] |
| 23 | B11220 | Japan | II | [2,3,13,17] |

References
- Satoh, K.; Makimura, K.; Hasumi, Y.; Nishiyama, Y.; Uchida, K.; Yamaguchi, H. Candida auris Sp. Nov., A Novel Ascomycetous Yeast Isolated from the External Ear Canal of an Inpatient in a Japanese Hospital. Microbiol. Immunol. 2009, 53, 41–44. [Google Scholar] [CrossRef] [PubMed]
- Chow, N.A.; Muñoz, J.F.; Gade, L.; Berkow, E.; Li, X.; Welsh, R.M.; Forsberg, K.; Lockhart, S.R.; Adam, R.; Alanio, A.; et al. Tracing the Evolutionary History and Global Expansion of Candida auris Using Population Genomic Analyses. MBio 2020, 11, e03364-19. [Google Scholar] [CrossRef] [PubMed]
- Muñoz, J.F.; Gade, L.; Chow, N.A.; Loparev, V.N.; Juieng, P.; Berkow, E.L.; Farrer, R.A.; Litvintseva, A.P.; Cuomo, C.A. Genomic Insights into Multidrug-Resistance, Mating and Virulence in Candida auris and Related Emerging Species. Nat. Commun. 2018, 9, 5346. [Google Scholar] [CrossRef] [Green Version]
- Kordalewska, M.; Perlin, D.S. Identification of Drug Resistant Candida auris. Front. Microbiol. 2019, 10, 1918. [Google Scholar] [CrossRef] [PubMed]
- Dewaele, K.; Lagrou, K.; Frans, J.; Hayette, M.-P.; Vernelen, K. Hospital Laboratory Survey for Identification of Candida auris in Belgium. J. Fungi 2019, 5, 84. [Google Scholar] [CrossRef] [Green Version]
- Buil, J.B.; van der Lee, H.A.; Curfs-Breuker, I.; Verweij, P.E.; Meis, J.F. External Quality Assessment Evaluating the Ability of Dutch Clinical Microbiological Laboratories to Identify Candida auris. J. Fungi 2019, 5, 94. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- CDC. Identification of Candida auris. Available online: https://www.cdc.gov/fungal/candida-auris/identification.html#identify (accessed on 10 August 2021).
- Caceres, D.H.; Forsberg, K.; Welsh, R.M.; Sexton, D.J.; Lockhart, S.R.; Jackson, B.R.; Chiller, T. Candida auris: A Review of Recommendations for Detection and Control in Healthcare Settings. J. Fungi 2019, 5, 111. [Google Scholar] [CrossRef] [Green Version]
- Mahmoudi, S.; Afshari, S.A.K.; Gharehbolagh, S.A.; Mirhendi, H.; Makimura, K. Methods for Identification of Candida auris, the Yeast of Global Public Health Concern: A review. J. Mycol. Med. 2019, 29, 174–179. [Google Scholar] [CrossRef] [PubMed]
- Sattler, J.; Noster, J.; Brunke, A.; Plum, G.; Wiegel, P.; Kurzai, O.; Meis, J.F.; Hamprecht, A. Comparison of Two Commercially Available qPCR Kits for the Detection of Candida auris. J. Fungi 2021, 7, 154. [Google Scholar] [CrossRef]
- Martínez-Murcia, A.; Navarro, A.; Bru, G.; Chowdhary, A.; Hagen, F.; Meis, J.F. Internal Validation of GPS™ MONODOSE CanAur Dtec-qPCR Kit Following the UNE/EN ISO/IEC 17025: 2005 for Detection of the Emerging Yeast Candida auris. Mycoses 2018, 61, 877–884. [Google Scholar] [CrossRef] [Green Version]
- Chow, N.A.; de Groot, T.; Badali, H.; Abastabar, M.; Chiller, T.M.; Meis, J.F. Potential Fifth Clade of Candida auris, Iran, 2018. Emerg. Infect. Dis. 2019, 25, 1780–1781. [Google Scholar] [CrossRef] [Green Version]
- Lockhart, S.R.; Etienne, K.A.; Vallabhaneni, S.; Farooqi, J.; Chowdhary, A.; Govender, N.P.; Colombo, A.L.; Calvo, B.; Cuomo, C.A.; Desjardins, C.A.; et al. Simultaneous Emergence of Multidrug-Resistant Candida auris on 3 Continents Confirmed by Whole-Genome Sequencing and Epidemiological Analyses. Clin. Infect. Dis. 2017, 64, 134–140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- CDC. Tracking Candida auris. Available online: https://www.cdc.gov/fungal/candida-auris/tracking-c-auris.html (accessed on 10 August 2021).
