Genomic and Chemical Profiling of B9, a Unique Penicillium Fungus Derived from Sponge
Abstract
:1. Introduction
2. Materials and Methods
2.1. Stains
2.2. Species Identification
2.3. Intraspecies Comparison
2.4. Whole Genome Sequencing
2.5. Functional Genome Annotation
2.6. Extractions of Secondary Metabolites (SMs)
2.7. In Vitro PD-1/PD-L1 Binding Assay
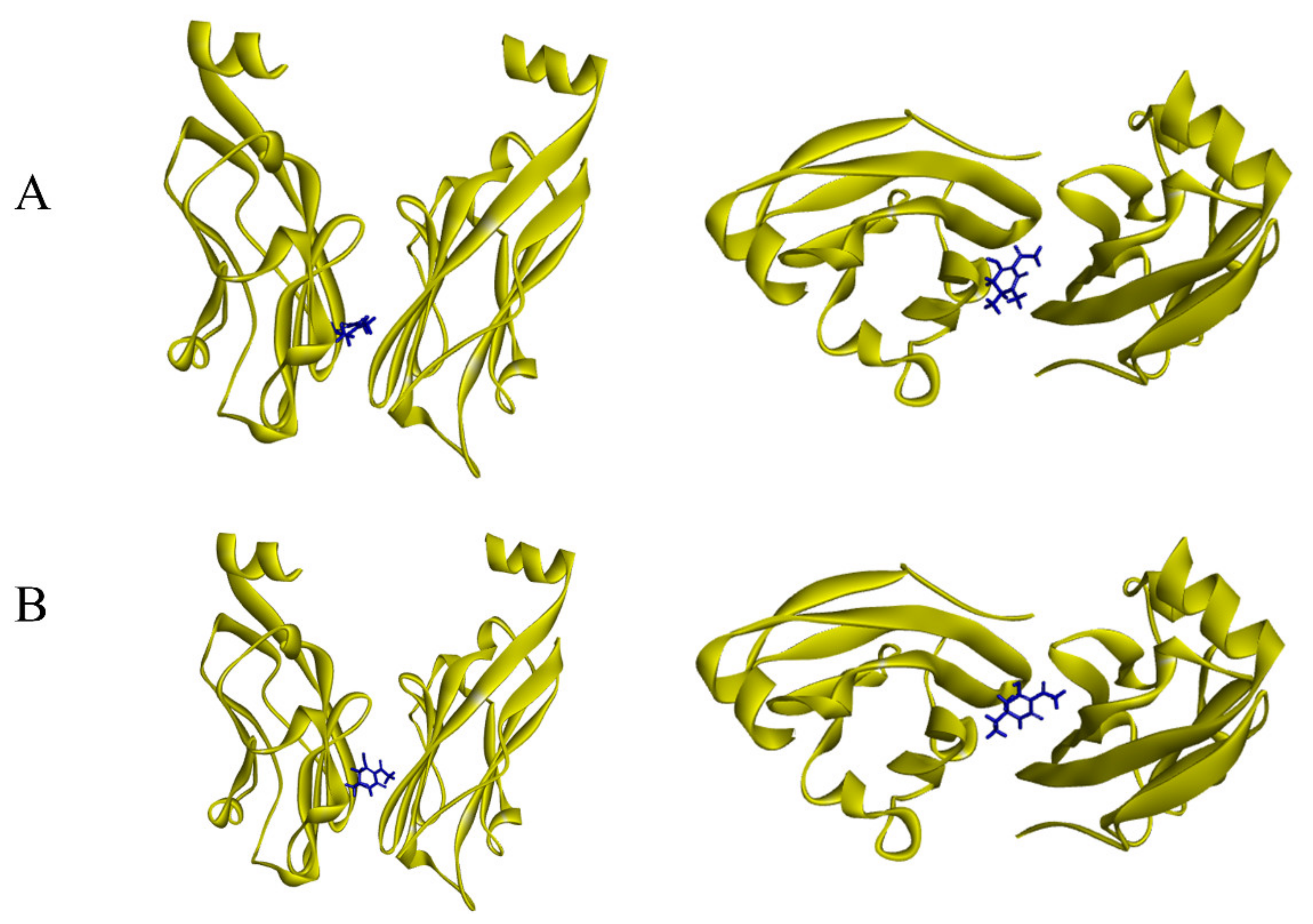
2.8. Docking Studies
3. Results
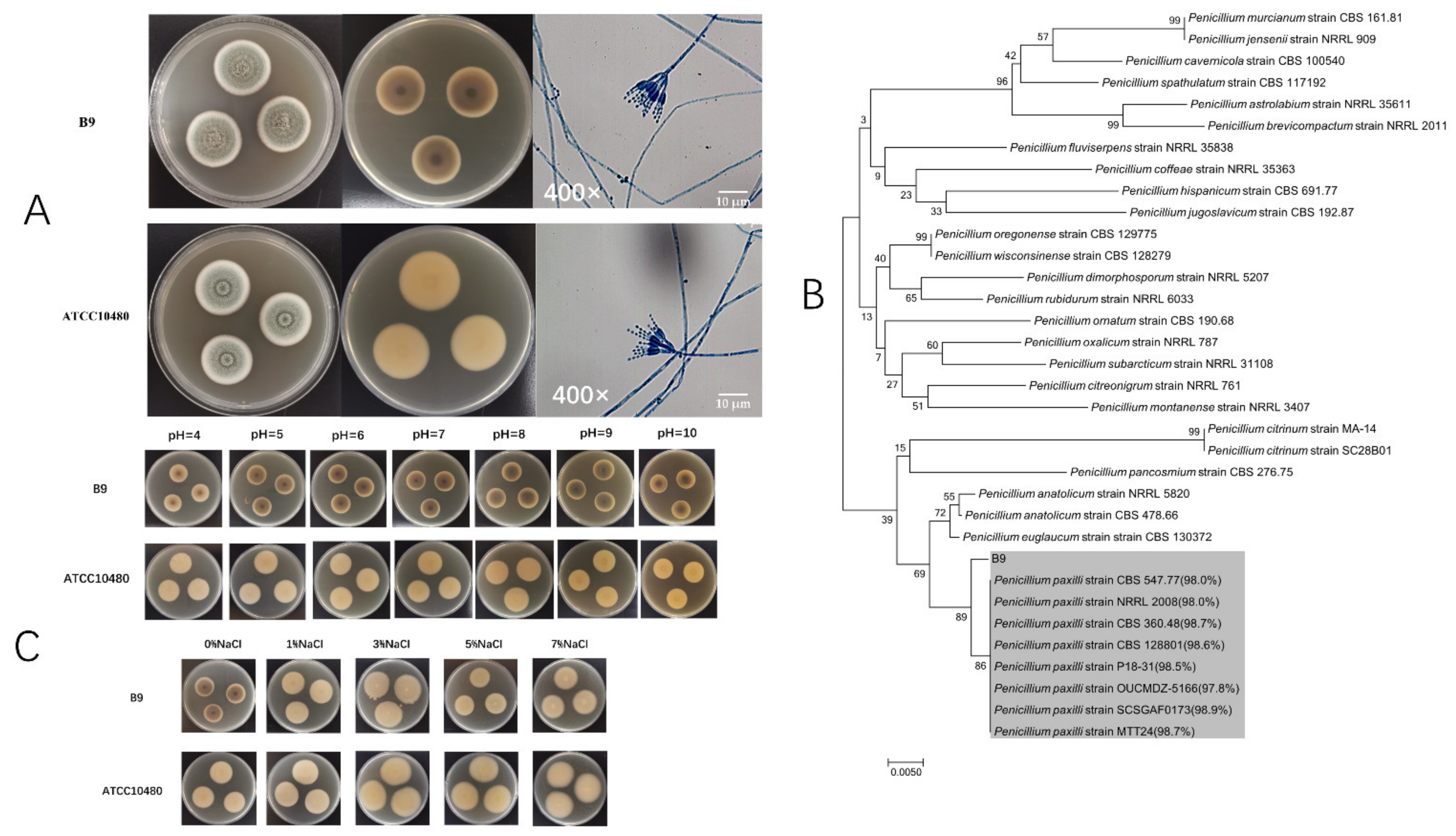
3.1. Species Identification of Strain B9
3.2. Genome Sequencing
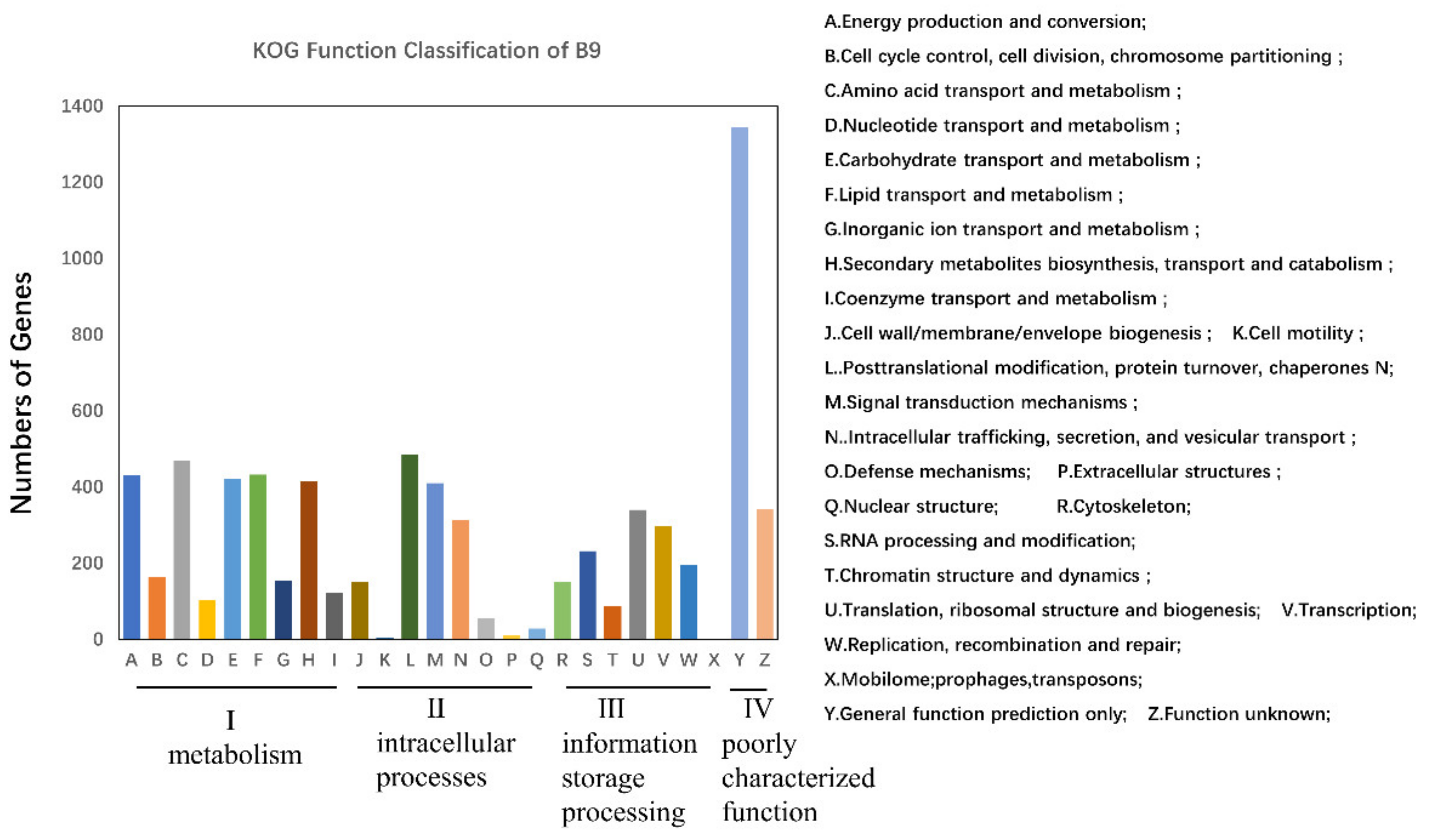
3.3. Genomic Functional Annotation
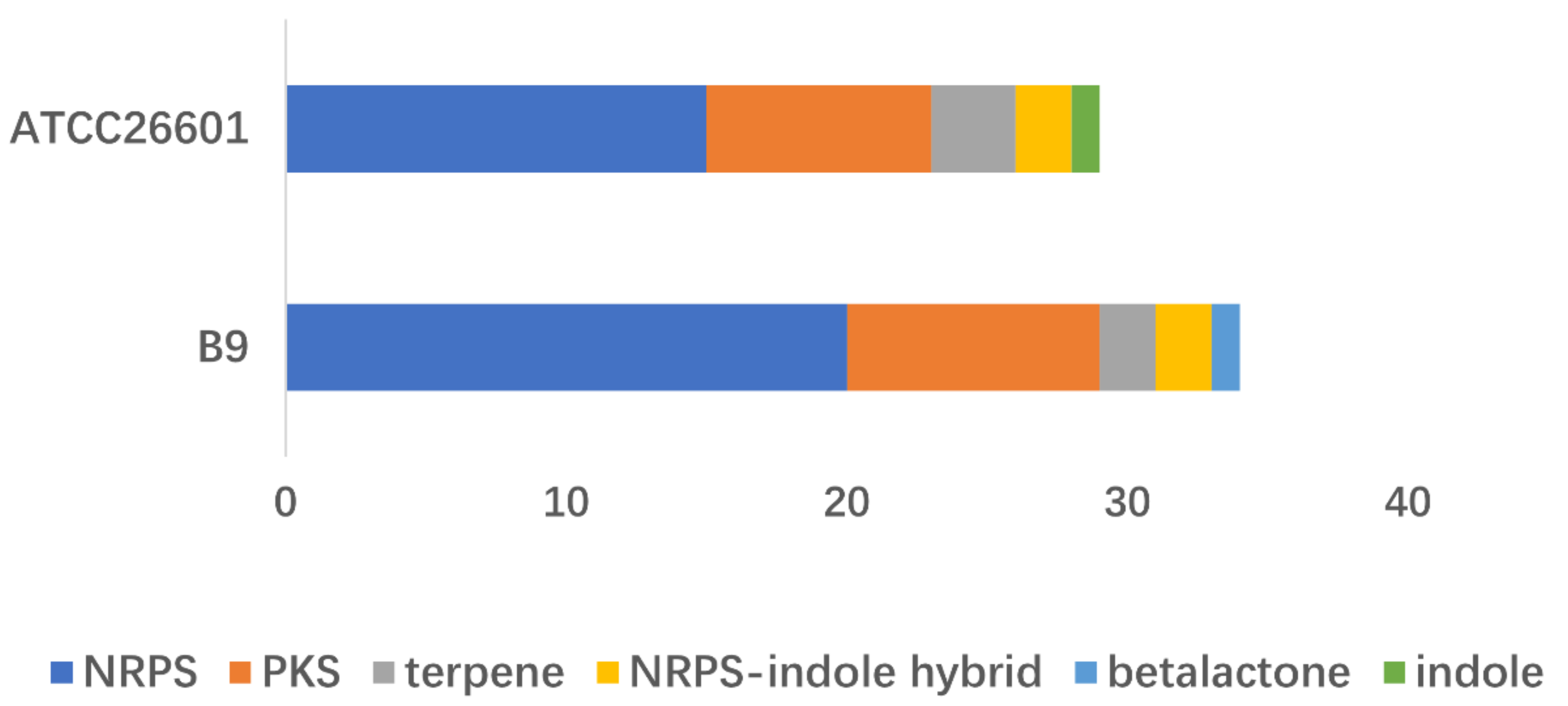
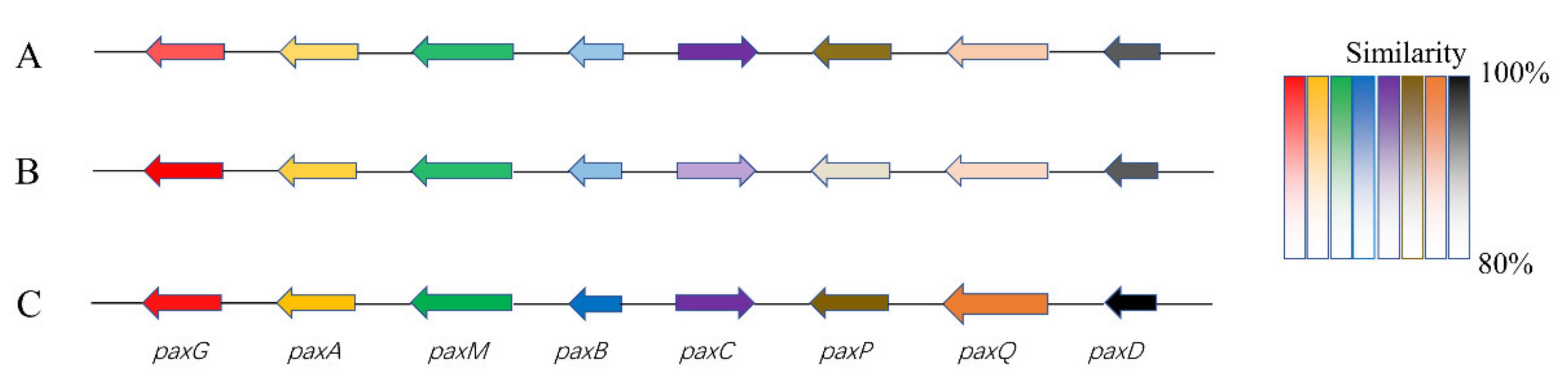
3.4. Prediction of the Biosynthetic Gene Clusters (BGCs)
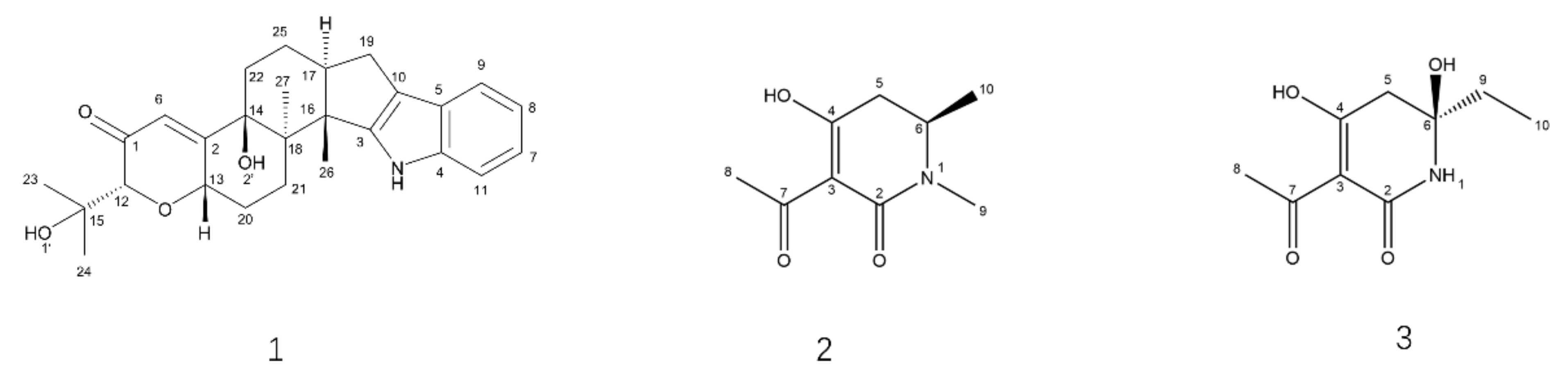
3.5. SMs Isolation and Chemical Characterization
3.6. Biological Activities of New Compounds
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MEA | malt extract agar |
| BGC | biosynthetic gene cluster |
| NMR | nuclear magnetic resonance |
| MS | mass spectrometry |
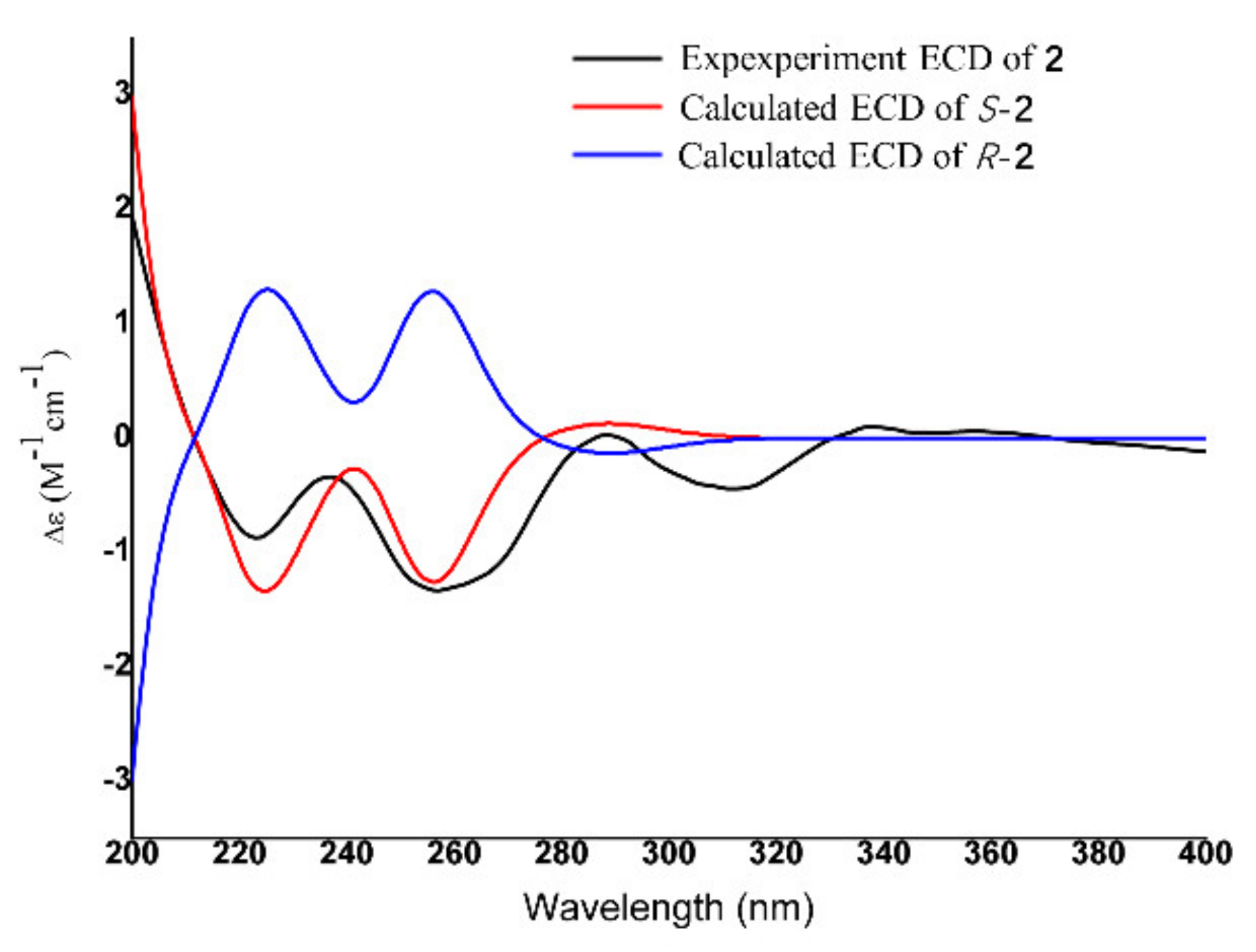
| ECD | electronic circular dichroism |
| PD-1 | programmed cell death protein 1 |
| PD-L1 | programmed cell death 1 ligand 1 |
| SM | secondary metabolite |
| BK | big potassium |
| ITS | internal transcribed spacer |
