Use of Defensins to Develop Eco-Friendly Alternatives to Synthetic Fungicides to Control Phytopathogenic Fungi and Their Mycotoxins
Abstract
:1. Introduction
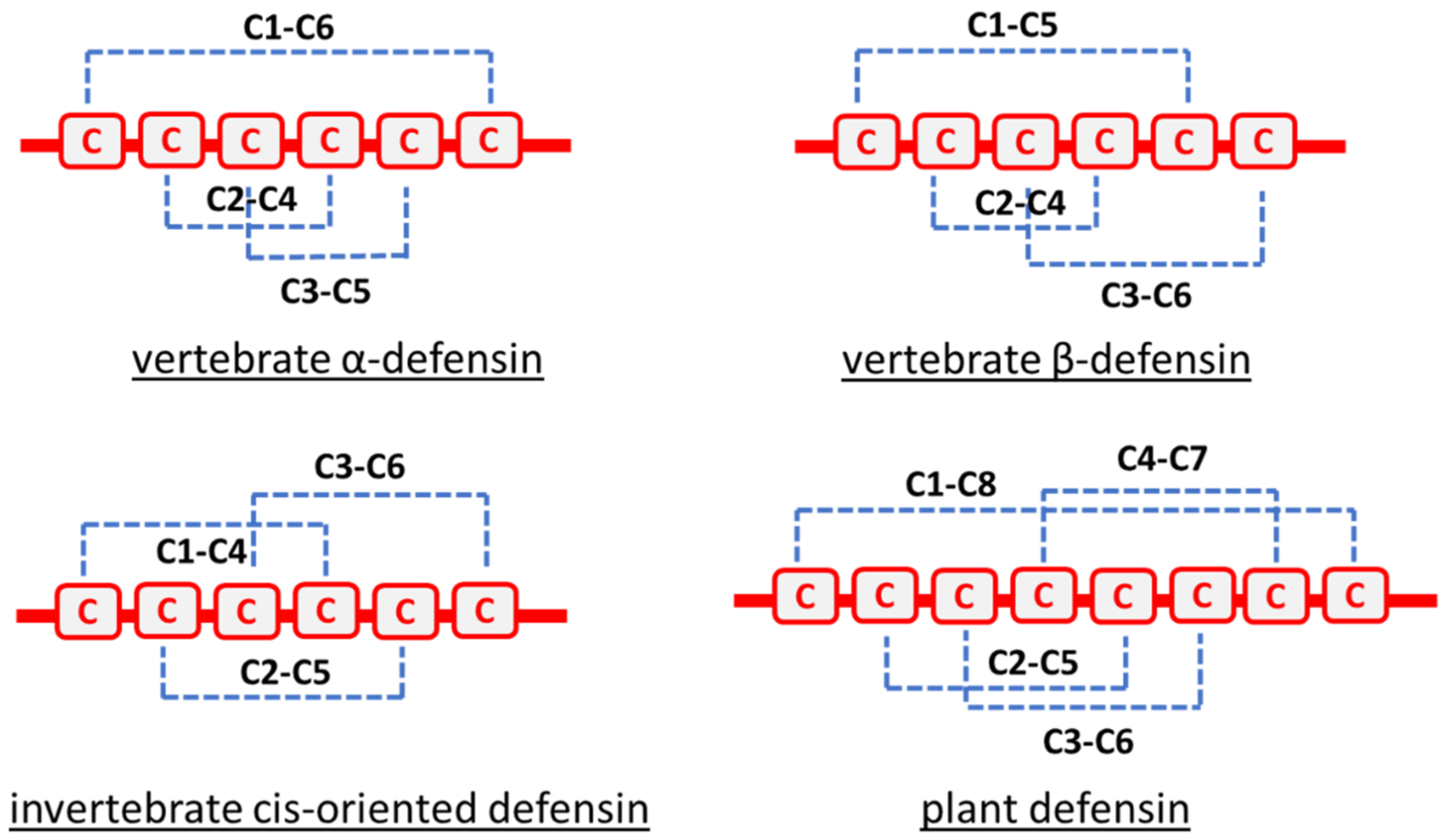
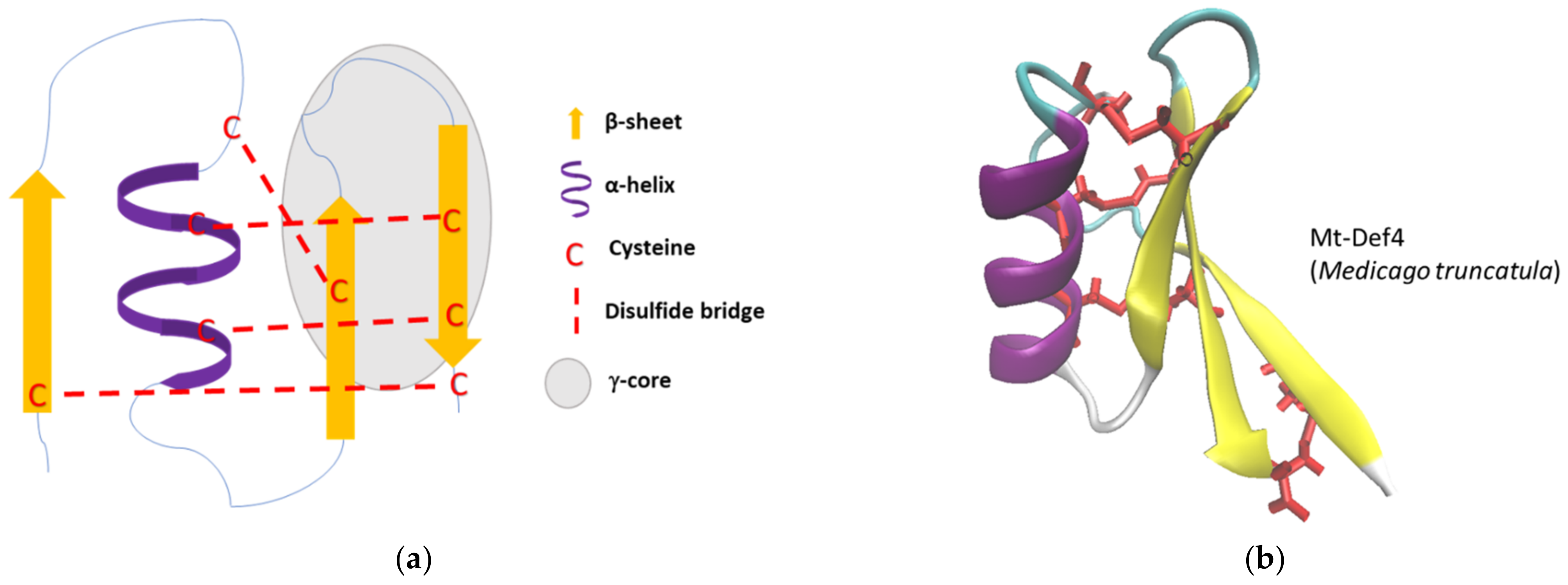
2. Origin and Characteristics of Defensins
3. Activity of Defensins against Fungal Phytopathogens
4. Antifungal Mechanism of Action of Defensins
4.1. Interactions with Host Membrane Components and Induction of Fungal Membranes Disorders
4.2. Induction of Oxidative Stress and Apoptosis
4.3. Internalization and Intracellular Targets
5. Exploiting Defensins to Protect Crops from Phytopathogenic Fungi and Mycotoxin Contamination
5.1. Transgenic Plants Overexpressing Defensin for an Enhanced Resistance to Phytopathogenic Fungi
5.2. Developing Defensin-Based Plant Protection Products for the Control of Phytopathogenic Fungi
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Pennisi, E. Armed and Dangerous. Science 2010, 327, 804–805. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Godfray, H.C.J.; Mason-D’Croz, D.; Robinson, S. Food System Consequences of a Fungal Disease Epidemic in a Major Crop. Philos. Trans. R. Soc. B Biol. Sci. 2016, 371, 20150467. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.; McRoberts, N.; Nelson, A. The Global Burden of Pathogens and Pests on Major Food Crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Fisher, M.C.; Henk, D.A.; Briggs, C.J.; Brownstein, J.S.; Madoff, L.C.; McCraw, S.L.; Gurr, S.J. Emerging Fungal Threats to Animal, Plant and Ecosystem Health. Nature 2012, 484, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Velásquez, A.C.; Castroverde, C.D.M.; He, S.Y. Plant–Pathogen Warfare under Changing Climate Conditions. Curr. Biol. 2018, 28, R619–R634. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fones, H.N.; Bebber, D.P.; Chaloner, T.M.; Kay, W.T.; Steinberg, G.; Gurr, S.J. Threats to Global Food Security from Emerging Fungal and Oomycete Crop Pathogens. Nat. Food 2020, 1, 332–342. [Google Scholar] [CrossRef]
- Dean, R.; Van Kan, J.A.L.; Pretorius, Z.A.; Hammond-Kosack, K.E.; Di Pietro, A.; Spanu, P.D.; Rudd, J.J.; Dickman, M.; Kahmann, R.; Ellis, J.; et al. The Top 10 Fungal Pathogens in Molecular Plant Pathology: Top 10 Fungal Pathogens. Mol. Plant Pathol. 2012, 13, 414–430. [Google Scholar] [CrossRef] [Green Version]
- Perincherry, L.; Lalak-Kańczugowska, J.; Stępień, Ł. Fusarium-Produced Mycotoxins in Plant-Pathogen Interactions. Toxins 2019, 11, 664. [Google Scholar] [CrossRef] [Green Version]
- Marin, S.; Ramos, A.J.; Cano-Sancho, G.; Sanchis, V. Mycotoxins: Occurrence, Toxicology, and Exposure Assessment. Food Chem. Toxicol. 2013, 60, 218–237. [Google Scholar] [CrossRef]
- Bhat, R.; Rai, R.V.; Karim, A.A. Mycotoxins in Food and Feed: Present Status and Future Concerns. Compr. Rev. Food Sci. Food Saf. 2010, 9, 57–81. [Google Scholar] [CrossRef]
- Awuchi, C.G.; Ondari, E.N.; Ogbonna, C.U.; Upadhyay, A.K.; Baran, K.; Okpala, C.O.R.; Korzeniowska, M.; Guiné, R.P.F. Mycotoxins Affecting Animals, Foods, Humans, and Plants: Types, Occurrence, Toxicities, Action Mechanisms, Prevention, and Detoxification Strategies—A Revisit. Foods 2021, 10, 1279. [Google Scholar] [CrossRef] [PubMed]
- Eskola, M.; Kos, G.; Elliott, C.T.; Hajšlová, J.; Mayar, S.; Krska, R. Worldwide Contamination of Food-Crops with Mycotoxins: Validity of the Widely Cited “FAO Estimate” of 25. Crit. Rev. Food Sci. Nutr. 2020, 60, 2773–2789. [Google Scholar] [CrossRef] [PubMed]
- Arya, A.; Perelló, A.E. (Eds.) Management of Fungal Plant Pathogens; CABI: Wallingford, UK, 2010; ISBN 978-1-84593-603-7. [Google Scholar]
- Ghanney, N. Management of Fungal Plants Diseases. Eur. J. Biol. Res. 2017, 7, 309–4314. [Google Scholar] [CrossRef]
- Agrios, G.N. Plant Pathology; Elsevier: Amsterdam, The Netherlands, 1988; ISBN 978-0-12-044563-9. [Google Scholar]
- Oliver, R.P.; Hewitt, H.G. Fungicides in Crop Protection, 2nd ed.; CABI: Boston, MA, USA, 2014; ISBN 978-1-78064-166-9. [Google Scholar]
- Fisher, M.C.; Hawkins, N.J.; Sanglard, D.; Gurr, S.J. Worldwide Emergence of Resistance to Antifungal Drugs Challenges Human Health and Food Security. Science 2018, 360, 739–742. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Steinberg, G.; Gurr, S.J. Fungi, Fungicide Discovery and Global Food Security. Fungal Genet. Biol. 2020, 144, 103476. [Google Scholar] [CrossRef]
- Pellegrino, E.; Bedini, S. Enhancing Ecosystem Services in Sustainable Agriculture: Biofertilization and Biofortification of Chickpea (Cicer Arietinum L.) by Arbuscular Mycorrhizal Fungi. Soil Biol. Biochem. 2014, 68, 429–439. [Google Scholar] [CrossRef]
- Mahanty, T.; Bhattacharjee, S.; Goswami, M.; Bhattacharyya, P.; Das, B.; Ghosh, A.; Tribedi, P. Biofertilizers: A Potential Approach for Sustainable Agriculture Development. Environ. Sci. Pollut. Res. 2017, 24, 3315–3335. [Google Scholar] [CrossRef]
- Balasubramanian, P.; Karthickumar, P. Biofertilizers and Biopesticides: A Holistic Approach for Sustainable Agriculture. In Sustainable Utilization of Natural Resources; Mondal, P., Dalai, A.K., Eds.; CRC Press: Boca Raton, FL, USA, 2017; pp. 255–284. ISBN 978-1-315-15329-2. [Google Scholar]
- Wang, G. Human Antimicrobial Peptides and Proteins. Pharmaceuticals 2014, 7, 545–594. [Google Scholar] [CrossRef] [Green Version]
- Zasloff, M. Antimicrobial Peptides of Multicellular Organisms. Nature 2002, 415, 389–395. [Google Scholar] [CrossRef]
- Lei, J.; Sun, L.; Huang, S.; Zhu, C.; Li, P.; He, J.; Mackey, V.; Coy, D.H.; He, Q. The Antimicrobial Peptides and Their Potential Clinical Applications. Am. J. Transl. Res. 2019, 11, 3919–3931. [Google Scholar]
- Xhindoli, D.; Pacor, S.; Benincasa, M.; Scocchi, M.; Gennaro, R.; Tossi, A. The Human Cathelicidin LL-37—A Pore-Forming Antibacterial Peptide and Host-Cell Modulator. Biochim. Biophys. Acta—Biomembr. 2016, 1858, 546–566. [Google Scholar] [CrossRef] [PubMed]
- Tanner, J.; Deplazes, E.; Mancera, R. The Biological and Biophysical Properties of the Spider Peptide Gomesin. Molecules 2018, 23, 1733. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vergis, J.; Malik, S.S.; Pathak, R.; Kumar, M.; Ramanjaneya, S.; Kurkure, N.V.; Barbuddhe, S.B.; Rawool, D.B. Antimicrobial Efficacy of Indolicidin Against Multi-Drug Resistant Enteroaggregative Escherichia Coli in a Galleria Mellonella Model. Front. Microbiol. 2019, 10, 2723. [Google Scholar] [CrossRef] [PubMed]
- Rebuffat, S. Microcins. In Handbook of Biologically Active Peptides; Elsevier: Amsterdam, The Netherlands, 2013; pp. 129–137. ISBN 978-0-12-385095-9. [Google Scholar]
- Terras, F.R.G.; Torrekens, S.; Van Leuven, F.; Osborn, R.W.; Vanderleyden, J.; Cammue, B.P.A.; Broekaert, W.F. A New Family of Basic Cysteine-Rich Plant Antifungal Proteins from Brassicaceae Species. FEBS Lett. 1993, 316, 233–240. [Google Scholar] [CrossRef] [Green Version]
- Boulanger, N.; Lowenberger, C.; Volf, P.; Ursic, R.; Sigutova, L.; Sabatier, L.; Svobodova, M.; Beverley, S.M.; Späth, G.; Brun, R.; et al. Characterization of a Defensin from the Sand Fly Phlebotomus Duboscqi Induced by Challenge with Bacteria or the Protozoan Parasite Leishmania Major. Infect. Immun. 2004, 72, 7140–7146. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilson, S.S.; Wiens, M.E.; Smith, J.G. Antiviral Mechanisms of Human Defensins. J. Mol. Biol. 2013, 425, 4965–4980. [Google Scholar] [CrossRef]
- Cabezas-Cruz, A.; Tonk, M.; Bouchut, A.; Pierrot, C.; Pierce, R.J.; Kotsyfakis, M.; Rahnamaeian, M.; Vilcinskas, A.; Khalife, J.; Valdés, J.J. Antiplasmodial Activity Is an Ancient and Conserved Feature of Tick Defensins. Front. Microbiol. 2016, 7, 1682. [Google Scholar] [CrossRef]
- Kudryashova, E.; Seveau, S.M.; Kudryashov, D.S. Targeting and Inactivation of Bacterial Toxins by Human Defensins. Biol. Chem. 2017, 398, 1069–1085. [Google Scholar] [CrossRef]
- Couto, J.; Tonk, M.; Ferrolho, J.; Antunes, S.; Vilcinskas, A.; de la Fuente, J.; Domingos, A.; Cabezas-Cruz, A. Antiplasmodial Activity of Tick Defensins in a Mouse Model of Malaria. Ticks Tick-Borne Dis. 2018, 9, 844–849. [Google Scholar] [CrossRef]
- Sathoff, A.E.; Velivelli, S.; Shah, D.M.; Samac, D.A. Plant Defensin Peptides Have Antifungal and Antibacterial Activity Against Human and Plant Pathogens. Phytopathology 2019, 109, 402–408. [Google Scholar] [CrossRef] [Green Version]
- Tonk, M.; Cabezas-Cruz, A.; Valdés, J.J.; Rego, R.O.M.; Grubhoffer, L.; Estrada-Peña, A.; Vilcinskas, A.; Kotsyfakis, M.; Rahnamaeian, M. Ixodes Ricinus Defensins Attack Distantly-Related Pathogens. Dev. Comp. Immunol. 2015, 53, 358–365. [Google Scholar] [CrossRef]
- De Souza Cândido, E.; e Silva Cardoso, M.H.; Sousa, D.A.; Viana, J.C.; de Oliveira-Júnior, N.G.; Miranda, V.; Franco, O.L. The Use of Versatile Plant Antimicrobial Peptides in Agribusiness and Human Health. Peptides 2014, 55, 65–78. [Google Scholar] [CrossRef] [PubMed]
- Falanga, A.; Nigro, E.; De Biasi, M.; Daniele, A.; Morelli, G.; Galdiero, S.; Scudiero, O. Cyclic Peptides as Novel Therapeutic Microbicides: Engineering of Human Defensin Mimetics. Molecules 2017, 22, 1217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wei, H.; Movahedi, A.; Xu, C.; Sun, W.; Li, L.; Li, D.; Zhuge, Q. Characterization, Expression Profiling, and Functional Analysis of a Populus Trichocarpa Defensin Gene and Its Potential as an Anti-Agrobacterium Rooting Medium Additive. Sci. Rep. 2019, 9, 15359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ganz, T.; Selsted, M.E.; Szklarek, D.; Harwig, S.S.; Daher, K.; Bainton, D.F.; Lehrer, R.I. Defensins. Natural Peptide Antibiotics of Human Neutrophils. J. Clin. Investig. 1985, 76, 1427–1435. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.M.; Gonçalves, S.; Santos, N.C. Defensins: Antifungal Lessons from Eukaryotes. Front. Microbiol. 2014, 5, 97. [Google Scholar] [CrossRef] [Green Version]
- Lay, F.; Anderson, M. Defensins—Components of the Innate Immune System in Plants. Curr. Protein Pept. Sci. 2005, 6, 85–101. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Lu, W. Defensins: A Double-Edged Sword in Host Immunity. Front. Immunol. 2020, 11, 764. [Google Scholar] [CrossRef]
- Shafee, T.M.A.; Lay, F.T.; Hulett, M.D.; Anderson, M.A. The Defensins Consist of Two Independent, Convergent Protein Superfamilies. Mol. Biol. Evol. 2016, 33, 2345–2356. [Google Scholar] [CrossRef] [Green Version]
- Shafee, T.M.A.; Lay, F.T.; Phan, T.K.; Anderson, M.A.; Hulett, M.D. Convergent Evolution of Defensin Sequence, Structure and Function. Cell. Mol. Life Sci. 2017, 74, 663–682. [Google Scholar] [CrossRef]
- Gerdol, M.; Schmitt, P.; Venier, P.; Rocha, G.; Rosa, R.D.; Destoumieux-Garzón, D. Functional Insights From the Evolutionary Diversification of Big Defensins. Front. Immunol. 2020, 11, 758. [Google Scholar] [CrossRef] [PubMed]
- Lehrer, R.I.; Cole, A.M.; Selsted, M.E. θ-Defensins: Cyclic Peptides with Endless Potential. J. Biol. Chem. 2012, 287, 27014–27019. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Dias, R.O.; Franco, O.L. Cysteine-Stabilized Aβ Defensins: From a Common Fold to Antibacterial Activity. Peptides 2015, 72, 64–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dimarcq, J.L.; Bulet, P.; Hetru, C.; Hoffmann, J. Cysteine-Rich Antimicrobial Peptides in Invertebrates. Biopolymers 1998, 47, 465–477. [Google Scholar] [CrossRef]
- Dorin, J.R.; McHugh, B.J.; Cox, S.L.; Davidson, D.J. Mammalian Antimicrobial Peptides; Defensins and Cathelicidins. In Molecular Medical Microbiology; Elsevier: Amsterdam, The Netherlands, 2015; pp. 539–565. ISBN 978-0-12-397169-2. [Google Scholar]