- Kean, R.; Brown, J.; Gulmez, D.; Ware, A.; Ramage, G. Candida auris: A Decade of Understanding of an Enigmatic Pathogenic Yeast. J. Fungi 2020, 6, 30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bruno, M.; Kersten, S.; Bain, J.M.; Jaeger, M.; Rosati, D.; Kruppa, M.D.; Lowman, D.W.; Rice, P.J.; Graves, B.; Ma, Z.; et al. Transcriptional and Functional Insights into the Host Immune Response against the Emerging Fungal Pathogen Candida auris. Nat. Microbiol. 2020, 5, 1516–1531. [Google Scholar] [CrossRef]
- Muñoz, J.F.; Welsh, R.M.; Shea, T.; Batra, D.; Gade, L.; Howard, D.; Rowe, L.A.; Meis, J.F.; Litvintseva, A.P.; Cuomo, C.A. Clade-Specific Chromosomal Rearrangements and Loss of Subtelomeric Adhesins in Candida auris. Genetics 2021, 218, iyab029. [Google Scholar] [CrossRef] [PubMed]
- Chowdhary, A.; Prakash, A.; Sharma, C.; Kordalewska, M.; Kumar, A.; Sarma, S.; Tarai, B.; Singh, A.; Upadhyaya, G.; Upadhyay, S.; et al. A Multicentre Study of Antifungal Susceptibility Patterns among 350 Candida auris Isolates (2009–2017) in India: Role of the ERG11 and FKS1 Genes in Azole and Echinocandin Resistance. J. Antimicrob. Chemother. 2018, 73, 891–899. [Google Scholar] [CrossRef]
- Maphanga, T.G.; Naicker, S.D.; Kwenda, S.; Muñoz, J.F.; van Schalkwyk, E.; Wadula, J.; Nana, T.; Ismail, A.; Coetzee, J.; Govind, C. In-Vitro Antifungal Resistance of Candida auris Isolates from Bloodstream Infections, South Africa. Antimicrob. Agents Chemother. 2021, 65, e00517-21. [Google Scholar] [CrossRef] [PubMed]
- Welsh, R.M.; Sexton, D.J.; Forsberg, K.; Vallabhaneni, S.; Litvintseva, A. Insights into the Unique Nature of the East Asian Clade of the Emerging Pathogenic Yeast Candida auris. J. Clin. Microbiol. 2019, 57, e00007-19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Safari, F.; Madani, M.; Badali, H.; Kargoshaie, A.; Fakhim, H.; Kheirollahi, M.; Meis, J.F.; Mirhendi, H. Autochthonous Fifth Clade Case of Candida auris Otomycosis in Iran. Mycopathologia 2021, in press. [Google Scholar]