| MEGA7 | Molecular Evolutionary Genetics Analysis 7 |
| KOG | Cluster of Orthologous Groups of proteins |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| antiSMASH | antibiotics & Secondary Metabolite Analysis Shell |
| NRPS | nonribosomal peptide synthase |
| PKS | polyketide synthase |
| AChE | acetylcholine esterase |
| MIBiG | Minimum Information about a Biosynthetic Gene cluster (MIBiG) |
| HRESIMS | high-resolution electrospray ionization mass spectroscopy |
| HMBC | heteronuclear multiple bond correlation |
| HTRF | homogeneous time-resolved fluorescence |
| PDA | potato dextrose agar |
| LPCB | lactophenol cotton blue |
| BLAST | Basic Local Alignment Search Tool |
| PBS | phosphate buffered saline |
| HPLC | high-performance liquid chromatography |
| CaM | calmodulin |
References
- Kaleem, S.; Qin, L.; Yi, W.; Lian, X.Y.; Zhang, Z. Bioactive Metabolites from the Mariana Trench Sediment-Derived Fungus Penicillium sp. SY2107. Mar. Drugs 2020, 18, 258. [Google Scholar] [CrossRef]
- Liu, S.; Su, M.; Song, S.J.; Jung, J.H. Marine-Derived Penicillium Species as Producers of Cytotoxic Metabolites. Mar. Drugs 2017, 15, 329. [Google Scholar] [CrossRef] [Green Version]
- Robson, G.D.; van West, P.; Gadd, G. Exploitation of Fungi; Cambridge University Press: Cambridge, UK, 2007; Volume 26. [Google Scholar]