- Picart, P.; Pirttilä, A.; Raventos, D.; Kristensen, H.-H.; Sahl, H.-G. Identification of Defensin-Encoding Genes of Picea Glauca: Characterization of PgD5, a Conserved Spruce Defensin with Strong Antifungal Activity. BMC Plant Biol. 2012, 12, 180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chan, Y.S.; Ng, T.B. Northeast Red Beans Produce a Thermostable and PH-Stable Defensin-Like Peptide with Potent Antifungal Activity. Cell Biochem. Biophys. 2013, 66, 637–648. [Google Scholar] [CrossRef]
- Maemoto, A.; Qu, X.; Rosengren, K.J.; Tanabe, H.; Henschen-Edman, A.; Craik, D.J.; Ouellette, A.J. Functional Analysis of the α-Defensin Disulfide Array in Mouse Cryptdin-4. J. Biol. Chem. 2004, 279, 44188–44196. [Google Scholar] [CrossRef] [Green Version]
- White, S.H.; Wimley, W.C.; Selsted, M.E. Structure, Function, and Membrane Integration of Defensins. Curr. Opin. Struct. Biol. 1995, 5, 521–527. [Google Scholar] [CrossRef]
- De Samblanx, G.W.; Goderis, I.J.; Thevissen, K.; Raemaekers, R.; Fant, F.; Borremans, F.; Acland, D.P.; Osborn, R.W.; Patel, S.; Broekaert, W.F. Mutational Analysis of a Plant Defensin from Radish (Raphanus sativus L.) Reveals Two Adjacent Sites Important for Antifungal Activity. J. Biol. Chem. 1997, 272, 1171–1179. [Google Scholar] [CrossRef] [Green Version]
- Sagaram, U.S.; Pandurangi, R.; Kaur, J.; Smith, T.J.; Shah, D.M. Structure-Activity Determinants in Antifungal Plant Defensins MsDef1 and MtDef4 with Different Modes of Action against Fusarium graminearum. PLoS ONE 2011, 6, e18550. [Google Scholar] [CrossRef] [Green Version]
- De Coninck, B.; Cammue, B.P.A.; Thevissen, K. Modes of Antifungal Action and in Planta Functions of Plant Defensins and Defensin-like Peptides. Fungal Biol. Rev. 2013, 26, 109–120. [Google Scholar] [CrossRef]
- Contreras, G.; Shirdel, I.; Braun, M.S.; Wink, M. Defensins: Transcriptional Regulation and Function beyond Antimicrobial Activity. Dev. Comp. Immunol. 2020, 104, 103556. [Google Scholar] [CrossRef]
- Wilson, C.L.; Schmidt, A.P.; Pirilä, E.; Valore, E.V.; Ferri, N.; Sorsa, T.; Ganz, T.; Parks, W.C. Differential Processing of α- and β-Defensin Precursors by Matrix Metalloproteinase-7 (MMP-7). J. Biol. Chem. 2009, 284, 8301–8311. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lay, F.T.; Poon, S.; McKenna, J.A.; Connelly, A.A.; Barbeta, B.L.; McGinness, B.S.; Fox, J.L.; Daly, N.L.; Craik, D.J.; Heath, R.L.; et al. The C-Terminal Propeptide of a Plant Defensin Confers Cytoprotective and Subcellular Targeting Functions. BMC Plant Biol. 2014, 14, 41. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hegedüs, N.; Marx, F. Antifungal Proteins: More than Antimicrobials? Fungal Biol. Rev. 2013, 26, 132–145. [Google Scholar] [CrossRef]
- Kragh, K.M. Characterization and Localization of New Antifungal Cysteine-Rich Proteins from Beta vulgaris. Mol. Plant Microb. Interact. 1995, 8, 424. [Google Scholar] [CrossRef] [PubMed]
- Osborn, R.W.; De Samblanx, G.W.; Thevissen, K.; Goderis, I.; Torrekens, S.; Van Leuven, F.; Attenborough, S.; Rees, S.B.; Broekaert, W.F. Isolation and Characterisation of Plant Defensins from Seeds of Asteraceae, Fabaceae, Hippocastanaceae and Saxifragaceae. FEBS Lett. 1995, 368, 257–262. [Google Scholar] [CrossRef] [Green Version]
- Marquès, L.; Oomen, R.J.F.J.; Aumelas, A.; Le Jean, M.; Berthomieu, P. Production of an Arabidopsis Halleri Foliar Defensin in Escherichia Coli. J. Appl. Microbiol. 2009, 106, 1640–1648. [Google Scholar] [CrossRef]
- Vriens, K.; Peigneur, S.; De Coninck, B.; Tytgat, J.; Cammue, B.P.A.; Thevissen, K. The Antifungal Plant Defensin AtPDF2.3 from Arabidopsis Thaliana Blocks Potassium Channels. Sci. Rep. 2016, 6, 32121. [Google Scholar] [CrossRef] [Green Version]
- De Beer, A.; Vivier, M.A. Four Plant Defensins from an Indigenous South African Brassicaceae Species Display Divergent Activities against Two Test Pathogens despite High Sequence Similarity in the Encoding Genes. BMC Res. Notes 2011, 4, 459. [Google Scholar] [CrossRef] [Green Version]
- Terras, F.R.; Schoofs, H.M.; De Bolle, M.F.; Van Leuven, F.; Rees, S.B.; Vanderleyden, J.; Cammue, B.P.; Broekaert, W.F. Analysis of Two Novel Classes of Plant Antifungal Proteins from Radish (Raphanus sativus L.) Seeds. J. Biol. Chem. 1992, 267, 15301–15309. [Google Scholar] [CrossRef]
- Terras, F.R.; Eggermont, K.; Kovaleva, V.; Raikhel, N.V.; Osborn, R.W.; Kester, A.; Rees, S.B.; Torrekens, S.; Van Leuven, F.; Vanderleyden, J. Small Cysteine-Rich Antifungal Proteins from Radish: Their Role in Host Defense. Plant Cell 1995, 7, 573–588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saitoh, H.; Kiba, A.; Nishihara, M.; Yamamura, S.; Suzuki, K.; Terauchi, R. Production of Antimicrobial Defensin in Nicotiana Benthamiana with a Potato Virus X Vector. Mol. Plant-Microbe Interact. 2001, 14, 111–115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Slavokhotova, A.A.; Odintsova, T.I.; Rogozhin, E.A.; Musolyamov, A.K.; Andreev, Y.A.; Grishin, E.V.; Egorov, T.A. Isolation, Molecular Cloning and Antimicrobial Activity of Novel Defensins from Common Chickweed (Stellaria media L.) Seeds. Biochimie 2011, 93, 450–456. [Google Scholar] [CrossRef]
- Segura, A.; Moreno, M.; Molina, A.; García-Olmedo, F. Novel Defensin Subfamily from Spinach (Spinacia oleracea). FEBS Lett. 1998, 435, 159–162. [Google Scholar] [CrossRef] [Green Version]
- Ye, X.Y.; Ng, T.B. Peptides from Pinto Bean and Red Bean with Sequence Homology to Cowpea 10-KDa Protein Precursor Exhibit Antifungal, Mitogenic, and HIV-1 Reverse Transcriptase-Inhibitory Activities. Biochem. Biophys. Res. Commun. 2001, 285, 424–429. [Google Scholar] [CrossRef]
- Huang, J.; Wong, K.H.; Tay, S.V.; Serra, A.; Sze, S.K.; Tam, J.P. Astratides: Insulin-Modulating, Insecticidal, and Antifungal Cysteine-Rich Peptides from Astragalus membranaceus. J. Nat. Prod. 2019, 82, 194–204. [Google Scholar] [CrossRef]
- Ngai, P.H.K.; Ng, T.B. Coccinin, an Antifungal Peptide with Antiproliferative and HIV-1 Reverse Transcriptase Inhibitory Activities from Large Scarlet Runner Beans. Peptides 2004, 25, 2063–2068. [Google Scholar] [CrossRef]
- Ngai, P.H.K.; Ng, T.B. Phaseococcin, an Antifungal Protein with Antiproliferative and Anti-HIV-1 Reverse Transcriptase Activities from Small Scarlet Runner Beans. Biochem. Cell Biol. 2005, 83, 212–220. [Google Scholar] [CrossRef]
- Wong, J.H.; Ng, T.B. Gymnin, a Potent Defensin-like Antifungal Peptide from the Yunnan Bean (Gymnocladus Chinensis Baill). Peptides 2003, 24, 963–968. [Google Scholar] [CrossRef]
- Shenkarev, Z.O.; Gizatullina, A.K.; Finkina, E.I.; Alekseeva, E.A.; Balandin, S.V.; Mineev, K.S.; Arseniev, A.S.; Ovchinnikova, T.V. Heterologous Expression and Solution Structure of Defensin from Lentil Lens Culinaris. Biochem. Biophys. Res. Commun. 2014, 451, 252–257. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.H.; Ng, T.B. Limenin, a Defensin-like Peptide with Multiple Exploitable Activities from Shelf Beans. J. Pept. Sci. Off. Publ. Eur. Pept. Soc. 2006, 12, 341–346. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Rao, P.; Ye, X. Isolation and Biochemical Characterization of a Novel Leguminous Defense Peptide with Antifungal and Antiproliferative Potency. Appl. Microbiol. Biotechnol. 2009, 82, 79–86. [Google Scholar] [CrossRef]
- Spelbrink, R.G.; Dilmac, N.; Allen, A.; Smith, T.J.; Shah, D.M.; Hockerman, G.H. Differential Antifungal and Calcium Channel-Blocking Activity among Structurally Related Plant Defensins. Plant Physiol. 2004, 135, 2055–2067. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, A.G.; Hakimi, S.M.; Mittanck, C.A.; Wu, Y.; Woerner, B.M.; Stark, D.M.; Shah, D.M.; Liang, J.; Rommens, C.M. Fungal Pathogen Protection in Potato by Expression of a Plant Defensin Peptide. Nat. Biotechnol. 2000, 18, 1307–1310. [Google Scholar] [CrossRef] [PubMed]
- Almeida, M.S.; Cabral, K.M.S.; Zingali, R.B.; Kurtenbach, E. Characterization of Two Novel Defense Peptides from Pea (Pisum Sativum) Seeds. Arch. Biochem. Biophys. 2000, 378, 278–286. [Google Scholar] [CrossRef] [PubMed]
- Games, P.D.; dos Santos, I.S.; Mello, É.O.; Diz, M.S.S.; Carvalho, A.O.; de Souza-Filho, G.A.; Da Cunha, M.; Vasconcelos, I.M.; dos Ferreira, B.S.; Gomes, V.M. Isolation, Characterization and Cloning of a CDNA Encoding a New Antifungal Defensin from Phaseolus Vulgaris L. Seeds. Peptides 2008, 29, 2090–2100. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.H.; Zhang, X.Q.; Wang, H.X.; Ng, T.B. A Mitogenic Defensin from White Cloud Beans (Phaseolus Vulgaris). Peptides 2006, 27, 2075–2081. [Google Scholar] [CrossRef]
- Wong, J.H.; Ng, T.B. Sesquin, a Potent Defensin-like Antimicrobial Peptide from Ground Beans with Inhibitory Activities toward Tumor Cells and HIV-1 Reverse Transcriptase. Peptides 2005, 26, 1120–1126. [Google Scholar] [CrossRef]
- Song, X.; Wang, J.; Wu, F.; Li, X.; Teng, M.; Gong, W. CDNA Cloning, Functional Expression and Antifungal Activities of a Dimeric Plant Defensin SPE10 from Pachyrrhizus Erosus Seeds. Plant Mol. Biol. 2005, 57, 13–20. [Google Scholar] [CrossRef]
- Vijayan, S.; Guruprasad, L.; Kirti, P.B. Prokaryotic Expression of a Constitutively Expressed Tephrosia Villosa Defensin and Its Potent Antifungal Activity. Appl. Microbiol. Biotechnol. 2008, 80, 1023–1032. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.-H.; Hsu, M.-P.; Tan, C.-H.; Sung, H.-Y.; Kuo, C.G.; Fan, M.-J.; Chen, H.-M.; Chen, S.; Chen, C.-S. Cloning and Characterization of a Plant Defensin VaD1 from Azuki Bean. J. Agric. Food Chem. 2005, 53, 982–988. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.-J.; Chen, G.-H.; Hsu, H.-C.; Li, S.-S.; Chen, C.-S. Cloning and Functional Expression of a Mungbean Defensin VrD1 in Pichia Pastoris. J. Agric. Food Chem. 2004, 52, 2256–2261. [Google Scholar] [CrossRef] [PubMed]
- Kovaleva, V.; Krynytskyy, H.; Gout, I.; Gout, R. Recombinant Expression, Affinity Purification and Functional Characterization of Scots Pine Defensin 1. Appl. Microbiol. Biotechnol. 2011, 89, 1093–1101. [Google Scholar] [CrossRef] [PubMed]
- Odintsova, T.I.; Rogozhin, E.A.; Baranov, Y.; Musolyamov, A.K.; Yalpani, N.; Egorov, T.A.; Grishin, E.V. Seed Defensins of Barnyard Grass Echinochloa Crusgalli (L.) Beauv. Biochimie 2008, 90, 1667–1673. [Google Scholar] [CrossRef]
- Fujimura, M.; Ideguchi, M.; Minami, Y.; Watanabe, K.; Tadera, K. Amino Acid Sequence and Antimicrobial Activity of Chitin-Binding Peptides, Pp -AMP 1 and Pp -AMP 2, from Japanese Bamboo Shoots (Phyllostachys pubescens). Biosci. Biotechnol. Biochem. 2005, 69, 642–645. [Google Scholar] [CrossRef] [Green Version]
- De-Paula, V.S.; Razzera, G.; Medeiros, L.; Miyamoto, C.A.; Almeida, M.S.; Kurtenbach, E.; Almeida, F.C.L.; Valente, A.P. Evolutionary Relationship between Defensins in the Poaceae Family Strengthened by the Characterization of New Sugarcane Defensins. Plant Mol. Biol. 2008, 68, 321–335. [Google Scholar] [CrossRef]
- Kerenga, B.K.; McKenna, J.A.; Harvey, P.J.; Quimbar, P.; Garcia-Ceron, D.; Lay, F.T.; Phan, T.K.; Veneer, P.K.; Vasa, S.; Parisi, K.; et al. Salt-Tolerant Antifungal and Antibacterial Activities of the Corn Defensin ZmD32. Front. Microbiol. 2019, 10, 795. [Google Scholar] [CrossRef]
- Balandín, M.; Royo, J.; Gómez, E.; Muniz, L.M.; Molina, A.; Hueros, G. A Protective Role for the Embryo Surrounding Region of the Maize Endosperm, as Evidenced by the Characterisation of ZmESR-6, a Defensin Gene Specifically Expressed in This Region. Plant Mol. Biol. 2005, 58, 269–282. [Google Scholar] [CrossRef]