- Taghizadeh Armaki, M.; Mahdavi Omran, S.; Kiakojuri, K.; Khojasteh, S.; Jafarzadeh, J.; Tavakoli, M.; Badali, H.; Haghani, I.; Shokohi, T.; Hedayati, M.T. First Fluconazole-Resistant Candida auris Isolated from Fungal Otitis in Iran. Curr. Med. Mycol. 2021, 7, 51–54. [Google Scholar]
- Sexton, D.J.; Welsh, R.M.; Bentz, M.L.; Forsberg, K.; Jackson, B.; Berkow, E.L.; Litvintseva, A.P. Evaluation of Nine Surface Disinfectants against Candida auris Using a Quantitative Disk Carrier Method: EPA SOP-MB-35. Infect. Control Hosp. Epidemiol. 2020, 41, 1219–1221. [Google Scholar] [CrossRef]
- Chatterjee, P.; Choi, H.; Ochoa, B.; Garmon, G.; Coppin, J.D.; Allton, Y.; Lukey, J.; Williams, M.D.; Navarathna, D.; Jinadatha, C. Clade-Specific Variation in Susceptibility of Candida auris to Broad-Spectrum Ultraviolet C light (UV-C). Infect. Control Hosp. Epidemiol. 2020, 41, 1384–1387. [Google Scholar] [CrossRef] [PubMed]
- De Groot, T.; Chowdhary, A.; Meis, J.F.; Voss, A. Killing of Candida auris by UV-C: Importance of Exposure Time and Distance. Mycoses 2019, 62, 408–412. [Google Scholar] [CrossRef] [Green Version]
- Abe, M.; Katano, H.; Nagi, M.; Higashi, Y.; Sato, Y.; Kikuchi, K.; Hasegawa, H.; Miyazaki, Y. Potency of Gastrointestinal Colonization and Virulence of Candida auris in a Murine Endogenous Candidiasis. PLoS ONE 2020, 15, e0243223. [Google Scholar] [CrossRef] [PubMed]
- Forgács, L.; Borman, A.M.; Prépost, E.; Tóth, Z.; Kardos, G.; Kovács, R.; Szekely, A.; Nagy, F.; Kovacs, I.; Majoros, L. Comparison of In Vivo Pathogenicity of Four Candida auris Clades in a Neutropenic Bloodstream Infection Murine Model. Emerg. Microbes Infect. 2020, 9, 1160–1169. [Google Scholar] [CrossRef] [PubMed]
- Short, B.; Brown, J.; Delaney, C.; Sherry, L.; Williams, C.; Ramage, G.; Kean, R. Candida auris Exhibits Resilient Biofilm Characteristics In Vitro: Implications for Environmental Persistence. J. Hosp. Infect. 2019, 103, 92–96. [Google Scholar] [CrossRef]