- Ariantari, N.P.; Ancheeva, E.; Wang, C.; Mándi, A.; Knedel, T.O.; Kurtán, T.; Chaidir, C.; Müller, W.E.G.; Kassack, M.U.; Janiak, C.; et al. Indole Diterpenoids from an Endophytic Penicillium sp. J. Nat. Prod. 2019, 82, 1412–1423. [Google Scholar] [CrossRef]
- Ludlow, E.J.; Vassiliadis, S.; Ekanayake, P.N.; Hettiarachchige, I.K.; Reddy, P.; Sawbridge, T.I.; Rochfort, S.J.; Spangenberg, G.C.; Guthridge, K.M. Analysis of the Indole Diterpene Gene Cluster for Biosynthesis of the Epoxy-Janthitrems in Epichloë Endophytes. Microorganisms 2019, 7, 560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Imlach, W.L.; Finch, S.C.; Dunlop, J.; Meredith, A.L.; Aldrich, R.W.; Dalziel, J.E. The molecular mechanism of “ryegrass staggers,” a neurological disorder of K+ channels. J. Pharmacol. Exp. Ther. 2008, 327, 657–664. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, Y.; Xia, X.M.; Lingle, C.J. The functionally relevant site for paxilline inhibition of BK channels. Proc. Natl. Acad. Sci. USA 2020, 117, 1021–1026. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Du, X.; Li, H.; Wang, Y. Genome sequence of Aspergillus flavus A7, a marine-derived fungus with antibacterial activity. Genome 2021, 64, 719–733. [Google Scholar] [CrossRef]