- Fujimura, M.; Minami, Y.; Watanabe, K.; Tadera, K. Purification, Characterization, and Sequencing of a Novel Type of Antimicrobial Peptides, Fa -AMP1 and Fa -AMP2, from Seeds of Buckwheat (Fagopyrum esculentum Moench.). Biosci. Biotechnol. Biochem. 2003, 67, 1636–1642. [Google Scholar] [CrossRef]
- Rogozhin, E.A.; Oshchepkova, Y.I.; Odintsova, T.I.; Khadeeva, N.V.; Veshkurova, O.N.; Egorov, T.A.; Grishin, E.V.; Salikhov, S.I. Novel Antifungal Defensins from Nigella sativa L. Seeds. Plant Physiol. Biochem. 2011, 49, 131–137. [Google Scholar] [CrossRef]
- Bleackley, M.R.; Payne, J.A.E.; Hayes, B.M.E.; Durek, T.; Craik, D.J.; Shafee, T.M.A.; Poon, I.K.H.; Hulett, M.D.; van der Weerden, N.L.; Anderson, M.A. Nicotiana alata Defensin Chimeras Reveal Differences in the Mechanism of Fungal and Tumor Cell Killing and an Enhanced Antifungal Variant. Antimicrob. Agents Chemother. 2016, 60, 6302–6312. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dracatos, P.M.; Weerden, N.L.; Carroll, K.T.; Johnson, E.D.; Plummer, K.M.; Anderson, M.A. Inhibition of Cereal Rust Fungi by Both Class I and II Defensins Derived from the Flowers of NIcotiana alata. Mol. Plant Pathol. 2014, 15, 67–79. [Google Scholar] [CrossRef] [PubMed]
- Lay, F.T.; Brugliera, F.; Anderson, M.A. Isolation and Properties of Floral Defensins from Ornamental Tobacco and Petunia. Plant Physiol. 2003, 131, 1283–1293. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Beer, A.; Vivier, M.A. Vv-AMP1, a Ripening Induced Peptide from Vitis vinifera Shows Strong Antifungal Activity. BMC Plant Biol. 2008, 8, 75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Landon, C. Lead Optimization of Antifungal Peptides with 3D NMR Structures Analysis. Protein Sci. 2004, 13, 703–713. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fehlbaum, P.; Bulet, P.; Michaut, L.; Lagueux, M.; Broekaert, W.F.; Hetru, C.; Hoffmann, J.A. Insect Immunity. Septic Injury of Drosophila Induces the Synthesis of a Potent Antifungal Peptide with Sequence Homology to Plant Antifungal Peptides. J. Biol. Chem. 1994, 269, 33159–33163. [Google Scholar] [CrossRef]
- Cytryńska, M.; Mak, P.; Zdybicka-Barabas, A.; Suder, P.; Jakubowicz, T. Purification and Characterization of Eight Peptides from Galleria Mellonella Immune Hemolymph. Peptides 2007, 28, 533–546. [Google Scholar] [CrossRef]
- Lamberty, M.; Ades, S.; Uttenweiler-Joseph, S.; Brookhart, G.; Bushey, D.; Hoffmann, J.A.; Bulet, P. Insect Immunity. Isolation from the Lepidopteran Heliothis Virescens of a Novel Insect Defensin with Potent Antifungal Activity. J. Biol. Chem. 1999, 274, 9320–9326. [Google Scholar] [CrossRef] [Green Version]
- Lamberty, M.; Zachary, D.; Lanot, R.; Bordereau, C.; Robert, A.; Hoffmann, J.A.; Bulet, P. Insect Immunity. Constitutive Expression of a Cysteine-Rich Antifungal and a Linear Antibacterial Peptide in a Termite Insect. J. Biol. Chem. 2001, 276, 4085–4092. [Google Scholar] [CrossRef] [Green Version]
- Xu, X.-X.; Zhang, Y.-Q.; Freed, S.; Yu, J.; Gao, Y.-F.; Wang, S.; Ouyang, L.-N.; Ju, W.-Y.; Jin, F.-L. An Anionic Defensin from Plutella Xylostella with Potential Activity against Bacillus Thuringiensis. Bull. Entomol. Res. 2016, 106, 790–800. [Google Scholar] [CrossRef] [PubMed]
- Bíliková, K.; Wu, G.; Simúth, J. Isolation of a Peptide Fraction from Honeybee Royal Jelly as a Potential Antifoulbrood Factor. Apidologie 2001, 32, 275–283. [Google Scholar] [CrossRef] [Green Version]
- Gueguen, Y.; Herpin, A.; Aumelas, A.; Garnier, J.; Fievet, J.; Escoubas, J.-M.; Bulet, P.; Gonzalez, M.; Lelong, C.; Favrel, P.; et al. Characterization of a Defensin from the Oyster Crassostrea Gigas: Recombinant Production, Folding, Solution Structure, Antimicrobial Activities, and Gene Expression. J. Biol. Chem. 2006, 281, 313–323. [Google Scholar] [CrossRef] [Green Version]
- Romestand, B.; Molina, F.; Richard, V.; Roch, P.; Granier, C. Key Role of the Loop Connecting the Two Beta Strands of Mussel Defensin in Its Antimicrobial Activity. Eur. J. Biochem. 2003, 270, 2805–2813. [Google Scholar] [CrossRef] [PubMed]
- Cabezas-Cruz, A.; Tonk, M.; Bleackley, M.R.; Valdés, J.J.; Barrero, R.A.; Hernández-Jarguín, A.; Moutailler, S.; Vilcinskas, A.; Richard-Forget, F.; Anderson, M.A.; et al. Antibacterial and Antifungal Activity of Defensins from the Australian Paralysis Tick, Ixodes Holocyclus. Ticks Tick-Borne Dis. 2019, 10, 101269. [Google Scholar] [CrossRef]
- Ayroza, G.; Ferreira, I.L.C.; Sayegh, R.S.R.; Tashima, A.K.; da Silva Junior, P.I. Juruin: An Antifungal Peptide from the Venom of the Amazonian Pink Toe Spider, Avicularia Juruensis, Which Contains the Inhibitory Cystine Knot Motif. Front. Microbiol. 2012, 3, 324. [Google Scholar] [CrossRef] [Green Version]
- Tonk, M.; Cabezas-Cruz, A.; Valdés, J.J.; Rego, R.O.; Chrudimská, T.; Strnad, M.; Šíma, R.; Bell-Sakyi, L.; Franta, Z.; Vilcinskas, A.; et al. Defensins from the Tick Ixodes Scapularis Are Effective against Phytopathogenic Fungi and the Human Bacterial Pathogen Listeria grayi. Parasit. Vectors 2014, 7, 554. [Google Scholar] [CrossRef]
- Yang, M.; Chen, B.; Cai, J.-J.; Peng, H.; Ling-Cai; Yuan, J.-J.; Wang, K.-J. Molecular Characterization of Hepcidin AS-Hepc2 and AS-Hepc6 in Black Porgy (Acanthopagrus schlegelii): Expression Pattern Responded to Bacterial Challenge and in Vitro Antimicrobial Activity. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2011, 158, 155–163. [Google Scholar] [CrossRef]
- Lauth, X.; Babon, J.J.; Stannard, J.A.; Singh, S.; Nizet, V.; Carlberg, J.M.; Ostland, V.E.; Pennington, M.W.; Norton, R.S.; Westerman, M.E. Bass Hepcidin Synthesis, Solution Structure, Antimicrobial Activities and Synergism, and in Vivo Hepatic Response to Bacterial Infections. J. Biol. Chem. 2005, 280, 9272–9282. [Google Scholar] [CrossRef] [Green Version]
- Yamane, E.S.; Bizerra, F.C.; Oliveira, E.B.; Moreira, J.T.; Rajabi, M.; Nunes, G.L.C.; de Souza, A.O.; da Silva, I.D.C.G.; Yamane, T.; Karpel, R.L.; et al. Unraveling the Antifungal Activity of a South American Rattlesnake Toxin Crotamine. Biochimie 2013, 95, 231–240. [Google Scholar] [CrossRef] [Green Version]
- Thouzeau, C.; Le Maho, Y.; Froget, G.; Sabatier, L.; Le Bohec, C.; Hoffmann, J.A.; Bulet, P. Spheniscins, Avian β-Defensins in Preserved Stomach Contents of the King Penguin, Aptenodytes patagonicus. J. Biol. Chem. 2003, 278, 51053–51058. [Google Scholar] [CrossRef] [Green Version]
- Simon, A.; Kullberg, B.J.; Tripet, B.; Boerman, O.C.; Zeeuwen, P.; van der Ven-Jongekrijg, J.; Verweij, P.; Schalkwijk, J.; Hodges, R.; van der Meer, J.W.M.; et al. Drosomycin-Like Defensin, a Human Homologue of Drosophila Melanogaster Drosomycin with Antifungal Activity. Antimicrob. Agents Chemother. 2008, 52, 1407–1412. [Google Scholar] [CrossRef] [Green Version]
- Theis, T.; Wedde, M.; Meyer, V.; Stahl, U. The Antifungal Protein from Aspergillus Giganteus Causes Membrane Permeabilization. Antimicrob. Agents Chemother. 2003, 47, 588–593. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huber, A.; Hajdu, D.; Bratschun-Khan, D.; Gáspári, Z.; Varbanov, M.; Philippot, S.; Fizil, Á.; Czajlik, A.; Kele, Z.; Sonderegger, C.; et al. New Antimicrobial Potential and Structural Properties of PAFB: A Cationic, Cysteine-Rich Protein from Penicillium Chrysogenum Q176. Sci. Rep. 2018, 8, 1751. [Google Scholar] [CrossRef] [PubMed]
- Gun Lee, D.; Shin, S.Y.; Maeng, C.Y.; Jin, Z.Z.; Kim, K.L.; Hahm, K.S. Isolation and Characterization of a Novel Antifungal Peptide from Aspergillus niger. Biochem. Biophys. Res. Commun. 1999, 263, 646–651. [Google Scholar] [CrossRef]
- Skouri-Gargouri, H.; Gargouri, A. First Isolation of a Novel Thermostable Antifungal Peptide Secreted by Aspergillus Clavatus. Peptides 2008, 29, 1871–1877. [Google Scholar] [CrossRef]
- Kovács, L.; Virágh, M.; Takó, M.; Papp, T.; Vágvölgyi, C.; Galgóczy, L. Isolation and Characterization of Neosartorya fischeri Antifungal Protein (NFAP). Peptides 2011, 32, 1724–1731. [Google Scholar] [CrossRef]
- Mielniczuk, E.; Skwaryło-Bednarz, B. Fusarium Head Blight, Mycotoxins and Strategies for Their Reduction. Agronomy 2020, 10, 509. [Google Scholar] [CrossRef] [Green Version]
- Varga, J.; Baranyi, N.; Muthusamy, C.; Vágvölgyi, C.; Kocsubé, S. Mycotoxin Producers in the Aspergillus Genus: An Update. Acta Biol. Szeged. 2015, 59, 151–167. [Google Scholar]
- Lacerda, A.F.; Vasconcelos, Ã.A.R.; Pelegrini, P.B.; Grossi de Sa, M.F. Antifungal Defensins and Their Role in Plant Defense. Front. Microbiol. 2014, 5, 116. [Google Scholar] [CrossRef] [Green Version]
- Leannec-Rialland, V.; Cabezas-Cruz, A.; Atanasova, V.; Chereau, S.; Ponts, N.; Tonk, M.; Vilcinskas, A.; Ferrer, N.; Valdés, J.J.; Richard-Forget, F. Tick Defensin γ-Core Reduces Fusarium Graminearum Growth and Abrogates Mycotoxins Production with High Efficiency. Sci. Rep. 2021, 11, 7962. [Google Scholar] [CrossRef]
- Zhang, Z.-T.; Zhu, S.-Y. Drosomycin, an Essential Component of Antifungal Defence in Drosophila. Insect Mol. Biol. 2009, 18, 549–556. [Google Scholar] [CrossRef] [PubMed]
- Tian, C.; Gao, B.; del Rodriguez, M.C.; Lanz-Mendoza, H.; Ma, B.; Zhu, S. Gene Expression, Antiparasitic Activity, and Functional Evolution of the Drosomycin Family. Mol. Immunol. 2008, 45, 3909–3916. [Google Scholar] [CrossRef]
- Broekaert, W.F.; Terras, F.; Cammue, B.; Osborn, R.W. Plant Defensins: Novel Antimicrobial Peptides as Components of the Host Defense System. Plant Physiol. 1995, 108, 1353–1358. [Google Scholar] [CrossRef] [PubMed]
- Thomma, B.P.H.J.; Cammue, B.P.A.; Thevissen, K. Mode of Action of Plant Defensins Suggests Therapeutic Potential. Curr. Drug Targets Infect. Disord. 2003, 3, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Vriens, K.; Cammue, B.; Thevissen, K. Antifungal Plant Defensins: Mechanisms of Action and Production. Molecules 2014, 19, 12280–12303. [Google Scholar] [CrossRef] [Green Version]
- Aumer, T.; Voisin, S.N.; Knobloch, T.; Landon, C.; Bulet, P. Impact of an Antifungal Insect Defensin on the Proteome of the Phytopathogenic Fungus Botrytis Cinerea. J. Proteome Res. 2020, 19, 1131–1146. [Google Scholar] [CrossRef] [PubMed]
- Parisi, K.; Shafee, T.M.A.; Quimbar, P.; van der Weerden, N.L.; Bleackley, M.R.; Anderson, M.A. The Evolution, Function and Mechanisms of Action for Plant Defensins. Semin. Cell Dev. Biol. 2019, 88, 107–118. [Google Scholar] [CrossRef]
- Struyfs, C.; Cammue, B.P.A.; Thevissen, K. Membrane-Interacting Antifungal Peptides. Front. Cell Dev. Biol. 2021, 9, 706. [Google Scholar] [CrossRef]
- El-Mounadi, K.; Islam, K.T.; Hernández-Ortiz, P.; Read, N.D.; Shah, D.M. Antifungal Mechanisms of a Plant Defensin MtDef4 Are Not Conserved between the Ascomycete Fungi Neurospora crassa and Fusarium graminearum. Mol. Microbiol. 2016, 100, 542–559. [Google Scholar] [CrossRef] [Green Version]
- Thevissen, K.; Francois, I.E.; Aerts, A.; Cammue, B. Fungal Sphingolipids as Targets for the Development of Selective Antifungal Therapeutics. Curr. Drug Targets 2005, 6, 923–928. [Google Scholar] [CrossRef] [PubMed]
- Aerts, A.M.; François, I.E.J.A.; Bammens, L.; Cammue, B.P.A.; Smets, B.; Winderickx, J.; Accardo, S.; De Vos, D.E.; Thevissen, K. Level of M(IP)2C Sphingolipid Affects Plant Defensin Sensitivity, Oxidative Stress Resistance and Chronological Life-Span in Yeast. FEBS Lett. 2006, 580, 1903–1907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramamoorthy, V.; Cahoon, E.B.; Li, J.; Thokala, M.; Minto, R.E.; Shah, D.M. Glucosylceramide Synthase Is Essential for Alfalfa Defensin-Mediated Growth Inhibition but Not for Pathogenicity of Fusarium graminearum. Mol. Microbiol. 2007, 66, 771–786. [Google Scholar] [CrossRef] [PubMed]