- Chatzimoschou, A.; Giampani, A.; Meis, J.F.; Roilides, E. Activities of Nine Antifungal Agents Against Candida auris Biofilms. Mycoses 2021, 64, 381–384. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.L.; Delaney, C.; Short, B.; Butcher, M.C.; McKloud, E.; Williams, C.; Kean, R.; Ramage, G. Candida auris Phenotypic Heterogeneity Determines Pathogenicity In Vitro. Msphere 2020, 5, e00371-20. [Google Scholar] [CrossRef]
- Vatanshenassan, M.; Boekhout, T.; Mauder, N.; Robert, V.; Maier, T.; Meis, J.F.; Berman, J.; Then, E.; Kostrzewa, M.; Hagen, F. Evaluation of Microsatellite Typing, ITS Sequencing, AFLP Fingerprinting, MALDI-TOF MS, and Fourier-Transform Infrared Spectroscopy Analysis of Candida auris. J. Fungi 2020, 6, 146. [Google Scholar] [CrossRef] [PubMed]
- De Groot, T.; Puts, Y.; Berrio, I.; Chowdhary, A.; Meis, J.F. Development of Candida auris Short Tandem Repeat Typing and Its Application to a Global Collection of Isolates. MBio 2020, 11, e02971-19. [Google Scholar] [CrossRef] [Green Version]
- Rhodes, J.; Fisher, M.C. Global Epidemiology of Emerging Candida auris. Curr. Opin. Microbiol. 2019, 52, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Huang, S.; Sun, M.; Liu, S.; Liu, Y.; Wang, W.; Zhang, X.; Wang, H.; Hua, W. An Improved Allele-Specific PCR Primer Design Method for SNP Marker Analysis and Its Application. Plant Methods 2012, 8, 34. [Google Scholar] [CrossRef] [Green Version]
- Carolus, H.; Pierson, S.; Muñoz, J.F.; Subotić, A.; Cruz, R.B.; Cuomo, C.A.; Van Dijck, P. Genome-Wide Analysis of Experimentally Evolved Candida auris Reveals Multiple Novel Mechanisms of Multidrug Resistance. Mbio 2021, 12, e03333-20. [Google Scholar] [CrossRef]