- Du, X.; Li, H.; Qi, J.; Chen, C.; Lu, Y.; Wang, Y. Genome mining of secondary metabolites from a marine-derived Aspergillus terreus B12. Arch. Microbiol. 2021, 203, 5621–5633. [Google Scholar] [CrossRef]
- Leck, A. Preparation of lactophenol cotton blue slide mounts. Community Eye Health 1999, 12, 24. [Google Scholar]
- Wu, B.; Wiese, J.; Schmaljohann, R.; Imhoff, J.F. Biscogniauxone, a New Isopyrrolonaphthoquinone Compound from the Fungus Biscogniauxia mediterranea Isolated from Deep-Sea Sediments. Mar. Drugs 2016, 14, 204. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Stanke, M.; Keller, O.; Gunduz, I.; Hayes, A.; Waack, S.; Morgenstern, B. AUGUSTUS: Ab initio prediction of alternative transcripts. Nucleic Acids Res. 2006, 34, W435–W439. [Google Scholar] [CrossRef] [Green Version]
- Galperin, M.Y.; Wolf, Y.I.; Makarova, K.S.; Vera Alvarez, R.; Landsman, D.; Koonin, E.V. COG database update: Focus on microbial diversity, model organisms, and widespread pathogens. Nucleic Acids Res. 2021, 49, D274–D281. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Kloosterman, A.M.; Charlop-Powers, Z.; van Wezel, G.P.; Medema, M.H.; Weber, T. antiSMASH 6.0: Improving cluster detection and comparison capabilities. Nucleic Acids Res. 2021, 49, W29–W35. [Google Scholar] [CrossRef]
- Fox, E.M.; Howlett, B.J. Biosynthetic gene clusters for epipolythiodioxopiperazines in filamentous fungi. Mycol. Res. 2008, 112, 162–169. [Google Scholar] [CrossRef]