- De Paula, V.S.; Razzera, G.; Barreto-Bergter, E.; Almeida, F.C.L.; Valente, A.P. Portrayal of Complex Dynamic Properties of Sugarcane Defensin 5 by NMR: Multiple Motions Associated with Membrane Interaction. Structure 2011, 19, 26–36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thevissen, K.; de Mello Tavares, P.; Xu, D.; Blankenship, J.; Vandenbosch, D.; Idkowiak-Baldys, J.; Govaert, G.; Bink, A.; Rozental, S.; de Groot, P.W.J.; et al. The Plant Defensin RsAFP2 Induces Cell Wall Stress, Septin Mislocalization and Accumulation of Ceramides in Candida albicans: RsAFP2 Affects C. Albicans Cell Wall and Septin. Mol. Microbiol. 2012, 84, 166–180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fernandes, C.M.; de Castro, P.A.; Singh, A.; Fonseca, F.L.; Pereira, M.D.; Vila, T.V.M.; Atella, G.C.; Rozental, S.; Savoldi, M.; Del Poeta, M.; et al. Functional Characterization of the A Spergillus nidulans Glucosylceramide Pathway Reveals That LCB Δ8-Desaturation and C9-Methylation Are Relevant to Filamentous Growth, Lipid Raft Localization and Ps D1 Defensin Activity: Aspergillus nidulans Glucosylceramide Pathway. Mol. Microbiol. 2016, 102, 488–505. [Google Scholar] [CrossRef] [Green Version]
- Sagaram, U.S.; El-Mounadi, K.; Buchko, G.W.; Berg, H.R.; Kaur, J.; Pandurangi, R.S.; Smith, T.J.; Shah, D.M. Structural and Functional Studies of a Phosphatidic Acid-Binding Antifungal Plant Defensin MtDef4: Identification of an RGFRRR Motif Governing Fungal Cell Entry. PLoS ONE 2013, 8, e82485. [Google Scholar] [CrossRef] [Green Version]
- Poon, I.K.; Baxter, A.A.; Lay, F.T.; Mills, G.D.; Adda, C.G.; Payne, J.A.; Phan, T.K.; Ryan, G.F.; White, J.A.; Veneer, P.K.; et al. Phosphoinositide-Mediated Oligomerization of a Defensin Induces Cell Lysis. eLife 2014, 3, e01808. [Google Scholar] [CrossRef]
- Baxter, A.A.; Richter, V.; Lay, F.T.; Poon, I.K.H.; Adda, C.G.; Veneer, P.K.; Phan, T.K.; Bleackley, M.R.; Anderson, M.A.; Kvansakul, M.; et al. The Tomato Defensin TPP3 Binds Phosphatidylinositol (4,5)-Bisphosphate via a Conserved Dimeric Cationic Grip Conformation To Mediate Cell Lysis. Mol. Cell. Biol. 2015, 35, 1964–1978. [Google Scholar] [CrossRef] [Green Version]
- De Medeiros, L.N.; Angeli, R.; Sarzedas, C.G.; Barreto-Bergter, E.; Valente, A.P.; Kurtenbach, E.; Almeida, F.C.L. Backbone Dynamics of the Antifungal Psd1 Pea Defensin and Its Correlation with Membrane Interaction by NMR Spectroscopy. Biochim. Biophys. Acta—Biomembr. 2010, 1798, 105–113. [Google Scholar] [CrossRef] [Green Version]
- Aerts, A.M.; François, I.E.J.A.; Cammue, B.P.A.; Thevissen, K. The Mode of Antifungal Action of Plant, Insect and Human Defensins. Cell. Mol. Life Sci. 2008, 65, 2069–2079. [Google Scholar] [CrossRef]
- Schneider, T.; Kruse, T.; Wimmer, R.; Wiedemann, I.; Sass, V.; Pag, U.; Jansen, A.; Nielsen, A.K.; Mygind, P.H.; Raventos, D.S.; et al. Plectasin, a Fungal Defensin, Targets the Bacterial Cell Wall Precursor Lipid II. Science 2010, 328, 1168–1172. [Google Scholar] [CrossRef] [Green Version]
- Thevissen, K.; Terras, F.R.; Broekaert, W.F. Permeabilization of Fungal Membranes by Plant Defensins Inhibits Fungal Growth. Appl. Environ. Microbiol. 1999, 65, 5451–5458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van der Weerden, N.L.; Lay, F.T.; Anderson, M.A. The Plant Defensin, NaD1, Enters the Cytoplasm of Fusarium Oxysporum Hyphae. J. Biol. Chem. 2008, 283, 14445–14452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Islam, K.T.; Velivelli, S.L.S.; Berg, R.H.; Oakley, B.; Shah, D.M. A Novel Bi-Domain Plant Defensin MtDef5 with Potent Broad-Spectrum Antifungal Activity Binds to Multiple Phospholipids and Forms Oligomers. Sci. Rep. 2017, 7, 16157. [Google Scholar] [CrossRef]
- Lay, F.T.; Mills, G.D.; Poon, I.K.H.; Cowieson, N.P.; Kirby, N.; Baxter, A.A.; van der Weerden, N.L.; Dogovski, C.; Perugini, M.A.; Anderson, M.A.; et al. Dimerization of Plant Defensin NaD1 Enhances Its Antifungal Activity. J. Biol. Chem. 2012, 287, 19961–19972. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brogden, K.A. Antimicrobial Peptides: Pore Formers or Metabolic Inhibitors in Bacteria? Nat. Rev. Microbiol. 2005, 3, 238–250. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, Y.; Ishibashi, J.; Yukuhiro, F.; Asaoka, A.; Taylor, D.; Yamakawa, M. Antibacterial Activity and Mechanism of Action of Tick Defensin against Gram-Positive Bacteria. Biochim. Biophys. Acta—Gen. Subj. 2003, 1624, 125–130. [Google Scholar] [CrossRef]
- Xiang, F.; Xie, Z.; Feng, J.; Yang, W.; Cao, Z.; Li, W.; Chen, Z.; Wu, Y. Plectasin, First Animal Toxin-Like Fungal Defensin Blocking Potassium Channels through Recognizing Channel Pore Region. Toxins 2015, 7, 34–42. [Google Scholar] [CrossRef]
- Almeida, M.S.; Cabral, K.M.S.; Kurtenbach, E.; Almeida, F.C.L.; Valente, A.P. Solution Structure of Pisum sativum Defensin 1 by High Resolution NMR: Plant Defensins, Identical Backbone with Different Mechanisms of Action. J. Mol. Biol. 2002, 315, 749–757. [Google Scholar] [CrossRef]
- Kushmerick, C.; de Souza Castro, M.; Santos Cruz, J.; Bloch, C.; Beirão, P.S.L. Functional and Structural Features of γ-Zeathionins, a New Class of Sodium Channel Blockers. FEBS Lett. 1998, 440, 302–306. [Google Scholar] [CrossRef] [Green Version]
- Zhu, S.; Peigneur, S.; Gao, B.; Umetsu, Y.; Ohki, S.; Tytgat, J. Experimental Conversion of a Defensin into a Neurotoxin: Implications for Origin of Toxic Function. Mol. Biol. Evol. 2014, 31, 546–559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, L.; Xie, Z.; Zhang, Q.; Li, Y.; Yang, F.; Chen, Z.; Li, W.; Cao, Z.; Wu, Y. Scorpion Potassium Channel-Blocking Defensin Highlights a Functional Link with Neurotoxin. J. Biol. Chem. 2016, 291, 7097–7106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aerts, A.M.; François, I.E.J.A.; Meert, E.M.K.; Li, Q.-T.; Cammue, B.P.A.; Thevissen, K. The Antifungal Activity of RsAFP2, a Plant Defensin from Raphanus Sativus, Involves the Induction of Reactive Oxygen Species in Candida albicans. J. Mol. Microbiol. Biotechnol. 2007, 13, 243–247. [Google Scholar] [CrossRef]
- Aerts, A.M.; Carmona-Gutierrez, D.; Lefevre, S.; Govaert, G.; François, I.E.J.A.; Madeo, F.; Santos, R.; Cammue, B.P.A.; Thevissen, K. The Antifungal Plant Defensin RsAFP2 from Radish Induces Apoptosis in a Metacaspase Independent Way in Candida albicans. FEBS Lett. 2009, 583, 2513–2516. [Google Scholar] [CrossRef]
- Hayes, B.M.E.; Bleackley, M.R.; Wiltshire, J.L.; Anderson, M.A.; Traven, A.; van der Weerden, N.L. Identification and Mechanism of Action of the Plant Defensin NaD1 as a New Member of the Antifungal Drug Arsenal against Candida Albicans. Antimicrob. Agents Chemother. 2013, 57, 3667–3675. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hayes, B.M.E.; Bleackley, M.R.; Anderson, M.A.; van der Weerden, N.L. The Plant Defensin NaD1 Enters the Cytoplasm of Candida Albicans via Endocytosis. J. Fungi 2018, 4, 20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aerts, A.M.; Bammens, L.; Govaert, G.; Carmona-Gutierrez, D.; Madeo, F.; Cammue, B.P.A.; Thevissen, K. The Antifungal Plant Defensin HsAFP1 from Heuchera Sanguinea Induces Apoptosis in Candida Albicans. Front. Microbiol. 2011, 2, 47. [Google Scholar] [CrossRef] [Green Version]
- Ochiai, A.; Ogawa, K.; Fukuda, M.; Ohori, M.; Kanaoka, T.; Tanaka, T.; Taniguchi, M.; Sagehashi, Y. Rice Defensin OsAFP1 Is a New Drug Candidate against Human Pathogenic Fungi. Sci. Rep. 2018, 8, 11434. [Google Scholar] [CrossRef]
- Montibus, M.; Ducos, C.; Bonnin-Verdal, M.-N.; Bormann, J.; Ponts, N.; Richard-Forget, F.; Barreau, C. The BZIP Transcription Factor Fgap1 Mediates Oxidative Stress Response and Trichothecene Biosynthesis But Not Virulence in Fusarium Graminearum. PLoS ONE 2013, 8, e83377. [Google Scholar] [CrossRef]
- Hale, J.D.; Hancock, R.E. Alternative Mechanisms of Action of Cationic Antimicrobial Peptides on Bacteria. Expert Rev. Anti Infect. Ther. 2007, 5, 951–959. [Google Scholar] [CrossRef]
- Do Nascimento, V.V.; de Mello, É.O.; Carvalho, L.P.; de Melo, E.J.T.; de O. Carvalho, A.; Fernandes, K.V.S.; Gomes, V.M. PvD1 Defensin, a Plant Antimicrobial Peptide with Inhibitory Activity against Leishmania amazonensis. Biosci. Rep. 2015, 35, e00248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lobo, D.S.; Pereira, I.B.; Fragel-Madeira, L.; Medeiros, L.N.; Cabral, L.M.; Faria, J.; Bellio, M.; Campos, R.C.; Linden, R.; Kurtenbach, E. Antifungal Pisum Sativum Defensin 1 Interacts with Neurospora Crassa Cyclin F Related to the Cell Cycle †. Biochemistry 2007, 46, 987–996. [Google Scholar] [CrossRef] [PubMed]
- Montesinos, E. Antimicrobial Peptides and Plant Disease Control. FEMS Microbiol. Lett. 2007, 270, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sher Khan, R.; Iqbal, A.; Malak, R.; Shehryar, K.; Attia, S.; Ahmed, T.; Ali Khan, M.; Arif, M.; Mii, M. Plant Defensins: Types, Mechanism of Action and Prospects of Genetic Engineering for Enhanced Disease Resistance in Plants. 3 Biotech. 2019, 9, 192. [Google Scholar] [CrossRef]
- Abdallah, N.A.; Shah, D.; Abbas, D.; Madkour, M. Stable Integration and Expression of a Plant Defensin in Tomato Confers Resistance to Fusarium Wilt. GM Crops 2010, 1, 344–350. [Google Scholar] [CrossRef]
- Deb, D.; Shrestha, A.; Sethi, L.; Das, N.C.; Rai, V.; Das, A.B.; Maiti, I.B.; Dey, N. Transgenic Tobacco Expressing Medicago Sativa Defensin (Msdef1) Confers Resistance to Various Phyto-Pathogens. Nucleus 2020, 63, 179–190. [Google Scholar] [CrossRef]
- Zhu, Y.J.; Agbayani, R.; Moore, P.H. Ectopic Expression of Dahlia Merckii Defensin DmAMP1 Improves Papaya Resistance to Phytophthora palmivora by Reducing Pathogen Vigor. Planta 2007, 226, 87–97. [Google Scholar] [CrossRef]
- Jha, S.; Tank, H.G.; Prasad, B.D.; Chattoo, B.B. Expression of Dm-AMP1 in Rice Confers Resistance to Magnaporthe Oryzae and Rhizoctonia Solani. Transgenic Res. 2009, 18, 59–69. [Google Scholar] [CrossRef]
- Kostov, K.; Christova, P.; Slavov, S.; Batchvarova, R. Constitutive Expression of a Radish Defensin Gene Rs-AFP2 in Tomato Increases the Resisstance to Fungal Pathogens. Biotechnol. Biotechnol. Equip. 2009, 23, 1121–1125. [Google Scholar] [CrossRef]
- Li, Z.; Zhou, M.; Zhang, Z.; Ren, L.; Du, L.; Zhang, B.; Xu, H.; Xin, Z. Expression of a Radish Defensin in Transgenic Wheat Confers Increased Resistance to Fusarium graminearum and Rhizoctonia cerealis. Funct. Integr. Genom. 2011, 11, 63–70. [Google Scholar] [CrossRef]
- Portieles, R.; Ayra, C.; Gonzalez, E.; Gallo, A.; Rodriguez, R.; Chacón, O.; López, Y.; Rodriguez, M.; Castillo, J.; Pujol, M.; et al. NmDef02, a Novel Antimicrobial Gene Isolated from Nicotiana megalosiphon Confers High-Level Pathogen Resistance under Greenhouse and Field Conditions: NmDef02, a Novel Antimicrobial Protein. Plant Biotechnol. J. 2010, 8, 678–690. [Google Scholar] [CrossRef]
- Soto, N.; Hernández, Y.; Delgado, C.; Rosabal, Y.; Ortiz, R.; Valencia, L.; Borrás-Hidalgo, O.; Pujol, M.; Enríquez, G.A. Field Resistance to Phakopsora Pachyrhizi and Colletotrichum Truncatum of Transgenic Soybean Expressing the NmDef02 Plant Defensin Gene. Front. Plant Sci. 2020, 11, 562. [Google Scholar] [CrossRef] [PubMed]
- Kanzaki, H.; Nirasawa, S.; Saitoh, H.; Ito, M.; Nishihara, M.; Terauchi, R.; Nakamura, I. Overexpression of the Wasabi Defensin Gene Confers Enhanced Resistance to Blast Fungus (Magnaporthe grisea) in Transgenic Rice. Theor. Appl. Genet. 2002, 105, 809–814. [Google Scholar] [CrossRef]
- Khan, R.S.; Nishihara, M.; Yamamura, S.; Nakamura, I.; Mii, M. Transgenic Potatoes Expressing Wasabi Defensin Peptide Confer Partial Resistance to Gray Mold (Botrytis cinerea). Plant Biotechnol. 2006, 23, 179–183. [Google Scholar] [CrossRef] [Green Version]