- Hou, X.; Lee, A.; Jiménez-Ortigosa, C.; Kordalewska, M.; Perlin, D.S.; Zhao, Y. Rapid Detection of ERG11-Associated Azole Resistance and FKS-Associated Echinocandin Resistance in Candida auris. Antimicrob. Agents Chemother. 2019, 63, e01811-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ding, X.; Wang, Z.; Zhou, K.; Xu, L.; Xu, H.; Wang, Y. Allele-Specific Primers for Diagnostic PCR Authentication of Dendrobium officinale. Planta Med. 2003, 69, 587–588. [Google Scholar] [PubMed]
- Wu, D.Y.; Ugozzoli, L.; Pal, B.K.; Wallace, R.B. Allele-Specific Enzymatic Amplification of Beta-Globin Genomic DNA for Diagnosis of Sickle Cell Anemia. Proc. Natl. Acad. Sci. USA 1989, 86, 2757–2760. [Google Scholar] [CrossRef] [Green Version]
- Weber, B.; Meldgaard, P.; Hager, H.; Wu, L.; Wei, W.; Tsai, J.; Khalil, A.; Nexo, E.; Sorensen, B.S. Detection of EGFR Mutations in Plasma and Biopsies from Non-Small Cell Lung Cancer Patients by Allele-Specific PCR Assays. BMC Cancer 2014, 14, 294. [Google Scholar] [CrossRef] [Green Version]
- Sanchaisuriya, K.; Chunpanich, S.; Fucharoen, G.; Fucharoen, S. Multiplex Allele-Specific PCR Assay for Differential Diagnosis of Hb S, Hb D-Punjab and Hb Tak. Clin. Chim. Acta 2004, 343, 129–134. [Google Scholar] [CrossRef]
- Lone, S.A.; Ahmad, A. Candida auris-The Growing Menace to Global Health. Mycoses 2019, 62, 620–637. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mizusawa, M.; Miller, H.; Green, R.; Lee, R.; Durante, M.; Perkins, R.; Hewitt, C.; Simner, P.J.; Carroll, K.C.; Hayden, R.T.; et al. Can Multidrug-Resistant Candida auris Be Reliably Identified in Clinical Microbiology Laboratories? J. Clin. Microbiol. 2017, 55, 638–640. [Google Scholar] [CrossRef] [Green Version]
- Snayd, M.; Dias, F.; Ryan, R.W.; Clout, D.; Banach, D.B.; Warnock, D.W. Misidentification of Candida auris by RapID Yeast Plus, a Commercial, Biochemical Enzyme-Based Manual Rapid Identification System. J. Clin. Microbiol. 2018, 56, e00080-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Theill, L.; Dudiuk, C.; Morales-Lopez, S.; Berrio, I.; Rodríguez, J.Y.; Marin, A.; Gamarra, S.; Garcia-Effron, G. Single-tube classical PCR for Candida auris and Candida haemulonii Identification. Rev. Iberoam Micol 2018, 35, 110–112. [Google Scholar] [CrossRef]