- Wheadon, M.J.; Townsend, C.A. Evolutionary and functional analysis of an NRPS condensation domain integrates β-lactam, d-amino acid, and dehydroamino acid synthesis. Proc. Natl. Acad. Sci. USA 2021, 118, e2026017118. [Google Scholar] [CrossRef]
- Gsaller, F.; Blatzer, M.; Abt, B.; Schrettl, M.; Lindner, H.; Haas, H. The first promoter for conditional gene expression in Acremonium chrysogenum: Iron starvation-inducible mir1(P). J. Biotechnol. 2013, 163, 77–80. [Google Scholar] [CrossRef]
- Shrestha, S.; Seong, S.H.; Paudel, P.; Jung, H.A.; Choi, J.S. Structure Related Inhibition of Enzyme Systems in Cholinesterases and BACE1 In Vitro by Naturally Occurring Naphthopyrone and Its Glycosides Isolated from Cassia obtusifolia. Molecules 2017, 23, 69. [Google Scholar] [CrossRef] [Green Version]
- Baxter, A.; Fitzgerald, B.J.; Hutson, J.L.; McCarthy, A.D.; Motteram, J.M.; Ross, B.C.; Sapra, M.; Snowden, M.A.; Watson, N.S.; Williams, R.J.; et al. Squalestatin 1, a potent inhibitor of squalene synthase, which lowers serum cholesterol in vivo. J. Biol. Chem. 1992, 267, 11705–11708. [Google Scholar] [CrossRef]
- Oikawa, H.; Yokota, T.; Sakano, C.; Suzuki, Y.; Naya, A.; Ichihara, A. Solanapyrones, Phytotoxins Produced by Alternaria solani: Biosynthesis and Isolation of Minor Components. Biosci. Biotechnol. Biochem. 1998, 62, 2016–2022. [Google Scholar] [CrossRef] [Green Version]