- Ntui, V.O.; Thirukkumaran, G.; Azadi, P.; Khan, R.S.; Nakamura, I.; Mii, M. Stable Integration and Expression of Wasabi Defensin Gene in “Egusi” Melon (Colocynthis citrullus L.) Confers Resistance to Fusarium Wilt and Alternaria Leaf Spot. Plant Cell Rep. 2010, 29, 943–954. [Google Scholar] [CrossRef] [PubMed]
- Kong, K.; Ntui, V.O.; Makabe, S.; Khan, R.S.; Mii, M.; Nakamura, I. Transgenic Tobacco and Tomato Plants Expressing Wasabi Defensin Genes Driven by Root-Specific LjNRT2 and AtNRT2.1 Promoters Confer Resistance against Fusarium Oxysporum. Plant Biotechnol. 2014, 31, 89–96. [Google Scholar] [CrossRef] [Green Version]
- François, I.E.J.A.; Dwyer, G.I.; De Bolle, M.F.C.; Goderis, I.J.W.M.; Van Hemelrijck, W.; Proost, P.; Wouters, P.; Broekaert, W.F.; Cammue, B.P.A. Processing in Transgenic Arabidopsis Thaliana Plants of Polyproteins with Linker Peptide Variants Derived from the Impatiens Balsamina Antimicrobial Polyprotein Precursor. Plant Physiol. Biochem. 2002, 40, 871–879. [Google Scholar] [CrossRef]
- Jha, S.; Chattoo, B.B. Transgene Stacking and Coordinated Expression of Plant Defensins Confer Fungal Resistance in Rice. Rice 2009, 2, 143–154. [Google Scholar] [CrossRef] [Green Version]
- Coca, M.; Bortolotti, C.; Rufat, M.; Peñas, G.; Eritja, R.; Tharreau, D.; del Pozo, A.M.; Messeguer, J.; San Segundo, B. Transgenic Rice Plants Expressing the Antifungal AFP Protein from Aspergillus Giganteus Show Enhanced Resistance to the Rice Blast Fungus Magnaporthe grisea. Plant Mol. Biol. 2004, 54, 245–259. [Google Scholar] [CrossRef]
- Banzet, N.; Latorse, M.-P.; Bulet, P.; François, E.; Derpierre, C.; Dubald, M. Expression of Insect Cystein-Rich Antifungal Peptides in Transgenic Tobacco Enhances Resistance to a Fungal Disease. Plant Sci. 2002, 162, 995–1006. [Google Scholar] [CrossRef]
- Zarinpanjeh, N.; Motallebi, M.; Zamani, M.R.; Ziaei, M. Enhanced Resistance to Sclerotinia sclerotiorum in Brassica napus by Co-Expression of Defensin and Chimeric Chitinase Genes. J. Appl. Genet. 2016, 57, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, K.; Kuwabara, C.; Umeki, N.; Fujioka, M.; Saburi, W.; Matsui, H.; Abe, F.; Imai, R. The Cold-Induced Defensin TAD1 Confers Resistance against Snow Mold and Fusarium Head Blight in Transgenic Wheat. J. Biotechnol. 2016, 228, 3–7. [Google Scholar] [CrossRef]
- Kaur, J.; Thokala, M.; Robert-Seilaniantz, A.; Zhao, P.; Peyret, H.; Berg, H.; Pandey, S.; Jones, J.; Shah, D. Subcellular Targeting of an Evolutionarily Conserved Plant Defensin MtDef4.2 Determines the Outcome of Plant–Pathogen Interaction in Transgenic Arabidopsis. Mol. Plant Pathol. 2012, 13, 1032–1046. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.K.; Pothana, A.; Prasad, K.; Shah, D.; Kaur, J.; Bhatnagar, D.; Chen, Z.-Y.; Raruang, Y.; Cary, J.W.; Rajasekaran, K.; et al. Peanuts That Keep Aflatoxin at Bay: A Threshold That Matters. Plant Biotechnol. J. 2018, 16, 1024–1033. [Google Scholar] [CrossRef] [Green Version]
- Turrini, A.; Sbrana, C.; Pitto, L.; Ruffini Castiglione, M.; Giorgetti, L.; Briganti, R.; Bracci, T.; Evangelista, M.; Nuti, M.P.; Giovannetti, M. The Antifungal Dm-AMP1 Protein from Dahlia Merckii Expressed in Solanum melongena Is Released in Root Exudates and Differentially Affects Pathogenic Fungi and Mycorrhizal Symbiosis. New Phytol. 2004, 163, 393–403. [Google Scholar] [CrossRef]
- Kaur, J.; Fellers, J.; Adholeya, A.; Velivelli, S.L.S.; El-Mounadi, K.; Nersesian, N.; Clemente, T.; Shah, D. Expression of Apoplast-Targeted Plant Defensin MtDef4.2 Confers Resistance to Leaf Rust Pathogen Puccinia triticina but Does Not Affect Mycorrhizal Symbiosis in Transgenic Wheat. Transgenic Res. 2017, 26, 37–49. [Google Scholar] [CrossRef] [Green Version]
- Stotz, H.U.; Spence, B.; Wang, Y. A Defensin from Tomato with Dual Function in Defense and Development. Plant Mol. Biol. 2009, 71, 131–143. [Google Scholar] [CrossRef]
- Allen, A.; Snyder, A.K.; Preuss, M.; Nielsen, E.E.; Shah, D.M.; Smith, T.J. Plant Defensins and Virally Encoded Fungal Toxin KP4 Inhibit Plant Root Growth. Planta 2007, 227, 331–339. [Google Scholar] [CrossRef]
- Audenaert, K.; Vanheule, A.; Höfte, M.; Haesaert, G. Deoxynivalenol: A Major Player in the Multifaceted Response of Fusarium to Its Environment. Toxins 2014, 6, 1–19. [Google Scholar] [CrossRef] [Green Version]
- Ponts, N. Mycotoxins Are a Component of Fusarium graminearum Stress-Response System. Front. Microbiol. 2015, 6, 1234. [Google Scholar] [CrossRef] [Green Version]
- Paul, P.A.; Bradley, C.A.; Madden, L.V.; Lana, F.D.; Bergstrom, G.C.; Dill-Macky, R.; Esker, P.D.; Wise, K.A.; McMullen, M.; Grybauskas, A.; et al. Meta-Analysis of the Effects of QoI and DMI Fungicide Combinations on Fusarium Head Blight and Deoxynivalenol in Wheat. Plant Dis. 2018, 102, 2602–2615. [Google Scholar] [CrossRef] [Green Version]
- Fisher, M.C.; Gurr, S.J.; Cuomo, C.A.; Blehert, D.S.; Jin, H.; Stukenbrock, E.H.; Stajich, J.E.; Kahmann, R.; Boone, C.; Denning, D.W.; et al. Threats Posed by the Fungal Kingdom to Humans, Wildlife, and Agriculture. mBio 2020, 11, e00449-20. [Google Scholar] [CrossRef]
- Mahlapuu, M.; Håkansson, J.; Ringstad, L.; Björn, C. Antimicrobial Peptides: An Emerging Category of Therapeutic Agents. Front. Cell. Infect. Microbiol. 2016, 6, 194. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Assoni, L.; Milani, B.; Carvalho, M.R.; Nepomuceno, L.N.; Waz, N.T.; Guerra, M.E.S.; Converso, T.R.; Darrieux, M. Resistance Mechanisms to Antimicrobial Peptides in Gram-Positive Bacteria. Front. Microbiol. 2020, 11, 593215. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Liu, S.; Wang, H. Invasive Fungi-Derived Defensins Kill Drug-Resistant Bacterial Pathogens. Peptides 2018, 99, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Kaewklom, S.; Wongchai, M.; Petvises, S.; Hanpithakphong, W.; Aunpad, R. Structural and Biological Features of a Novel Plant Defensin from Brugmansia × candida. PLoS ONE 2018, 13, e0201668. [Google Scholar] [CrossRef] [PubMed]
- Benítez-Chao, D.F.; León-Buitimea, A.; Lerma-Escalera, J.A.; Morones-Ramírez, J.R. Bacteriocins: An Overview of Antimicrobial, Toxicity, and Biosafety Assessment by in Vivo Models. Front. Microbiol. 2021, 12, 677. [Google Scholar] [CrossRef]
- Andersson, L.; Blomberg, L.; Flegel, M.; Lepsa, L.; Nilsson, B.; Verlander, M. Large-Scale Synthesis of Peptides. Biopolymers 2000, 55, 227–250. [Google Scholar] [CrossRef]
- Skalska, J.; Andrade, V.M.; Cena, G.L.; Harvey, P.J.; Gaspar, D.; Mello, É.O.; Henriques, S.T.; Valle, J.; Gomes, V.M.; Conceição, K.; et al. Synthesis, Structure, and Activity of the Antifungal Plant Defensin PvD1. J. Med. Chem. 2020, 63, 9391–9402. [Google Scholar] [CrossRef] [PubMed]
- Li, Y. Recombinant Production of Antimicrobial Peptides in Escherichia coli: A Review. Protein Expr. Purif. 2011, 80, 260–267. [Google Scholar] [CrossRef]
- Druzinec, D.; Salzig, D.; Brix, A.; Kraume, M.; Vilcinskas, A.; Kollewe, C.; Czermak, P. Optimization of Insect Cell Based Protein Production Processes—Online Monitoring, Expression Systems, Scale Up. In Yellow Biotechnology II; Vilcinskas, A., Ed.; Advances in Biochemical Engineering/Biotechnology; Springer: Berlin/Heidelberg, Germany, 2013; Volume 136, pp. 65–100. ISBN 978-3-642-39901-5. [Google Scholar]
- Kozlov, S.; Vassilevski, A.; Grishin, E. Antimicrobial Peptide Precursor Structures Suggest Effective Production Strategies. Recent Pat. Inflamm. Allergy Drug Discov. 2008, 2, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Deng, T.; Ge, H.; He, H.; Liu, Y.; Zhai, C.; Feng, L.; Yi, L. The Heterologous Expression Strategies of Antimicrobial Peptides in Microbial Systems. Protein Expr. Purif. 2017, 140, 52–59. [Google Scholar] [CrossRef]
- Li, Y. Carrier Proteins for Fusion Expression of Antimicrobial Peptides in Escherichia coli. Biotechnol. Appl. Biochem. 2009, 54, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Butt, T.R.; Edavettal, S.C.; Hall, J.P.; Mattern, M.R. SUMO Fusion Technology for Difficult-to-Express Proteins. Protein Expr. Purif. 2005, 43, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Bommarius, B.; Jenssen, H.; Elliott, M.; Kindrachuk, J.; Pasupuleti, M.; Gieren, H.; Jaeger, K.-E.; Hancock, R.E.W.; Kalman, D. Cost-Effective Expression and Purification of Antimicrobial and Host Defense Peptides in Escherichia Coli. Peptides 2010, 31, 1957–1965. [Google Scholar] [CrossRef] [Green Version]
- Sadr, V.; Saffar, B.; Emamzadeh, R. Functional Expression and Purification of Recombinant Hepcidin25 Production in Escherichia Coli Using SUMO Fusion Technology. Gene 2017, 610, 112–117. [Google Scholar] [CrossRef]
- Mo, Q.; Fu, A.; Lin, Z.; Wang, W.; Gong, L.; Li, W. Expression and Purification of Antimicrobial Peptide AP2 Using SUMO Fusion Partner Technology in Escherichia Coli. Lett. Appl. Microbiol. 2018, 67, 606–613. [Google Scholar] [CrossRef]
- Kant, P.; Liu, W.-Z.; Pauls, K.P. PDC1, a Corn Defensin Peptide Expressed in Escherichia Coli and Pichia Pastoris Inhibits Growth of Fusarium graminearum. Peptides 2009, 30, 1593–1599. [Google Scholar] [CrossRef]
- Al Kashgry, N.A.T.; Abulreesh, H.H.; El-Sheikh, I.A.; Almaroai, Y.A.; Salem, R.; Mohamed, I.; Waly, F.R.; Osman, G.; Mohamed, M.S.M. Utilization of a Recombinant Defensin from Maize (Zea Mays L.) as a Potential Antimicrobial Peptide. AMB Express 2020, 10, 208. [Google Scholar] [CrossRef]
- Kovaleva, V.; Bukhteeva, I.; Kit, O.Y.; Nesmelova, I.V. Plant Defensins from a Structural Perspective. Int. J. Mol. Sci. 2020, 21, 5307. [Google Scholar] [CrossRef] [PubMed]
- Tleuova, A.B.; Wielogorska, E.; Talluri, V.S.S.L.P.; Štěpánek, F.; Elliott, C.T.; Grigoriev, D.O. Recent Advances and Remaining Barriers to Producing Novel Formulations of Fungicides for Safe and Sustainable Agriculture. J. Control. Release Off. J. Control. Release Soc. 2020, 326, 468–481. [Google Scholar] [CrossRef]
- Gao, X.; Ding, J.; Liao, C.; Xu, J.; Liu, X.; Lu, W. Defensins: The Natural Peptide Antibiotic. Adv. Drug Deliv. Rev. 2021, 179, 114008. [Google Scholar] [CrossRef] [PubMed]
- Markets and Markets. Biofungicides Market by Type (Microbial Species, Botanical), Mode of Application (Soil Treatment, Foliar Application, Seed Treatment), Species (Bacillus, Trichoderma, Streptomyces, Pseudomonas), Crop Type, Formulation, and Region—Global Forecast to 2025; Markets and Markets: Pune, India, 2020. [Google Scholar]
- Mateo-Sagasta, J.; Zadeh, S.M.; Turral, H.; Burke, J. Water Pollution from Agriculture: A Global Review; Executive, Summary; FAO: Rome, Italy; International Water Management Institute (IWMI); CGIAR Research Program on Water, Land and Ecosystems (WLE): Colombo, Sri Lanka, 2017. [Google Scholar]
- Barratt, B.I.P.; Moran, V.C.; Bigler, F.; van Lenteren, J.C. The Status of Biological Control and Recommendations for Improving Uptake for the Future. BioControl 2018, 63, 155–167. [Google Scholar] [CrossRef] [Green Version]
- NAFTA Technical Working Group on Pesticides North American Free Trade Agreement Technical Working Group on Pesticides—Updated Procedures for the Joint Review of Biopesticides (i.e., Microbials and Biochemicals). 2010. Available online: https://www.canada.ca/en/health-canada/services/consumer-product-safety/pesticides-pest-management/public/protecting-your-health-environment/public-registry/north-american-free-trade-agreement-technical-working-group-pesticides-updated-procedures-joint-review-biopesticides-microbials-biochemicals.html (accessed on 23 January 2022).