- Turner, S.A.; Butler, G. The Candida Pathogenic Species Complex. Cold Spring Harb. Perspect. Med. 2014, 4, a019778. [Google Scholar] [CrossRef] [Green Version]
- Ambaraghassi, G.; Dufresne, P.J.; Dufresne, S.F.; Vallières, É.; Muñoz, J.F.; Cuomo, C.A.; Berkow, E.L.; Lockhart, S.R.; Luong, M.-L. Identification of Candida auris by Use of the Updated Vitek 2 Yeast Identification System, Version 8.01: A Multilaboratory Evaluation Study. J. Clin. Microbiol. 2019, 57, e00884-19. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. PCR Protoc. Guide Methods Appl. 1990, 18, 315–322. [Google Scholar]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W.; Consortium, F.B. Nuclear Ribosomal Internal Transcribed Spacer (ITS) Region as a Universal DNA Barcode Marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leach, L.; Zhu, Y.; Chaturvedi, S. Development and Validation of a Real-Time PCR Assay for Rapid Detection of Candida auris from Surveillance Samples. J. Clin. Microbiol. 2018, 56, e01223-17. [Google Scholar] [CrossRef] [Green Version]
- Walchak, R.C.; Buckwalter, S.P.; Zinsmaster, N.M.; Henn, K.M.; Johnson, K.M.; Koelsch, J.M.; Herring, S.A.; Steinmetz, L.K.; Reed, K.A.; Barth, J.E. Candida auris Direct Detection from Surveillance Swabs, Blood, and Urine Using a Laboratory-Developed PCR Method. J. Fungi 2020, 6, 224. [Google Scholar] [CrossRef]
- Prakash, A.; Sharma, C.; Singh, A.; Singh, P.K.; Kumar, A.; Hagen, F.; Govender, N.; Colombo, A.; Meis, J.; Chowdhary, A. Evidence of Genotypic Diversity Among Candida auris Isolates by Multilocus Sequence Typing, Matrix-Assisted Laser Desorption Ionization Time-of-Flight Mass Spectrometry and Amplified Fragment Length Polymorphism. Clin. Microbiol. Infect. 2016, 22, 271–277. [Google Scholar] [CrossRef] [Green Version]