- Scott, B.; Young, C.A.; Saikia, S.; McMillan, L.K.; Monahan, B.J.; Koulman, A.; Astin, J.; Eaton, C.J.; Bryant, A.; Wrenn, R.E.; et al. Deletion and gene expression analyses define the paxilline biosynthetic gene cluster in Penicillium paxilli. Toxins 2013, 5, 1422–1446. [Google Scholar] [CrossRef]
- Nicoletti, R.; Trincone, A. Bioactive Compounds Produced by Strains of Penicillium and Talaromyces of Marine Origin. Mar. Drugs 2016, 14, 37. [Google Scholar] [CrossRef] [Green Version]
- Vu, D.; Groenewald, M.; de Vries, M.; Gehrmann, T.; Stielow, B.; Eberhardt, U.; Al-Hatmi, A.; Groenewald, J.Z.; Cardinali, G.; Houbraken, J.; et al. Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud. Mycol. 2019, 92, 135–154. [Google Scholar] [CrossRef]
- Garnica, S.; Schön, M.E.; Abarenkov, K.; Riess, K.; Liimatainen, K.; Niskanen, T.; Dima, B.; Soop, K.; Frøslev, T.G.; Jeppesen, T.S.; et al. Determining threshold values for barcoding fungi: Lessons from Cortinarius (Basidiomycota), a highly diverse and widespread ectomycorrhizal genus. FEMS Microbiol. Ecol. 2016, 92, fiw045. [Google Scholar] [CrossRef] [Green Version]