- Neelabh; Singh, K.; Rani, J. Sequential and Structural Aspects of Antifungal Peptides from Animals, Bacteria and Fungi Based on Bioinformatics Tools. Probiotics Antimicrob. Proteins 2016, 8, 85–101. [Google Scholar] [CrossRef]


| Defensin | Amino Acid (AA) Sequence (Accession n°) | AA n°ber | MM (Da)/pI | Organism (Family) | Targeted Species (IC50 and MIC) | Ref. |
|---|---|---|---|---|---|---|
| Plant defensin | ||||||
| AX1 | AICKKPSKFFKGACGRDADCEKACDQENWPGGVCVPFLRCECQRSC (P81493) | 44 | 4895.72/7.14 | Beta vulgaris L (Amaran-thaceae) | Cercospora beticola (IC50 = 0.79 µM *) | [62] |
| AX2 | ATCRKPSMYFSGACFSDTNCQKACNREDWPNGKCLVGFKCECQRPC (P82010) | 46 | 5185.01/7.31 | C. beticola (IC50 = 0.39 µM *) | ||
| Dm-AMP1 | ELCEKASKTWSGNCGNTGHCDNQCKSWEGAAHGACHVRNGKHMCFCYFNC (P0C8Y4) | 46 | 4997.63/6.87 | Dahlia merckii (Asteraceae) | B. cinerea K1147 (IC50 = 2.17 µM *); Cladosporium sphaerospermum K0791 (IC50 = 0.54 µM *); F. culmorum K0311 (IC50 = (0.18–0.9) µM *); Leptosphaeria maculans LM36uea (IC50 = 0.27 µM *); Penicillium digitatum K0879 (IC50 = 0.36 µM *); Trichoderma viride K1127 (IC50 = 18.1 µM *); Septoria tritici K1097 (IC50 = 0.18 µM *); Verticillium alboatrum K0937 (IC50 = 0.72 µM *) | [63] |
| Dm-AMP2 | EVCEKASKTWSGNCGNTGHC | 20 | 2111.33/6.36 | B. cinerea K1147 (IC50 = 1.81 µM *); C. sphaerospermum K0791 (IC50 = 0.54 µM *); F. culmorum K0311 (IC50 = 0.54 µM *); L. maculans LM36uea (IC50 = 0.18 µM *); P. digitatum K0879 (IC50 = 0.36 µM *); T. viride K1127 (IC50 = 18.1 µM *); S. tritici K1097 (IC50 = 0.18 µM *); V. albo-atrum K0937 (IC50 = 0.36 µM *) | ||
| AhPDF1.1 | QRLCEKPSGTWSGVCGNNGACRNQCIRLEKARHGS | 51 | 5707.65/7.74 | Arabidopsis helleri (Brassicaceae) | F. oxysporum (MIC = 0.6 µM) | [64] |
| At-AFP1 | KLCERPSGTWSGVCGNSNACKNQCINLEKARHGSCNYVFPAHKCICYFPC (P30224) | 50 | 5539.44/7.53 | Arabidopsis thaliana (Brassicaceae) | Alternaria brassicicola MUCL 20,297 (IC50 =1.8 µM *); B. cinerea MUCL 30,158 (IC50 = 0.7 µM *); F. culmorum IMI 180,420 (IC50 = 0.54 µM *); F. oxysporum f. sp. lycopersici MUCL 909 (IC50 = 0.54 µM *); Pyricularia oryzae MUCL 30,166 (IC50 = 0.05 µM *); Verticillium dahliae MUCL 6963 (IC50 = 0.27 µM *) | [29] |
| AtPDF2.3 | RTCESKSHRFKGPCVSTHNCANVCHNEGFGGGKCRGFRRRCYCTRHC (Q9ZUL7) | 49 | 5348.15/8.49 | B. cinerea B05-10 (IC50 = 5.8 µM); B. cinerea R16 (IC50 = 5.8 µM); F. oxysporum 5176 (IC50 = 4.4 µM); F. culmorum MUCL 30,162 (IC50 = 1.0 µM); V. dahliae MUCL 19,210 (IC50 = 4.4 µM); F. graminearum PH-1 (IC50 = 1.4 µM) | [65] | |
| Hc-AFP1 | RYCERSSGTWSGVCGNSGKCSNQCQRLEGAAHGSCNYVFPAHKCICYYPC (G8GZ62) | 50 | 5483.21/7.33 | Heliophila coronopifolia (Brassicaceae) | B. cinerea (IC50 = 4.56 µM *); Fusarium solani (IC50 = 4.56 µM *) | [66] |
| Hc-AFP2 | QKLCERPSGTWSGVCGNNNACRNQCINLEKARHGSCNYVFPAHKCICYFPC (G8GZ63) | 51 | 5722.61/7.54 | B. cinerea (IC50 = (1.75–2.62) µM *); F. solani (IC50 = (1.75–2.62) µM *) | ||
| Hc-AFP3 | RYCERSSGTWSGVCGNTDKCSSQCQRLEGAAHGSCNYVFPAHKCICYYPC (G8GZ64) | 50 | 5528.24/7.09 | B. cinerea (IC50 = (3.62–4.52) µM *); F. solani (IC50 = 4.52 µM *) | ||
| Hc-AFP4 | QKLCERPSGTWSGVCGNNGACRNQCIRLERARHGSCNYVFPAHKCICYFPC (G8GZ65) | 51 | 5735.66/7.75 | B. cinerea (IC50 = (2.61–3.49) µM *); F. solani (IC50 = (0.87–1.74) µM *) | ||
| Rs-AFP1 | QKLCERPSGTWSGVCGNNNACKNQCINLEKARHGSCNYVFPAHKCICYFPC (P69241) | 51 | 5694.60/7.53 | R. sativus (Brassicaceae) | A. brassicicola MUCL 20,297 (IC50 = 2.64 µM *); B. cinerea MUCL 30,158 (IC50 = 1.41 µM *); F. culmorum IMI 180,420 (IC50 = 0.88 µM *); F. oxysporum f. sp. lycopersici MUCL 909 (IC50 = 5.28 µM *); P. oryzae MUCL 30,166 (IC50 = 0.05 µM *); V. dahliae MUCL 6963 (IC50 = 0.88 µM *) | [29] |
| Ascochyta pisi (IC50 = 0.88 µM *); C. beticola (IC50 = 0.35 µM *); Colletotrichum lindemuthianum (IC50 = 17.61 µM *); F. oxysporum f. sp. pisi (IC50 = 2.64 µM *); Mycosphaerella fijiensis var. fijiensis (IC50 = 0.7 µM *); Nectria haematococca (IC50 = 1.06 µM *); Phoma betae (IC50 = 0.35 µM *); Pyrenophora tritici-repentis (IC50 = 0.53 µM *); P. oryzae (IC50 = 0.05 µM *); Rhizoctonia solani (IC50 = 17.61 µM *); Sclerotinia sclerotiorum (IC50 = 3.52 µM *); Septoria nodorum (IC50 = 3.52 µM *); Trichoderma hamatum (IC50 = 1.06 µM *); V. dahliae (IC50 = 0.88 µM *) | [67] | |||||
| Rs-AFP2 | QKLCQRPSGTWSGVCGNNNACKNQCIRLEKARHGSCNYVFPAHKCICYFPC (P30230) | 51 | 5735.70/7.94 | B. cinerea K1147 (IC50 = 1.75 µM *); C. sphaerospermum K0791 (IC50 = 0.52 µM *); F. culmorum K0311 (IC50 = 0.26 µM *); F. culmorum K0311 (IC50 = 0.87 µM *); L. maculans LM36uea (IC50 = 2.1 µM *); P. digitatum K0879 (IC50 = 0.26 µM *); T. viride K1127 (IC50 = 5.25 µM *); S. tritici K1097 (IC50 = 0.26 µM *); V. albo-atrum K0937 (IC50 = 2.1 µM *) | [63] | |
| A. brassicicola MUCL 20,297 (IC50 = 0.35 µM *); B. cinerea MUCL 30,158 (IC50 = 0.35 µM *); F. culmorum IMI 180,420 (IC50 = 0.35 µM *); F. oxysporum f. sp. lycopersici MUCL 909 (IC50 = 0.35 µM *); P. oryzae MUCL 30,166 (IC50 = 0.7 µM *); V. dahliae MUCL 6963 | [29] | |||||
| A. pisi (IC50 = 0.7 µM *); C. beticola (IC50 = 0.35 µM *); C. lindemuthianum (IC50 = 0.52 µM *); F. oxysporum f. sp. Pisi (IC50 = 0.35 µM *); M. fijiensis var. fijiensis (IC50 = 0.26 µM *); N. haematococca (IC50 = 0.35 µM *); P. betae (IC50 = 0.17 µM *); P. tritici-repentis (IC50 = 0.26 µM *); R. solani (IC50 = 17.49 µM *); S. sclerotiorum (IC50 = 17.49 µM *); S. nodorum (IC50 = 2.62 µM *); T. hamatum (IC50 = 0.35 µM *); V. dahliae (IC50 = 0.26 µM *); Venturia inaequalis (IC50 = 4.37 µM *) | [67] | |||||
| Defensin-like protein 4 | QKLCERSSGTWSGVCGNNNACKNQCINLEGARHGSCNYIFPYHRCICYFPC (O24331) | 51 | 5747.58/7.33 | A. brassicicola (IC50 = 0.87 µM *); B. cinerea (IC50 = 1.57 µM *); F. culmorum (IC50 = 1.92 µM *) | [68] | |
| Defensin-like protein 3 | KLCERSSGTWSGVCGNNNACKNQCIRLEGAQHGSCNYVFPAHKCICYFPC (O24332) | 50 | 5499.34/7.33 | A. brassicicola (IC50 = 0.36 µM *); B. cinerea (IC50 = 0.36 µM *); F. culmorum (IC50 = 0.36 µM *) | ||
| Sa-AFP2 | QKLCQRPSGTWSGVCGNNNACRNQCINLEKARHGSCNYVFPAHKCICYFPC (P30232) | 51 | 5721.63/7.74 | S. alba (Brassicaceae) | A. brassicicola MUCL 20,297 (IC50 = 0.79 µM *); B. cinerea MUCL 30,158 (IC50 = 0.61 µM *); F. culmorum IMI 180,420 (IC50 = 0.4 µM *); F. oxysporum f. sp. lycopersici MUCL 909 (IC50 = 0.4 µM *); P. oryzae MUCL 30,166 (IC50 = 0.05 µM *); V. dahliae MUCL 6963 (IC50 = 0.21 µM *) | [29] |
| WT1 | QKLCEKSSGTWSGVCGNNNACKNQCINLEGARHGSCNYIFPYHRCICYFPC (Q9FS38) | 51 | 5719.56/7.33 | Eutrema japonicum (Brassicaceae) | Magnaporthe grisea (IC50 = 0.87 µM *); B. cinerea (IC50 = 3.5 µM *) | [69] |
| Sm-AMP-D1 | KICERASGTWKGICIHSNDCNNQCVKWENAGSGSCHYQFPNYMCFCYFDC (C0HL82) | 50 | 5763.55/6.28 | Stellaria media L. (Caryophyllaceae) | Bipolaris sorokiniana 6/10 (IC50 = 0.5 µM); F. oxysporum 16/10 (IC50 = 0.35 µM); F. graminearum VKM F-1668 (IC50 = 0.52 µM); Fusarium avenaceum VKM F-2303 (IC50 = 0.52 µM); B. cinerea SGR-1 (IC50 = 1.0 µM); P. betae VKM F-2532 (IC50 = 0.52 µM); Pythium debaryanum VKM F-1505 (IC50 = 1.0 µM) | [70] |
| Sm-AMP-D2 | KICERASGTWKGICIHSNDCNNQCVKWENAGSGSCHYQFPNYMCFCYFNC (C0HL83) | 50 | 5762.57/6.77 | B. sorokiniana 6/10 (IC50 = 0.5 µM); F. oxysporum 16/10 (IC50 = 0.35 µM); F. graminearum VKM F-1668 (IC50 = 0.52 µM); F. avenaceum VKM F-2303 (IC50 = 0.52 µM); B. cinerea SGR-1 (IC50 = 1.0 µM); P. betae VKM F-2532 (IC50 = 0.52 µM); P. debaryanum VKM F-1505 (IC50 = 1.0 µM) | ||
| So-D2 | GIFSSRKCKTPSKTFKGICTRDSNCDTSCRYEGYPAGDCKGIRRRCMCSKPC | 52 | 5803.79/8.34 | S. oleracea (Chenopodiaceae) | F. culmorum (IC50 = 0.2 µM); F. solani (IC50 = 11 µM); Colletotrichum lagenarium (IC50 = 11 µM); Bipolaris maydis (IC50 = 6 µM) | [71] |
| AB2 | RTCENLANTYRGPCITTGSCDDHCKNKEHLRSGRCRDDFRCW | 47 | 5469.18/7.33 | Adzuckia angularia (Fabaceae) | B. cinerea (IC50 = 3.5 µM) | [72] |
| Beta-astratide bM1 | CEKPSKFFSGPCIGSSGKTQCAYLCRRGEGLQDGNCKGLKCVCAC | 45 | 4734.58/7.52 | Astragalus membranaceus (Fabaceae) | F. oxysporum CICC 2532 (IC50 = 4.92 µM *); Alternaria alternata CICC 2465 (IC50 = 4.75 µM *); R. solani CICC 40,259 (IC50 = 27.52 µM *); Curvularia lunata CICC 40,301 (IC50 = 0.57 µM *) | [73] |
| Coccinin | KQTENLADTY (P84785) | 10 | 1182.25/4.19 | Phaseolus coccineus cv. ‘Major’ (Fabaceae) | F. oxysporum (MIC = 81 µM); B. cinerea (MIC = 109 µM); R. solani (MIC = 134 µM); Mycosphaerella arachidicola (MIC = 75 µM) | [74] |
| Phaseococcin | KTCENLADTYKGPPPFFTTG | 20 | 2187.46/6.03 | P. coccineus cv. ‘Minor’ (Fabaceae) | F. oxysporum (MIC = 89 µM); B. cinerea (MIC = 102 µM); R. solani (MIC = 140 µM); M. arachidicola (MIC = 70 µM) | [75] |
| Ct-AMP1 | NLCERASLTWTGNCGNTGHCDTQCRNWESAKHGACHKRGNWKCFCYFNC (Q7M1F2) | 49 | 5613.32/7.33 | Clitoria ternatea (Fabaceae) | B. cinerea K1147 (IC50 = 3.56 µM *); C. sphaerospermum K0791 (IC50 = 1.07 µM *); F. culmorum K0311 (PDB medium) (IC50 = 1.78 µM *); F. culmorum K0311 (SMF medium) (IC50 = 0.11 µM *); L. maculans LM36uea (IC50 = =1.07 µM *); P. digitatum K0879 (IC50 = 3.56 µM *); T. viride K1127 (IC50 = 17.81 µM *); S. tritici K1097 (IC50 = 0.36 µM *); V. albo-atrum K0937 (IC50 = 0.36 µM *) | [63] |
| Gymnin | KTCENLADDY (P84200) | 10 | 1171.25/3.8 | Gymnocladus chinensis (Fabaceae) | F. oxysporum (IC50 = 2 µM); Cercospora arachidicola (IC50 = 10 µM) | [76] |
| Lc-def | KTCENLSDSFKGPCIPDGNCNKHCKEKEHLLSGRCRDDFRCWCTRNC (B3F051) | 47 | 5449.23/7.08 | Lens culinaris (Fabaceae) | Aspergillus niger VKM F-2259 (IC50 = 18.5 µM); Aspergillus versicolor VKM F-1114 (IC50 = 18.5 µM); B. cinerea VKM F-3700 (IC50 = 9.25 µM); F. culmorum VKM F-844 (IC50 = (18.5–37.0) µM) | [77] |
| Limenin | KTCENLADTYKGPCFTTGGCDDHCKNKEHLLSGRCRDDFRCWCTRNC | 47 | 5403.12/6.77 | Phaseolus limensis (Fabaceae) | B. cinerea (MIC = 2.9 µM); F. oxysporum (MIC = 2.1 µM); M. arachidicola (MIC = 0.34 µM) | [78] |