- Szekely, A.; Borman, A.M.; Johnson, E.M. Candida auris Isolates of the Southern Asian and South African Lineages Exhibit Different Phenotypic and Antifungal Susceptibility Profiles In Vitro. J. Clin. Microbiol. 2019, 57, e02055-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
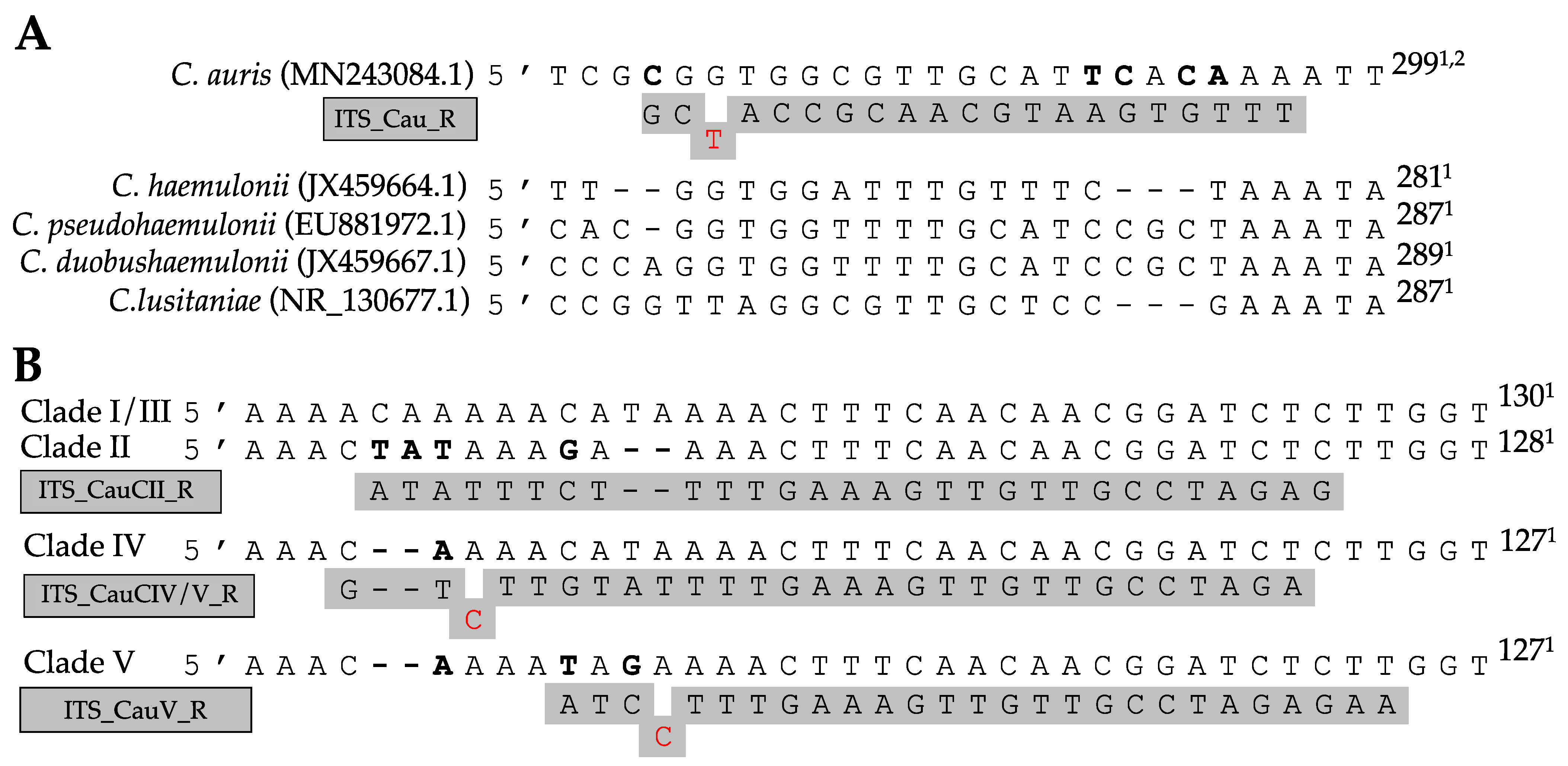
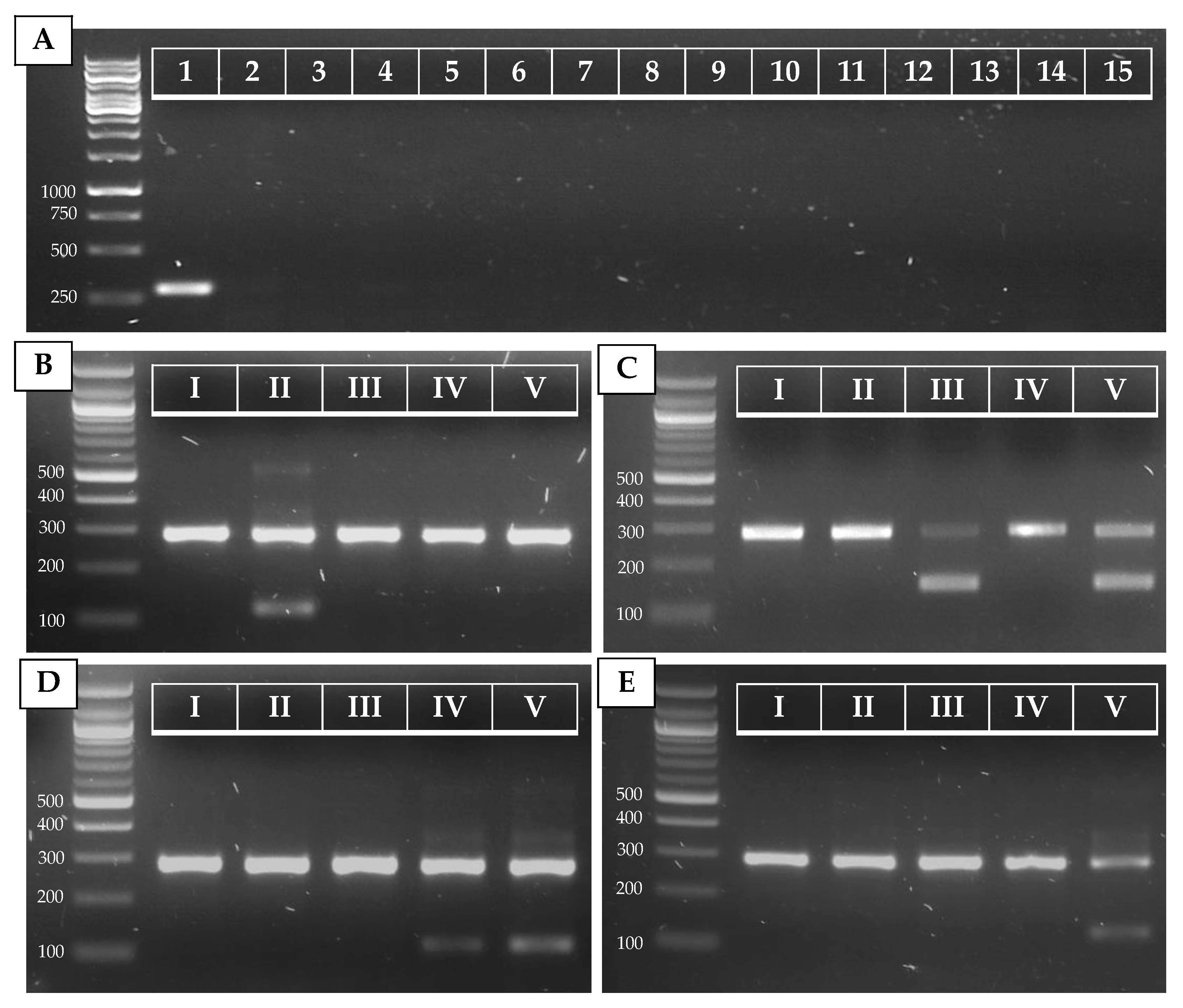
| Primer Name | Sequence | Use (Ta) | Target | Reference |
|---|---|---|---|---|
| ITS1 | TCCGTAGGTGAACCTGCGG | Sequencing (59) AS-PCR (68 or 78) 1 | Fungi | [47] |
| ITS4 | TCCTCCGCTTATTGATATGC | Sequencing (60) | Fungi | [47] |
| ITS_Cau_R | TTTGTGAATGCAACGCCATCG | AS-PCR (78) | C. auris | This study |
| ITS_CauCII_R | GAGATCCGTTGTTGAAAGTTTTCTTTATA | AS-PCR (68) | Clade II | This study |
| RHA1_CauCIII/V_F | TTGCGGTTGAAATGGGTGCT | AS-PCR (68) | Clade III/V | This study |
| RHA1_CauCIII/V_R | TGGCATGTTTCCGGCTTAGA | AS-PCR (68) | Clade III/V | This study |
| ITS_CauCIV/V_R | AGATCCGTTGTTGAAAGTTTTATGTTCTG | AS-PCR (68) | Clade IV/V | This study |
| ITS_CauCV_R | AAGAGATCCGTTGTTGAAAGTTTCCTA | AS-PCR (68) | Clade V | This study |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carolus, H.; Jacobs, S.; Lobo Romero, C.; Deparis, Q.; Cuomo, C.A.; Meis, J.F.; Van Dijck, P. Diagnostic Allele-Specific PCR for the Identification of Candida auris Clades. J. Fungi 2021, 7, 754. https://doi.org/10.3390/jof7090754
Carolus H, Jacobs S, Lobo Romero C, Deparis Q, Cuomo CA, Meis JF, Van Dijck P. Diagnostic Allele-Specific PCR for the Identification of Candida auris Clades. Journal of Fungi. 2021; 7(9):754. https://doi.org/10.3390/jof7090754
Chicago/Turabian StyleCarolus, Hans, Stef Jacobs, Celia Lobo Romero, Quinten Deparis, Christina A. Cuomo, Jacques F. Meis, and Patrick Van Dijck. 2021. "Diagnostic Allele-Specific PCR for the Identification of Candida auris Clades" Journal of Fungi 7, no. 9: 754. https://doi.org/10.3390/jof7090754