- Tagami, K.; Liu, C.; Minami, A.; Noike, M.; Isaka, T.; Fueki, S.; Shichijo, Y.; Toshima, H.; Gomi, K.; Dairi, T.; et al. Reconstitution of biosynthetic machinery for indole-diterpene paxilline in Aspergillus oryzae. J. Am. Chem. Soc. 2013, 135, 1260–1263. [Google Scholar] [CrossRef]
- Guzik, K.; Tomala, M.; Muszak, D.; Konieczny, M.; Hec, A.; Błaszkiewicz, U.; Pustuła, M.; Butera, R.; Dömling, A.; Holak, T.A. Development of the Inhibitors that Target the PD-1/PD-L1 Interaction-A Brief Look at Progress on Small Molecules, Peptides and Macrocycles. Molecules 2019, 24, 2071. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zavareh, R.B.; Spangenberg, S.H.; Woods, A.; Martínez-Peña, F.; Lairson, L.L. HSP90 Inhibition Enhances Cancer Immunotherapy by Modulating the Surface Expression of Multiple Immune Checkpoint Proteins. Cell Chem. Biol. 2021, 28, 158–168. [Google Scholar] [CrossRef] [PubMed]


| Genome | Value |
|---|---|
| Assembly size (Mp) | 32.96 |
| G + C (%) | 47.7 |
| Assembled scaffolds | 185 |
| N50 length (bp) | 1,131,010 |
| average length (bp) | 186,810 |
| Predicted Protein-Coding Genes | 11,110 |
| average length of Predicted Protein-Coding Genes (bp) | 1545.92 |
| average depth of reads cover | 189.27 |
| total non-coding RNA | 269 |
| Sequencing Method | Illumina HiSeq |
| No. | 13C NMR | 1H NMR | HMBC |
|---|---|---|---|
| 1 | |||
| 2 | 170.7 | ||
| 3 | 109.7 | ||