| Limyin | KTCENLATYYRGPCF | 15 | 1766.03/7.51 | F. solani (IC50 = 8.6 µM) | [79] | |
| Ms-Def1 (alfAFP) | RTCENLADKYRGPCFSGCDTHCTTKENAVSGRCRDDFRCWCTKRC (Q4G3V1) | 45 | 5194.90/7.32 | Medicago sativa (Fabaceae) | F. graminearum (IC50 = (1.2–2.3) µM *) | [80] |
| F. graminearum PH-1 (IC50 = (2–4) µM *); F. graminearum PH-1 (MIC > 6 µM) | [56] | |||||
| V. dahliae (MIC = 1 µM *) | [81] | |||||
| Mt-Def2 | KTCENLADKYRGPCFSGCDTHCTTKENAVSGRCRDDFRCWCTKRC (Q5YLG8) | 45 | 5166.89/7.32 | M. Truncatula (Fabaceae) | F. graminearum PH-1 (IC50 = (0.75–1) µM) | [56] |
| F. oxysporum f. sp. medicaginis 7F-3 (IC50 = 0.7 µM); F. oxysporum f. sp. medicaginis 31F-3 (IC50 = 1.9 µM); Phoma medicaginis STC (IC50 = 0.3 µM); P. medicaginis WS-2 (IC50 = 2.6 µM); Clavibacter insidiosus (IC50 = 0.1 µM) | [35] | |||||
| Mt-Def4 | RTCESQSHKFKGPCASDHNCASVCQTERFSGGRCRGFRRRCFCTTHC (G7L736) | 47 | 5343.08/7.97 | F. graminearum PH-1 (IC50 = (0.75–1) µM) | [56] | |
| F. oxysporum f. sp. medicaginis 7F-3 (IC50 = 0.7 µM); F. oxysporum f. sp. medicaginis 31F-3 (IC50 = 1.9 µM); P. medicaginis STC (IC50 = 0.3 µM); P. medicaginis WS-2 (IC50 = 2.6 µM) | [35] | |||||
| PsD1 | KTCEHLADTYRGVCFTNASCDDHCKNKAHLISGTCHNWKCFCTQNC (P81929) | 46 | 5208.93/6.81 | Pisum sativum (Fabaceae) | A. niger EK0197 (IC50 = 2.3 µM *); A. versicolor 40028LMR/INCQS (IC50 = 1 µM *); Fusarium moniliforme 2414UFPe (IC50 = 4.2 µM *); F. oxysporum 2665UFPe (IC50 = 19.2 µM *); F. solani 2389UFPe (IC50 = 2.3 µM *) | [82] |
| PsD2 | KTCENLSGTFKGPCIPDGNCNKHCRNNEHLLSGRCRDDFRCWCTNRC (P81930) | 47 | 5404.15/7.33 | A. niger EK0197 (IC50 = 1.9 µM *); A. versicolor 40028LMR/INCQS (IC50 = 0.06 µM *); F. moniliforme 2414UFPe (IC50 = 1.85 µM *); F. oxysporum 2665UFPe (IC50 = 18.5 µM *); F. solani 2389UFPe (IC50 = 1.57 µM *) | ||
| PvD1_PTA2c | KTCENLADTYKGPCFTTGSCDDHCKNKEHLRSGRCRDDFRCWCTKNC (F8QXP9) | 47 | 5448.16/7.08 | Phaseolus vulgaris (Fabaceae) | F. solani (IC50 = 18.35 µM *); Fusarium laterithium (IC50 = 18.35 µM *); R. solani (IC50 = 18.35 µM *); F. oxysporum (IC50 = 18.35 µM *) | [83] |
| B. cinerea (IC50 = 1 µM) | [72] | |||||
| P. vulagris white cloud defensin | KTCENLADTFRGPCFATSNCDDHCKNKEHLLSGRCRDDFRCWCTRNC | 47 | 5472.18/6.77 | P. vulgaris cv. “white cloud bean” (Fabaceae) | B. cinerea (MIC = 2.8 µM); F. oxysporum (MIC = 2.3 µM); M. arachidicola (MIC = 0.72 µM) | [84] |
| Sesquin | KTCENLADTY (P84868) | 10 | 1157.27/4.19 | Vigna unguiculate (Fabaceae) | B. cinerea (IC50 = 2.5 µM); F. oxysporum (IC50 = 1.4 µM); M. arachidicola (IC50 = 0.15 µM) | [85] |
| SPE10 | KTCENLADTFRGPCFTDGSCDDHCKNKEHLIKGRCRDDFRCWCTRNC (Q6B519) | 47 | 5500.24/6.77 | Pachyrhizus erosus (Fabaceae) | Aspergillus flavus (IC50 = 5.45 µM *); A. niger (IC50 = 8.18 µM *); B. maydis (IC50 = 2.73 µM *); B. cinerea (IC50 = 18.18 µM *); Colletotrichum gloeosporides (IC50 = 18.18 µM *); F. oxysporum f.sp. lycopersic (IC50 = 18.18 µM *); F. oxysporum f.sp. vasinfectum (IC50 = 18.18 µM *); Penicillium spp. (IC50 = 18.18 µM *); Rhizopus stolonifer (IC50 = 18.18 µM *); V. dahliae (IC50 = 18.18 µM *) | [86] |
| TvD1 | KTCENLADTYRGPCFTTGSCDDHCKNKEHLLSGRCRDDFRCWCTKRC (Q2KM12) | 47 | 5475.23/7.09 | Tephrosia villosa (Fabaceae) | Nothopassalora personata (MIC = 2.05 µM *); F. oxysporum (MIC = 5.12 µM *); Fusarium verticillioides (MIC = 5.12 µM *); B. cinerea (MIC = 5.12 µM *); Curvularia sp (MIC = 5.12 µM *); R. solani (MIC = 7.78 µM *) | [87] |
| VaD1 | KTCMTKKEGWGRCLIDTTCAHSCRKQGYKGGNCKGMRRTCYCLLDC (A0A0S3QXX7) | 46 | 5209.23/8.12 | Vigna angularis (Fabaceae) | F. oxysporum (IC50 = 5.76 µM *); F. oxysporum f. sp. pisi (IC50 = 10.21 µM *) | [88] |
| VrD1 | RTCMIKKEGWGKCLIDTTCAHSCKNRGYIGGNCKGMTRTCYCLVNC (Q6T418) | 46 | 5122.15/7.92 | Vigna radiata (Fabaceae) | F. oxysporum (IC50 = 1.1 µM *); F. oxysporum CCRC 35,270 (IC50 = 3.4 µM *); F. oxysporum f. sp. Pisi (IC50 = 2.4 µM *); P. oryzae (IC50 = 4 µM *); R. solani (IRTCENLADKYRGPCFSGCDTHCTTKENAVSGRCRDDFRCWCTKRCC50 = 17.7 µM *) | [89] |
| PgD1 | RTCKTPSGKFKGVCASSNNCKNVCQTEGFPSGSCDFHVANRKCYCSKPCP (Q6RSS6) | 50 | 5377.21/7.91 | Picea glauca (Pinaceae) | Nectria galligena (MIC = 2.6 µM *); F. oxysporum (MIC = 2.6 µM *) | [51] |
| PgD5 | RMCESQSHKFKGYCASSSNCKVVCQTEKFLTGSCRDTHFGNRRCFCEKPC | 50 | 5729.62/7.72 | F. oxysporum (MIC = 1.92 µM *); V. dahliae (MIC = 0.35 µM *); B. cinerea (MIC = 0.7 µM *) | ||
| PsDef1 | RMCKTPSGKFKGYCVNNTNCKNVCRTEGFPTGSCDFHVAGRKCYCYKPCP (A4L7R7) | 50 | 5601.58/8.12 | Pinus sylvestris (Pinaceae) | F. solani UKM F-50639 (IC50 = 0.16 µM *); F. oxysporum UKM F-52897 (IC50 = 0.52 µM *); B. cinerea UKM F-16753 (IC50 = 0.07 µM *) | [90] |
| Ec-AMP-D1 | RECQSQSHRYKGACVHDTNCASVCQTEGFSGGKCVGFRGRCFCTKAC (P86518) | 47 | 5107.82/7.54 | Echinochloa crusgalli (Poaceae) | F. graminearum (IC50 = 2.94 µM *); F. verticillioides (IC50 = 1.66 µM *); Diplodia maydis (IC50 = 2.45 µM *); F. oxysporum (IC50 = 19.97 µM *) | [91] |
| Ec-AMP-D2 | RECQSQSHRYKGACVHDTNCASVCQTEGFSGGKCVGFRGRCFCTKHC (P86519) | 47 | 5173.89/7.54 | F. oxysporum (IC50 = 19.71 µM *) | ||
| Pp-AMP1 | KSCCRSTQARNIYNAPRFAGGSRPLCALGSGCKIVDDKKTPPND | 44 | 4697.39/8.61 | Phyllostachys pubescens (Poaceae) | F. oxysporum IFO 6384 (IC50 = 0.43 µM *) | [92] |
| Pp-AMP2 | KSCCRSTTARTARVPCAKKSNIYNGCRVPGGCKIQEAKKCEPPYD | 45 | 4919.76/8.52 | F. oxysporum IFO 6384 (IC50 = 0.41 µM *) | ||
| Sd1 | RYCLSQSHRFKGLCMSSSNCANVCQTENFPGGECKADGATRKCFCKKIC (B2CNV2) | 49 | 5412.32/7.72 | Saccharum officinarum (Poaceae) | A. niger (IC50 = 2.0 µM); F. solani (IC50 = 1.0 µM) | [93] |
| Sd3 | RHRHCFSQSHKFVGACLRESNCENVCKTEGFPSGECKWHGIVSKCHCKRIC | 51 | 5864.82/7.73 | A. niger (IC50 = 1.0 µM); F. solani (IC50 > 20 µM) | ||
| Sd5 | HTPTPTPICKSRSHEYKGRCIQDMDCNAACVKESESYTGGFCNGRPPFKQCFCTKPCKRERAAATLRWPGL (A0A1B3B2K6) | 71 | 7967.21/7.91 | A. niger (IC50 > 20 µM); F. solani (IC50 = 10 µM) | ||
| SI alpha-1 | RVCMGKSQHHSFPCISDRLCSNECVKEEGGWTAGYCHLRYCRCQKAC (P21923) | 47 | 5382.26/7.33 | Sorghum bicolor (Poaceae) | B. cinerea K1147 (IC50 = 18.58 µM *); C. sphaerospermum K0791 (IC50 = 14.86 µM *); F. culmorum K0311 (IC50 = 37.16 µM *); P. digitatum K0879 (IC50 = 37.16 µM *); T. viride K1127 (IC50 = 9.29 µM *) | [63] |
| Tk-AMP-D1 | RTCQSQSHKFKGACFSDTNCDSVCRTENFPRGQCNQHHVERKCYCERDC (P84963) | 49 | 5744.40/7.1 | Triticum kiharae (Poaceae) | F. graminearum (IC50 = 5.22 µM *); F. verticillioides (IC50 = 5.22 µM *) | [91] |
| ZmD32 | RTCQSQSHRFRGPCLRRSNCANVCRTEGFPGGRCRGFRRRCFCTTHC (A0A317Y7J2) | 47 | 5466.33/10.85 | Zea mays (Poaceae) | F. graminearum PH-1 (IC50 = 1 µM) | [94] |
| ZmESR6 | KLCSTTMDLLICGGAIPGAVNQACDDTCRNKGYTGGGFCNMKIQRCVCRKPC (D1MAH4) | 52 | 5516.57/7.52 | F. oxysporum f.sp. Conglutinans (IC50 = 3 µM); F. oxysporum f.sp.lycopersici (IC50 = 3 µM); Plectosphaerella cucumerina (IC50 = 2 µM) | [95] | |
| Fa-AMP1 | AQCGAQGGGATCPGGLCCSQWGWCGSTPKYCGAGCQSNCK (P0DKH7) | 40 | 3887.42/7.07 | Fagopyrum esculentum (Polygonaceae) | F. oxysporum IFO 6384 (IC50 = 4.89 µM *) | [96] |
| Fa-AMP2 | AQCGAQGGGATCPGGLCCSQWGWCGSTPKYCGAGCQSNCR (P0DKH8) | 40 | 3915.44/7.07 | F. oxysporum IFO 6384 (IC50 = 7.41 µM *) | ||
| Ns-D1 | KFCEKPSGTWSGVCGNSGACKDQCIRLEGAKHGSCNYKPPAHRCICYYEC (P86972) | 50 | 5487.32/7.32 | Nigella sativa (Ranunculaceae) | A. niger VKM F-33 (IC50 = 0.64 µM *); B. sorokiniana VKM F-1446 (IC50 = 0.55 µM *); F. oxysporum (IC50 = 1.73 µM *); F. graminearum VKM F-1668 (IC50 = 1.26 µM *; F. culmorum VKM F-2303 (IC50 = 1.26 µM *); B. cinerea (IC50 = 4.99 µM *) | [97] |
| Ns-D2 | KFCEKPSGTWSGVCGNSGACKDQCIRLEGAKHGSCNYKLPAHRCICYYEC (P86973) | 50 | 5503.36/7.32 | A. niger VKM F-33 (IC50 = 0.64 µM *); B. sorokiniana VKM F-1446 (IC50 = 0.33 µM *); F. oxysporum (IC50 = 0.96 µM *); F. graminearum VKM F-1668 (IC50 = 1.25 µM *); F. culmorum VKM F-2303 (IC50 = 1.25 µM *); B. cinerea (IC50 = 2.49 µM *) | ||
| Ah-AMP1 | LCNERPSQTWSGNCGNTAHCDKQCQDWEKASHGACHKRENHWKCFCYFNC (Q7M1F3) | 50 | 5863.53/6.82 | Aesculus hippocastanum (Sapindaceae) | B. cinerea K1147 (IC50 = 4.26 µM *); C. sphaerospermum K0791 (IC50 = 0.85 µM *); F. culmorum K0311 (IC50 = 2.05 µM *); F. culmorum K0311 (IC50 = 0.12 µM *); L. maculans LM36uea (IC50 = 0.09 µM *); P. digitatum K0879 (IC50 = 1.02 µM *); T. viride K1127 (IC50 = 17.05 µM *); S. tritici K1097 (IC50 = 0.85 µM *); V. albo-atrum K0937 (IC50 = 1.02 µM *) | [63] |
| Hs-AFP1 | DGVKLCDVPSGTWSGHCGSSSKCSQQCKDREHFAYGGACHYQFPSVKCFCKRQC (P0C8Y5) | 54 | 5948.76/7.32 | Heuchera sanguinea (Saxifragaceae) | B. cinerea K1147 (IC50 = 1 µM *); C. sphaerospermum K0791 (IC50 = 0.2 µM *); F. culmorum K0311 (IC50 = 0.2 µM *); L. maculans LM36uea (IC50 = 4.2 µM *); P. digitatum K0879 (IC50 = 0.2 µM *); T. viride K1127 (IC50 = 2.5 µM *); S. tritici K1097 (IC50 = 0.1 µM *); V. albo-atrum K0937 (IC50 = 1 µM *) | |
| NaD1 | RECKTESNTFPGICITKPPCRKACISEKFTDGHCSKILRRCLCTKPC (Q8GTM0) | 47 | 5304.37/7.91 | Nicotiana alata (Solanaceae) | A. niger 5181 (IC50 = 2.1 ± 0.76 µM); A. flavus 5310 (IC50 > 10 µM); F. oxysporum f.sp. Vasinfectum (IC50 = 1.5 ± 0.25 µM); F. graminearum (IC50 = 0.4 ± 0.3 µM); Colletotrichum graminicola (IC50 = 4.4 ± 0.1 µM); Aspergillus parasiticus 4467 (IC50 = 4.5 ± 0.27 µM) | [98] |
| V. dahliae (IC50 = 0.75 µM); Thielaviopsis basicola (IC50 = 1 µM); Aspergillus nidulans (IC50 = 0.8 µM); Puccinia coronata f.sp. Avenae (IC50 = 2.5 µM); Puccinia sorghi (IC50 = 2 µM) | [99] | |||||
| NaD2 | RTCESQSHRFKGPCARDSNCATVCLTEGFSGGDCRGFRRRCFCTRPC (A0A1B2YLI5) | 47 | 5264.02/7.76 | F. oxysporum f.sp. Vasinfectum (IC50 = 8.3 µM); F. graminearum (IC50 = 2 µM); V. dahliae (IC50 > 10 µM); T. basicola (IC50 = 7 µM); A. nidulans (IC50 = 5 µM); P. coronata f.sp. Avenae (IC50 = 4 µM); P. sorghi (IC50 = 5 µM) | ||
| PhD1 | ATCKAECPTWDSVCINKKPCVACCKKAKFSDGHCSKILRRCLCTKEC (Q8H6Q1) | 47 | 5211.33/7.67 | Petunia hybrida (Solanaceae) | F. oxysporum (MIC = (0–0.38) µM *); B. cinerea (MIC = (0.38–1.92) µM *) | [100] |