| 4 | 191.8 | ||
| 5 | 42.8 | 2.49 (1H, dd, 15.9, 5.2); 2.34 (1H, dd, 15.9, 6.2) | C10, C6, C4, C3 |
| 6 | 47.0 | 3.76 (m) | C10, C5, C4, C2 |
| 7 | 198.9 | ||
| 8 | 31.2 | 2.41 (3H, s) | C7 |
| 9 | 21.2 | 2.37 (3H, s) | C2 |
| 10 | 17.8 | 1.32 (3H, d, 6.6) | C6, C5, C4 |
| No. | 13C NMR | 1H NMR | HMBC |
|---|---|---|---|
| 1 | |||
| 2 | 179.7 | ||
| 3 | 107.9 | ||
| 4 | 201.6 | ||
| 5 | 39.2 | 2.87 (1H, d, 15.2); 2.62 (1H, d, 15.2) | C4, C5, C6, C9 |
| 6 | 77.3 | ||
| 7 | 197.1 | ||
| 8 | 27.3 | 2.42 (3H, s) | C3, C7 |
| 9 | 30.1 | 1.66 (1H, dt, 17.5, 6,2); 1.73 (1H, dt, 17.5,6.2) | C4, C5, C6, C10 |
| 10 | 6.85 | 0.89 (3H, t, 6.2) | C6, C9 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, C.; Qi, J.; He, Y.; Lu, Y.; Wang, Y. Genomic and Chemical Profiling of B9, a Unique Penicillium Fungus Derived from Sponge. J. Fungi 2022, 8, 686. https://doi.org/10.3390/jof8070686
Chen C, Qi J, He Y, Lu Y, Wang Y. Genomic and Chemical Profiling of B9, a Unique Penicillium Fungus Derived from Sponge. Journal of Fungi. 2022; 8(7):686. https://doi.org/10.3390/jof8070686
Chicago/Turabian StyleChen, Chaoyi, Jiangfeng Qi, Yajing He, Yuanyuan Lu, and Ying Wang. 2022. "Genomic and Chemical Profiling of B9, a Unique Penicillium Fungus Derived from Sponge" Journal of Fungi 8, no. 7: 686. https://doi.org/10.3390/jof8070686