| PhD2 | GTCKAECPTWEGICINKAPCVKCCKAQPEKFTDGHCSKILRRCLCTKPC (Q8H6Q0) | 49 | 5403.55/7.52 | F. oxysporum (MIC = (0.38–1.92) µM *); B. cinerea (MIC = (0.38–1.92) µM *) | ||
| Vv-AMP1 | RTCESQSHRFKGTCVRQSNCAAVCQTEGFHGGNCRGFRRRCFCTKHC (D7TAI4) | 47 | 5355.13/8.24 | Vitis vinifera (Vitaceae) | F. oxysporum ATCC 10,913 (IC50 = 1.12 µM *); V. dahliae ATCC 96,522 (IC50 = 0.34 µM *); F. solani (IC50 = 1.79 µM *); B. cinerea (IC50 = 2.43 µM *) | [101] |
| Invertebrate defensin | ||||||
| AgDef1 | ATCDLASGFGVGSSLCAAHCIARRYRGGYCNSKAVCVCRN (B2FZB7) | 40 | 4141.80/7.82 | Anopheles gambiae (Insecta) | F. culmorum (MIC = (3–6) µM); F. oxysporum (MIC = (1.5–3) µM) | [30] |
| Defensin ARD1 | DKLIGSCVWGAVNYTSNCNAECKRRGYKGGHCGSFANVNCWCET (P84156) | 44 | 4803.43/7.24 | Archaeoprepona demophon (Insecta) | Aspergillus fumigatus GASP 4707 (MIC = 2.6 µM *) | [102] |
| DEFC | ATCDLLSGFGVGDSACAAHCIARRNRGGYCNAKKVCVCRN (P81603) | 40 | 4161.84/7.81 | Aedes aegypti (Insecta) | F. culmorum (MIC = (50–100) µM) | [30] |
| Drosomycin | DCLSGRYKGPCAVWDNETCRRVCKEEGRSSGHCSPSLKCWCEGC (P41964) | 44 | 4897.59/6.75 | Drosophila melanogaster (Insecta) | B. cinerea MUCL 30,158 (IC50 = 1.2 µM); F. culmorum IMI 180,420 (IC50 = 1.0 µM); A. brassicicola MUCL 20,297 (IC50 = 0.9 µM); Alternaria longipes CBS 62,083 (IC50 = 1.4 µM); N. haematococca CollectionVanEtten160-2-2 (IC50 = 1.8 µM); F. oxysporum MUCL 909 (IC50 = 4.2 µM); A. pisi MUCL 30,164 (IC50 = 3.2 µM) | [103] |
| Gm defensin-like peptide | DKLIGSCVWGATNYTSDCNAECKRRGYKGGHCGSFWNVNCWCEE (P85215) | 44 | 4949.53/6.21 | Galleria mellonella (Insecta) | A. niger (MIC = (1.4–2.9) µM); Trichoderma harzianum (MIC = (1.4–2.9) µM) | [104] |
| Galleria defensin | DTLIGSCVWGATNYTSDCNAECKRRGYKGGHCGSFLNVNCWCE (P85213) | 43 | 4720.29/6.2 | F. oxysporum (MIC = (8.5–16.9) µM); A. niger (MIC = (1.1–2.1) µM); T. harzianum (MIC = (2.1–4.2) µM) | ||
| Heliomicin | DKLIGSCVWGAVNYTSDCNGECKRRGYKGGHCGSFANVNCWCET (D3G9G5) | 44 | 4790.39/6.87 | Heliothis virescens (Insecta) | F. culmorum IMI 180,420 (MIC = (0.2–0.4) µM); F. oxysporum MUCL 909 (MIC = (1.5–3.0) µM); N. haematococca 160.2.2 (MIC = (0.4–0.8) µM); A. fumigatus (MIC = (6–12) µM); T. viride MUCL 19,724 (MIC = (1.5–3) µM) | [105] |
| PduDef | ATCDLLSAFGVGHAACAAHCIGHGYRGGYCNSKAVCTCRR (P83404) | 40 | 4101.74/7.55 | Phlebotomus duboscqi (Insecta) | A. fumigatus (MIC = 12.5–25 µM); F. culmorum (MIC = 1.56–3.12 µM); F. oxysporum (MIC = 3.12–6.25 µM); T. viride (MIC = 3.12–6.25 µM) | [30] |
| Phormicin | ATCDLLSGTGINHSACAAHCLLRGNRGGYCNGKGVCVCRN (P10891) | 40 | 4066.69/7.55 | Protophormia terraenovae (Insecta) | F. culmorum IMI 180,420 (MIC = 3 µM); F. oxysporum MUCL 909 (MIC = 6 µM) | [105] |
| F. culmorum (MIC = (1.5–3.0) µM); F. oxysporum (MIC = (3–6) µM); N. haematococca (MIC = (0.8–1.5) µM); T. viride (MIC = (6–12) µM) | [106] | |||||
| PxDef | RIPCQYEDATEDTICQQHCLPKGYSYGICVSYRCSCV | 37 | 4233.84/5.27 | Plutella xylostella (Insecta) | B. cinerea (MIC = 15.0 µM); Penicillium crustosum (MIC = 13.0 µM); Colletotrichum gloeosporioides Penz. (MIC = 17.3 µM); Colletotrichum orbiculare (MIC = 12.5 µM); F. oxysporum (MIC = 8.0 µM) | [107] |
| Royalisin | VTCDLLSFKGQVNDSACAANCLSLGKAGGHCEKGVCICRKTSFKDLWDKRF (P17722) | 51 | 5525.45/7.5 | Apis mellifera (Insecta) | B. cinerea (MIC = 4.9 µM *) | [108] |
| Termicin | ACNFQSCWATCQAQHSIYFRRAFCDRSQCKCVFVRG (P82321) | 36 | 4221.89/7.82 | Pseudacanthotermes springer (Insecta) | F. culmorum (MIC = (0.2–0.4) µM); F. oxysporum (MIC = (0.8–1.5) µM); N. haematococca (MIC = (0.05–0.1) µM); Trichoderma viridae (MIC = (6–12) µM) | [106] |
| Cg-Def | GFGCPGNQLKCNNHCKSISCRAGYCDAATLWLRCTCTDCNGKK (Q4GWV4) | 43 | 4642.40/7.53 | Crassostrea gigas (Bivalvia) | B. cinerea (MIC > 20 µM); P. crustosum (MIC > 20 µM); F. oxysporum (MIC = 9 µM) | [109] |
| MGD-1 | GFGCPNNYQCHRHCKSIPGRCGGYCGGWHRLRCTCYRC (P80571) | 38 | 4351.07/7.99 | Mytilus galloprovincialis (Bivalvia) | F. oxysporum (MIC = 5 µM) | [110] |
| DefMT3 | GYYCPFRQDKCHRHCRSFGRKAGYCGNFLKRTCICVKK (A0A089VRA3) | 38 | 4531.39/9.09 | Ixodes ricinus (Arachnida) | F. culmorum (IC50 = 4 µM); F. graminearum 8/1 (IC50 = 4 µM) | [36] |
| DefMT5 | GFFCPYNGYCDRHCRKKLRRRGGYCGGRWKLTCICIMN | 38 | 4533.43/9.11 | F. culmorum (IC50 = 4 µM); F. graminearum 8/1 (IC50 = 4 µM) | ||
| DefMT6 | GFGCPLNQGACHNHCRSIKRRGGYCSGIIKQTCTCYRK | 38 | 4217.95/8.76 | F. culmorum (IC50 = 12 µM); F. graminearum 8/1 (IC50 = 2 µM) | ||
| Holosin 2 | GFGCPLNQRACHRHCRSIGRRGGFCAGLIKQTCTCYRK (A0A5C1Z8V5) | 38 | 4256.05/9.39 | Ixodes holocyclus (Arachnida) | F. graminearum PH-1 (MIC = 5 µM) | [111] |
| Holosin 3 | GFGCPNEWRCNAHCKRNRFRGGYCDSWFRRRCHCYG (A0A5C1ZAY3) | 36 | 4400.01/8.59 | F. graminearum PH-1 (MIC = 5 µM) | ||
| Juruin | FTCAISCDIKVNGKPCKGSGEKKCSGGWSCKFNVCVKV (B3EWQ0) | 38 | 4012.80/7.99 | Avicularia juruensis (Arachnida) | A. niger (MIC= (5–10) µM) | [112] |
| Scapularisin-3 | AFGCPFDQGTCHSHCRSIRRRGERCSGFAKRTCTCYQK (B7Q4Z2) | 38 | 4355.00/8.49 | Ixodes scapularis (Arachnida) | F. culmorum (IC50 = 0.5 µM); F. graminearum 8/1 (IC50 = 1 µM) | [113] |
| Scapularisin-6 | GFGCPFDQGACHRHCQSIGRRGGYCAGFIKQTCTCYHN (Q5Q979) | 38 | 4180.76/7.55 | F. culmorum (IC50 = 1 µM); F. graminearum 8/1 (IC50 = 2 µM) | ||
| Vertebrate defensin | ||||||
| Hepcidin-1 (Hepcidin-6) | CRFCCRCCPRMRGCGLCCRF | 20 | 2374.04/7.77 | Acanthopagrus schlegelii (Sparidae) | A. niger CGMCC 3.316 (MIC = 20–40 µM); F. graminearum CGMCC 3.3490 (MIC = 20–40 µM); F. solani CGMCC 3.5840 (MIC = 20–40 µM) | [114] |
| Hepcidin-2 | SPAGCRFCCGCCPNMRGCGVCCRF (Q68M56) | 24 | 2531.12/7.33 | A. niger CGMCC 3.316 (MIC= 40–60 µM); F. graminearum CGMCC 3.3490 (MIC > 60 µM); F. solani CGMCC 3.5840 (MIC > 60 µM) | ||
| Hepcidin | GCRFCCNCCPNMSGCGVCCRF (P82951) | 21 | 2263.79/7.08 | A. niger (MIC = 44 µM) | [115] | |
| Crotamine | YKQCHKKGGHCFPKEKICLPPSSDFGKMDCRWRWKCCKKGSG (Q9PWF3) | 42 | 4889.85/8.58 | Crotalus durissus terrificus (Viperidae) | A. fumigatus IOC 4526 (MIC >25.5 µM *) | [116] |
| Spheniscin-2 | SFGLCRLRRGFCARGRCRFPSIPIGRCSRFVQCCRRVW (P83430) | 38 | 4507.47/11.47 | Aptenodytes patagonicus (Spheniscidae) | A. fumigatus (MIC= (3–6) µM) | [117] |
| Human drosomycin-like defensin | CLAGRLDKQCTCRRSQPSRRSGHEVGRPSPHCGPSRQCGCHMD | 43 | 4751.43/8.15 | Homo sapiens (Hominidae) | A. fumigatus ATCC MYA1163 (MIC = 6.25 µM); A. nidulans AZN 2867 (MIC = 6.25 µM); Aspergillus ustus (MIC = 12.5 µM); F. solani AZN 6836 (MIC = 25 µM); F. oxysporum (MIC = 6.25 µM) | [118] |
| Fungus defensin | ||||||
| AFP | ATYNGKCYKKDNICKYKAQSGKTAICKCYVKKCPRDGAKCEFDSYKGKCYC (P17737) | 51 | 5805.86/8.34 | Aspergillus giganteus (Trichocomaceae) | Fusarium sporotrichioides IfGB 39/1601 (MIC = 0.02 µM *); F. moniliforme IfGB 39/1402 (MIC = 0.02 µM *); A. niger ATCC 9029 (MIC = 0.17 µM *); A. niger NRRL 372 (MIC = 0.17 µM *); A. niger IfGB 15/1803 (MIC = 0.17 µM *); Fusarium equiseti IfGB 39/0701 (MIC = 0.17 µM *); Fusarium lactis IfGB 39/0701 (MIC = 0.17 µM *); F. oxysporum IfGB 39/1201 (MIC = 0.17 µM *); Fusarium proliferatum IfGB 39/1501 (MIC = 0.17 µM *); Fusarium sp. IfGB 39/1101 (MIC = 0.17 µM *); Aspergillus awamori ATCC 22,342 (MIC = 0.34 µM *); F. oxysporum f.sp. lini IfGB 39/0801 (MIC = 1.38 µM *); Fusarium bulbigenum IfGB 39/0301 (MIC = 1.72 µM *); F. oxysporum f.sp. vasinfectum IfGB 39/1301 (MIC = 1.72 µM *); F. solani IfGB 39/1001 (MIC = 20.67 µM *); Fusarium poae IfGB 39/0901 (MIC = 31 µM *); A. nidulans DSM 969 (MIC = 34.45 µM *); A. nidulans G191 (MIC = 34.45 µM *); A. giganteus IfGB 15/0903 (MIC = 68.90 µM *); A. giganteus MDH 18,894 (MIC = 68.90 µM *); Fusarium aquaeductuum IfGB 39/0101 (MIC = 68.90 µM *); F. culmorum IfGB 39/0403 (MIC = 68.90 µM *) | [119] |
| PAFB | LSKFGGECSLKHNTCTYLKGGKNHVVNCGSAANKKCKSDRHHCEYDEHHKRVDCQTPV (D0EXD3) | 58 | 6500.36/7.74 | Penicillium chrysogenum (Trichocomaceae) | A. fumigatus (MIC = 0.25 µM); A. niger (MIC = 0.50 µM); Aspergillus terreus (MIC = 1 µM) | [120] |
| PAF | AKYTGKCTKSKNECKYKNDAGKDTFIKCPKFDNKKCTKDNNKCTVDTYNNAVDCD (Q01701) | 55 | 6250.08/7.89 | A. fumigatus (MIC = 1 µM); A. niger (MIC = 0.25 µM); A. terreus (MIC = 32 µM) | ||
| AnAFP | LSKYGGECSVEHNTCTYLKGGKDHIVSCPSAANLRCKTERHHCEYDEHHKTVDCQTPV (A2QM98) | 58 | 6517.29/6.24 | A. niger (Trichocomaceae) | A. flavus KCTC 1375 (MIC = 8 µM); A. fumigatus KCTC 6145 (MIC = (4–8) µM); F. oxysporum KCTC 6076 (MIC = (8–15) µM); F. solani KCTC 6326 (MIC = 8 µM) | [121] |
| AcAFP | ATYDGCKCYKKDNICKYKAQSGKT (D3Y2M3) | 24 | 2717.14/8.43 | Aspergillus clavatus (Trichocomaceae) | F. oxysporum (MIC = 8.57 µM *); F. oxysporum (IC50 = 1.25 µM *) | [122] |
| NFAP | LEYKGECFTKDNTCKYKIDGKTYLAKCPSAANTKCEKDGNKCTYDSYNRKVKCDFRH (A1D8H8) | 57 | 6625.56/7.92 | Neosartorya fischeri (Trichocomaceae) | A. niger (MIC = (3.77–15.09) µM *); A. nidulans (MIC = 30.19 µM *) | [123] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leannec-Rialland, V.; Atanasova, V.; Chereau, S.; Tonk-Rügen, M.; Cabezas-Cruz, A.; Richard-Forget, F. Use of Defensins to Develop Eco-Friendly Alternatives to Synthetic Fungicides to Control Phytopathogenic Fungi and Their Mycotoxins. J. Fungi 2022, 8, 229. https://doi.org/10.3390/jof8030229
Leannec-Rialland V, Atanasova V, Chereau S, Tonk-Rügen M, Cabezas-Cruz A, Richard-Forget F. Use of Defensins to Develop Eco-Friendly Alternatives to Synthetic Fungicides to Control Phytopathogenic Fungi and Their Mycotoxins. Journal of Fungi. 2022; 8(3):229. https://doi.org/10.3390/jof8030229
Chicago/Turabian StyleLeannec-Rialland, Valentin, Vessela Atanasova, Sylvain Chereau, Miray Tonk-Rügen, Alejandro Cabezas-Cruz, and Florence Richard-Forget. 2022. "Use of Defensins to Develop Eco-Friendly Alternatives to Synthetic Fungicides to Control Phytopathogenic Fungi and Their Mycotoxins" Journal of Fungi 8, no. 3: 229. https://doi.org/10.3390/jof8030229







